Abstract
Endophytic fungal-based biopesticides are sustainable and ecologically-friendly biocontrol agents of several pests and diseases. However, their potential in managing tomato fusarium wilt disease (FWD) remains unexploited. This study therefore evaluated effectiveness of nine fungal isolates against tomato fusarium wilt pathogen, Fusarium oxysporum f. sp. lycopersici (FOL) in vitro using dual culture and co-culture assays. The efficacy of three potent endophytes that inhibited the pathogen in vitro was assessed against FWD incidence, severity, and ability to enhance growth and yield of tomatoes in planta. The ability of endophytically-colonized tomato (Solanum lycopersicum L.) plants to systemically defend themselves upon exposure to FOL were also assessed through defence genes expression using qPCR. In vitro assays showed that endophytes inhibited and suppressed FOL mycelial growth better than entomopathogenic fungi (EPF). Endophytes Trichoderma asperellum M2RT4, Hypocrea lixii F3ST1, Trichoderma harzianum KF2R41, and Trichoderma atroviride ICIPE 710 had the highest (68.84–99.61%) suppression and FOL radial growth inhibition rates compared to EPF which exhibited lowest (27.05–40.63%) inhibition rates. Endophytes T. asperellum M2RT4, H. lixii F3ST1 and T. harzianum KF2R41 colonized all tomato plant parts. During the in planta experiment, endophytically-colonized and FOL-infected tomato plants showed significant reduction of FWD incidence and severity compared to non-inoculated plants. In addition, these endophytes contributed to improved growth promotion parameters and yield. Moreover, there was significantly higher expression of tomato defence genes in T. asperellum M2RT4 colonized than in un-inoculated tomato plants. These findings demonstrated that H. lixii F3ST1 and T. asperellum M2RT4 are effective biocontrol agents against FWD and could sustainably mitigate tomato yield losses associated with fusarium wilt.
Similar content being viewed by others
Introduction
Globally, the agriculture industry is endangered by numerous plant pests and diseases, and their prevalence is increasing1. Farmers spend more than 40% of their crop production cost on pest control, while up to 50–82% reduction in attainable yield in the major crop has been attributed to both pests and diseases2,3,4. Tomato (Solanum lycopersicum L.), being the second most cultivated vegetable worldwide is among threatened commodities5. For instance, in Kenya, tomato production plays a vital role in improving lives, particularly for smallholder farmers, through employment and household income generation6,7. The crop accounts for approximately 14% and 6.72% of Kenya’s total vegetable and horticultural production, respectively8. In terms of local market sales, tomato makes up approximately 20% of total vegetable production in Kenya and contributes nearly 158.4 million US dollars annually to the Kenyan economy9. The potential yield of this crop in Kenya can reach up to 30.7 t ha−110. However, pests, diseases, and physiological disorders reduce the crop's achievable productivity by up to 12 tonnes per hectare8,11. According to Singh et al.12 and Ma et al.13, tomato is prone to approximately more than 200 diseases of which soil-borne diseases are the most prevalent.
Fusarium wilt is the most frequent soil-borne disease hindering tomato production13. It could cause up to 10–80% loss in the total production of tomatoes thereby threatening the livelihood of small-scale farmers8,13. The pathogenic fungus Fusarium oxysporum f. sp. lycopersici, which causes this disease enters the host roots and invades its vascular tissues, preventing water and nutrient uptake14,15. This results in wilting of one side of the plant, starting with the older leaves and progressing to the death of the entire host plant15,16. Moreover, the brown vascular darkening discolouration in cross sections of the stem tissue and stem decay are observed, especially in wet conditions17. Furthermore, the pathogen produces resting structures “chlamydospores” that can survive in soil for a decade, making its control extremely difficult15.
The management options of fusarium wilt range from cultural, resistant, chemical, and botanical to biological control16. Initially, uprooting infected plants and the use of resistant varieties was the safe, effective, and affordable strategy for farmers16. However, uprooting is laborious and finding resistant cultivars is difficult owing to the breakdown of resistance due to the various races of this pathogen16,18. Soil solarization and application of chemical pesticides such as methyl bromide, benomyl, or carbendazim are used to control the pathogen. However, significant concerns about their harmful effects on humans, animals, the environment and biodiversity have been raised11,19. To address these challenges, eco-friendly and sustainable strategies such as biological control that can effectively manage fusarium wilt in tomatoes are paramount13,20.
Biological control involves the use of living organisms (biocontrol agents) such as entomopathogenic fungi (EPF) and endophyte-based biopesticides in the sustainable management of crop pests and diseases21,22,23. These biological control agents (BCAs) like endophytes, grow faster and occupy the root zone of the plant before the pathogenic fungi are established thereby promoting plant health, triggering plant defence system and resistance to pathogen attack13,24,25,26. Recently, several EPF and endophytes such as Metarhizium anisopliae, Beauveria bassiana, Trichoderma spp. isolates and Hypocrea lixii have been shown to endophytically colonize tomato plants, promote growth, and protect the host plants against pests and pathogens27,28. Furthermore, yield improvement and systemic resistance to fusarium wilt were reported in tomato plants endophytically colonized by BCAs Trichoderma asperellum and B. bassiana26.
In addition, these BCAs especially endophytes benefit the host plants by activating their defence genes in endophytically colonized tomato plants and this was evidenced by higher expression levels of pathogenesis-related proteins and enzymes in comparison to untreated tomato plants24,26,29,30. For instance, the increased level of expression of pathogenesis-related proteins- PR-1 was seen in FOL-infected tomato plants pre-treated with T. asperellum strains TS-12 and TS-3924. Induction of defence-related SWRKY gene transcripts was found in tomato plants endophytically colonized with Trichoderma erinaceum and infected with FOL31. Tomato lipoxygenase (TomLoxC), which is responsible for jasmonic acid (JA) biosynthesis, was found to be highly expressed in tomato plants pre-treated with Trichoderma isolates and inoculated with Pseudomonas syringae pv. Tomato, and Botrytis cinerea, respectively32,33. Furthermore Prasetyawan et al.34 and Ueki et al.35, reported the role of pathogenesis-related proteins-PR-2 (β-1,3-glucanase) in suppressing a variety of fusarium wilt pathogens. Although several Trichoderma species' defence mechanisms have been studied, there is a gap in the proper understanding of this topic, as the effectiveness of these BCAs/endophytes varies from species to species, from one strain to another within a species, and is influenced by environmental factors36,37.
