Abstract
BCR::ABL1-negative myeloproliferative neoplasms are hematopoietic disorders characterized by panmyelosis. JAK2 V617F is a frequent variant in these diseases and often occurs in the 46/1 haplotype. The G allele of rs10974944 has been shown to be associated with this variant, specifically its acquisition, correlations with familial cases, and laboratory alterations. This study evaluated the association between the 46/1 haplotype and JAK2 V617F in patients with myeloproliferative neoplasms in a population from the Brazilian Amazon. Clinical, laboratory and molecular sequencing analyses were considered. Carriers of the G allele of rs10974944 with polycythemia vera showed an increase in mean corpuscular volume and mean corpuscular hemoglobin, while in those with essential thrombocythemia, there was an elevation in red blood cells, hematocrit, and hemoglobin. Associations were observed between rs10974944 and the JAK2 V617F, in which the G allele (OR 3.4; p < 0.0001) and GG genotype (OR 4.9; p = 0.0016) were associated with JAK2 V617F + and an increase in variant allele frequency (GG: OR 15.8; p = < 0.0001; G: OR 6.0; p = 0.0002). These results suggest an association between rs10974944 (G) and a status for JAK2 V617F, JAK2 V617F + _VAF ≥ 50%, and laboratory alterations in the erythroid lineage.
Similar content being viewed by others
Introduction
Myeloproliferative neoplasms (MPNs) are clonal diseases characterized by hyperplasia of the myeloid lineage with effective maturation, which results in leukocytosis in peripheral blood, increased erythrocyte mass and possible progression to medullary fibrosis or leukemic transformation1. They have an incidence rate of 6 cases per 100,000 individuals and mostly affect white males between 60 and 70 years of age2. Polycythemia vera (PV), essential thrombocythemia (ET), and primary myelofibrosis (MF) are the most common BCR::ABL1-negative MPNs, though differ in signs, symptoms, hematological and clinical alterations, and genetic findings3.
JAK2 V617F (dbSNP ID: rs77375493) is the main genetic finding in MPNs and has a frequency of 95% in PV cases and between 50–60% in ET and MF cases4. This somatic variant triggers the substitution of valine by phenylalanine at codon 617, which alters the pseudo-kinase domain of the JAK2 protein and conditions a constitutive activation of the JAK/STAT signaling pathway5.
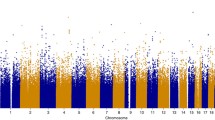
Studies have shown a significant correlation between JAK2 V617F and the 46/1 haplotype, a set of germline genetic variations distributed along chromosome 9p.24.1. This haplotype covers regions with a high number of genetic variants in JAK2 (exons 12 and 14) and is in linkage disequilibrium with the variant rs10974944 (C > G), located in intron 12 of the same gene6 (Fig. 1). Studies indicate that this genetic alteration is a factor that favors the acquisition of JAK2 V617F by increasing the mutational rate of JAK2, which can lead to DNA damage and replication errors7,8,9. In addition to being identified in MPN patients of various populations, this haplotype has also been associated with more pronounced alterations in laboratory exams, presence of splenomegaly, inflammatory dysregulation, familial cases of MPNs (increasing the risk of developing any myeloproliferative neoplasm by 5 to 7 times) and abnormal methylation of the gene promoter10,11,12,13. Therefore, the JAK2 46/1 haplotype confers predisposition to the development of myeloproliferative neoplasms associated with the JAK2 V617F mutation (OR = 3.7; 95% CI = 3.1–4.3) and provides a conceptual framework in which a constitutional genetic component is associated with a substantial increase in the risk of acquiring a specific somatic mutation14.
In this study, we performed genetic sequencing of intron 12 of the JAK2 gene to identify the rs10974944 variant (C > G), in strong linkage disequilibrium with the 46/1 haplotype, in 100 patients with BCR::ABL1-negative myeloproliferative neoplasms (polycythemia vera: n = 39; essential thrombocythemia: n = 61) for whom clinical and laboratory information was available for clinical and laboratory characterization.
Results
Characterization of the study population
The study included individuals clinically diagnosed with polycythemia vera (PV) (n = 39) or essential thrombocythemia (ET) (n = 61), whose clinical-laboratory characteristics are presented in the supplementary material. The female gender was more prevalent among individuals diagnosed with ET (n = 48, p = 0.002). The median age of the participants ranged between the fifth and sixth decades of life (p = 0.441).
Regarding hematological results, the medians of overall red blood cell count (RBC), hematocrit (Ht), hemoglobin (Hb), and total white blood cell count (WBC) were significantly higher in the PV group compared to the ET group (p < 0.05) (see Table SI). Other hematological markers, such as mean corpuscular volume (103.9 pg, p < 0.0001), mean corpuscular hemoglobin (33.5 fL, p < 0.0001), and overall platelet count (467,000 × cells/mm3, p < 0.0001), were also significantly elevated in the ET group compared to the PV group. Hemorrhagic events were more frequent in patients with ET compared to PV (p = 0.003), while the frequency of splenomegaly and thrombotic events did not differ significantly between PV and ET (p > 0.05) groups.
