Abstract
Lower-grade gliomas (LGGs) exhibit highly variable clinical behaviors, while classic histology characteristics cannot accurately reflect the authentic biological behaviors, clinical outcomes, and prognosis of LGGs. In this study, we carried out analyses of whole exome sequencing, RNA sequencing and DNA methylation in primary vs. recurrent LGG samples, and also combined the multi-omics data to construct a prognostic prediction model. TCGA-LGG dataset was searched for LGG samples. 523 samples were used for whole exome sequencing analysis, 532 for transcriptional analysis, and 529 for DNA methylation analysis. LASSO regression was used to screen genes with significant association with LGG survival from the frequently mutated genes, differentially expressed genes, and differentially methylated genes, whereby a prediction model for prognosis of LGG was further constructed and validated. The most frequently mutated diver genes in LGGs were IDH1 (77%), TP53 (48%), ATRX (37%), etc. Top significantly up-regulated genes were C6orf15, DAO, MEOX2, etc., and top significantly down-regulated genes were DMBX1, GPR50, HMX2, etc. 2077 genes were more and 299 were less methylated in recurrent vs. primary LGG samples. Thirty-nine genes from the above analysis were included to establish a prediction model of survival, which showed that the high-score group had a very significantly shorter survival than the low-score group in both training and testing sets. ROC analysis showed that AUC was 0.817 for the training set and 0.819 for the testing set. This study will be beneficial to accurately predict the survival of LGGs to identify patients with poor prognosis to take specific treatment as early, which will help improve the treatment outcomes and prognosis of LGG.
Similar content being viewed by others
Introduction
Brain and nervous system tumors are fatal tumors that have an estimation of over 300,000 new cases (1.6%) and over 250,000 cancer death (2.5%) globally in 20201. A recent nationwide epidemiological investigation in China revealed that in 2016, there were about 109,000 new cases and 585,000 cancer-death of brain tumor2. Malignant primary brain tumors belong to the most difficult-to-treat cancer, with a 5-year overall survival of under 35%3. In adults, gliomas are the most common type of malignant brain tumors, making up 80% of all the malignant brain tumors3,4, with a universally fatal outcome due to common recurrence and progression. Current routine glioma treatment includes surgery, radiation, chemotherapy, immunotherapy5, targeted therapy6, etc., which results in unsatisfactory outcomes and prognosis.
Lower-grade gliomas (LGGs) are defined as World Health Organization (WHO) grades II and III gliomas, including diffuse low-grade and intermediate-grade gliomas, accounting for approximately 20% of glioma cases7. LGG has a relative better survival compared with high grade glioma, with an average survival of about 7 years8,9. However, all LGGs will eventually progress to glioblastoma and death8. Due to their highly invasive nature, it is impossible to give complete neurosurgical resection, and thus the residual tumors lead to recurrence and malignant progression to grade IV glioblastomas. LGGs exhibit highly variable clinical behaviors in that some subsets of gliomas remain stable for years while some will quickly progress to glioblastoma (WHO grade IV gliomas) with high extend of malignancy within months. Accurate classification of LGGs based on prediction of prognosis is necessary and paramount for early and precise treatment to achieve satisfactory outcomes and prognosis. However, classic histology characteristics cannot accurately reflect the authentic biological behaviors, clinical outcomes, and prognosis of LGGs.
Genomics identifies mutations related to carcinogenesis and cancer progression. Transcriptomics defines genomics regulation particularly related to hallmarks of cancers10. DNA methylation as a major epigenetic signature participates in the regulation of pathogenesis and progression of cancer11,12,13. Some studies employed uni-omics data to predict the prognosis of LGGs, with unsatisfactory results. For example, Kang et al. combined five lncRNAs and three immune cell types to construct a risk model for prediction of survival of LGG14. Lin et al. identified differentially expressed lncRNAs in radiosensitive vs radioresistant patients to select and construct a three-lncRNA (contributing to pathogenic processes) signature to predict overall survival of LGGs after radiotherapy15. Tan et al. combined six immune associated genes (CD163, FPR3, LPAR5, P2ry12, PLAUR, SIGLEC1) to construct a prognostic model for LGG, where the calculated risk score could be used to differentiate the overall survival rates of LGG16.