Recent studies reported for the antagonistic effects of fungal isolates from various genera such as Bacillus, Streptomyces, Trichoderma, Hypocrea, Beauveria, and Metarhizium against plants pests and pathogens13,26,38,39,40,41,42,43. However, knowledge of the effectiveness of locally isolated EPF and endophytes to sustainably manage fusarium wilt is largely unknown. Additionally, the mechanisms used by these microbial biocontrol agents to stimulate tomato plant growth and yield and confer resistance against fusarium wilt have not been explored. Hence, this study aimed to contribute to the sustainable management of fusarium wilt disease of tomato crops in Kenya using entomopathogenic fungi and endophyte-based biopesticides. Consequently, an in vitro assessment of some selected entomopathogenic and endophytic fungal isolates against Fusarium oxysporum lycopersici fungal pathogen was conducted. In addition, the efficacy of the potent fungal isolates on the occurrence and severity of fusarium wilt disease in tomato crops under greenhouse conditions was evaluated. Furthermore, the study assessed the expression levels of some selected defence genes in tomato plants endophytically colonized by T. asperellum M2RT4 when exposed to Fusarium oxysporum lycopersici.
Results
Identification of Fusarium wilt pathogen for bioassays
The isolated fungus was identified as Fusarium oxysporum f. sp. lycopersici (FOL) based on key morphological features described by Leslie and Summerell44, such as white colony color and purplish-pink pigmentation (Fig. S1A and B). Three types of spores—microconidia, macroconidia, and chlamydospores—were observed under a microscope (40× magnification). The pathogen was successfully re-isolated from inoculated tomato roots and stems, with 100% and 80% colonization rates, respectively. Besides, the re-isolated pathogen exhibited similar morphological features to the inoculated FOL mother plates following slide preparation for comparison/confirmation test. Furthermore, using ITS 4 and ITS 5, EF1 α, and RPB1 primers, the molecular testing confirmed the identification of the pathogen as FOL with 100%, 99.23%, and 98.64% nucleotide similarity to the FOL sequences deposited to GenBank of accession numbers MT530269.1, XM_018381269.1, and XM_018377523.1, respectively.
Antagonistic effect of the selected fungal biocontrol agents against Fusarium oxysporum lycopersici
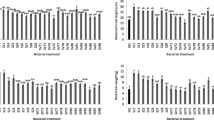
The endophytic fungal isolates showed high potential in inhibiting and suppressing the pathogen compared to EPF (Fig. 1). A significant difference (χ2 = 2277.7, df = 8, P < 0.001) was observed among the nine screened fungal BCAs against FOL (Fig. 2). Of all the four endophytic fungal isolates, T. asperellum M2RT4 had the highest inhibition and suppression of radial growth of FOL, while T. atroviride ICIPE 710 had the lowest. Trichoderma asperellum M2RT4, H. lixii F3ST1, T. harzianum KF2R41, and T. atroviride ICIPE 710 achieved 99.61%, 90.5%, 86.84%, and 68.84% FOL mycelia growth inhibition, respectively. Among EPF, M. anisopliae ICIPE 20 had the highest inhibition rate (40.63%) of FOL followed by B. bassiana ICIPE 273 (38.10%), M. anisopliae ICIPE 18 (35.53%), M. anisopliae ICIPE 655 (35.23%), while the lowest inhibition rate (27.05%) of FOL mycelia growth was obtained in the dual culture of FOL and B. bassiana ICIPE 706 (Fig. 2).
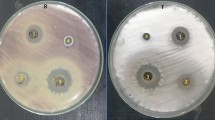
Inhibitory and suppressive effects of selected biocontrol agents against Fusarium oxysporum lycopersici mycelia growth on the 12th day of their dual culture assay: (a) Trichoderma asperellum M2RT4, (b) Hypocrea lixii F3ST1, (c) Trichoderma harzianum KF2R41, (d) Trichoderma atroviride ICIPE 710, and (e) Beauveria bassiana ICIPE 706, (f) Beauveria bassiana ICIPE 273, (g) Metarhizium anisopliae ICIPE 20, (h) Control, (i) Metarhizium anisopliae ICIPE 18, (j) Metarhizium anisopliae ICIPE 665.
Average percentage (mean ± SE) of mycelial growth inhibition of four endophytic and five entomopathogenic fungal isolates against Fusarium oxysporum lycopersici on the 12th day of their dual culture assay. Different small letters above the error bars denote significant differences across the treatment means.
Co-culture of fungal biocontrol agents and test pathogen
All the endophytic fungal isolates were dominant in terms of mycelia and pigments produced in their co-culture with FOL on the 7th day post-inoculation. In contrast, FOL mycelia and pigment predominated on co-cultured EPF and FOL plates. Among all fungal BCAs, only T. asperellum M2RT4 suppressed/inhibited the growth and expression of the mycelia of FOL in their co-cultured plates on day 14 after incubation. Similarly, dominance in mycelial morphology was observed in co-cultures of T. harzianum KF2R41, H. lixii F3ST1, and T. atroviride ICIPE 710 with FOL; however, a small amount of FOL mycelia were also observed in these co-cultures. On the other hand, FOL mycelia and its pigmentation were entirely dominant in its co-culture with M. anisopliae isolates ICIPE 18, ICIPE 20, and ICIPE 655, although mycelia were more abundant in the co-culture of FOL and M. anisopliae ICIPE 665. Furthermore, white spores of B. bassiana were observed in the co-cultured plates of B. bassiana isolates ICIPE 273 and ICIPE 706 and FOL; however, FOL mycelia and pigmentation were dominant (Fig. S2).
