Abstract
Genetic counseling and testing are more accessible than ever due to reduced costs, expanding indications and public awareness. Nonetheless, many patients missed the opportunity of genetic counseling and testing due to barriers that existed at that time of their cancer diagnoses. Given the identified implications of pathogenic mutations on patients’ treatment and familial outcomes, an opportunity exists to utilize a ‘traceback’ approach to retrospectively examine their genetic makeup and provide consequent insights to their disease and treatment. In this study, we identified living patients diagnosed with breast cancer (BC) between July 2007 and January 2022 who would have been eligible for testing, but not tested. Overall, 422 patients met the eligibility criteria, 282 were reached and invited to participate, and germline testing was performed for 238, accounting for 84.4% of those invited. The median age (range) was 39.5 (24–64) years at BC diagnosis and 49 (31–75) years at the date of testing. Genetic testing revealed that 25 (10.5%) patients had pathogenic/likely pathogenic (P/LP) variants; mostly in BRCA2 and BRCA1. We concluded that long overdue genetic referral through a traceback approach is feasible and effective to diagnose P/LP variants in patients with history of BC who had missed the opportunity of genetic testing, with potential clinical implications for patients and their relatives.
Similar content being viewed by others
Introduction
Breast cancer (BC) is the most reported cancer in the Kingdom of Jordan, and represents almost 20% of all reported cancers in the country. It also represents the second most common cancer-related death, after lung cancer1. Generally, it is estimated that around 5–10% of all BC cases are hereditary2,3; i.e., are attributed to pathogenic germline variants in cancer predisposing genes2,4. Such rate can be higher among younger patients and those with significant family history of breast or other cancers3,5. In fact, 25% of BC patients have been reported to have a family history of the disease, and having a first-degree relative with the disease is associated with a 1.7–fourfold increased risk of developing it6,7. BRCA1 and BRCA2 mutations are the most common pathogenic variants (PVs) associated with hereditary BC8. Other genes that also are associated with hereditary BC include the high-penetrance genes TP53, CDH1, PTEN, PALB2, and STK11, and low/moderate-penetrance genes ATM, CHEK2, BRIP1, and RAD51C, and RAD51D9,10,11. The cumulative incidence rate of BC related to the high-penetrance genes can be as high as 80%, and yet, significant proportion of “familial” clustering of breast cancer remains unexplained12,13.
In this regard, genetic testing has become a revolutionary medical practice for BC patients due to its therapeutic implications on systemic treatment, surveillance14,15 and surgical decisions16,17. These include recommending annual mammograms and MRI screenings of the remaining breast tissue for BRCA1/2 mutation carriers, or risk-reducing surgeries to prevent contralateral BC in BRCA1/2 PV-carriers8,18, use of mastectomy rather than lumpectomy with radiotherapy for BC patients with PVs in TP5319, and utilizing risk-reducing salpingo-oophorectomy for BRCA1/2, BRIP1, and RAD51 C/D- carrying patients as a preventive approach of subsequent ovarian cancer development, due to its increased association with these PVs8,20,21.
Despite the important implications of targeted therapy and surgical interventions, genetic testing and referrals remain underutilized for BC patients22,23. A recent study has shown that in the United States, 53% of breast cancer patients who are at high risk for genetic mutations underwent genetic testing, with only 25% of BC survivors reporting undergoing genetic testing at diagnosis22,24. The reported reasons for inadequate testing include lack of physician recommendations24, reduced awareness among physicians, inadequate time to fully assess family history, and inaccessibility of genetic counseling and testing due to financial restrains25,26,27. Such issues may be even more pronounced in health care systems of developing countries, where genetic testing services are not well established despite refinements in genome sequencing technology and the development of affordable multigene panels for clinical genetic testing28,29,30,31,32.