The carcinogenesis and cancer progression are attributable to multiple gene effects17,18,19. Therefore, multiple omics data can more accurately provide clues for the mechanistic studies, prediction of treatment outcomes and prognosis, etc. of cancers. Multi-omics analysis is a more promising way to predict the LGGs prognosis. A study combined analyses of whole exome sequence (WES), DNA copy number, transcriptional analysis (messenger RNA expression, microRNA expression), DNA methylation, and targeted protein expression to classify three nonoverlapping, prognostically significant subtypes of LGGs; IDH mutation and 1p/19q codeletion conferred LGGs most favorable clinical outcomes, and most LGGs without IDH mutation molecularly and clinically exhibited similarity to glioblastoma20. Pan et al. developed an i-Modern model, which combined multi-omics, including somatic mutations, copy number variation, transcription profile, miRNA, DNA methylation, etc. with deep learning network to predict the prognosis (high vs low-risk glioma), and thus accurately stratified gliomas patients21. However, these studies were in term of all types of gliomas instead of the sole LGGs. Until now, the analysis integrating multi-omics data to predict prognosis of LGGs is still lacking.
In this study, we collected 523, 532, 529 samples of LGGs from TCGA database to carry out analysis of WES, RNA sequencing and DNA methylation to profile the frequently mutated genes, differentially expressed genes (DEGs) and differentially methylated genes (DMGs) in primary vs. recurrent LGG samples and then to combine these multi-omics data to establish a prediction model for the survival of LGGs. This study will be beneficial to accurately predict the survival of LGGs to identify patients with poor prognosis so as to take specific treatment as early, which will be helpful improve the treatment outcomes and prognosis of LGG.
Results
WES sequencing
523 samples of LGGs underwent WES sequencing, which showed 504 (96.37%) had mutations in driver genes. The most frequently mutated diver genes were IDH1 (77%), TP53 (48%), ATRX (37%), CIC (22%), TTN (11%), FUBP1 (9%), PIK3CA (9%), NOTCH1 (7%), MUC16 (6%), EGFR (6%), etc. (Fig. S1). Missense mutation was the most common mutations type (Fig. S1). In addition, a few LGG samples showed very high tumoral burden.
The mutation profile of LGG samples was shown in Fig. 1. Missense mutation was the most common mutations type (Fig. 1A, F–I), and single nucleotide polymorphism (SNP) was the most common variant type among the SNP, insertion (INS) and deletion (Del) in LGG samples (Fig. 1B). For SNV, C > T was the most common subtype (Fig. 1C, G). The mutation number per sample was shown. Normally, the mutation number in LGG patients ranged from tens to hundreds, while a few samples had extremely high mutation number (up to 9000, Fig. 1D, E). The ratios of transition (Ti) and transversion (Tv) were shown, and the Ti percentage was about 75% (Fig. 1H).
Mutation profile in LGG tissues. (A, B). the distribution of various variant classification and mutational events (SNP, insertion, and deletion) in all the LGG cases, respectively. (C). the number of SNV subtypes (i.e., T > G, T > A, T > C, C > T, C > G, and C > A) in all the LGG cases. (D). the average mutation number in each LGG sample. (E). the average number of the classified variants in all the LGG cases. (F). ten top frequently mutated genes and the components of the involved mutation types. (G). the percentages of each type of mutation in LGG samples; (H). the percentages of transition (Ti) and transversion (Tv). (I). the distribution of mutation types in each LGG sample.
The mutation signature in each LGG sample were shown. Among the 30 COSMIC signatures, Signature 1 had the most obvious contribution to mutations in each sample, followed by Signature 14 (Fig. S2A). The correlation heat map about the mutation profile in each sample was shown, where the red and yellow indicated the higher correlation samples while the blue showed the lower correlation samples (Fig. S2B).
KEGG enrichment analysis revealed that the frequently mutated genes were predominantly enriched in neuron function-related signal pathways, such as ‘focal adhesion’, ‘axon guidance’, and ‘glutamatergic synapse’22 (Fig. 2A). GO analysis showed that the frequently mutated genes were enriched in actin cytoskeleton, cell leading edge, etc. by cellular component (CC) enrichment, in axon development, axonogenesis, synapse organization, cell junction assembly, etc. by biological process (BP) enrichment, in GTPase regulator activity, nucleoside-triphosphatase regulator activity, etc. by molecular function (MF) enrichment (Fig. 2B–D).