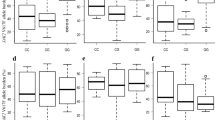
Regarding genetic findings, the presence of JAK2 V617F+ was more frequent in patients with PV (58.9%, p = 0.020) (Fig. 2a), and a variant allele frequency (VAF) of ≥ 50% was also more common in patients with this hematologic condition (41%, p = 0.005) (Fig. 2b).
A greater frequency of patients with ET (95.1%, p < 0.0001) received cytoreductive treatment in comparison to PV patients (66.6%).
Identified genetic variants
Data on the allelic and genotypic frequency of rs10974944 (C > G) are presented in Figs. 2c and d. Of all the individuals included in the study, 63% exhibited the rs10974944 variant (G): 26% in homozygosity (GG) and 37% in heterozygosity (CG). The GG genotype of rs10974944 was more prevalent in the PV group (36%), whereas CG was more homogeneous between the groups (33.3% in PV and 39.3% in ET). Regarding allelic frequency, the G allele was more frequent in the PV (53.6%) group, and the wild-type allele proved to be more prevalent in the ET (60.7%) group.
Table 1 presents the hematological data of individuals with polycythemia vera and essential thrombocythemia stratified according to the absence or presence of the rs10974944 (CC and G carriers, respectively). In PV, G carriers showed significantly increased values for MCV and MCH (p = 0.030 and p = 0.041, respectively), while in ET, patients with the variant exhibited elevated indices of RBC, Ht, and Hb with demonstrated statistical significance (p < 0.05).
Data on the correlation between the G allele and clinical characteristics are shown in Table 2. A significant correlation was observed between the G allele and thrombotic events in patients with PV (p = 0.041) and a similar trend in ET, however, without significance statistics (p = 0.073). These data suggest that the G allele of rs10974944 may be associated with an increased risk of thrombotic events in patients with PV.
Distribution of variants in patients stratified according to JAK2 V617F status and variant allele frequency
Considering the possible association of rs10974944 with JAK2 V617F, the genotypic frequency analysis of rs10974944 (C > G) was performed according to the positive (+) or negative (−) status of JAK2 V617F and its variant allele frequency (VAF), with data described in Table 3.
Homozygous individuals for rs10974944 (GG) showed a significantly higher frequency of JAK2 V617F+ status and a higher likelihood of being positive for this variant when compared to the CC genotype (42.2% vs 12.5%; OR 4.9; 95% CI 1.8–13.9; p = 0.00016) (Table 3). We emphasize the correlation of the rs10974944 G allele with the V617F variant, which demonstrated a 3.4-fold higher probability of being present in JAK2 V617F+ individuals compared to individuals carrying the C allele (61.1% vs 38.9%; OR 3.4; 95% CI 1.9–6.2; p < 0.0001).
Additionally, the analyses revealed that individuals with the GG genotype of rs10974944 had a 13.1-fold higher probability of having a VAF greater than 50% when compared to individuals with the CC genotype (75% vs 15%; OR 13.1; 95% CI 1.8–72.3; p = 0.004). Regarding the allele, carriers of the G allele showed a sixfold higher risk of having a VAF of ≥ 50% compared to the wild-type allele (C) (82.5% vs 17.5%; OR 6.0; 95% CI 2.1–14.8; p = 0.0002). These results demonstrate an association between rs10974944 and the variation in VAF in JAK2 V617F.
Analysis of rs10119004 and rs10815151 was also performed. The homozygosity for rs10119004 AA genotype demonstrated a higher prevalence of JAK2 V617F + status in comparison to AG and GG genotypes (60.0% vs 33.3% and 6.7%, respectively; OR 2.1; 95% CI 0.9–4.8; p = 0.077). Additionally, individuals with the AA genotype exhibited a significantly elevated likelihood of VAF ≥ 50% compared to AG carriers (80% vs 15%; OR 5.1; 95% CI 1.3–19.6; p = 0.017). The allele A showed a higher frequency in JAK2 V617F + individuals compared to allele G carriers (76.6% vs 23.4%; OR 2.1; 95% CI 1.1–3.9; p = 0.025) and carriers of allele A had a significantly elevated likelihood of VAF ≥ 50% compared to allele G carriers (87.5% vs 32%; OR 3.3; 95% CI 1.1–10.0; p = 0.043).
Regarding rs10815151, the CC genotype displayed a higher frequency of JAK2 V617F+ status compared to CT and TT genotypes (73.4% vs 15.5% and 11.1%; OR 2.8; 95% CI 1.2–6.6; p = 0.021). Moreover, individuals with the CC genotype showed a higher probability of having VAF ≥ 50% compared to those with CT genotype (85% vs 10%; OR 3.2; 95% CI 0.7–13.9; p = 0.176). The C allele exhibited a higher prevalence in individuals with JAK2 V617F+ status compared to T allele carriers (81% vs 19%; OR 1.8; 95% CI 0.9–3.5; p = 0.1019), and significantly increased probability of having VAF ≥ 50% compared to those with the T allele (90% vs 74%; OR 5.5; 95% CI 1.7–18.2; p = 0.0032).