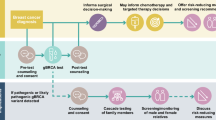
For patients of our facility, genetic counseling and testing for eligible patients have become routinely practiced since 2017. However, it is not offered in other public health care facilities in the country. This presented a potentially life-saving opportunity for our patients through a “traceback” approach, which is a strategy, through the retrospective identification, aimed at identifying BC patients and survivors who were eligible for genetic testing but had not been previously tested. This method involves offering genetic testing to these individuals, identifying probands, tracing their family members, and subsequently offering them genetic counseling and testing33,34,35. The emerging data from our country shows a high rate of PVs/LPVs in our population36,37. A higher prevalence of PVs could reflect the younger median age of BC diagnosis, which is 51 years, 10 years younger than the median age of diagnosis in Western countries38. Utilizing a traceback approach may provide further insights on the genetic makeup of our unique patient population, in addition to helping identify relatives with PVs who may benefit from cascade testing, and providing BC patients and survivors with certain PVs/LPVs with life-altering risk management treatment options.
Materials and methods
Participant identification
The study team identified living BC patients who had been eligible for genetic testing at the time of diagnosis, using our hospital-based cancer registry which compiles records of all patients diagnosed since July 2007. Patient eligibility was based on our institution’s latest guidelines for genetic testing, which conform to the National Comprehensive Cancer Network guidelines for Genetic/Familial High-Risk Assessment Version 2.202139. Accordingly, eligible patients included patients diagnosed with BC at an age of 45 years or younger, patients who were diagnosed with triple-negative BC at 60 years-old or younger, and BC patients diagnosed at age 50 or younger with a first-blood relative with history of BC, between July 2007 and January 2022. We excluded any patient who underwent genetic testing or who had been previously offered genetic counseling and testing.
Participant recruitment
Eligible patients were contacted by one of the study team members using the contact information available in their electronic medical records, explained the objectives of genetic counseling, obtained verbal agreement for enrollment in a counseling session and testing, and scheduled an in-person visit at our institution's Genetic Counseling Clinic. A 45-min in-person interview was arranged at the clinic as a single visit for each patient, during which a genetic counselor would explain the traceback approach, answer all questions, provide the patient with genetic counseling, conduct a family pedigree, and obtain their written informed consent to have a blood sample drawn for genetic testing.
Genetic testing methodology
Genetic testing was performed utilizing a peripheral blood sample, DNA was extracted, and testing was performed using a next-generation sequencing (NGS) panel at a reference laboratory. A multi-gene panel was utilized which was comprised of 20 genes: ATM, BARD1, BRCA1, BRCA2, BRIP1, CDH1, CHEK2, EPCAM, MLH1, MSH2, MSH6, NBN, NF1, PALB2, PMS2, PTEN, RAD51C, RAD51D, STK11 and TP53. The genetic testing and sequencing report followed the American College of Medical Genetics and Genomics (ACMG) classification guidance for pathogenic (P), likely pathogenic (LP), variants of uncertain significance (VUS), likely benign, and benign variants40.
Disclosure of genetic test results
Study participants who had a negative genetic testing result were informed by the genetic counselor over the phone. If a patient’s test result revealed P/LPs or VUS, an in-person clinic appointment was arranged to explain the results and their significance and recommend mitigative interventions. Cascade testing for unaffected family members was offered at significantly discounted rates to cover only the cost of specimen shipment and handling.
Ethics declaration
The study was conducted according to the guidelines of the Declaration of Helsinki, and approved by the Institutional Review Board of King Hussein Cancer Center. All subjects gave their written informed consent for inclusion before they participated in the study.
Statistical analysis
Clinical and pathological characteristics of patients who underwent genetic testing were tabulated and described by percentages, medians, and ranges. The results of the genetic tests were reported in numbers and percentages. Kruskal–Wallis, Pearson, Wilcoxon testing, and median interquartile ranges were used to compare rates of P/LP and VUS rates among the different study groups. A p value < 0.05 was considered statistically significant in all statistical analyses.
Results
A total of 422 patients with BC met the eligibility criteria through chart review, and all were females. Of which, 140 (33.2%) unreachable or unparticipated, this was due to 58 patients not having updated contact information or being unreachable, 31 who did not follow up at our institution, 27 being out of the country at the time of conducting the study, and 24 missing multiple appointments (Fig. 1).