Endophytic colonization assessment
Before the in planta bioassay, the endophytic colonization of the identified potent endophytes which suppressed the pathogen in vitro was assessed or confirmed through seed inoculation. Significant differences in the tomato seedling colonization by the various endophytic isolates were observed on roots (χ2 = 6, df = 2, P = 0.04979), stems (χ2 = 26.571, df = 2, P < 0.001), and leaves (χ2 = 33.273, df = 2, P < 0.001) (Fig. 3). Trichoderma asperellum M2RT4, T. harzianum KF2R41, and H. lixii F3ST1 colonized all the tomato plant parts and were effectively re-isolated from inoculated tomato seedlings. However, the rate of colonization varied among isolates and plant sections/tissues. Among the isolates, T. asperellum M2RT4 demonstrated high potential of colonizing all tomato plant parts at high rates; however, the roots exhibited the highest rate of colonization rate (100%) compared to stems (90%) and leaves (80%) (Fig. 3). A similar trend was seen in the colonization rate of T. harzianum KF2R41, where the colonization rates of roots, stems, and leaves were 100%, 75%, and 70%, respectively. Furthermore, H. lixii F3ST1 colonized tomato roots, stems, and leaves at the rates of 95, 70, and 60%, respectively (Fig. 3).
Effect of the selected potent fungal endophytes on fusarium wilt disease incidence and severity in planta
Tomato plants inoculated with endophytes T. asperellum M2RT4, H. lixii F3ST1, and T. harzianum KF2R41 significantly-reduced the incidence and severity of fusarium wilt disease at 15-, 30-, 45- and 60-days post-inoculation (dpi) compared to the FOL-infected plants (P < 0.001). Of all the treatments, T. asperellum M2RT4 proved to be the most effective with the lowest disease incidence (1.66%) and severity (0.04/4) (Fig. 4A and B). The highest incidence (80%) and severity (2.7/4) of fusarium wilt disease were recorded in FOL-inoculated plants. The continuous increase in disease incidence up to 30 dpi and disease severity up to 60 dpi were also recorded in FOL-inoculated plants (Tables 1 and 2). Only moderate symptoms and decreased fusarium disease incidence (6.6%, 32.5%) and severity (0.5/4, 1.1/4) were observed in H. lixii F3ST1 and T. harzianum KF2R41 inoculated plants, respectively. The greatest Area Under Disease Progress Curve (AUPDC) was obtained in tomato plants infected with FOL without pre-treatment with endophytes. This was followed by FOL-infected tomato plants pretreated with endophytes T. harzianum KF2R41, H. lixii F3ST1, T. asperellum M2RT4, showing AUPDC values of 1250, 562.5, and 78.125, respectively (Table 3). Furthermore, there was no incidence nor disease severity recorded in non-inoculated plants (Fig. 4A and B).
Effects of selected potent fungal endophytes in planta on fusarium wilt disease. Mean percentage and score of fusarium wilt incidence and severity at 15-, 30-, 45- and 60-days post-inoculation (dpi). (A) disease incidence, (B) disease severity. Each value represents the mean ± SE. Means followed by different letters are significantly different across the treatments at 95% CI (SNK test, P ≤ 0.05, n = 8).
Growth enhancement in endophytically-colonized and FOL-infected tomato plants
There were significant differences in all evaluated growth parameters of tomato plants pre-treated with the selected potent endophytes, infected with FOL, endophytically-colonized and FOL- infected, and un-inoculated tomato plants across all the assessment periods: 15-, 30-, 45- and 60-days post-inoculation (dpi) (Fig. 5). Significant differences in the number of fully developed leaves were observed among the treatments at 15, 30, 45, and 60 dpi (χ2 = 221.09, df = 7, P < 0.001; χ2 = 238.26, df = 7, P < 0.001; χ2 = 320.55, df = 7, P < 0.001; χ2 = 802.98, df = 7, P < 0.001), respectively (Fig. 5A). The highest number of fully developed leaves (18.83 ± 0.51) was recorded in T. asperellum M2RT4 endophytically-colonized tomato plants, while FOL-infected tomato plants had the lowest (6.16 ± 0.42). Furthermore, from 45 to 60 dpi, the number of fully developed leaves decreased from 6.75 to 6.16 (± 0.42) in FOL-infected plants. Time series analysis revealed significant differences (χ2 = 1780.391, df = 7, P < 0.001; χ2 = 3522.2, df = 3, P < 0.01) in number of fully developed leaves across treatments and all sampling periods, respectively.
Effect of the selected potent fungal endophytes on the growth of FOL-infected tomato plants at 15-, 30-, 45- and 60-days post-inoculation (dpi). (A) Number of fully developed leaves, (B) Leaf width, (C) Leaf length, (D) Plant height. The bars represent mean ± standard error. Means followed by different small letters are significant differences across the treatment at 95% CI (SNK test, P ≤ 0.05, n = 8).
The leaf width differed significantly in response to the various treatments across all the post-inoculation periods (15, 30, 45 and 60 dpi) as the tomato plants were growing (χ2 = 22.361, df = 7, P < 0.001; χ2 = 25.463, df = 7, P < 0.001; χ2 = 589.11, df = 7, P < 0.001; χ2 = 834.65, df = 7, P < 0.001), respectively (Fig. 5B). The result of time series analysis showed significant differences in leaf width and all the post-inoculation periods (χ2 = 563.1, df = 7, P < 0.001; χ2 = 603.59, df = 3, P < 0.001), respectively. A similar trend was observed in leaf length in the endophytically-colonized, FOL-infected, endophytically-colonized and FOL-infected, and un-inoculated tomato plants at 15, 30, 45, 60 dpi (χ2 = 26.1, df = 7, P < 0.001; χ2 = 27.079, df = 7, P < 0.001; χ2 = 508.91, df = 7, P < 0.001; χ2 = 635.65, df = 7, P < 0.001), respectively (Fig. 5C). However, the leaf length did not differ at 15 and 30 dpi. Besides, significant differences (χ2 = 760.61, df = 7, P < 0.001; χ2 = 2404.78, df = 3, P < 0.001) in leaf length and all sampling periods were observed among the treatments, respectively.