RNA sequencing
Recurrence is an important characteristics of tumor malignancy, therefore, we observed the transcriptomics in 514 primary versus 18 recurrent LGG tissues through RNA sequencing. The differentially expressed RNAs with adjusted p values (padj) less than 0.05 and log2 fold change > 2 were selected. The significantly up- and down-regulated genes in recurrent vs. primary LGG tissues were shown in Tables S1and S2, most of which were protein-coding genes, and the remaining were mostly lncRNAs. Top 50 significantly up-regulated DEGs were C6orf15, DAO, MEOX2, LINC02587, LTF, MIR3976HG, LINC00507, LINC00588, LGR6, LINC02822, etc. (Table 1). Top 50 significantly down-regulated DEGs were DMBX1, GPR50, HMX2, MAFA, COL2A1, TLX3, KC877982.1, BARX1, LBX1-AS1, MEOX1, etc. (Table 2).
The relationship between the RNA samples from the primary and recurrent LGG tissues was primarily analyzed through the inter-sample PCA dimension-reduction cluster analysis. A plot was profiled based on the first and second primary components (PC1 and PC2) to show the gathering among samples (Fig. 3A). The correlation matrix across all samples according to the gene expression of each sample, whereby the heat map of the correlation matrix after clustering was mapped (Fig. 3B).
The inter-sample PCA dimension-reduction cluster analysis of the gene expression in primary and recurrent samples. (A). plot based on the first and second primary components (PC1 and PC2) showed the gathering among the primary (shown in red) and recurrent tumor (shown in blue) tissues. (B). the correlation matrix across all samples according to the gene expression of each sample, whereby the heat map of the correlation matrix after clustering was mapped.
The Trans Per Million (TPM) and Fragments Per Kilobase Million (FPKM) of recurrent vs. primary LGG tissues were shown in Fig. S3. The distribution of the expression in primary and recurrent samples were similar, without obvious bias, which meant that they were both suitable for the subsequent analysis. Particularly, TPM is more suitable for comparison between these LGG samples due to the standardization of sequencing depth by TPM.
Heat-map of DEGs showed the primary LGG samples were mainly clustered in the middle and the recurrent samples were in both sides (Fig. 4A). Volcano Plot showed DEGs with padj < 0.05 and log2 fold change > 2 (Fig. 4B). The chromosomal distribution analysis showed that the recurrence-related DEGs were lest aggregated in Chromosome 13 (Fig. 4C).
Functional enrichment analysis revealed that the DEGs were mainly enriched in neuroactive ligand-receptor interaction by KEGG enrichment23 (Fig. 5A), in collagen-containing extracellular matrix, chromosomal region, condensed chromosome, etc. by GO-CC enrichment (Fig. 5B), in organelle fission, nuclear division, extracellular matrix organization, etc. by GO-BP enrichment (Fig. 5C), and in channel activity, passive transmembrane transporter activity, etc. by GO-MF enrichment (Fig. 5D). In addition, the general expression variation trend of the entries in each sample in four databases of HALLMARK, KEGG, GO-BP and REACTOME underwent GSEA analysis. It could be seen that the enrichment pathways in HALLMARK and KEGG mainly exhibited a down-regulation trend, while those in GO-BP and REACTOME mainly showed an up-regulation trend (Fig. 5E–H).
DNA methylation
After exclusion of the samples with incomplete information, 529 samples were analyzed for DNA methylation, including 515 primary and 14 recurrent LGG tissues. PCA dimension-reduction analysis demonstrated some profile difference between the recurrent and primary samples (Fig. 6A). The heatmap of the differential methylation of each sample in recurrent vs. primary LGG samples were shown (Fig. 6B).
Volcanic plot of differential methylation between the primary and recurrent LGG samples showed 2077 genes were more methylated and 299 were less methylated in recurrent vs. primary samples (Fig. S4A). The distribution analysis showed the differential methylation probes were mainly within genes (including gene body and IGR) (Fig. S4B).
The DMEs were mainly enriched in neuroactive ligand-receptor interaction, cAMP signaling pathway, focal adhesion, axon guidance, etc. by KEGG enrichment24, in protein serine/threonine/tyrosine-kinase activity, channel activity, passive transmembrane transportor activity, protein serine/threonine kinase activity, etc. by GO MF enrichment, in axon development, axonogenesis, synapse organization, regulation of neuron projection development, etc. by GO BP enrichment, and in presynapse, cell- cell junction, collagen-containing extracellular matrix, synaptic membrane, etc. by GO CC enrichment (Fig. 7).