Of the 282 individuals who could be tested, 44 (15.6%) declined to provide a sample for testing due to: patient disinterest, physical burden, inactive insurance at our institution, social fears, and emotional stress (as reported by 13, 12, 8, 6, and 5 patients, respectively), Table 1.
Of the 238 individuals participated in genetic testing (Fig. 1), the most common indication for study inclusion was being diagnosed with BC at 45 years or younger, which was the case for the majority of the participants (n = 185, 77.7%), followed by a triple-negative BC diagnosis at the age of 60 or younger for 70 (29.4%) patients, while 44 (18.5%) patients were included due to having one or more first-degree relatives with BC and were 50 years or younger at diagnosis. Sixty-one (25.6%) patients met more than one eligibility criterion.
The median age for the study population was 39.5 (24–64) years at BC diagnosis, and 49 (31–75) years at the time of genetic testing, resulting in a median time of 8 (2–20) years between diagnosis and genetic testing as part of this study. The majority (97.5%, n = 232) of the patients were disease-free at the time of genetic testing.
The chart review of the BC pathology reports for the patients who underwent genetic testing revealed that 225 (94.5%) had invasive ductal carcinoma, and 83 (34.9%) had high-grade pathology at the time of their initial diagnosis. In addition, 147 (61.8%) of the patients had estrogen receptor positive disease, 40 (16.8%) HER2/neu positive, and 73 (30.7%) had triple-negative disease. The characteristics of the patients tested are presented in Table 2.
Genetic testing results
P/LP variants were identified in 25 (10.5%) patients, namely, BRCA2 (n = 11), BRCA1 (n = 9), TP53 (n = 2), PALB2 (n = 2) and one case with ATM. On the other hand, 69 (29.0%) patients expressed VUS (Fig. 2). The frequency of P/LP-positive results significantly varied among the different age groups of BC diagnosis. The highest rate (22.7%) was among patients who were younger than 39 at diagnosis (p = 0.003), while age at BC diagnosis was statistically nonsignificant for patients who had VUS (p = 0.52).
Diagnosis with triple-negative breast cancer was not significantly associated with P/LP nor VUS (p = 0.55 and 0.72, respectively). Additionally, while having a record of any family history of cancer among first-, second-, or third-degree relatives was not significantly different among the P/LP and VUS cohorts, patients with two first-degree relatives who had positive BC diagnoses were significantly more likely to present with positive P/LP results (p = 0.0002). Statistical analyses results are summarized in Table 3.
Discussion
To our knowledge, this is the first study in the Middle East and North African (MENA) region that evaluated the feasibility and effectiveness of using the traceback approach to diagnose P/LP variants in individuals with a history of BC who had otherwise missed the opportunity of genetic counseling and testing at the time of diagnosis. Despite a median interval of 8 years between the initial cancer diagnosis and genetic testing, the rate of P/LP variants among our patients was 10.5%, comparable to the P/LP rates observed in upfront testing for breast cancer patients deemed eligible at diagnosis. These findings affirm that delayed genetic assessments can still be clinically relevant and effective36,37. In addition, the study results highlighted important patterns to patients with P/LPs and VUS and provided an opportunity for further testing and risk reduction strategies for relatives of patients who were otherwise unaware of the highly consequential mutations they may be carrying.