Identified haplotypes
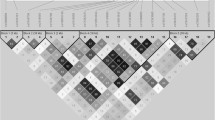
The linkage disequilibrium (LD) of rs10974944 and JAK2 V617F (rs77375493) is demonstrated in Fig. 3. The variants identified in the analyzed region were included in the haplotype analysis. When these genetic changes are paired, they give rise to nine haplotypes (Table 4). Haplotype analysis revealed that haplotype 2 (rs10974944G/rs10815151C/rs1011004A/rs77375493T) was more prevalent in individuals with JAK2 V617F+ (46.5%; OR 19.6; 95% CI 3.1–208; p = < 0,0001), which indicates a strong correlation between the variants. This information is in accordance with that contained in supplementary figure I and supplementary table III, where it is possible to note the same haplotype 2 more frequent in patients with PV, a neoplasm that presented a higher frequency of the JAK2 V617F variant.
Linkage disequilibrium (LD) structure of JAK2 intron 12 in patients JAK2 V617 positive (JAK2 V617F+) and JAK2 V617 negative (JAK2 V617-). Numbers in the boxes indicate the value of the LD correlation coefficient (r2) multiplied by 100. Lighter shades of boxes indicate a decreased r2 value, strong LD is represented by the dark gray box.
Discussion
Myeloproliferative neoplasms have characteristic alterations in laboratory exams, as well as genetic findings that permit their identification and differentiation. Findings involving genetic alterations in introns are not yet fully understood, but this scenario is becoming of increasing interest for understanding the etiopathogenic aspects and the role of these DNA regions in these diseases.
Essential thrombocythemia proved to be the most frequent myeloproliferative neoplasm, which are findings that align with the premises established by Torres15, who studied a population with BCR::ABL1-negative myeloproliferative neoplasms in the state of Amazonas (Brazil). Similar data were described by Macedo16, who reported a similar scenario in patients from the states of Paraná and São Paulo who had the same hematologic malignancy, and these data converge with descriptions found in other countries17,18.
The age range of individuals was between the fifth and seventh decades of life, which is consistent with what is stated in other studies19,20. The progressive accumulation of genetic variations in hematopoietic stem cells and the biological machinery of the DNA repair system21,22, an increase or decrease in telomeres23,24 and cumulative exposure to risk factors throughout life, such as smoking and obesity25,26, may explain the prevalence of this age group in the context of myeloproliferative neoplasms.
Regarding clinical characteristics, polycythemia vera (PV) showed an equal proportion of men and women, while essential thrombocythemia (ET) revealed a majority of cases involving women, and these data are in line with the literature27,28. Some studies have demonstrated that women have an increased risk of developing myeloproliferative neoplasms29 and a higher likelihood of developing cardiovascular complications and splenomegaly26. The reason for this risk is uncertain, but changes in sex chromosomes, hormonal factors and gene expression may be possible contributors to this process28. Laboratory data, and thrombotic and hemorrhagic events presented as expected for each neoplasm: PV demonstrated a higher prevalence of increased erythrogram values and ET showed changes in the megakaryocytic series, with a higher risk of hemorrhagic events, as described by the World Health Organization3, and in other studies on the subject27,30.
Regarding the genetic findings, PV demonstrates a higher prevalence of positive cases for the JAK2 V617F variant, since it is directly associated with the specific pathogenesis of this hematologic malignancy36 and plays a role in the constitutive activation of the JAK-STAT pathway5. It is interesting to note that 58% of our PV population was positive for the variant, which may initially differ from findings commonly described in the literature that point to JAK2 V617F frequencies of over 70% in Brazilian, Korean, Chinese, Japanese, and European patients31,32,33,34,35.
Our analysis reveals a notable specificity in our population compared to the data documented in the literature, especially in patients with PV, where 42% of these patients did not present the JAK2 V617F variant or other pathogenic genetic alterations along the coding region of JAK2, as established by WHO diagnostic criteria3. This atypical behavior suggests significant gaps in our understanding of the genetic factors underlying the etiopathogenesis of myeloproliferative neoplasms in the Amazonian population. This gap underscores the pressing need for further studies to achieve a more comprehensive understanding of the genetic profile of these diseases and other contributing factors. Therefore, additional studies in our population are recommended, exploring other genes relevant to myelopoiesis and epigenetic regulation, such as DNMT3A (DNA Methyltransferase 3 Alpha), NFE2 (Nuclear factor erythroid 2), SF3B1 (Splicing Factor 3b Subunit 1), TET2 (Tet Methylcytosine Dioxygenase 2), ASXL1 (ASXL Transcriptional Regulator 1) and EZH2 (Enhancer Of Zeste 2 Polycomb Repressive Complex 2 Subunit)21,52. Analysis of these genes may provide valuable insights into the genetic behavior of myeloproliferative neoplasms in the Amazonian population and elucidate other factors involved in PV pathophysiology, beyond the known variants in JAK2 V617F and JAK2 exons 12 and 14.
