Abstract
Osteoporosis is a common skeletal disease affecting millions of individuals world-wide, with an increased risk of fracture, and a decreased quality of life. Despite its well-known consequences, the etiology of osteoporosis and optimal treatment methods are not fully understood. Human genetic studies have identified genetic variants within the FMN2/GREM2 locus to be associated with trabecular volumetric bone mineral density (vBMD) and vertebral and forearm fractures, but not with cortical bone parameters. GREM2 is a bone morphogenetic protein (BMP) antagonist. In this study, we employed Grem2-deficient mice to investigate whether GREM2 serves as the plausible causal gene for the fracture signal at the FMN2/GREM2 locus. We observed that Grem2 is moderately expressed in bone tissue and particularly in osteoblasts. Complete Grem2 gene deletion impacted mouse survival and body growth. Partial Grem2 inactivation in Grem2+/− female mice led to increased trabecular BMD of femur and increased trabecular bone mass in tibia due to increased trabecular thickness, with an unchanged cortical thickness, as compared with wildtype littermates. Furthermore, Grem2 inactivation stimulated osteoblast differentiation, as evidenced by higher alkaline phosphatase (Alp), osteocalcin (Bglap), and osterix (Sp7) mRNA expression after BMP-2 stimulation in calvarial osteoblasts and osteoblasts from the long bones of Grem2−/− mice compared to wildtype littermates. These findings suggest that GREM2 is a possible target for novel osteoporotic treatments, to increase trabecular bone mass and prevent osteoporotic fractures.
Similar content being viewed by others
Introduction
Osteoporosis is a disease that affects millions of individuals across the globe. One in three men and every other woman will at some point suffer from an osteoporotic fracture, with subsequent great personal suffering and a high societal cost1. As the human population is getting increasingly older2,3, the number of osteoporosis patients will increase dramatically. While the consequences of this disease are relatively well-known, there remains a knowledge gap concerning its etiology and the most safe and effective treatment methods. In recent years, the development of genome-wide association studies (GWAS) has emerged as a crucial tool for identifying potential therapeutic avenues for osteoporosis4.
Separate GWAS meta-analyses for cortical and trabecular volumetric bone mineral density (vBMD) in the tibia have revealed a genetic variant (rs9287237) within the FMN2/GREM2 locus associated with trabecular vBMD, but not cortical bone parameters5. Another GWAS identified a correlated SNP (rs9661787) in the GREM2 locus as significantly associated with trabecular vBMD in the lumbar spine6. The genetic variant (rs9287237) in the FMN2/GREM2 locus was also associated with fracture risk and X-ray verified vertebral fractures5. Furthermore, expression quantitative trait locus (eQTL) analyses in human osteoblasts showed a significant association between the trabecular vBMD associated SNP in the FMN2/GREM2 locus and the nearby GREM2 gene's expression, suggesting GREM2 as a potential mediator of the trabecular vBMD association5. Notably, most recently, we also demonstrated that the rs9287237 SNP in the FMN2/GREM2 locus is associated with forearm fractures at a genome-wide significant (GWS) level7.
GREM2, also known as Gremlin2 and PRDC (protein related to DAN and cerberus), is a bone morphogenetic protein (BMP) antagonist8,9. BMPs have effects in many different tissues, regulating cell proliferation, differentiation, and death10, and it has been suggested that BMP signaling is essential for limb bud outgrowth11. BMPs regulate proliferation and differentiation of osteoblasts and chondrocytes8,10. In vitro studies have demonstrated that reduced Grem2 expression in osteoblasts elevates osteogenesis and during osteoblast differentiation, BMP-2 induces the expression of Grem2 that then acts as a negative regulator of BMP-2 signaling12.
Despite the compelling evidence from different GWAS, the causal gene responsible for the associations with areal bone mineral density (aBMD), estimated bone mineral density (eBMD, obtained from quantitative ultrasound of the heel), vertebral fractures, and forearm fractures in the FMN2/GREM2 locus remains to be identified. The objective of this study is to, by using Grem2-deficient mice, investigate whether GREM2 serves as the plausible causal gene for the fracture signal at the FMN2/GREM2 locus. Our hypothesis posits that Grem2-deficient mice will exhibit increased bone mass preferably at trabecular bone sites.
Materials and methods
Animal experiment
All animal procedures were approved by the Gothenburg Animal Research Ethics Committee, Sweden, and the animals were cared for according to their and the ARRIVE guidelines. The mice were housed under a controlled temperature (20 °C), and photoperiod (12 h of light and 12 h of darkness), at a humidity of 45–70% in an animal house at University of Gothenburg. The mice were housed in individually ventilated cages (airflow 5 l/h under a balanced pressure) in groups of five or ten. Pellet diet (Envigo Harlan Teklad 2016) and water were available ad libitum.The mice were euthanized using Ketador (Richter Pharma) mixed with Dexdomitor (Orion Pharma), followed by exsanguination and cervical dislocation. Tibia, femur, and vertebra were dissected, fixed in formalin, and stored in ethanol. Soft tissues were snap-frozen in liquid nitrogen following dissection. Cortical bone from cleaned and flushed tibia shafts and trabecular bone from the vertebral body were stored in RNAlater (76106, Qiagen) and frozen.