Prediction model for LGG survival
LASSO analysis showed that among the frequently mutated genes, IDH2, CIC, IDH1, MUC16, EGFR and SMARCA4 were most associated with LGG survival. These gene were further used to build a model, which showed that in the training set, high model score group had a very significant shorter survival than the low-score group (p < 0.0001), with a ROC AUC of 0.715 (Fig. S5A,B), which was validated in the testing set, with a ROC AUC of 0.721 (Fig. S5C,D).
Among the DEGs, 22 genes, including BNC1, LINC02587, HOXC4, H2BC12, IGF2BP2, ABCC3, HMX1, AGMO, OTP, CMYA5, WEE1, MND1, AL390755.1, TNFRSF11B, SCNN1B, SRY, IRX5, HOXA7, LINC01965, ISL2, DMRTA2, and IGFBP2 were most associated with LGG survival. A model combination of these DEGs showed in the training set, high model score group had a very significant shorter survival than the low-score group (p < 0.0001), with a ROC AUC of 0.859 (Fig. S6A,B), which was validated in the testing set, with a ROC AUC of 0.765 (Fig. S6C,D).
DMEs analysis by LASSO showed that KIAA1598, TUBA1B, WEE1, PARK2, CRYGD, PDGFB, E2F2, GLT25D2, BDNFOS, DHX36, and SRRM4 methylation was significantly associated with LGG survival. A model combining these DMEs indicated that in the training set, the high model score group had a very significantly shorter survival than the low-score group (p < 0.0001, Fig. S7A), which was also found in the testing set (Fig. S7C). The ROC analysis showed an AUC of 0.742 for training set (Fig. S7B) and 0.722 for testing set (Fig. S7D).
Eventually, 39 genes from the analysis of genomic, RNA sequencing and DNA methylation were combined to establish a prediction model for survival of LGG patients. In the training set, the high model score group had a very significantly shorter survival than high-score group (p < 0.0001, Fig. 8A). ROC analysis showed that AUC for the model was 0.817 (Fig. 8A–B). Similar result was observed in the testing set, with an AUC for ROC analysis of 0.819 (Fig. 8C–D).
Establishment of nomogram prognostic model
After constructing the risk scoring model, we developed single-omics and multi-omics Nomogram prognostic models to analyze the predictive capabilities of various feature genes for prognosis. The Nomogram model from the single-omics data revealed, consistent with clinical outcomes, that IDH2 was the most powerful predictor in the WES model (Fig. S8A). In the transcriptomics model, the most influential genes were BNC1 (ENSG00000169594.13) and AGMO (ENSG00000187546.14) (Fig. S8C). For the methylation model, SRRM4 emerged as the strongest predictor (Fig. S8E). Upon integrating information from all three omics, the genes with the highest predictive capabilities were the transcriptomic expressions of WEE1 (ENSG00000166483.11), BNC1, and AGMO (Fig. S8G). Survival ROC analysis for both single-omics and multi-omics showed that, similar to the risk scoring model, integrating the three omics data improved the accuracy and stability of the predictive models (Figs. S8B, S8D, S8F and S8H).
Discussion
The carcinogenesis and cancer progression are related to comprehensive effects of multiple genes. Multi-omics analysis is a more promising way to predict the LGGs prognosis. However, the analysis integrating multi-omics data to predict prognosis of LGGs is still lacking. In this study, we collected LGGs samples from TCGA database to carry out analysis of WES, transcription, and DNA methylation and then to combine these multi-omics data to establish a prediction model for the survival of LGGs. This study will be beneficial to accurately predict the survival of LGGs to identify patients with poor prognosis to take specific treatment as early, which will be helpful improve the treatment outcomes and prognosis of LGG.
Mutations in IDH1 characterize the majority of LGG, which is associated with a favorable prognosis25. Consistently, we observed a high mutation frequency (77%) in IDH1. ATRX is also a frequently mutated gene. Liu et al. detected 33% ATRX mutations in grade II gliomas and 46% in grade III gliomas–grade II and III gliomas belong to LGG26. Similarly, we reported similar mutation frequency (37%) in ATRX. In addition, we observed frequent mutations in TP53 (48%), CIC (22%), TTN (11%), et al. This result will be beneficial to explore the mechanisms of the carcinogenesis and progression of LGG as well as the prevention and management of LGG.