In the literature, the germline haplotype 46/1, identified by the rs10974944 (C > G) variant, has a well-documented association with JAK2 V617F14,36,37,38 as also observed in our study. The data regarding the frequency of the minor allele of rs10974944 in the Brazilian population and the Amazon region remain scarce, making this study pioneering in this investigation. The absence of previous studies on this variant in the Amazonian population underscores the importance of the current work in filling this gap in the genetic knowledge of this population. However, the frequency of the 46/1 haplotype, associated with rs10974944, has been linked to a higher prevalence in patients with myeloproliferative neoplasms, especially those harboring JAK2 V617F+. This association has been observed not only in other Brazilian populations as described by Macedo et al.16 but also in studies conducted across various populations worldwide, including Asian, European, and North American populations, as discussed in one of our previous integrative reviews13. Additionally, the ancestral contribution to the Brazilian population, particularly in the Amazon region, is characterized by a mixture of three main ethnic groups: Native Americans (NAM), Europeans (EUR), and Africans (AFR)39. Therefore, it is plausible to infer that the genetic behavior of the variant in these populations, as described previously, is similar, thus strengthening the discussion regarding similar behavior in our population.
The high frequency of the G allele of rs10974944 in individuals positive for JAK2 V617F contributes to discussions about the non-random correlation between these two genetic alterations13,40 This relationship is in line with another finding from our study, haplotype 2 (rs10974944G/rs10815151C/rs1011004A/rs77375493T), which strengthens concepts based on the interaction between rs10974944 (C > G) and JAK2 V617F (rs77375493—G > T). These propositions are in agreement with findings involving haplotype 46/1 in other Brazilian, Taiwanese, European, Chinese, and Japanese populations16,32,33,34,41, indicating that the possible mechanisms preceding the acquisition of JAK2 V617F are not limited to a specific ethnic group; therefore, its evolutionary basis can be considered as a genetic predisposition factor for the disease8.
Studies report a higher risk of individuals with the GG genotype of rs10974944 being positive for JAK2 V617F14,40,42. Consistent with the results of the aforementioned studies, our population exhibited a four-fold increase in the risk of positive JAK2 V617F in individuals with the GG genotype of rs10974944 (OR 4.1; 95% CI 8–13.9). These findings support the hypothesis of hypermutability, which establishes haplotype 46/1 as a dysregulating agent of the JAK2 gene, which increases the risk of DNA replication errors and conditions a mutagenic scenario for the acquisition of variants with selective advantages, such as JAK2 V617F43,44,45.
The association of rs10974944 (G) and the JAK2 V617F VAF suggests a possible involvement of haplotype 46/1 in clonal expansion. We identified a six-fold higher risk of individuals carrying the G allele of rs10974944 and JAK2 V617F VAF of ≥ 50%. Our data indicate that the marker of haplotype 46/1 may play a role not only in the acquisition of JAK2 V617F but is also attributed to clonal expansion, maintenance, and survival. Tefferi46 suggests that JAK2 V617F is not the initial clonogenic event in MPNs but rather one of several subclones derived from an ancestral clone. This is in accordance with the notes of Pardanani et al.47, which support the hypothesis that this haplotype is located in a favorable cis regulatory environment, which facilitates the acquisition of JAK2 V617F, and which, in turn, is responsible for clonal expansion and the development of MPNs.
Furthermore, the possible role of acquired uniparental disomy, a genetic event that leads to mitotic recombination associated with neutral loss of heterozygosity of chromosome 9p in MPN patients, reducing both the haplotype and JAK2 V617F to a homozygous state14,48,49, cannot be ruled out. In this context, cells with both variants theoretically have a selective advantage, which conditions greater myeloproliferative potential and favors the establishment of variant cells over healthy cells, thus explaining the increased VAF in individuals with the combination rs10974944 (G) + rs77375493 (T) (JAK2 V617F) in homozygosity.
Association between changes in hematological indices, clinical characteristics and the presence of 46/1 is observed in the literature16,33,50; however, this is not a consensus among the scientific community8. Our data show significant differences in MCV, MCH values in the PV group, and RBC, Hb, and Ht in TE carriers of the G allele of rs10974944, which has been observed in previous studies7,42,51. The significant demonstration of rs10974944 with thrombotic events strengthens the use of this variant as a tool for monitoring patients and investigating clinical findings of polycythemia vera. For a more reliable correlation of this correlation, new studies are needed, with more robust populations, to observe the behavior of the variant in relation to clinical and hematological characteristics in PV patients.