Generation of Grem2-deficient mice
Grem2-deficient mice were generated by breeding female and male Grem2+/− mice commercially available from Taconic (TF3196, Hudson, NY), on a C57/BL6N background. Grem2+/+ mice were used as littermate controls. In brief, the Grem2−/− mice have a deletion of the coding exon 213. Presence of the intact Grem2 allele was detected using real-time qPCR (StepOnePlus Real-Time PCR System, Thermo Fisher Scientific) with the following primers and FAM-labelled probe: 5′-ATCCTCAACCGCTTCTGCTA-3′, 5′-GGAGTCCTCCTCCTTCTTCA-3′, and 5′-FAM-TCCTTCTACATCCCGCGACACG-3′, whereas absence of the intact Grem2 allele was detected by the following primers and VIC-labelled probe: 5′-TCTGGATTCATCGACTGTGG-3′, 5′-TCAGCAATATCACGGGTAGC-3′, and 5′-VIC-CCTGATAGCGGTCCGCCACA-3′.
Assessment of bone parameters
Dual X-ray absorptiometry
Total body bone mineral density (BMD) was measured using the Lunar PIXImus densitometer (Wipro GE Healthcare).
High-resolution microcomputed tomography (µCT)
High-resolution microcomputed tomography (µCT) analyses were performed on proximal tibia and vertebra L5 using the 1172 µCT (Bruker MicroCT), as previously described14. The trabecular bone parameters were analyzed in the metaphyseal region of tibia, starting approximately 650 µm from the proximal growth plate, continuing for roughly 134 µm in distal direction. Cortical thickness was measured in the diaphyseal region of the tibia, starting approximately 5.2 mm from the growth plate, continuing for roughly 134 µm in distal direction. For trabecular parameters from vertebra L5, trabecular bone caudal of the pedicles were analyzed starting at a distance of approximately 4.5 µm caudal of the lower end of the pedicles, continuing in a longitudinal direction for 230 µm in the caudal direction. The data was analyzed using the CTAn software (Bruker MicroCT).
Peripheral quantitative computed tomography (pQCT)
Computed tomography (pQCT) analyses were performed using the XCT Research M (v.4.5B, Norland Stratec) at a resolution of 70 µm15. In order to measure trabecular volumetric BMD, the scans were positioned in the metaphysis of femur at a distance corresponding to 3.4% of the total femur length, proximal from the distal growth plate. The trabecular region was defined as the inner 45% of the total cross-sectional area16.
Bone histomorphometry
For dynamic histomorphometry, the mice were labeled intraperitoneally using the fluorochromes calcein (C0875, Merck GmbH) and alizarin (A3882, Merck GmbH) 8 and 1 day prior to termination, respectively. After dissection, tibia was fixated in formalin, dehydrated in ethanol, and imbedded in methyl methacrylate. Dynamic trabecular bone parameters were measured in unstained 8-µm-thick sections, and static trabecular parameters were determined in 4-µm-thick plastic sections stained in Masson-Goldner’s Trichrome. All parameters were analyzed using the OsteoMeasure7 histomorphometry system (OsteoMetrics), according to the guidelines of the American Society for Bone and Mineral Research17.
Real-time quantitative PCR
Total RNA from cortical (shafts from flushed tibia) and trabecular-rich (vertebral body) bone, brain cortex, muscle gastrocnemius, brown fat, retroperitoneal fat, and gonadal fat was prepared using TRIzol Reagent (15596018, Thermo Fisher Scientific) followed by the RNeasy Mini Kit (74116, Qiagen). Total RNA from ovary, hypothalamus, lung, liver, aorta, kidney, spleen, uterus, and heart was prepared using the RNeasy Mini Kit (74116, Qiagen). The RNA was reversed transcribed into cDNA (4368814, Applied Biosystems) and real-time PCR analyses were performed using the StepOnePlus Real-Time PCR System (Thermo Fisher Scientific). Grem2 expression was measured using the following custom-made primer and probe sets located within the deleted region of exon 2: forward primer: 5′-TGGCTGTGCTGGTAAAGGTA-3′, reverse primer: 5′-TTGATCTGGTGATGCCACCT-3′, and probe: FAM-5′-CGCCCGCAGGCCGGTTCTTC-3′. The following Assay-on-Demand primer and probe sets were used: Alp, Mm00475834_m1; Bglap (encoding osteocalcin), Mm03413826_mH; and Sp7 (encoding osterix), Mm04209856_m1. For Grem2 expression analyses in liver and gonadal fat of Grem2+/− heterozygote and wildtype littermates, Mm00501909_m1 Assay-on-Demand primer and probe set was used. Relative gene expression was calculated by the 2–∆∆Ct method using the expression of the 18S ribosomal subunit (4310893E, Thermo Fisher Scientific) as internal standard.
Primary bone cell cultures
Primary calvarial bone cells
Primary calvarial bone cells were isolated from 4 to 6 day old mice by consequential collagenase treatments18. Cells in collagenase fractions 6–10 were cultured in complete αMEM (α-MEM medium, 22561-021, Gibco, supplemented with 10% heat inactivated fetal bovine serum (FBS, F7524, Sigma), 2 mM GlutaMAX (35050-038, Gibco), 50 µg/ml gentamicin (15750-037, Gibco), 100 U/ml penicillin and 100 µg/ml streptomycin (15140-148, Gibco)) for 4 days prior to re-seeding at 20,000 cells per square centimeter in 48-well plates. The cells were cultured in complete osteogenic αMEM (complete α-MEM medium with 2 mM β-glycerophosphate (G9422, Sigma), and 0.2 mM l-Ascorbic acid 2-phosphate sesquimagnesium salt hydrate (A8960, Sigma)) with change of half of the media volume every 2–3 days. For mRNA expression analyses, the cells were lysed in RNeasy lysis (RLT) buffer with β-mercaptoethanol (M6250, Sigma-Aldrich) followed with RNeasy Micro RNA kit (74004, Qiagen), cDNA synthesis, and real-time PCR as described above. To study osteoblast differentiation, cells were treated with or without BMP-2 (100 ng/ml, 355-BEC, R&D Systems).