Significant differences (χ2 = 52.809, df = 7, P < 0.001; χ2 = 43.178, df = 7, P < 0.001; χ2 = 1206.4, df = 7, P < 0.001; χ2 = 480.45, df = 7, P < 0.001) in plant height were observed among treatments at 15, 30, 45 and 60 dpi, respectively (Fig. 5D). The highest plant height (84.75 ± 1.38 cm) was recorded in H. lixii F3ST1-colonized tomato plants, while the lowest was recorded in FOL-infected plants (23.66 ± 2.57 cm). There were also significant differences in plant heights of endophytically-colonized, FOL-infected, endophytically colonized and FOL-infected, as well as un-inoculated tomato plants across all sampling periods (χ2 = 760.61, df = 7, P < 0.001; χ2 = 2404.78, df = 3, P < 0.001), respectively (Fig. 5D). Furthermore, the interaction of treatments and sampling periods was significantly different (P < 0.001) in all evaluated tomato growth parameters.
The results of plant biomass accumulation are provided in Table 4. There was a significant difference (χ2 = 603.54, df = 7, P < 0.001) in the dry weight of the roots of endophytically-colonized, FOL-infected, endophytically-colonized, FOL-infected, and un-inoculated tomato plants. Similarly, a significant difference (χ2 = 780.72, df = 7, P < 0.001) was observed in the dry weight of shoots across various treatments. The dry weight of the whole plant also differed significantly (χ2 = 179.01, df = 7, P < 0.001) among the treatments. Overall, tomato plants pre-treated with endophytes displayed an increased dry weight of roots and shoots. However, the dry weight of the whole plant was higher in H. lixii F3ST1 and T. asperellum M2RT4 pre-treated plants.
Effects of selected endophytes on FOL-infected tomato plants yield
The results of the effects of endophytes on the yield performance of the various treatments of tomato plants are shown in Table 5. There were significant differences in the number of fruits per plant, individual fruit weight, and yield (T ha−1) of endophytically colonized, FOL-infected, endophytically-colonized and FOL-infected, and un-inoculated tomato plants (χ2 = 173.42, df = 7, P < 0.001; χ2 = 550.87, df = 7, P < 0.001; χ2 = 619.88, df = 7, P < 0.001), respectively. Number of fruits per plant, individual fruit weight and yields were increased by 38.10%, 33.94%, and 59.02%, respectively, in FOL-infected tomato plants pre-treated with endophyte T. asperellum M2RT4 compared to control plants (Table 6). A similar trend was also observed in the evaluated yield parameters of FOL-infected tomato plants pre-treated with endophyte T. harzianum KF2R41, where percent increase in the number of fruits plants per plant, individual fruit weight, and yield were 25.19%, 20.09%, and 40.17%, respectively (Table 6). Furthermore, a percentage increase number of fruits per plant (26.77%), fruit weight (43.74%), and yield (58.74%) were recorded in FOL-infected tomato plants pre-treated with H. lixii F3ST1 (Table 6). On the other hand, Fusarium oxysporum lycopersici had a negative impact on all of the examined yield metrics in infected tomato plants by lowering the number of fruits per plant, individual fruit weight, and yield.
In general, tomato plants endophytically colonised by T. asperellum M2RT4 recorded the highest number of fruits, while the highest fruit weights were obtained in H. lixii F3ST1 colonized tomato plants. There were no significant differences in mean yield between T. asperellum M2RT4 and H. lixii F3ST1 endophytically-colonized and FOL-infected tomato plants and their positive controls in all the evaluated yield parameters. Furthermore, tomato yield did not differ considerably when T. harzianum KF2R41 endophytically-colonized and FOL-infected tomato plants were compared to non-inoculated tomato plants.
Defence gene expression in endophytically-colonized tomato plants and or plants infected by Fusarium oxysporum lycopersici
The expression level of the four selected tomato defence genes: TomPR1, TomPR2, TomLoxC, and SIWRKY4 in roots, stems and leaves of T. asperellum M2RT4 endophytically colonized, FOL-infected, T. asperellum M2RT4 endophytically-colonized and FOL- infected, as well as un-inoculated tomato plants are presented in (Fig. 6). Tomato GAPDH was used to normalise the gene expressions utilising the 2−ΔΔCT method45 and the selected genes did not have influence in the leaf treatments (df = 3, P = 0.3404), the stem treatments (df = 3, P = 0.1198) and the root treatments (df = 3, P = 0.08529). Consequently, there was an observed upregulation of the 4 selected defence genes (pathogenesis-related genes (TomPR1 and TomPR2), Loxigenase C (TomLoxC) and SWIRKY4) in the leaves (Fig. 6A), with the mean relative gene expression of SIWRKY4 gene (2.60 ± 0.20) being highest from the T. asperellum M2RT4 treatment, followed by TomPR2 gene (2.55 ± 0.23), TomPR1 (1.54 ± 0.22) and the least observed in TomLoxC gene (0.82 ± 0.12) in the control treatment.
The upregulation of the assessed defence genes was also observed in stems of T. asperellum M2RT4 endophytically colonized, FOL-infected, T. asperellum M2RT4 endophytically-colonized and FOL- infected, as well as un-inoculated tomato plants (Fig. 6B), with the mean relative gene expression of TomPR2 gene (4.65 ± 0.83) being highest from the Trichoderma asperellum M2RT4 treatment, followed by SIWRKY4 gene (4.10 ± 0.57) and the least observed in TomLoxC gene (1.08 ± 0.07) in the control treatment. In terms of treatments, the mean relative gene expression of the T. asperellum M2RT4 treatment was most elevated in the TomPR2 gene (4.65 ± 0.83) and SIWRKY4 gene (4.10 ± 0.57) compared to 2.35 ± 0.17 and 2.00 ± 0.17, respectively in the control treatment. In all four treatments, the mean relative gene expression of TomPR2 and SIWRKY4 seemed to be equal while was least in the TomLoxC gene (Fig. 6B). However, the upregulation observed was not statistically significant across all four treatments (df = 9, P = 1.0000).