It is worth first noting the relatively young age of our patient cohort, who had a median age of 39.5 years-old at BC diagnosis. This observation aligns with the trend noted among Jordanian women, who are typically diagnosed with BC at a median age of 51 years, a decade earlier than their counterparts in Western nations38. Age appeared to have an impact on test result outcomes among our patients, with notable variations in patients with P/LP observed across different age groups. P/LP variants were particularly prevalent among individuals aged 39 years old or younger. This is consistent with previous reports associating younger BC patients with a higher predisposition for hereditary etiology. Reports have estimated that 12% of BC patients who developed the disease at ≤ 40 years were linked with PVs in BRCA1 or BRCA2 genes41,42,43, compared to global estimates of 3–4% of all women with BC18,44,45. Moreover, an analysis by Daly et al. of 130,151 BC patients using a 25-gene panel revealed that young age (≤ 45 years) at diagnosis was strongly associated with presence of PVs for BRCA1 (OR 3.95, 95% CI 3.64–4.29), moderate for BRCA2 (OR 1.98, 95% CI 1.84–2.14), modest associations for ATM (OR 1.22, 95% CI 1.08–1.37) and CHEK2 (OR 1.34, 95% CI 1.21–1.47) genes, and not associated with PVs for PALB2 (OR 1.12, 95% CI 0.98–1.27)46. Future prospective analyses of BC patients from our region could provide further insights to the prevalence of specific P/LPs among early-onset cancer patients.
The vast majority of the study participants (97.5%) presented with non-metastatic disease at diagnosis, and all individuals identified as carriers of P/LP variants were disease-free at the time of genetic testing. These carriers stand to gain from personalized screening, risk-reduction strategies, and family planning. Furthermore, cascade testing is clearly warranted, offering their family members the opportunity of genetic counseling and testing, and potentially earlier detection and prevention of cancer47. Most importantly, the identification of P/LP variants among our cohort of patients with history of BC provided an opportunity for significant risk mitigation procedures. Since P/LPs variants in BRCA1, BRCA2, CHEK2, or PALB2 substantially increase the risk of contralateral BC, the patients may benefit from additional aggressive surveillance specifically for contralateral BC with supplemental magnetic resonance imaging (MRI) and mammograms, as per international guidelines48. In addition, since P/LPs in BRCA1/2 have been associated with increased risk of subsequent ovarian cancer in BC survivors, our patients with positive P/LPs in these genes may benefit from prophylactic salpingo-oophorectomy20. The findings of this study could also guide the type of risk reduction surgery a patient might undergo, as is the case for BC patients with TP53 PVs, for which mastectomies are preferred over lumpectomies19. On the other hand, VUS were found in 69 (29%) of our subjects. Although the majority of VUS reclassification downgrade variants to benign or likely benign49,50, the result is noteworthy as VUS could place an additional burden on individuals and the health care system, by requiring regular reviews for potentially reinterpreted genetic test results.
Furthermore, the characteristics of our patient population addressed the feasibility of identifying eligible individuals for genetic testing based on age and pathology (i.e., triple-negative BC) as per the NCCN guidelines followed in the study design. Indeed, of the 238 identified eligible BC patients, 185 (77.7%) were diagnosed with breast cancer at aged 45 years or younger, and only 70 (29.4%) had triple-negative disease aged 60 years or younger. However, major challenges were observed in identifying eligible patients based on family history alone, as this process usually relies on a patient’s medical records, which tended to be less extensive in the early years of the study period. This was exacerbated by the fact that physicians rarely updated the family history records of patients who had completed active cancer treatment despite potential changes to their relatives’ health status over time. In fact, only 14 (0.6%) of the eligible patients were identified based on family history records alone. This observation presents yet another example to the importance of comprehensive noting and regular review of cancer patients’ family history records51.
Limitations
While this study provides promising insights into the feasibility and efficacy of the traceback approach for genetic testing in BC patients and survivors who were eligible for genetic testing but had not been previously tested, several limitations must be considered; the study only included patients who were still alive and willing to undergo genetic testing, which may introduce a survival bias. Patients who have died since their initial diagnosis or those who were not reachable or declined participation might have different genetic test results, potentially skewing the prevalence and spectrum of P/LP variants observed. The retrospective nature of the study limits the ability to control for variables that might influence the results, such as changes in eligibility criteria for genetic testing over time; this may affect the generalizability of the findings to current patients.