Primary osteoblast cell cultures from long bones
Primary osteoblast cell cultures from tibia from 12-week-old female Grem2+/+ and Grem2−/− mice were isolated by outgrowth of cells from collagenase-treated diaphyseal cortical bone, as previously described19. Primary bone cells from tibia were incubated in complete osteogenic αMEM, and the experiments were conducted as described for primary calvarial bone cells above. To study osteoblast differentiation, cells were treated with or without BMP-2 (100 ng/ml, 355-BEC, R&D Systems). For mineralization analyses, the cells were cultured for 10 days and then washed in PBS, fixed in 2.5% glutaric aldehyde in 70% ethanol, washed three times in 70% ethanol and air dried. Mineralized nodules were visualized by staining with 1% alizarin red in 50% ethanol for 5 min, followed by three washes in 50% ethanol and air dried. Whole well stitched images were taken using a Jenoptik Gryphax camera on a Nikon Eclipse 80i microscope. The percentage of the well surface covered by mineralized matrix was determined using the Bioquant Osteo software (version 2023 v23.5.60).
Primary osteoclast cell cultures
Bone marrow cells from wildtype mice were cultured in suspension culture discs (430591, Corning) in complete αMEM with 30 ng/ml macrophage-stimulating factor (M-CSF, 416-ML-050, R&D Systems), for 2 days. The adherent bone marrow macrophages (BMMs) were used as osteoclast progenitors20,21. The cells were detached and spot-seeded in 24-well plates (20,000 cells per well). The cells were cultured in complete αMEM with 30 ng/ml M-CSF with or without the addition of 4 ng/ml receptor activator of nuclear factor κ-B ligand (RANKL, 462-TEC, R&D Systems) to induce osteoclast differentiation. After 3 days, the medium was changed, and 4 days after seeding, the cells were harvested for RNA preparation by lysis in RLT buffer with β-mercaptoethanol.
Statistics
Results are presented as mean ± standard error of the mean. Genotype ratios were tested against a Mendelian ratio (1:2:1) with a one sample chi-square goodness of fit test (SPSS, version 29.0.0.0). One-way ANOVA followed by Dunnett’s multiple comparisons test (GraphPad Prism, version 10.0.3) was used to assess the effect of genotype on body weight between Grem2−/− versus Grem2+/+ and Grem2+/− versus Grem2+/+ mice. To calculate the difference in Grem2 expression over time in wildtype calvarial osteoblast, one-way ANOVA followed by Dunnett’s multiple comparisons test (GraphPad Prism) was used, using the expression of day 1 as the reference. To evaluate the effect of genotype (Grem2+/− and Grem2+/+), age (10 or 12 weeks), and the interaction thereof, two-way ANOVA was used (GraphPad Prism). For analyses of primary osteoblast cell cultures, Student’s t test was used to evaluate the difference in gene expression between two groups (Grem2+/+ mice vs. Grem2+/− mice, Grem2+/+ cells vs. Grem2−/− cells, or Ctrl vs. BMP-2). Two-way ANOVA (GraphPad Prism) was used to assess overall interaction between BMP-2 effect in Grem2+/+ cells vs. BMP-2 effect in Grem2−/− cells. Power analysis was performed to determine group sizes. The analysis suggested that the use of eleven Grem2+/+ and eleven Grem2+/− mice would give 80% power to detect a biological significant effect (where P < 0.05) with a 1.26 s.d. change in trabecular bone mass. We therefore aimed to use at least eleven mice per group in the in vivo studies.
Result
Grem2 is moderately expressed in bone tissue
To study the expression pattern of Grem2, we compared the Grem2 expression in 16 tissues from mice (female, aged 12 weeks) and observed high expression in ovary, certain brain structures, such as hypothalamus and brain cortex, and lung, and moderate expression in cortical bone and vertebral body, rich in trabecular bone (Fig. 1a). Analyses of isolated calvarial cells, containing a large proportion of osteoblasts, indicated that cultured primary osteoblasts express Grem2, but the expression decreases with time in these cultures (Fig. 1b). No expression of Grem2 was present in cultured bone marrow macrophages (BMM) or RANKL-differentiated osteoclasts (OCL) (Fig. 1a). To further elucidate expression of Grem2 in bone cells, single cell RNA-sequencing (scRNAseq) datasets of freshly isolated primary mouse calvarial cells and cultured calvarial cells were used22. Analysis of the datasets demonstrated expression of Grem2 both in freshly isolated primary osteoblasts and in cultured osteoblasts from the calvaria (Fig. 1c, d). Grem2 was modestly expressed in osteoblasts in scRNAseq datasets of mouse bone marrow cells (Supplementary Fig. 1)23. FMN2 is expresssed in brain tissues but no expression of FMN2 was observed in freshly isolated primary mouse calvarial cells or in cultured calvarial cells (Supplementary Fig. 2).
Grem2 is expressed in osteoblasts but not in osteoclasts in mice. (a) Grem2 mRNA expression pattern in various tissues of 12-week-old female, wildtype mice (n = 6), bone marrow macrophages (BMM), and osteoclasts (OCL). Data presented as % of the expression in trabecular-rich vertebral body and shown as individual values with the mean presented as horizontal lines and ± standard error of the mean as vertical lines. N.D. not detectable. (b) Grem2 mRNA expression in primary calvarial osteoblasts from wildtype mice decreases over time in culture. Data presented as % of the expression at day 1 and shown as individual values with the mean presented as horizontal lines and ± standard error of the mean as vertical lines. Difference in expression over time was analyzed using one-way ANOVA followed by Dunnett’s multiple comparisons test with day 1 as the reference group. n = 4 per timepoint. The experiment was repeated three times. (c,d) Cluster analysis (c), and feature plots of Grem2 expression (d) in freshly isolated primary mouse calvarial cells and cultured calvarial cells published by Ayturk et al.22, publicly available at Single Cell Portal (https://singlecell.broadinstitute.org/single_cell/study/SCP1337). Freshly isolated cells and cultured cells form separate clusters as indicated in the figure22.