A similar trend was observed in the root treatments (Fig. 6C) whereby the mean relative gene expression in the Trichoderma asperellum M2RT4 treatment was highest in TomPR2 gene (6.05 ± 0.95) followed by SIWRKY4 gene (5.69 ± 0.80) with least expression observed in TomLoxC gene (2.35 ± 0.29) as compared to 2.58 ± 0.68, 1.80 ± 0.58 and 1.08 ± 0.31, respectively from the control treatment. Although there was upregulation of the four selected genes in the roots, the variances were not statistically significant (df = 9, P = 0.99988) across all the treatments.
Discussion
The use of entomopathogenic (EPF) and endophytic fungi (EF) as a safe and effective biocontrol strategy to combat plant pests and diseases while enhancing crop performance has gained global attention13,16,26. Nevertheless, knowledge of the effectiveness of locally isolated EPF and EF to sustainably manage fusarium wilt in tomato plants is largely unknown16,41. Additionally, the mechanisms used by these biological control agents (BCAs) to improve tomato growth and yield while conferring resistance against fusarium wilt pathogen are not well explored37. In this study, the causal agent of fusarium wilt of tomato was isolated, identified and used as a test pathogen to assess the antagonistic effects of nine BCAs in Metarhizium, Beauveria, Hypocrea and Trichoderma genera under in vitro, as well as their in planta ability to control this disease and enhance tomato growth and yield. Furthermore, the mechanism used by T. asperellum M2RT4, the most potent isolate, to overcome the negative effects of FOL on infected tomato plants was evaluated using q-PCR.
Our morphological and molecular identification results revealed that the isolated pathogenic fungus was Fusarium oxysporum f. sp. lycopersici. This confirmation was based on physical observation of white colour of the colony and a purplish-pink pigmentation produced by this isolate on the PDA medium. In addition to the physical observations, the morphological features showed typical spore topologies of FOL as described by Leslie and Summerell44, in their Fusarium spp. identification key, as reported by Mwangi et al.46 and Nagaraj et al.47. Furthermore, the identity of the FOL isolate was confirmed by molecular markers where the similarity of the pathogen from this study was linked to FOL isolates deposited in GenBank at 98.64 to 100%. Pathogenicity test was also conducted to assess if the identified fungus is avirulent or virulent in tomato cultivar Moneymaker which is known to be susceptible to fusarium wilt. The results showed stunting and yellowing of lower leaves in FOL-inoculated tomato seedlings; confirming that the isolated FOL was virulent in the host plant. These observations concurred with previous research reports on symptoms caused by FOL in infected tomato plants16,48.
The in vitro antagonistic effects assessment revealed that all the nine entomopathogenic and endophytic fungal isolates in this study were able to inhibit and/or suppress FOL mycelia growth. The entomopathogenic fungal isolates showed a lower FOL mycelia inhibition rate (less than 50%) compared to fungal endophytes whose inhibition rate was almost 100%. This could be due to slow growth rate of EPF compared to the pathogenic fungus, which could also be attributed to the inoculation technique. Ragavendran et al.49 demonstrated that, when Beauveria spp. and Metarhizium spp. are sub-cultured using streaking and spreading techniques, their growth and spread over the whole media increased. However, in this study, the EPF isolates were placed on the side of the plate using the agar block technique, following the protocol of dual culture assay. Furthermore, our observations also established that, in addition to the entomopathogenic effects of the tested EPF on insect pests17, they could also inhibit FOL pathogen at some extent. According to Shah et al.50, and Kumar et al.51, fungi in the order “Hypocreales” which encompass Trichoderma spp. and Hypocrea lixii, grow quickly and form concentric rings, giving them a strong competitive ability.
Although all the tested BCAs were able to inhibit FOL growth, three endophytic fungal isolates, T. asperellum M2RT4, T. harzianum KF2R41, and H. lixii F3ST1 outperformed all the other isolates with high inhibition rates. In this study, these isolates grew fast and hence overgrew, sporulated on the FOL colony, and impeded its growth. These results are in agreement with previous studies which reported the inhibitive and suppressive ability of Hypocrea lixii and Trichoderma species against various soil-borne pathogens36,52,53,54,55. In vitro studies conducted by20,56, Trichoderma spp. showed the potential of inhibiting Sclerotium rolfsii and Fusarium oxysporum lycopersici mycelial growth at almost 100% and 76.94% inhibition rates, respectively. In a similar study, four Trichoderma strains of T. asperellum TaspHu1, T. harzianum TharHu2, T. hamatum ThamHu3, and T. atroviride TatrHu4 were reported to cause more than 80% Fusarium oxysporum radial growth inhibition36. The ability of H. lixii and Trichoderma species to inhibit the growth of pathogens or plant pathogens has been attributed to their fast growth and competition for nutrients and space, as well as the production of enzymes responsible for degrading fungal cell walls such as α-1-3-glucanase, β-1-3-glucanase, and chitinase13,26,52,57.
The endophytic colonization assessment confirmed the presence of the artificially inoculated endophytes in all the tomato plants' parts. Endophytes are capable of forming symbiotic relationships with inoculated host plants by penetrating and colonizing the plant tissues, and this needs to be proven through their re-isolation from inoculated hosts58. In our study, endophytes T. asperellum M2RT4, T. harzianum KF2R41, and H. lixii F3ST1 colonized the roots, stem and leaves of tomato plants. However, the colonization differed across the isolates in various plant parts. For instance, T. asperellum M2RT4 had the highest colonization rate in inoculated tomato plants compared to T. harzianum KF2R41 and H. lixii F3ST1. These results were in agreement with previous studies that reported endophytic colonization of tomato and nightshade plants by various Trichoderma strains and H. lixii differed across the plant's parts28,59. Similarly, among tomato parts assessed, roots and leaves had the highest and lowest colonization rates, respectively. The low rate of colonization of leaves could likely be attributed to vertical transmission movement of endophytes in Phylum Ascomycota where the genus Trichoderma belongs60, and since our inoculation was done through seeds of the host plant. According to Robert and Brown61, endophytes develop within plant tissues from roots to leaves by ascending migration. This could explain the observed phenomena of low colonization rate in leaves compared to other plant parts. Furthermore, considering this study was conducted under controlled laboratory and greenhouse conditions where the seeds and soil were sterilized, to further confirm the colonization of the selected isolates under field conditions, we recommend field validation of the colonization of the selected isolates in future studies.