For the WES data, we showed a few samples had extremely high (up to over 9000) mutation number, while normally the mutation number in LGG patients ranged from tens to hundreds. This suggests very high TMB in some LGG cases, which might be helpful to the personalized treatment for LGG patients.
Through RNA sequencing, we showed most significantly up-regulated recurrence-related DEGs included C6orf15, DAO, MEOX2, LINC02587, LTF, MIR3976HG, LINC00507, LINC00588, LGR6, LINC02822, etc., and most significantly down-regulated DGEs included DMBX1, GPR50, HMX2, MAFA, COL2A1, TLX3, KC877982.1, BARX1, LBX1-AS1, MEOX1, etc. This result would provide clues to the mechanistic study on the recurrence of LGG.
Notably, we showed that the recurrence-related DEGs were lest aggregated in Chromosome 13. This implies that the upstream and downstream regulation of DEGs might be lost in genes locating in Chromosomal 13, suggesting the potential roles of Chromosomal 13 genes in the negative regulation of recurrence-related DEGs in LGG.
DNA methylation is a major epigenetic signature occurring in early cancer events, which exhibits great potential in the risk assessment, early identification and prognosis prediction of various cancers27,28,29. We showed 2077 genes were more methylated and 299 were less methylated in recurrent vs. primary samples, and that the differential methylation was mainly within genes, which might suggest the epigenetic alterations in recurrence of LGG.
We combined the screened 6 survival-related frequently mutated genes, 22 DEGs, and 11 DMEs to build a prediction model for LGG survival, which showed the potential to differentiate the subjects with shorter survival from those with longer survival in both the training and testing sets. In addition, the AUC for the combined model was higher than that for WES-, RNA sequencing or DNA methylation-based model, indicating the advantages of the multi-omics data-based model in the prediction of prognosis of LGG.
One limitation of this study is the use of relatively simple linear regression to construct predictive models. We also attempted to use DeepOmix30, a deep learning framework for multi-omicsdata, to construct predictive models, but encountered severe overfitting issues (Fig S9). While employing simple models helps to some extent in mitigating overfitting issues due to a limited sample size, it also constrains the model’s ability to improve accuracy with an increasing number of future samples. In future research, employing deep learning methods optimized for addressing overfitting issues would be a preferable option31.
This study analyzed the LGG samples from the TCGA database, where the genetic characteristics and clinical variables had some difference from those in Chinese population. Next, tissue samples will be collected from the Chinse LGG patients to validate this result.
Materials and methods
Data collation
The GEO and TCGA datasets were searched, while only TCGA-LGG dataset had a adequate sample number (more than 500) and simultaneously had WES, RNA and methylation data sets, which met the requirement of multi-omics modeling analysis. Therefore, the TCGA-LGG dataset was finally selected for subsequent analysis. The TCGAbiolinks R package was used to download WES and RNAseq data, and the processed methylation signal matrix data was downloaded from UCSC XENA. 523 samples were used for WES analysis, 532 for transcriptional analysis and 529 for DNA methylation analysis (based on signal values on 450 K methylation chip).
Multi-omics analysis
The downloaded WES data were kept as Maf files. The Maftools R was used for data reading, mutation information statistics, and oncoplot mapping, MutationPatters R package for signature analysis, and ClusterProfiler R package for functional enrichment analysis.
For the downloaded RNAseq read count data, the DESeq2 R package was used for data reading, standardization, and differential expression analysis. The vidger R package was used for mapping the volcano plot, the RCircos R package for screening the distribution of DEGs on the genome. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) functional enrichment analysis22,23,24 of DEGs was performed using the ClusterProfiler R package. The important pathways of MsigDB, KEGG, GO and REACTOME databases were enriched by GSVA R package.
For the downloaded methylation data, ChAMP R package was used for data reading, standardization, and calculation of the differential methylation probe and DMEs. Principal component analysis (PCA) dimension-reduction analysis was performed using the FactoMineR and factoextra R packages. Pheatmap and ggplot R packages were used to draw the related images. Functional enrichment analysis of DMEs was performed using the ClusterProfiler R package.