Complete Grem2 gene deletion affects mouse survival and growth
To obtain Grem2-deficient mice, we bred Grem2+/− female mice with Grem2+/− male mice. We observed that Grem2−/− mice were not born according to Mendel’s law of inheritance with the expected Mendelian ratio (1:2:1). From 320 pups, we obtained 88 Grem2+/+ mice, 200 Grem2+/− mice, and 32 Grem2−/− mice, demonstrating that a complete deletion of the Grem2 gene affects survival (Supplementary Fig. 3). A one sample chi-square goodness of fit test confirmed an unexpected Mendelian ratio (P = 2.5 × 10–9).
At 5 weeks of age, the few Grem2−/− mice born were small with a lower body weight compared to wildtype littermates (female mice: -13.3 ± 5.3%, P = 0.0046; male mice: −20.6 ± 5.8%, P = 3.0 × 10–4). No differences in body weight were found between Grem2+/− heterozygote and wildtype littermates (female mice: −2.6 ± 1.5%, P = 0.74; male mice: + 1.6 ± 1.9%, P = 0.88).
Increased trabecular bone mass in female mice with partial Grem2 inactivation
As the Grem2−/− mice born were severely growth restricted, it was not feasible to study their skeletal phenotype. Therefore, skeletal analyses on adult mice were performed only on the Grem2+/− heterozygote and wildtype littermates. Grem2 mRNA expression analysis demonstrated a reduction in Grem2 expression in the trabecular-rich vertebral body (−42.8 ± 8.2%, P = 0.0023) in the heterozygote Grem2+/− mice, compared with wildtype Grem2+/+ littermates (Fig. 2a). A similar decrease in Grem2 expression was seen in liver and gonadal fat (Supplementary Fig. 4).
Increased trabecular bone mass in mice with partial inactivation of Grem2. (a) mRNA expression analyses of Grem2 in trabecular-rich vertebral body of 12-week-old male Grem2+/+ (n = 11) and Grem2+/− (n = 11) mice. (b) Body weight in Grem2+/+ (10-week-old, n = 10; 12-week-old, n = 11), and Grem2+/− (10-week-old, n = 11; 12-week-old, n = 10) mice. (c,d) Length of tibia (c) and femur (d) in Grem2+/+ (10-week-old, n = 10; 12-week-old, n = 11), and Grem2+/− (10-week-old, n = 11; 12-week-old, n = 11) mice. (e) Whole body areal bone mineral density (aBMD) as measured by DXA in Grem2+/+ (10-week-old, n = 9; 12-week-old, n = 11), and Grem2+/− (10-week-old, n = 11; 12-week-old, n = 11) mice. (f–h) Trabecular bone volume over tissue volume (BV/TV (f)), trabecular number (Tb.N (g)), and trabecular thickness (Tb.Th (h)) in tibia as measured by µCT in Grem2+/+ (10-week-old, n = 10; 12-week-old, n = 11), and Grem2+/− (10-week-old, n = 11; 12-week-old, n = 11) mice. (i) Trabecular bone mineral density (BMD) in femur as measured by pQCT in Grem2+/+ (10-week-old, n = 10; 12-week-old, n = 11), and Grem2+/− (10-week-old, n = 11; 12-week-old, n = 11) mice. (j) Cortical thickness of tibia as measured by µCT in Grem2+/+ (10-week-old, n = 10; 12-week-old, n = 11), and Grem2+/− (10-week-old, n = 11; 12-week-old, n = 11) mice. Unless otherwise stated, the results refer to female mice. Individual values are presented in all graphs with the mean presented as horizontal lines and ± standard error of the mean presented as vertical lines. Statistical analyses were performed using two-sided Student’s t test (for (a)), and a mixed model two-way ANOVA were used to evaluate the genotype effect (Grem2+/+ or Grem2+/−), the age effect (10-week-old or 12-week-old), and the overall interaction effect between genotype and age (for (b–j)).
Similar to their condition at 5 weeks of age, 10- and 12-week-old Grem2+/− female mice appeared healthy when compared to Grem2+/+ littermate controls, with unchanged body weight (−1.5 ± 2.15%, P = 0.48; Fig. 2b). Furthermore, when analyzing the skeleton, Grem2+/− female mice had a similar length of long bones compared to Grem2+/+ littermate controls (tibia; −8.86 ± 0.51%, P = 0.10, Fig. 2c; femur; −0.54 ± 0.60%, P = 0.38, Fig. 2d). DXA analyses of the two independent 10- and 12-week-old Grem2+/− female mice cohorts compared to Grem2+/+ littermate controls revealed no differences in total body areal bone mineral density (aBMD; + 0.63 ± 1.14%, P = 0.59; Fig. 2e). However, a more detailed analysis of the tibia by μCT revealed an increase in trabecular bone volume fraction (BV/TV, + 10.0 ± 3.5%, P = 0.0063; Fig. 2f), as a result of increased trabecular number (Tb.N, + 10.6 ± 3.3%, P = 0.0038; Fig. 2g), whereas the trabecular thickness was unchanged (Tb.Th, −0.04 ± 1.80%, P = 0.98; Fig. 2h). pQCT analysis of femur displayed an increase in trabecular BMD in the Grem2+/− female mice cohorts compared to the Grem2+/+ littermate controls (+ 14.0 ± 4.5%, P = 0.0031; Fig. 2i), confirming the results from tibia. In contrast, there was no difference in the trabecular bone of the vertebra in the Grem2+/− female mice compared to Grem2+/+ littermate controls (Table 1). No difference in cortical thickness of the long bones was observed in the Grem2+/− female mice compared to Grem2+/+ littermate controls (Ct.Th, −0.19 ± 1.26%, P = 0.87; Fig. 2j). In order to determine the mechanisms for the increased trabecular BV/TV of the tibia, static and dynamic histomorphometry were performed. However, static histomorphometry did not reveal any effects on neither the number of osteoblasts nor the number of osteoclasts and the dynamic histomorphometry analyses of the trabecular bone of tibia did not reveal any differences in the bone formation rate in the Grem2+/− mice compared to Grem2+/+ littermate controls (Table 2), suggesting a new steady state of bone turnover has been reached.