The study lacks long-term follow-up data to evaluate how genetic findings affected patient treatment and family risk management. It also does not detail the identification of at-risk relatives via cascade testing, or the effectiveness of the risk-reduction strategies used. These limitations stem from the study starting before the Early Detection and Prevention Clinic was established at our center in late 2022. We did not report results for patients tested according to the guidelines since the availability of genetic counseling and testing since 2017 to compare with the traceback approach. Additionally, the economic effects of implementing traceback genetic testing were not assessed.
Conclusions
Tracing back individuals with history of BC who had missed the opportunity to undergo genetic testing to diagnose P/LP variants, is a feasible and effective method that can provide actionable information for both patients and their relatives, and guide personalized treatment decisions and preventative measures, even after long overdue genetic referral. P/LP rates were found to be comparable to the rate of P/LP in upfront testing for eligible breast cancer patients in our country52.
Data availability
Data generated during and analysed during the current study are available from the corresponding author upon request. All variants reported in this manuscript, are submitted regularly to ClinVar. Our most recent submission has been processed in March 2023. Details are available at: https://www.ncbi.nlm.nih.gov/clinvar/submitters/500031.
References
WHO. WHO Report on Cancer: Setting Priorities, Investing Wisely and Providing Care for All. Ganeva (2020). https://www.who.int/publications/i/item/9789240001299. Accessed 15 Oct 2023.
Couch, F. J. et al. Associations between cancer predisposition testing panel genes and breast cancer. JAMA Oncol. 3, 1190–1196 (2017).
Riahi, A., el Gourabi, M. & Chabouni-bouhamed, H. Dissimilarity between sporadic, non-BRCA1/2 families and hereditary breast cancer, linked to BRCA genes, in the Tunisian population. Breast Cancer 23, 807–812 (2016).
Marmolejo, D. H. et al. Overview of hereditary breast and ovarian cancer (HBOC) guidelines across Europe. Eur. J. Med. Genet. 64, 104350 (2021).
Evans, D. G. et al. High likelihood of actionable pathogenic variant detection in breast cancer genes in women with very early onset breast cancer. J. Med. Genet. 59, 115–121 (2022).
Mai Tran, T. X., Kim, S., Song, H. & Park, B. Family history of breast cancer, mammographic breast density and breast cancer risk: Findings from a cohort study of Korean women. Breast 65, 180–186 (2022).
Mukama, T. et al. Familial risk of breast cancer by dynamic, accumulative, and static definitions of family history. Cancer 126, 2837–2848 (2020).
Kuchenbaecker, K. B. et al. Risks of breast, ovarian, and contralateral breast cancer for BRCA1 and BRCA2 mutation carriers. JAMA 317, 2402–2416 (2017).
Vargas, A. C., Reis-Filho, J. S. & Lakhani, S. R. Phenotype-genotype correlation in familial breast cancer. J. Mammary Gland Biol. Neoplasia 16, 27–40 (2011).
Shiovitz, S. & Korde, L. A. Genetics of breast cancer: A topic in evolution. Ann. Oncol. 26, 1291–1299 (2015).
Lerner-Ellis, J., Khalouei, S., Sopik, V. & Narod, S. A. Genetic risk assessment and prevention: The role of genetic testing panels in breast cancer. Expert Rev. Anticancer Ther. https://doi.org/10.1586/14737140.2015.109087915,1315-1326 (2015).
Antoniou, A. et al. Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case series unselected for family history: A combined analysis of 22 studies. Am. J. Hum. Genet. 72, 1117–1130 (2003).
Schon, K. & Tischkowitz, M. Clinical implications of germline mutations in breast cancer: TP53. Breast Cancer Res. Treat. 167, 417 (2018).
Fong, P. C. et al. Inhibition of poly (ADP-ribose) polymerase in tumors from BRCA mutation carriers. N. Engl. J. Med. 361, 123–134 (2009).
Byrski, T. et al. Response to neoadjuvant therapy with cisplatin in BRCA1-positive breast cancer patients. Breast Cancer Res. Treat. 115, 359–363 (2009).