Beyond host colonization, the potential of H. lixii and Trichoderma spp. includes pest and disease control, increased plant biomass, as well as improved plant growth and yield37,60,62,63,64. In this study, endophytes T. asperellum M2RT4, T. harzianum KF2R41, and H. lixii F3ST1 were found to be highly effective in lessening incidence and severity of fusarium wilt disease caused by Fusarium oxysporum lycopersici in-planta, increasing dry weight of roots and shoots, as well as boosting growth and yield of colonized tomato plants. Results from this study demonstrated that tomato plants inoculated with the three potent endophytes were able to defend themselves against FOL as compared to control plants. This finding was also confirmed by the low incidence, severity and AUDPC of fusarium wilt in endophytically-colonized and FOL-infected tomato plants in comparison to FOL-infected tomato plants. For instance, fusarium wilt incidence was reduced by 96%, 88%, and 56% in FOL-infected tomato plants pre-treated with the three endophytes T. asperellum M2RT4, H. lixii F3ST1, and T. harzianum KF2R41, respectively (Table 1). Similar trends were observed in T. asperellum M2RT4, H. lixii F3ST1, and T. harzianum KF2R41, where disease severity reduced by 97.92%, 66.67%, and 44.44%, respectively (Table 2) through systemic resistance development of the host crop, and the effects of potential antifungal metabolites produced by the endophytes as well as natural mycoparasitism. These results are in line with previous studies' findings which reported reduced incidence and severity of Fusarium spp. in various host plants such as maize, melons and tomatoes24,65,66. Besides, the lower AUDPC were obtained in FOL infected tomato plants pre-treated with the selected endophytes compared to FOL-infected tomato plants untreated with endophytes (Fig S3). This observation indicated the effectiveness of endophytes against fusarium wilt. According to Sinno et al.37 and Baron and Rigobelo60, efficacy of H. lixii and Trichoderma spp. in disease suppression is related to indirect triggering of systemic defence in their host plants, as well as the production and release of secondary metabolites substances. This could be achieved through a priming mechanism which enables endophytically-colonized plants to respond quickly and defend themselves against pathogens62,63,67.
Significant increase in tomato plant growth parameters, as well as increased yield in FOL-infected tomato plants pre-treated with the three endophytes, concurs with previous studies that reported increased plant biomass and growth parameters in response to their treatment with Trichoderma spp.28,36,62. This growth stimulation might be attributed to endophytes producing phytohormones such as auxins and gibberellins which are important for the development of roots and shoots as well as fruit settings and maturity as reported by36,60. The three endophytes, T. asperellum M2RT4, H. lixii F3ST1, and T. harzianum KF2R41 were able to enhance plant height in FOL-infected tomato plants, with H. lixii F3ST1 recording the highest plant height. These findings concurred with the previously reported results by28. Our findings also agreed with previous studies that reported improved crop yield under Trichoderma spp. application regardless of the presence of the pathogen62.
Observing how potent endophyte T. asperellum M2RT4 is in colonizing tomato plants, lessening fusarium wilt incidence and severity while boosting tomato growth and yield, this study attempted to decipher the mechanism by which this isolate confers these benefits. To achieve this, the change in the expression levels of the TomPR1, TomPR2, TomloxC and SIWRKY in roots, stems and leaves of T. asperellum M2RT4-endophytically colonized, FOL-infected, T. asperellum M2RT4 endophytically colonized and FOL-infected tomato plants were assessed as compared to non-inoculated tomato plants. Our study revealed that the levels of expression of TomPR1, and TomPR2 genes were upregulated in the roots, stems and leaves of tomato plants after their exposure to the endophyte T. asperellum M2RT4 when compared to FOL-infected and un-exposed tomato plants. These findings agree with previous research studies that found enhanced expression of pathogenesis-related proteins PR-1 (TomPR1) and β-1,3-glucanases (TomPR2) in Arabidopsis thaliana and tomato plants treated with Trichoderma spp. in responses to bacterial and fungal pathogens such as Fusarium oxysporum and Pythium aphanidermatum invasions52,68,69,70. According to Win et al.70 and Soliman et al.71, TomPR1 and TomPR2 are responsible for defending host plant against fungal invasion by lysis of the pathogen cell wall thereby suppressing pathogenic fungal growth. In this study, we also observed a significant increase in expression level of TomLoxC gene in the roots, stems and leaves of T. asperellum M2RT4 endophytically-colonized tomato plants, as well as in T. asperellum M2RT4 endophytically-colonized and FOL-infected tomato plants compared to FOL-infected and un-inoculated tomato plants. Although previous studies only identified TomloxC expression in tomato fruits, our findings revealed that TomloxC activity began during the crop’s vegetative phase30,72,73. Furthermore, our results also showed that T. asperellum M2RT4 induced significant upregulation of the SIWRKY4 gene in both roots, stems and leaves of colonized tomato plants and in FOL-infected plants.
Conclusion
Our findings proved that the two endophytes H. lixii F3ST1 and T. asperellum M2RT4 could successfully control Fusarium oxysporum lycopersici while boosting the yield of tomato crop. Thus, these endophytes could be formulated into biopesticides to sustainably control FOL pathogen and consequently reduce fusarium wilt disease incidence and severity in tomato cropping systems. Furthermore, the endophyte T. asperellum M2RT4 triggered the expression of Pathogenesis Related Protein-1 (TomPR1), β-1,3-glucanases (TomPR2), TomloxC, and SIWRKY4 genes, whose main function is to protect host plants against fungal invasion for improved yield quantity and quality31,70,71. The upregulation of the studied genes was negatively correlated with reduced incidence and severity of fusarium wilt in T. asperellum M2RT4 endophytically-colonized tomato plants. Therefore, the higher expression of the assessed tomato defence genes and reduced severity of FOL disease in T. asperellum M2RT4 colonized than in un-inoculated tomato plants, indicate that the endophyte T. asperellum M2RT4 is able to confer systemic resistance in tomato plants against Fusarium oxysporum lycopersici. Further studies should also be warranted in exploring the host plant detection and reaction mechanisms to the beneficial fungi based on microbe-associated molecular patterns as previously demonstrated by Bittel and Robatzek74 and Boller and Felix75.