Adult 10-week-old Grem2+/− male mice also had unchanged body weight and length of the long bones as compared to their Grem2+/+ littermate controls (Table 3). DXA analysis revealed an increased aBMD in the Grem2+/− male mice compared to Grem2+/+ littermate controls (+ 5.36 ± 1.3%, P = 0.011; Table 3). When the trabecular and cortical bone was analyzed separately, an increasing trend in BV/TV was noticed (+ 9.42 ± 7.31%, P = 0.28), however, no other differences was detected (Table 3).
Grem2 inactivation stimulates osteoblast differentiation in primary bone cells
As GREM2 is an inhibitor of BMP, we aimed to investigate if downregulation of Grem2 affects osteoblast differentiation. To this end, we harvested primary calvarial bone cells from Grem2+/+ and Grem2−/− pups, and primary osteoblast cells from long bones of Grem2+/+ and Grem2−/− mice. As anticipated, the calvarial osteoblasts and osteoblasts isolated from long bones of Grem2−/− mice cultured in osteogenic media, did not display any Grem2 expression (Fig. 3a, b). Calvarial osteoblasts isolated from wildtype mice, cultured in osteogenic media and stimulated with BMP-2 for 7 days, showed an upregulation in Grem2 expression (+ 154 ± 18.9%, P = 1.9 × 10–4; Fig. 3a). Similarly, an increase in Grem2 expression was observed in BMP-2 stimulated osteoblasts isolated from long bones of wildtype mice (+ 258 ± 62.8%, P = 0.0064; Fig. 3b). As expected, BMP-2 stimulation had no effect on Grem2 expression in calvarial osteoblasts or osteoblasts isolated from the long bones of Grem2−/− mice (Fig. 3a, b).
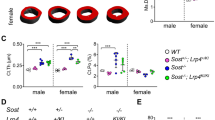
Grem2 inactivation stimulates osteoblast differentiation in primary bone cells. (a,b) mRNA expression levels of Grem2 in primary calvarial bone cells (a) and primary cells from long bones (b), after being cultured in osteogenic media and stimulated with or without bone morphogenetic protein-2 (BMP-2) for 7 days. (c,d) mRNA expression levels of Alp in primary calvarial bone cells (c) and primary cells from long bones (d), after being cultured in osteogenic media and stimulated with or without BMP-2 for 7 days. (e,f) mRNA expression levels of Bglap in primary calvarial bone cells (e) and primary cells from long bones (f), after being cultured in osteogenic media and stimulated with or without BMP-2 for 7 days. (g,h) mRNA expression levels of Sp7 in primary calvarial bone cells (g) and primary cells from long bones (h), after being cultured in osteogenic media and stimulated with or without BMP-2 for 7 days. (i) Alizarin red staining of mineralized nodules in osteoblast cultures of long bones from Grem2+/+ (left) and Grem2−/− (right) mice, after being cultured in osteogenic media and stimulated with BMP-2 for 10 days. Scale bar: 500 µm. (j) Quantification of mineralized nodules stained with alizarin red in osteoblast cultures of long bones from Grem2+/+ and Grem2−/− mice, after being cultured in osteogenic media and stimulated with or without BMP-2 for 10 days. Individual values are presented in all graphs with the mean presented as horizontal lines and ± standard error of the mean presented as vertical lines. Statistical analyses were performed using two-sided Student’s t test when comparing two groups (Grem2+/+ vs. Grem2−/−, or Ctrl vs. BMP-2 treatment), and a mixed model two-way ANOVA was used to evaluate the overall interaction effect between BMP-2 treatment in Grem2+/+ cells vs. BMP-2 treatment in Grem2−/− cells.