The present research is the first to analyze the 46/1 haplotype using the rs10974944 variant, present in intron 12 of JAK2, in a population from the Brazilian Amazon. The results of this study show that the rs10974944 (G) variant has a strong correlation with the JAK2 V617F+ variant, demonstrated especially in PV_JAK2 V617F+ patients. A correlation of the variant with a high allelic variant burden of JAK2 V617F, thrombotic events and hematological changes was also observed. The variant is a promising possibility for clinical use for investigating and monitoring laboratory changes and/or increased VAF in identified hematological malignancies.
Materials and methods
Population
One hundred individuals clinically diagnosed with BCR::ABL1-negative myeloproliferative neoplasms were included in the study. The study was conducted from February 2021 to January 2023. Laboratory analysis was performed at the Genomics Laboratory of the Foundation Hospital for Hematology and Hemotherapy of the State of Amazonas.
Ethical approval
The study was submitted to and approved by the Ethics Committee of the Foundation Hospital for Hematology and Hemotherapy of the State of Amazonas under opinion No. 4,450,813 and certificate of ethical appreciation No. 39991420.6.0000.0009. Written informed consent was obtained from patients. This study complied with Resolution No. 466/2012 of the National Health Council for research involving human subjects and followed the parameters determined by the Declaration of Helsinki.
Clinical and laboratory data
Clinical data (gender, age, splenomegaly, thrombotic and/or hemorrhagic events) and laboratory data were obtained from medical records.
Identification of JAK2 V617F
The variant was identified according to the parameters and specifications established by Torres et al.15 . The negative status for the JAK2 V617F variant was confirmed using the allele-specific PCR technique.
Biological sample and DNA extraction
Venous blood samples were collected in tubes containing EDTA, and DNA was extracted using Brazol (Lgcbio, Brazil), following the manufacturer’s instructions, and stored at − 80 °C.
Conventional PCR and PCR purification
For the amplification of the DNA region under analysis, a reaction with a final volume of 25 μL was used with 50–100 ng of genomic DNA, Buffer (1x), MgCl2 (1.5 mM), forward primer CCAACTGAGTTTCCTTGCAG and reverse primer CTAGGTTAAGAGTATGTGGTTCC (0.4 mM), dNTP mix (0.2 mM), and TAQ (1 U). The PCR products were separated on a 1.5% agarose gel. The PCR product, a 572 bp amplicon, was purified with polyethylene glycol (PEG 8000) (Promega).
Nucleotide sequencing and sequence analysis
Approximately 5–30 ng of purified PCR product was applied to the sequencing reaction. Nucleotide sequencing was performed using BigDye Terminator v3.1 (Applied Biosystems), following the manufacturer’s recommendations and the primers described above. The products were purified by the EDTA/Ethanol protocol and evaluated in the 3500 XL Genetic Analyzer automatic sequencer (Applied BioSystems, USA), with POP-7 polymer. The sequences were initially analyzed using the Sequencing Analysis software (Applied BioSystems [Thermo Fisher Scientific, São Paulo, Brazil]). Geneious 6.0.6 software (Biomatters, USA) was used to map the variants and obtain contigs for the comparison with the reference sequence Homo sapiens Janus kinase 2 (JAK2), (NCBI: NG_009904.1).
Haplotype analysis
Haplotype frequencies were calculated using Haploview software (v.4.2) as a measure of linkage disequilibrium (LD). Haplotypes with frequencies of < 1% were not considered relevant for comparisons. Pairwise degree between nucleotides was analyzed using the LD structure, considering r2 values of > 0.8 as strong LD, < 0.8 as weak, and < 0.1 as negative LD. Hardy–Weinberg equilibrium was calculated by comparing estimated and observed genotype frequencies using the χ2 test. SNVs with p-values of < 0.001 were considered to be out of Hardy–Weinberg equilibrium.
Statistical analysis
The obtained results were subjected to the Shapiro–Wilk normality test. Categorical variables were expressed as absolute value (n) and relative frequency (%) and were tested using the χ2 and Fisher’s exact test with a 95% confidence interval. Numerical variables were expressed as median (Md) and interquartile range [IQR] with 75th percentile through GraphPad Prism v.9.0.2 software. For non-parametric variables, the Kruskal–Wallis test was performed. For both analyses, Dunn’s post-test for multiple comparisons was also conducted using GraphPad Prism v.9.0.2 software. P-values of < 0.05 were considered statistically significant.
Data availability
The datasets used and/or analyzed during the present study are available from the corresponding author on reasonable request. The GenBank accession number for the nucleotide sequence is PP208825.
References
Grinfeld, J. et al. Classification and personalized prognosis in myeloproliferative neoplasms. N. Engl. J. Med. 379(15), 1416–1430. https://doi.org/10.1056/nejmoa1716614 (2018).
Swerdlow, S.H. et al. World Health Organization Classification of Tumours This Book and all Other Volumes of the Series Can be Purchased : From all Countries. World Health Organization. (4th ed). (2017).