Material and methods
Study site
In vitro and greenhouse experiments were conducted at the International Centre of Insect Physiology and Ecology (icipe), Duduville campus, Nairobi, Kenya (S 01° 13′ 14.6″; E 036° 53′ 44.5″), 1612 m above sea level. For the bioprospecting study of the plant pathogen, roots and stem samples with brown discolouration of the vessels in their cross-sections from tomato plants with yellowing of the lower leaves, typically confined to one side of the host plant were collected. The samples were collected in Mwea—Kirinyaga county, Kenya (S-0.628516 E37.374851, 1163 m) for Fusarium oxysporum lycopersici (FOL) isolation.
Test pathogen isolation and identification
The field-collected samples were cut into small pieces of 1 cm for roots and stems targeting disease leading edge which is mainly characterized by high pathogen activity76. Samples were surface sterilized by submerging them in 70% ethanol for 1 min and then 1.5% sodium hypochlorite for 2 min. They were then rinsed three times for one minute with distilled water to wash out the sodium hypochlorite and blot-dried on a paper towel under a laminar flow hood. Afterwards, using sterile forceps, each piece of the blot-dried sample was placed into Petri dishes containing Potato Dextrose Agar (PDA) medium supplemented with 0.1% chloramphenicol antibiotic. The Petri dishes were then sealed with parafilm and incubated at 25 ± 2 °C for 7 days. Fungal colonization was assessed in each isolated plant tissue and actively growing fungi were isolated and sub-cultured onto new PDA plates for purification. Several subcultures were conducted to acquire pure fungal isolates before their identification. The pure cultures were obtained by transferring the mycelia to Petri dishes containing PDA medium and incubated as mentioned above.
To confirm whether the isolated fungus was FOL, macroscopic, microscopic, and molecular identification were conducted. Macroscopically, FOL was identified through physical observation of the colony morphology and pigmentation of the 7-day pure culture plates of the isolates, and by morphological key described by Leslie and Summerell44, as reference. The same morphological key was used to identify the isolates microscopically based on the shape and size of the spores displayed with the ZEISS Primo Star microscope at 40× magnification. For molecular identification, total DNA was extracted from pure cultures of the fungi using the Isolate II Plant DNA Extraction Kit (Bioline, London, UK) as per the manufacturer’s instructions. The extracted DNA quality and quantity were assessed using a Nanodrop 2000/2000c spectrophotometer (Thermo Fischer Scientific, Wilmington, USA), and stored at − 20 °C, until further processing. The intergenic transcribed spacer region ITS 5 and ITS 4 primers, (5ʹ-TCCTCCGCTTATTGATATGC-3ʹ) and (3ʹ-GGAAGTAAAAGTCGTAACAAGG-5ʹ), elongation factor 1-α EF1-983 (5ʹ-GCY CCY GGH CAY CGT GAY TTY AT-3ʹ) and EF1-2218 (3ʹ-AT GAC ACC RAC RGC RAC RGT YTG-5ʹ), largest subunit of RNA polymerase RPB1-A (5ʹ-GARTGYCCDGGDCAYTTYGG-3ʹ) and RPB1-C (3ʹ-CCNGCDATNTCRTTRTCCATRTA-5ʹ) were used for amplification of the target regions. The samples were sent to Microgen for sequencing and the obtained sequences were edited and assembled using BioEdit Sequence Alignment Editor Version 7.2.577. The sequences were then matched with those found in GenBank of the National Center for Biotechnology Information (NCBI) through the Basic Local Alignment Search Tool (BLAST)78,79. The sample with a similarity index of 99–100% was considered as the positive FOL.
Preparation of biocontrol fungi
Both the endophytic and entomopathogenic fungal isolates were obtained from icipe’s Arthropod Pathology Unit (APU) Germplasm for the bioassays. The endophytic fungal isolates Hypocrea lixii F3ST1, Trichoderma asperellum M2RT4, T. atroviride ICIPE 710, and T. harzianum KF2R41 which were already reported as endophytes in tomato, nightshade and beans were used17,28,55. The entomopathogenic fungal isolates that were used were: Beauveria bassiana isolates (ICIPE 706 and ICIPE 273) and Metarhizium anisopliae isolates (ICIPE 18, ICIPE 20 and ICIPE 665) that have been previously reported to be effective against the tomato leafminer, Phthorimaea absoluta formerly known as Tuta absoluta59,80,81. The endophytic and EP fungal isolates have been selected based on their ability to colonize all tomato plant parts, control plant pests (T. absoluta (Meyrick), whiteflies, (Trialeurodes vaporariorum), and enhance crop performance, as reported by28,59,81.
These above isolates were subcultured on a PDA medium and incubated at 25 ± 2 °C for further use, except for Metarhizium anisopliae isolates which were maintained on Sabouraud dextrose agar (SDA) medium in complete darkness. The inoculum preparations for all the fungal isolates were done by harvesting conidia by scraping the surface of 2-week-old pure fungal cultures that were sporulating using a sterile spatula. The harvested conidia were mixed in 10 mL of sterile distilled water containing 0.05% (w/v) Triton X-100 (EMD Millipore Corporation, USA) in a universal bottle and vortexed for 5 min at about 700 rpm to produce homogenous conidial suspensions. Conidial counts were performed with an improved Neubauer Hemocytometer82. The inocula were adjusted to a concentration of 1 × 108 conidia mL−1.