To evaluate the role of GREM2 in osteoblast differentiation, we examined the expression levels of Alp, Bglap (which encodes osteocalcin), and Sp7 (which encodes osterix). Upon stimulation with BMP-2 for 7 days, both calvarial osteoblasts and osteoblasts from the long bones of wildtype mice exhibited a noteworthy increase in Alp (+ 1448 ± 311%, P = 0.0036, Fig. 3c; and + 2131 ± 339%, P = 7.6 × 10–4, Fig. 3d, respectively), Bglap (+ 197 ± 11.7%, P = 7.1 × 10–5, Fig. 3e; and + 765 ± 152%, P = 0.0025, Fig. 3f, respectively), and Sp7 expression (+ 537 ± 89.3%, P = 0.0011, Fig. 3g; and + 2049 ± 392%, P = 0.002, Fig. 3h, respectively). Similarly, calvarial osteoblasts and osteoblasts from the long bones of Grem2−/− mice also demonstrated a substantial increase in Alp (+ 299 ± 75.9%, P = 0.007, Fig. 3c; and + 2917 ± 163%, P = 2.5 × 10–6, Fig. 3d, respectively), Bglap (+ 447 ± 64.5%, P = 0.0021, Fig. 3e; and + 1910 ± 328, P = 0.0011, Fig. 3f, respectively), and Sp7 (+ 1366 ± 235%, P = 0.0044, Fig. 3g; and + 2296 ± 211%, P = 3.7 × 10–5, Fig. 3h, respectively) expression following the same BMP-2 stimulation period. Notably, the impact of BMP-2 stimulation on Alp, Bglap, and Sp7 expression was more pronounced in cells from Grem2−/− mice compared to those from Grem2+/+ littermates, as evident by interaction effects in both calvarial osteoblasts (Alp; F (1,11) = 12.5, P = 0.0047, Bglap; F (1,11) = 9.34, P = 0.011, and Sp7; F (1,11) = 10.4, P = 0.0082) and osteoblasts from the long bones (Alp; F (1,12) = 12.7, P = 0.0036, Bglap; F (1,12) = 7.59, P = 0.017, and Sp7; F (1,12) = 6.17, P = 0.029). To study mineralization, osteoblasts derived from long bones were stimulated with or without BMP-2 for 10 days and then stained with alizarin red (Fig. 3i, Supplementary Fig. 5). The percentage of mineralized surface increased following BMP-2 stimulation in osteoblasts from both Grem2+/+ (+ 6867 ± 2569%, P = 0.038) and Grem2−/− mice (+ 11,522 ± 1320%, P = 1.3 × 10–4) (Fig. 3j). However, the stimulatory response on mineralized surface was more pronounced in cells from Grem2−/− mice compared to those from littermate wildtype mice (interaction effect, F (1,12) = 38.81, P = 4.4 × 10–5).
Discussion
The FMN2/GREM2 locus was previously recognized as a bone-related locus through GWAS5. This study aimed to determine whether GREM2 functions as the plausible causal gene underlying the fracture signal within the FMN2/GREM2 locus. Utilizing a globally Grem2-deficient mouse model in our study, we scrutinized the model's phenotype, revealing evidence that establishes GREM2 as the plausible causative gene for the fracture signal at the FMN2/GREM2 locus. Primary bone cell cultures derived from Grem2−/− mice demonstrated a greater increase in Alp, Bglap, and Sp7 expression following BMP-2 stimulation compared to wildtype cells, indicating an increase in osteoblast differentiation as a result of the GREM2 inactivation.
By using mRNA expression analyses and publicly available databases containing analyses from single cell RNA sequencing data, we demonstrated that Grem2 is expressed in several tissues and cell types including osteoblasts but not osteoclasts. These results are consistent with a previous report indicating expression of Grem2 in pre-osteoblasts of embryonic day 18.5 mouse calvaria12, and our previous study, demonstrating GREM2 expression in human osteoblasts5. The expression of Grem2 in calvarial bone cells decreased over time in osteogenic media. However, since the time course gene expression analyses were made in total RNA preparations it cannot be concluded if the expression per osteoblast is changed or if the relative number of osteoblasts in the cultures decrease compared to other cells in the primary cell cultures. Several studies have shown that the percentage of macrophages increase during in vitro culture of calvarial bone cells 22,24.
The low number of born Grem2−/− mice implies that GREM2 directly affects crucial developmental processes. This observation is in line with a previous indication from the International Mouse Phenotyping Consortium (IMPC), suggesting preweaning lethality of Grem2−/− mice25. A previous study in zebrafish26 demonstrated that loss of grem2 leads to overexpression of pitx2 and downregulation of lefty2, two crucial regulators of asymmetric cardiac development27,28. Moreover, grem2 knockdown in zebrafish increases the levels of activated phosphorylated Smad and BMP signaling within cardiac cells, interfering with both ventricular and atrial differentiation, indicating that GREM2 is required for pharyngeal mesoderm patterning, proper cardiac differentiation, and atrial chamber formation during cardiac development26. Thus, the low number of Grem2 knockout mice born may be attributed to the pivotal role of GREM2 in cardiac development. Furthermore, the few Grem2−/− mice born in the present study were growth restricted, which might also be a consequence of impaired cardiac development. Given the growth restriction in Grem2−/− mice, further skeletal analyses were focused on Grem2+/− heterozygote mice and their wildtype littermates.
The Grem2+/− mice appeared healthy, with no differences in body weight or long bone length compared to wildtype controls. Detailed skeletal analyses using DXA and CT scans revealed intriguing findings. In female mice, no differences were observed in total body areal bone mineral density (aBMD), but Grem2+/− mice displayed an increase in trabecular BMD of femur and trabecular BV/TV of the tibia, attributed to an elevated trabecular number, whereas there was no change in cortical thickness.
As we previously demonstrated that a signal within the FMN2/GREM2 locus was associated with trabecular but not cortical bone parameters in humans5, our present findings, together with the report on large-scale whole exome sequencing data in the UK Biobank dataset revealing that missense variants in the GREM2 gene are strongly associated with estimated BMD in the heel29, support the notion that it is plausible that the GREM2 gene within this locus is linked to trabecular bone parameters5. Furthermore, our results, demonstrating that GREM2 is important for trabecular but not cortical bone parameters of the long bones, is in accordance with the fact that a signal within the FMN2/GREM2 locus has been linked to forearm fractures7, as it was recently demonstrated that distal forearm fractures are highly dependent on trabecular BMD30.