Multi-level modeling analysis
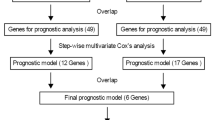
As illustrated in the flowchart (Fig S10), we initially applied LASSO for feature selection on the WES, transcriptomic, and methylation data from the TCGA-LGG dataset. Subsequently, the selected feature genes were employed to independently construct both single-omics and multi-omics prediction models.
In the feature screening stage, the highly frequently mutated genes, DMGs, and DMGs obtained in the above uni-omics analysis were evaluated. The LASSO method by glmnet R package was used to conduct feature selection with the survival status as the target values and the mutation, expression change folds of DEGs and methylation change folds of DMGs as the characteristic values. When using the LASSO model for feature selection, the alpha parameter of glmnet was set to 1, and the family parameter was set to ‘cox’. The minimum lambda of LASSO model was referred to screen the characteristic genes from each omics data. After that, each omics data-based survival model was constructed, using survival R package to group samples based on the median risk values of each sample and carry out the corresponding Kaplan–Meier (KM) survival analysis. The pROC R package was used to plot the Receiver Operating Characteristic Curve (ROC) and perform the area under the curve of ROC (AUC) calculations. The characteristic gene data of the above three omics were combined to build the multilevel model.
The relationship between the frequently mutated genes (with mutation frequency of greater than 10%), DEGs and DMGs and patient survival was calculated. Both single-omics and multi-omics models were built using samples containing the three types of omics data mentioned above. Before training the model, 30% of the samples were randomly selected from the dataset as the test set, and the remaining 70% of the samples were used for training. Then, the genes from each kind of omics data were selected to build a survival model based on the respective omics data. Finally, the genes selected from the three kinds of omics were combined to build a multi-omics survival model based on the three kinds of omics data. When training both single-omics and multi-omics survival models, the alpha parameter of glmnet was set to 0, the family parameter was set to ‘cox’, and the nfolds parameter was set to 10 to implement cross-validation. To evaluate the performance of each model, the trained model-related risk scores were calculated and used to observe the correlation with the survival, and ROC curve analysis was carried out to evaluate the performance of the model in the training and test data sets.
To further analyze the indicative role of the selected feature genes on prognosis as previously reported32, we employed the rms R package. We constructed both univariate and multi-omic Nomogram prognostic models for the feature genes identified in WES, transcriptome, and methylation datasets. Additionally, we utilized the survivalROC R package to assess the accuracy of each Nomogram model in predicting 1-year, 2-year, and 5-year survival rates.
Conclusion
We carried out WES, RNA sequencing and DNA methylation analysis to build a multi-omics-based model for prediction of the survival of LGG. This study will be useful to accurately predict the survival of LGGs to take specific treatment for those with poor expected prognosis as early, which will be helpful improve the treatment outcomes and prognosis of LGG.
Data availability
All data generated or analyzed during this study are included in this published article.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 Cancers in 185 countries. CA Cancer J Clin. 71(3), 209–249 (2021).
Zheng, R. S. et al. Cancer statistics in China. Chin. J Oncol. 45(3), 212–20 (2016).
Lapointe, S., Perry, A. & Butowski, N. A. Primary brain tumours in adults. Lancet 392(10145), 432–446 (2018).
Goodenberger, M. L. & Jenkins, R. B. Genetics of adult glioma. Cancer Genet. 205(12), 613–621 (2012).
Chheda, Z. S. et al. Novel and shared neoantigen derived from histone 3 variant H3.3K27M mutation for glioma T cell therapy. J. Exp. Med. 215(1), 141–157 (2018).
Lin, L., Cai, J. & Jiang, C. Recent advances in targeted therapy for glioma. Curr. Med. Chem. 24(13), 1365–1381 (2017).
Wang, T. & Mehta, M. P. Low-grade glioma radiotherapy treatment and trials. Neurosurg. Clin. N. Am. 30(1), 111–118 (2019).
Claus, E. B. et al. Survival and low-grade glioma: The emergence of genetic information. Neurosurg. Focus 38(1), E6 (2015).
Schomas, D. A. et al. Intracranial low-grade gliomas in adults: 30-Year experience with long-term follow-up at mayo clinic. Neuro. Oncol. 11(4), 437–445 (2009).
Hanahan, D. Hallmarks of cancer: New dimensions. Cancer Discov. 12(1), 31–46 (2022).
Zhang, Z. et al. N6-methyladenosine demethylase ALKBH5 suppresses colorectal cancer progression potentially by decreasing PHF20 mRNA methylation. Clin. Transl. Med. 12(8), e940 (2022).