Khoury, J. D. et al. The 5th the World Health Organization classification of haematolymphoid tumours: Myeloid and histiocytic/dendritic neoplasms. Leukemia https://doi.org/10.1038/s41375-022-01613-1 (2022).
Gou, P., Zhang, W. & Giraudier, S. Insights into the potential mechanisms of JAK2V617F somatic mutation contributing distinct phenotypes in myeloproliferative neoplasms. Int. J. Mol. Sci. https://doi.org/10.3390/ijms23031013 (2022).
Torres, D. G. et al. JAK2 variant signaling: Genetic, hematologic and immune implication in chronic myeloproliferative neoplasms. Biomolecules 12(2), 1–18. https://doi.org/10.3390/biom12020291 (2022).
Nielsen, C., Bojesen, S. E., Nordestgaard, B. G., Kofoed, K. F. & Birgens, H. S. JAK2V617F somatic mutation in the general population: Myeloproliferative neoplasm development and progression rate. Haematologica 99(9), 1448–1455. https://doi.org/10.3324/haematol.2014.107631 (2014).
Andrikovics, H. et al. JAK2 46/1 haplotype analysis in myeloproliferative neoplasms and acute myeloid leukemia. Leukemia 24(10), 1809–1813. https://doi.org/10.1038/leu.2010.172 (2010).
Jones, A. V. & Cross, N. C. P. Inherited predisposition to myeloproliferative neoplasms. Ther. Adv. Hematol. 4(4), 237–253. https://doi.org/10.1177/2040620713489144 (2013).
Tefferi, A. et al. JAK2 germline genetic variation affects disease susceptibility in primary myelofibrosis regardless of V617F mutational status: Nullizygosity for the JAK2 46/1 haplotype is associated with inferior survival. Leukemia 24(1), 105–109. https://doi.org/10.1038/leu.2009.225 (2010).
Mangaonkar, A. A. & Patnaik, M. M. Hereditary predisposition to hematopoietic neoplasms: When bloodline matters for blood cancers. Mayo Clin. Proc. 95(7), 1482–1498. https://doi.org/10.1016/j.mayocp.2019.12.013 (2020).
Olcaydu, D. et al. The role of the JAK2 GGCC haplotype and the TET2 gene in familial myeloproliferative neoplasms. Haematologica 96(3), 367–374. https://doi.org/10.3324/haematol.2010.034488 (2011).
Stolyar, M. A. et al. JAK2 haplotype 46/1 and JAK2 V617F allele burden in MPN: New evidence against the “hypermutability” hypothesis?. Int. J. Lab. Hematol. 40(1), e8–e10. https://doi.org/10.1111/ijlh.12765 (2018).
Paes, J., Silva, G. A., Tarragô, A. M. & Mourão, L. P. The contribution of JAK2 46/1 haplotype in the predisposition to myeloproliferative neoplasms. Int. J. Mol. Sci. 23(20), 12582 (2022).
Jones, A. V. et al. JAK2 haplotype is a major risk factor for the development of myeloproliferative neoplasms. Nat. Genet. 41(4), 446–449. https://doi.org/10.1038/ng.334 (2009).
Torres, D. G. et al. Molecular landscape of the JAK2 gene in chronic myeloproliferative neoplasm patients from the state of Amazonas Brazil. Biomed. Rep. 19(6), 1–5. https://doi.org/10.3892/br.2023.1680 (2023).
Macedo, L. C. et al. JAK2 46/1 haplotype is associated with JAK2 V617F—positive myeloproliferative neoplasms in Brazilian patients. Int. J. Lab. Hematol. 37(5), 654–660. https://doi.org/10.1111/ijlh.12380 (2015).
Harrison, C. N. et al. The impact of myeloproliferative neoplasms (MPNs) on patient quality of life and productivity: Results from the international MPN landmark survey. Ann. Hematol. 96(10), 1653–1665. https://doi.org/10.1007/s00277-017-3082-y (2017).
Varghese, C., Immanuel, T., Ruskova, A., Theakston, E. & Kalev-Zylinska, M. L. The epidemiology of myeloproliferative neoplasms in New Zealand between 2010 and 2017: Insights from the New Zealand cancer registry. Curr. Oncol. 28(2), 1544–1557. https://doi.org/10.3390/curroncol28020146 (2021).
Szuber, N. et al. Myeloproliferative neoplasms in the young: Mayo clinic experience with 361 patients age 40 years or younger. Am. J. Hematol. 93(12), 1474–1484. https://doi.org/10.1002/ajh.25270 (2018).
Büyükaşik, Y. et al. Polycythemia vera: Diagnosis, clinical course, and current management. Turk. J. Med. Sci. 48(4), 698–710. https://doi.org/10.3906/sag-1806-43 (2018).
Constantinescu, S. N., Vainchenker, W., Levy, G. & Papadopoulos, N. Functional consequences of mutations in myeloproliferative neoplasms. HemaSphere. https://doi.org/10.1097/HS9.0000000000000578 (2021).