As neither static nor dynamic histomorphometric analyses provided insights into the significance of GREM2 for the number of osteoclasts or osteblasts, or its impact on the bone formation process, possibly due to the establishment of a new equilibrium within the bone tissue in mice having a reduced expression of GREM2 from birth, additional in vitro studies were conducted. GREM2 is a BMP antagonist and we showed that stimulating calvarial osteoblasts and osteoblasts derived from the long bones with BMP-2, resulted in extensively enhanced expression of Grem2. These findings confirm and extend previous findings from both pre-osteoblasts from embryonic mouse calvaria and human bone marrow-derived mesenchymal stem cells among others, demonstrating an upregulation of Grem2 expression by BMP-2 stimulation12,31,32,33. These findings emphasize that expression of Grem2 is involved in a negative feedback mechanism. Importantly, the increase in expression of the osteogenic markers Alp, Bglap, and Sp7 was more pronounced in BMP-2 stimulated osteoblasts, from either calvaria or long bones, derived from Grem2-inactivated mice, further emphasizing the functional role of GREM2 as a BMP-2 antagonist. These results are in line with results from previous reports demonstrating that silencing Grem2 expression in cell cultures increases expression of Alp after BMP-2 stimulation, whereas overexpression of Grem2 in cell cultures decreases expression of Alp12,32. Mineralization studies on osteoblasts derived from long bones further confirmed the effect of Grem2 on osteoblast differentiation, as cells from mice lacking Grem2 had a higher percentage of mineralized surface after stimulation with BMP-2, when compared to cells from wildtype mice. Together with previous studies, our results demonstrate that suppression of Grem2 promotes osteoblast differentiation. The Grem2-deficient mouse model used in this study only captures changes in gene expression and does not account for other potential impacts, such as altered protein conformation or functional changes, which is a limitation of the study. Another limitation of the present study is that no data of FMN2 knockout mice are included. However, single cell RNA sequencing data demonstrated no expression of FMN2 in either freshly isolated primary mouse calvarial cells or cultured calvarial cells. Furthermore, a recent report on large-scale whole exome sequencing data in the UK Biobank dataset revealed that missense variants in the GREM2 gene, but not in the FMN2 gene, are strongly associated with estimated BMD in the heel, supporting the notion that GREM2 and not FMN2 is crucial for bone mass regulation29.
In conclusion, we functionally show that a partial deletion of Grem2 in mice leads to an increase in trabecular bone mass of long bones, with unchanged cortical bone. By using two different primary osteoblast cell cultures, we report mechanistic data showing that Grem2-inactivated cells display enhanced response to BMP-2 stimulation on osteoblast differentiation, as evident by an augmented increase in Alp, Bglap, and Sp7 expression. We believe that GREM2, being a BMP antagonist, is a promising new target for osteoporosis treatment. Further studies are warranted to explore the impact of GREM2 on bone mass and strength in neonatal and aged mice, and utilizing cell-specific and inducible knockout mouse models will provide a more comprehensive understanding of GREM2's involvement in bone biology.
Data availability
The datasets generated and analysed in the present study are available from the corresponding author upon reasonable request.
References
Harvey, N., Dennison, E. & Cooper, C. Osteoporosis: Impact on health and economics. Nat. Rev. Rheumatol. 6, 99–105 (2010).
Trajanoska, K. et al. Assessment of the genetic and clinical determinants of fracture risk: Genome wide association and mendelian randomisation study. BMJ (Clin. Res. Ed.) 362, k3225 (2018).
United Nations, D.o.E.a.S.A., Population Division. World population prospects: The 2015 revision, key findings and advance tables. In Working Paper No. ESA/P/WP.241. (2015).
Richards, J. B., Zheng, H. F. & Spector, T. D. Genetics of osteoporosis from genome-wide association studies: Advances and challenges. Nat. Rev. Genet. 13, 576–588 (2012).
Paternoster, L. et al. Genetic determinants of trabecular and cortical volumetric bone mineral densities and bone microstructure. PLoS Genet. 9, e1003247 (2013).
Nielson, C. M. et al. Novel genetic variants associated with increased vertebral volumetric BMD, reduced vertebral fracture risk, and increased expression of SLC1A3 and EPHB2. J. Bone Miner. Res. 31, 2085–2097 (2016).
Nethander, M. et al. An atlas of genetic determinants of forearm fracture. Nat. Genet. 55, 1820–1830 (2023).
Rosen, V. BMP and BMP inhibitors in bone. Ann. N. Y. Acad. Sci. 1068, 19–25 (2006).
Im, J., Kim, H., Kim, S. & Jho, E. H. Wnt/beta-catenin signaling regulates expression of PRDC, an antagonist of the BMP-4 signaling pathway. Biochem. Biophys. Res. Commun. 354, 296–301 (2007).
Katagiri, T. & Watabe, T. Bone morphogenetic proteins. Cold Spring Harbor Perspect. Biol. 8 (2016).
Salazar, V. S., Gamer, L. W. & Rosen, V. BMP signalling in skeletal development, disease and repair. Nat. Rev. Endocrinol. 12, 203–221 (2016).
Ideno, H. et al. Protein related to DAN and cerberus (PRDC) inhibits osteoblastic differentiation and its suppression promotes osteogenesis in vitro. Exp. Cell Res. 315, 474–484 (2009).
Vogel, P. et al. Malformation of incisor teeth in Grem2−/− mice. Vet. Pathol. 52, 224–229 (2015).
Moverare-Skrtic, S. et al. Osteoblast-derived WNT16 represses osteoclastogenesis and prevents cortical bone fragility fractures. Nat. Med. 20, 1279–1288 (2014).