Heemskerk-Gerritsen, B. A. M. et al. Improved overall survival after contralateral risk-reducing mastectomy in BRCA1/2 mutation carriers with a history of unilateral breast cancer: A prospective analysis. Int. J. Cancer 136, 668–677 (2015).
Evans, D. G. R. et al. Contralateral mastectomy improves survival in women with BRCA1/2-associated breast cancer. Breast Cancer Res. Treat. 140, 135–142 (2013).
Tung, N. M. et al. Management of hereditary breast cancer: American society of clinical oncology, American society for radiation oncology, and society of surgical oncology guideline. J. Clin. Oncol. 38, 2080–2106 (2020).
Heymann, S. et al. Radio-induced malignancies after breast cancer postoperative radiotherapy in patients with Li-Fraumeni syndrome. Radiat. Oncol. 5, 1–5 (2010).
Liu, Y. L. et al. Risk-reducing bilateral salpingo-oophorectomy for ovarian cancer: A review and clinical guide for hereditary predisposition genes. JCO Oncol. Pract. 18, 201–209 (2022).
Sessa, C. et al. Risk reduction and screening of cancer in hereditary breast-ovarian cancer syndromes: ESMO Clinical Practice Guideline. Ann. Oncol. 34, 33–47 (2023).
Febbraro, T. et al. Adherence patterns to National Comprehensive Cancer Network (NCCN) guidelines for referral to cancer genetic professionals. Gynecol. Oncol. 138, 109–114 (2015).
Childers, C. P., Childers, K. K., Maggard-Gibbons, M. & Macinko, J. National estimates of genetic testing in women with a history of breast or ovarian cancer. J. Clin. Oncol. 35, 3800 (2017).
Kurian, A. W. et al. Genetic testing and counseling among patients with newly diagnosed breast cancer. JAMA 317, 531–534 (2017).
Randall, L. M. et al. Multi-disciplinary summit on genetics services for women with gynecologic cancers: A Society of Gynecologic Oncology White Paper. Gynecol. Oncol. 146, 217–224 (2017).
Stuckey, A. et al. Adherence patterns to National Comprehensive Cancer Network Guidelines for referral of women with breast cancer to genetics professionals. Am. J. Clin. Oncol. 39, 363–367 (2016).
Offit, K. The future of clinical cancer genomics. Semin. Oncol. 43, 615–622 (2016).
Dorling, L. et al. Breast cancer risk genes—Association analysis in more than 113,000 women. N. Engl. J. Med. 384, 428–439 (2021).
Tuffaha, H. W. et al. Cost-effectiveness analysis of germ-line BRCA testing in women with breast cancer and cascade testing in family members of mutation carriers. Genet. Med. 20, 985–994 (2018).
Ladabaum, U. et al. Strategies to identify the Lynch syndrome among patients with colorectal cancer: A cost-effectiveness analysis. Ann. Intern. Med. 155, 69–79 (2011).
Guzauskas, G. F. et al. Cost-effectiveness of population-wide genomic screening for hereditary breast and ovarian cancer in the United States. JAMA Netw. Open 3, e2022874 (2020).
Manchanda, R. et al. Cost-effectiveness of population-based BRCA1, BRCA2, RAD51C, RAD51D, BRIP1, PALB2 mutation testing in unselected general population women. J. Natl. Cancer Inst. 110, 714–725 (2018).
Samimi, G. et al. Traceback: A proposed framework to increase identification and genetic counseling of BRCA1 and BRCA2 mutation carriers through family-based outreach. J. Clin. Oncol. 35, 2329–2337 (2017).
Schwartz, M. D. Identification of BRCA1 and BRCA2 mutation carriers through a traceback framework: Consent, privacy, and autonomy. J. Clin. Oncol. 35, 2226–2228 (2017).
Augustinsson, A. et al. Genetic testing in women with early-onset breast cancer: A Traceback pilot study. Breast Cancer Res. Treat. 190, 307–315 (2021).
Abdel-Razeq, H. et al. Abstract P3-07-06: Guideline-based multi-gene panel (MGP) testing for germline pathogenic variants among patients diagnosed with breast cancer: Regional perspectives. Cancer Res. 82, P3-07–06 (2022).