Azevedo, A. P. et al. DNA repair genes polymorphisms and genetic susceptibility to Philadelphia-negative myeloproliferative neoplasms in a Portuguese population: The role of base excision repair genes polymorphisms. Oncol. Lett. 13(6), 4641–4650. https://doi.org/10.3892/ol.2017.6065 (2017).
Barraco, D. et al. Gender effect on phenotype and genotype in patients with post-polycythemia vera and post-essential thrombocythemia myelofibrosis: Results from the MYSEC project. Blood Cancer J. 8(10), 10–13. https://doi.org/10.1038/s41408-018-0128-x (2018).
Ferrer, A., Mangaonkar, A. A. & Patnaik, M. M. Clonal hematopoiesis and myeloid neoplasms in the context of telomere biology disorders. Curr. Hematol. Malig. Rep. https://doi.org/10.1007/s11899-022-00662-8 (2022).
Duncombe, A. S. et al. Modifiable lifestyle and medical risk factors associated with myeloproliferative neoplasms. HemaSphere 4(1), 1–6 (2020).
Allahverdi, N., Yassin, M. & Ibrahim, M. Environmental factors, lifestyle risk factors, and host characteristics associated with philadelphia negative myeloproliferative neoplasm : A systematic review. Cancer Control 28, 1–12. https://doi.org/10.1177/10732748211046802 (2021).
Vannucchi, A. M. & Harrison, C. N. Emerging treatments for classical myeloproliferative neoplasms. Blood 129(6), 693–703. https://doi.org/10.1182/blood-2016-10-695965 (2017).
Geyer, H. L. et al. Associations between gender, disease features and symptom burden in patients with myeloproliferative neoplasms: An analysis by the MPN QOL international working group. Haematologica. 102, 85–93. https://doi.org/10.3324/haematol.2016.149559 (2017).
Patterson-Fortin, J. & Moliterno, A. R. Molecular pathogenesis of myeloproliferative neoplasms: Influence of age and gender. Curr. Hematol. Malig. Rep. 12(5), 424–431. https://doi.org/10.1007/s11899-017-0411-0 (2017).
Nangalia, J. & Green, A. R. Myeloproliferative neoplasms: From origins to outcomes. Blood. 130(23), 2475–2483. https://doi.org/10.1182/blood-2017-06-782037 (2017).
Porto-Soares, M. A. et al. Clinical and molecular profile of a Brazilian cohort of patients with classical BCR-ABL1-negative myeloproliferative neoplasms. Hematol. Trans. Cell Ther. 42(3), 238–244. https://doi.org/10.1016/j.htct.2019.07.008 (2020).
Zhang, X. et al. The JAK2 46/1 haplotype is a risk factor for myeloproliferative neoplasms in Chinese patients. Int. J. Hematol. 96(5), 611–616. https://doi.org/10.1007/s12185-012-1169-8 (2012).
Ohyashiki, J. H. et al. The C allele of JAK2 rs4495487 is an additional candidate locus that contributes to myeloproliferative neoplasm predisposition in the Japanese population. BMC Med. Genet. 13(1), 6. https://doi.org/10.1186/1471-2350-13-6 (2012).
Lighezan, D. L. et al. TET2 rs1548483 SNP associating with susceptibility to molecularly annotated polycythemia vera and primary myelofibrosis. J. Personal. Med. 10(4), 1–16. https://doi.org/10.3390/jpm10040259 (2020).
Lim, Y., Lee, J. O. & Bang, S. M. Incidence, survival and prevalence statistics of classical myeloproliferative neoplasm in Korea. J. Korean Med. Sci. 31(10), 1579–1585. https://doi.org/10.3346/jkms.2016.31.10.1579 (2016).
Olcaydu, D. et al. A common JAK2 haplotype confers susceptibility to myeloproliferative neoplasms. Nat. Genet. 41(4), 450–454. https://doi.org/10.1038/ng.341 (2009).
Anelli, L., Zagaria, A., Specchia, G. & Albano, F. The JAK2 GGCC (46/1) haplotype in myeloproliferative neoplasms: Causal or random?. Int. J. Mol. Sci. 19(4), 1–12. https://doi.org/10.3390/ijms19041152 (2018).
Kilpivaara, O. et al. A germline JAK2 SNP is associated with predisposition to the development of JAK2V617F -positive myeloproliferative neoplasms. Nat. Genet. 41(4), 455–459. https://doi.org/10.1038/ng.342.A (2009).
Leturiondo, A. L. et al. Association of nod2 and ifng single nucleotide polymorphisms with leprosy in the amazon ethnic admixed population. PLoS Negl. Trop. Dis. 14(5), 1–13. https://doi.org/10.1371/journal.pntd.0008247 (2020).
Trifa, A. P. et al. The G allele of the JAK2 rs10974944 SNP, part of JAK2 46/1 haplotype, is strongly associated with JAK2 V617F-positive myeloproliferative neoplasms. Ann. Hematol. 89(10), 979–983. https://doi.org/10.1007/s00277-010-0960-y (2010).