Li, Z. et al. DNMT1-mediated epigenetic silencing of TRAF6 promotes prostate cancer tumorigenesis and metastasis by enhancing EZH2 stability. Oncogene 41(33), 3991–4002 (2022).
Chiappinelli, K. B. & Baylin, S. B. Inhibiting DNA methylation improves antitumor immunity in ovarian cancer. J. Clin. Invest. https://doi.org/10.1172/JCI160186 (2022).
Kang, K. et al. Genomic instability in lower-grade glioma: Prediction of prognosis based on lncRNA and immune infiltration. Mol. Ther. Oncolytics 22, 431–443 (2021).
Lin, W. et al. A three-lncRNA signature predicts clinical outcomes in low-grade glioma patients after radiotherapy. Aging (Albany NY) 12(10), 9188–9204 (2020).
Tan, Y. Q. et al. Six immune associated genes construct prognostic model evaluate low-grade glioma. Front Immunol. 11, 606164 (2020).
Yurgelun, M. B. et al. Development and validation of the PREMMplus model for multigene hereditary cancer risk assessment. J. Clin. Oncol. 40(35), 4083–4094 (2022).
Skaugen, J. M. et al. Performance of a multigene genomic classifier in thyroid nodules with suspicious for malignancy cytology. Thyroid 32(12), 1500–1508 (2022).
Coughlin, S. E. et al. Multigene panel testing yields high rates of clinically actionable variants among patients with colorectal cancer. JCO Precis. Oncol. 6, e2200517 (2022).
Brat, D. J. et al. Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N. Engl. J. Med. 372(26), 2481–2498 (2015).
Pan, X. et al. i-Modern: Integrated multi-omics network model identifies potential therapeutic targets in glioma by deep learning with interpretability. Comput. Struct. Biotechnol. J. 20, 3511–3521 (2022).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28(11), 1947–1951 (2019).
Kanehisa, M. et al. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51(D1), D587–D592 (2023).
Yan, H. et al. IDH1 and IDH2 mutations in gliomas. N. Engl. J. Med. 360(8), 765–773 (2009).
Liu, X. Y. et al. Frequent ATRX mutations and loss of expression in adult diffuse astrocytic tumors carrying IDH1/IDH2 and TP53 mutations. Acta Neuropathol. 124(5), 615–625 (2012).
Li, D. et al. Discovery and validation of tissue-specific DNA methylation as noninvasive diagnostic markers for colorectal cancer. Clin Epigenetics 14(1), 102 (2022).
Guidry, K. et al. DNA methylation profiling identifies subgroups of lung adenocarcinoma with distinct immune cell composition, DNA methylation age, and clinical outcome. Clin. Cancer Res. 28(17), 3824–3835 (2022).
Drexler, R. et al. DNA methylation subclasses predict the benefit from gross total tumor resection in IDH-wildtype glioblastoma patients. Neuro Oncol. 25(2), 315–325 (2023).
Zhao, L. et al. DeepOmix: A scalable and interpretable multi-omics deep learning framework and application in cancer survival analysis. Comput. Struct. Biotechnol. J. 19, 2719–2725 (2021).
Lee, M. An ensemble deep learning model with a gene attention mechanism for estimating the prognosis of low-grade glioma. Bio. (Basel) 11(4), 586 (2022).
Guo, Y. et al. DNA methylation-driven genes for developing survival nomogram for low-grade glioma. Front Oncol. 11, 629521 (2021).
Funding
This project was supported by Nanjing Medical Technology Development Project (ZKX22026).
Author information
Authors and Affiliations
Contributions
Conceptualization, Z.D. and M.Y.; Data curation, Z.D.; Formal analysis, Z.D., Y.Y. and X.K.; Funding acquisition, M.Y.; Methodology, Z.D., Y.Y., X.K. and J.Y.; Project administration, M.Y.; Resources, Z.D. and X.K.; Software, B.L.; Supervision, M.Y.; Validation, M.Y.; Writing—original draft, Z.D.; Writing—review & editing, M.Y.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Du, Z., Jiang, Y., Yang, Y. et al. A multi-omics analysis-based model to predict the prognosis of low-grade gliomas. Sci Rep 14, 9427 (2024). https://doi.org/10.1038/s41598-024-58434-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-58434-8
- Springer Nature Limited