Abdel-Razeq, H. et al. Implementation of universal, pan-cancer germline genetic testing in patients with cancer in Jordan 40, 10580–10580 (2022). https://doi.org/10.1200/JCO.2022.40.16_suppl.10580.
Abdel-Razeq, H., Mansour, A. & Jaddan, D. Breast cancer care in Jordan. JCO Glob. Oncol. 6, 260–268 (2020).
Daly, M. B. et al. Genetic/familial high-risk assessment: Breast, ovarian, and pancreatic, version 2.2021, NCCN Clinical practice guidelines in oncology. J. Natl. Compr. Cancer Netw. 19, 77–102 (2021).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. https://doi.org/10.1038/gim.2015.30 (2015).
Rosenberg, S. M. et al. BRCA1 and BRCA2 mutation testing in young women with breast cancer. JAMA Oncol. 2, 730–736 (2016).
Lambertini, M. et al. Clinical behavior and outcomes of breast cancer in young women with germline BRCA pathogenic variants. NPJ Breast Cancer 7(1), 1–9 (2021).
Copson, E. R. et al. Germline BRCA mutation and outcome in young-onset breast cancer (POSH): A prospective cohort study. Lancet Oncol. 19, 169–180 (2018).
Ford, D., Easton, D. F. & Peto, J. Estimates of the gene frequency of BRCA1 and its contribution to breast and ovarian cancer incidence. Am. J. Hum. Genet. 57, 1457 (1995).
Cancer Genome Atlas Network, T. Comprehensive molecular portraits of human breast tumours (2012) . https://doi.org/10.1038/nature11412.
Daly, M. B. et al. The association between age at breast cancer diagnosis and prevalence of pathogenic variants. Breast Cancer Res. Treat. 199, 617–626 (2023).
Mandelker, D. et al. Germline-focussed analysis of tumour-only sequencing: Recommendations from the ESMO Precision Medicine Working Group. Ann. Oncol. 30, 1221–1231 (2019).
Yadav, S. et al. Contralateral breast cancer risk among carriers of germline pathogenic variants in ATM, BRCA1, BRCA2, CHEK2, and PALB2. J. Clin. Oncol. 41, 1703–1713 (2023).
Wright, M. et al. Factors predicting reclassification of variants of unknown significance. Am. J. Surg. 216, 1148–1154 (2018).
Makhnoon, S. et al. A multicenter study of clinical impact of variant of uncertain significance reclassification in breast, ovarian and colorectal cancer susceptibility genes. Cancer Med. 12, 2875–2884 (2023).
Clift, K. et al. Comparison of a focused family cancer history questionnaire to family history documentation in the Electronic Medical Record. J. Prim. Care Community Health 13, 215013192110697 (2022).
Kauffman, T. L. et al. Feasibility of a Traceback Approach for Using Pathology Specimens to Facilitate Genetic Testing in the Genetic Risk Analysis in Ovarian Cancer (GRACE) Study Protocol. J Pers Med 11, 94 (2021).
Acknowledgements
We would like to extend our gratitude to Mrs. Alice Haddadin from the King Hussein Cancer Center Medical Library for her assistance with the literature for this manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization, H.A.; methodology, H.A. and F.T.; validation, H.A. and H.B.; formal analysis, F.T.; data curation, B.S., O.S., S.A., S.E., H.B., K.A., and H.A.; writing—original draft preparation, H.A., S.I., F.T., H.B., S.A., and S.E.; supervision, H.A.; project administration, H.A., H.B., and B.S.; all authors reviewed and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abdel-Razeq, H., Tamimi, F., Iweir, S. et al. Genetic counseling and genetic testing for pathogenic germline mutations among high-risk patients previously diagnosed with breast cancer: a traceback approach. Sci Rep 14, 12820 (2024). https://doi.org/10.1038/s41598-024-63300-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-63300-8
- Springer Nature Limited