Chiang, Y., Chang, Y., Lin, H. & Huang, L. Germline variations at JAK2, TERT, HBS1L-MYB and MECOM and the risk of myeloproliferative neoplasms in Taiwanese population. Oncotarget. 8(44), 76204–76213 (2017).
Tapper, W. et al. Genetic variation at MECOM, TERT, JAK2 and HBS1L-MYB predisposes to myeloproliferative neoplasms. Nat. Commun. 6, 1–11. https://doi.org/10.1038/ncomms7691 (2015).
Masselli, E., Pozzi, G., Carubbi, C. & Vitale, M. The genetic makeup of myeloproliferative neoplasms: Role of germline variants in defining disease risk, phenotypic diversity and outcome. Cells https://doi.org/10.3390/cells10102597 (2021).
Campbell, P. J. Somatic and germline genetics at the JAK2 locus. Nat. Methods. 41(4), 385–386. https://doi.org/10.1038/ng0409-385 (2009).
Hermouet, S. & Vilaine, M. The JAK2 46/1 haplotype: A marker of inappropriate myelomonocytic response to cytokine stimulation, leading to increased risk of inflammation, myeloid neoplasm, and impaired defense against infection?. Haematologica 96(11), 1575–1579. https://doi.org/10.3324/haematol.2011.055392 (2011).
Tefferi, A. Molecular drug targets in myeloproliferative neoplasms: Mutant ABL1, JAK2, MPL, KIT, PDGFRA, PDGFRB and FGFR1. J. Cell. Mol. Med. 13(2), 215–237. https://doi.org/10.1111/j.1582-4934.2008.00559.x (2009).
Pardanani, A. et al. The JAK2 46/1 haplotype confers susceptibility to essential thrombocythemia regardless of JAK2V617F mutational statusclinical correlates in a study of 226 consecutive patients. Leukemia. 24(1), 110–114. https://doi.org/10.1038/leu.2009.226 (2010).
Kralovics, R., Guan, Y. & Prchal, J. T. Acquired uniparental disomy of chromosome 9p is a frequent stem cell defect in polycythemia vera. Exp. Hematol. 30(3), 229–236. https://doi.org/10.1016/S0301-472X(01)00789-5 (2002).
Sullivan, J. O. & Mead, A. J. Heterogeneity in myeloproliferative neoplasms: Causes and consequences. Adv. Biol. Regul. 71, 55–68. https://doi.org/10.1016/j.jbior.2018.11.007 (2018).
Martínez-Trillos, A. et al. Relationship between the 46/1 haplotype of the JAK2 gene and the JAK2 mutational status and allele burden, the initial findings, and the survival of patients with myelofibrosis. Ann. Hematol. 93(5), 797–802. https://doi.org/10.1007/s00277-013-1989-5 (2014).
Oddsson, A. et al. The germline sequence variant rs2736100-C in TERT associates with myeloproliferative neoplasms. Leukemia 28(6), 1371–1374. https://doi.org/10.1038/leu.2014.48 (2014).
Vainchenker, W. & Kralovics, R. Genetic basis and molecular pathophysiology of classical myeloproliferative neoplasms. Blood 129(6), 667–679. https://doi.org/10.1182/blood-2016-10-695940 (2017).
Acknowledgements
The authors would like to thank the Fundação de Amparo à Pesquisa do Estado do Amazonas (FAPEAM), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES). Our thanks also go to the Fundação Hospitalar de Hematologia e Hemoterapia do Amazonas (FHEMOAM) and the medical staff and laboratory professionals for their assistance in carrying out the work. Special thanks go to Dr Enedina Nogueira at the Genomics Laboratory CAM/UFAM for her support and the use of the infrastructure for the molecular analysis.
Funding
The present study was supported by the Fundação de Amparo à Pesquisa do Estado do Amazonas (Pro-Estado Program; grant nos. #002/2008, #007/2018 and #005/2019, and POSGRAD Program grant nos. #002/2023), Conselho Nacional de Desenvolvimento Científico e Tecnológico, and Coordenação de Aperfeiçoamento de Pessoal de Nivel Superior.
Author information
Authors and Affiliations
Contributions
Carried out the experiments: J.F.P; Population diagnosis: L.N.M.P., R.S.A., N.A.F.; Biological sample collection: J.F.P., D.G.T., D.C.A., M.A.S., E.V.B.A., E.A.M.; Original Draft—writing: J.F.P.; Figure production: J.F.P.; Statistical analysis—review: G.A.V., A.M.T., L.P.S.M; Manuscript—editing: J.F.P., G.A.V., A.M.T., L.P.S.M. All authors reviewed the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Paes, J.F., Torres, D.G., Aquino, D.C. et al. Exploring hematological alterations and genetics linked to SNV rs10974944 in myeloproliferative neoplasms among Amazon patients. Sci Rep 14, 9389 (2024). https://doi.org/10.1038/s41598-024-60090-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-60090-x
- Springer Nature Limited