Windahl, S. H., Vidal, O., Andersson, G., Gustafsson, J. A. & Ohlsson, C. Increased cortical bone mineral content but unchanged trabecular bone mineral density in female ERbeta(−/−) mice. J. Clin. Invest. 104, 895–901 (1999).
Vidal, O. et al. Estrogen receptor specificity in the regulation of skeletal growth and maturation in male mice. Proc. Natl. Acad. Sci. USA 97, 5474–5479 (2000).
Dempster, D. W. et al. Standardized nomenclature, symbols, and units for bone histomorphometry: A 2012 update of the report of the ASBMR Histomorphometry Nomenclature Committee. J. Bone Miner. Res. 28, 2–17 (2013).
Granholm, S., Henning, P., Lindholm, C. & Lerner, U. H. Osteoclast progenitor cells present in significant amounts in mouse calvarial osteoblast isolations and osteoclastogenesis increased by BMP-2. Bone 52, 83–92 (2013).
Bakker, A. D. & Klein-Nulend, J. Osteoblast isolation from murine calvaria and long bones. Methods Mol. Biol. (Clifton, N.J.) 816, 19–29 (2012).
Takeshita, S., Kaji, K. & Kudo, A. Identification and characterization of the new osteoclast progenitor with macrophage phenotypes being able to differentiate into mature osteoclasts. J. Bone Miner. Res. 15, 1477–1488 (2000).
Granholm, S., Lundberg, P. & Lerner, U. H. Calcitonin inhibits osteoclast formation in mouse haematopoetic cells independently of transcriptional regulation by receptor activator of NF-{kappa}B and c-Fms. J. Endocrinol. 195, 415–427 (2007).
Ayturk, U. M. et al. Single-cell RNA sequencing of calvarial and long-bone endocortical cells. J. Bone Miner. Res. 35, 1981–1991 (2020).
Dolgalev, I. & Tikhonova, A. N. Connecting the dots: Resolving the bone marrow niche heterogeneity. Front. Cell Dev. Biol. 9, 622519 (2021).
Chang, M. K. et al. Osteal tissue macrophages are intercalated throughout human and mouse bone lining tissues and regulate osteoblast function in vitro and in vivo. J. Immunol. 181, 1232–1244 (2008).
Birling, M. C. et al. A resource of targeted mutant mouse lines for 5,061 genes. Nat. Genet. 53, 416–419 (2021).
Müller, I. I. et al. Functional modeling in zebrafish demonstrates that the atrial-fibrillation-associated gene GREM2 regulates cardiac laterality, cardiomyocyte differentiation and atrial rhythm. Dis. Model Mech. 6, 332–341 (2013).
Bisgrove, B. W., Essner, J. J. & Yost, H. J. Regulation of midline development by antagonism of lefty and nodal signaling. Development (Cambridge, England) 126, 3253–3262 (1999).
Essner, J. J., Branford, W. W., Zhang, J. & Yost, H. J. Mesendoderm and left-right brain, heart and gut development are differentially regulated by pitx2 isoforms. Development (Cambridge, England) 127, 1081–1093 (2000).
Zhou, S. et al. Converging evidence from exome sequencing and common variants implicates target genes for osteoporosis. Nat. Genet. 55, 1277–1287 (2023).
Nethander, M. et al. BMD-related genetic risk scores predict site-specific fractures as well as trabecular and cortical bone microstructure. J. Clin. Endocrinol. Metab. 105, e1344-1357 (2020).
Suzuki, D. et al. BMP2 differentially regulates the expression of Gremlin1 and Gremlin2, the negative regulators of BMP function, during osteoblast differentiation. Calcif. Tissue Int. 91, 88–96 (2012).
Wang, C. L. et al. Gremlin2 suppression increases the BMP-2-induced osteogenesis of human bone marrow-derived mesenchymal stem cells via the BMP-2/Smad/Runx2 signaling pathway. J. Cell. Biochem. 118, 286–297 (2017).
Luo, X., Chang, H. M., Yi, Y., Sun, Y. & Leung, P. C. K. Bone morphogenetic protein 2 inhibits growth differentiation factor 8-induced cell signaling via upregulation of gremlin2 expression in human granulosa-lutein cells. Reprod. Biol. Endocrinol. 19, 173 (2021).
Acknowledgements
The authors thank Anna Westerlund and Biljana Aleksic from the University of Gothenburg for excellent technical assistance.
Funding
Open access funding provided by University of Gothenburg. This study was supported by the Swedish Research Council (grant number 2020-01392 and 2022-01156), the Swedish Foundation of Strategic Research, the Swedish state under the agreement between the Swedish government and the county councils, the ALF agreement in Gothenburg, the IngaBritt and Arne Lundberg Foundation, the Knut and Alice Wallenberg Foundation, and the Novo Nordisk Foundation.
Author information
Authors and Affiliations
Contributions
U.H.L., C.O., and S.M-S. conceived and designed the experiments; K.H.N, P.H., J.W., K.S., and S.M-S. performed experiments; K.H.N., C.O., and S.M-S. interpreted the results of the experiments; K.H.N. prepared figures; K.H.N. drafted the manuscript; K.H.N., P.H., J.W., K.S., U.H.L., C.O., and S.M-S. edited and revised the manuscript; K.H.N., P.H., J.W., K.S., U.H.L., C.O., and S.M-S. approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nilsson, K.H., Henning, P., Wu, J. et al. GREM2 inactivation increases trabecular bone mass in mice. Sci Rep 14, 12967 (2024). https://doi.org/10.1038/s41598-024-63439-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-63439-4
- Springer Nature Limited