Before any bioassay, the spore viability test was performed by spreading 0.1 mL of 3 × 106 conidial mL−1 onto 9 cm Petri dishes containing SDA or PDA. Plates were sealed with parafilm and incubated at 25 ± 2 °C in complete darkness for 15–18 h, depending on the fungal isolate as some germinate faster than others59. The isolates were set in quadruplets and arranged in a completely randomized design (CRD). After 15–18 h, the plates were unsealed, fixed with a drop of fixative lactophenol cotton blue, and randomly covered with four sterile microscope coverslips (2 × 2 cm)59. Percentage conidial germination was determined from 100 randomly selected conidia on the surface covered by each coverslip under a light microscope (40×)82. A conidium was considered to have germinated when the length of the germ tube was at least twice its diameter83.
In vitro assessment of fungal biocontrol agents against Fusarium oxysporum lycopersici
Using the dual culture assay, both EPF and endophytic fungal isolates were screened for their antagonistic activity against the FOL pathogen. Zero-point five centimeter (0.5 cm) of the mycelial disc from a 5-day-old culture of FOL was placed at 1 cm from the end of the 9 cm Petri dish with PDA. A similar disc of EPF or endophytic fungal isolates was placed opposite the mycelial disc of FOL. Plates solely inoculated with FOL served as control treatment. Each treatment was replicated four times, sealed and incubated in an inverted position at 25 ± 2 °C. The radius of mycelial growth of FOL, in both dual culture and in control, was measured with a ruler every 24 h until a pathogen completely covered the surface of the control plate. Percentage inhibition of radial growth was obtained using the formula \(\left[\left(\text{R}1-\text{R}2\right)\times 100/\text{R}1\right]\) as described by Myrchiang and Devi84, where R1 is the radius of radial growth of the pathogen towards the opposite side of the control plate, and R2 is the radius of radial growth of the pathogen towards the antagonist in dual culture.
To further screen for the antagonistic ability of the beneficial fungal isolates against FOL, the co-culture assay was also performed following the modified protocol described by Karuppiah et al.85. Zero-point one milliliter (0.1 mL) of 1 × 108 conidia mL−1 of FOL and fungal biocontrol agent (BCA) were spread together on a Petri dish with PDA. Single cultures of FOL served as control treatment. Plates were sealed with parafilm and incubated at 25 ± 2 °C for 14 days. Each Petri dish from the treatments and control was replicated four times. The experiment was set in a completely randomized design. Appearance, dominance of mycelia, and pigmentation produced by FOL/BCAs in single-cultured and dual-cultured plates were recorded by visual observation.
Suspension preparation and inoculation of the tomato seeds with the selected potent biocontrol agents
The tomato cultivar “Money-maker” seeds (SimLaw Seeds Ltd.) were bought in Agroduka, Nairobi, Kenya. The seeds were surface sterilized by soaking them in 70% ethanol for 2 min, followed by a 1.5% solution of sodium hypochlorite for 3 min with continuous shaking and rinsed three times in sterile distilled water for 1 min28. Seeds were dried at room temperature on a sterile paper towel for 3 h in a laminar flow hood86. The effectiveness of the surface sterilization procedure was done by plating the last rinse water using a sterile loop on a Petri dish containing PDA medium and also imprinting of surface sterilized seeds onto PDA (tissue imprint) supplemented with 100 mg/L Streptomycin and plates were incubated at 25 °C for 14 days19,24,55,86,87. Then, the absence of fungal growth after incubation indicated the successfulness of the sterilization procedure28. The seed inoculation was done by placing the surface-sterilized tomato seeds into the suspension of the potent endophytic fungal isolates at a concentration of 1 × 108 conidia mL−1 for 18 h before sowing them28. One portion of the seeds was left un-inoculated with antagonistic fungal isolates to serve as a control.
Endophytic colonization and pathogenicity assessment
The endophytes T. asperellum M2RT4, H. lixii F3ST1 and T. harzianum KF2R4 which exhibited more than 80% FOL mycelial growth inhibition during in vitro study, were selected for in-planta experiments. Both the inoculated and un-inoculated seeds were sown in pots of 8 cm diameter and 7 cm height filled with 1 kg of the sterile mixture ratio of 5:1 field soil and compost which were autoclaved at 121 °C for 2 h and left to cool for 72 h before their use. Five seeds were sown per pot and thinned to two after 2 weeks. The seedlings were maintained at room temperature in a greenhouse under natural light conditions with no additional fertilizer28,59,80. The seedlings were watered regularly once in 2 days, using sterile distilled water to make sure that nothing else would affect the experiment, except the planned treatments.
To confirm the presence of endophytes in tomato tissues, five randomly selected 3-week-old tomato seedlings were uprooted from their pots and washed with tap water to remove the soil. Plants were divided into three parts (root, stem, and leaves), cut into 1 cm root and stem pieces and 1 cm2 leaf pieces, and surface sterilized in a laminar flow hood as described by Paradza et al.28. Five pieces were placed on a 9 cm diameter Petri dish containing PDA enriched with antibiotic (25 μg/ml chloramphenicol) and incubated at 25 ± 2 °C for 2 to 3 weeks28. The number of plant pieces showing fungal outgrowth divided by the total number of plant pieces plated was used to compute the proportion of the plant parts colonized by the inoculated fungal isolate for each treatment28. The assessment was based on the morphological characteristics of the inoculated fungus that colonized the incubated plant part, and only the colonization by the inoculated fungus was recorded as positive28.
To confirm the virulence of isolated FOL, each pot with 4-week-old tomato seedlings was soil-drenched with 10 mL of FOL inoculum at a concentration of 1 × 108 conidia mL−1. The control treatment pots were soil-drenched with a solution of Triton X-100 and sterile distilled water without FOL. Pots were kept in the greenhouse for further inspection of the occurrence and severity of fusarium wilt disease symptoms as described by Seo and Kim88 and Andrade-Hoyos et al.55, respectively. Afterwards, FOL was re-isolated from diseased tissues to complete Koch’s postulates1.
In planta evaluation of the selected potent antagonists against Fusarium wilt
Both inoculated and un-inoculated tomato seeds were sown in pots of 16.5 cm base diameter, 18 cm tall, and 22.5 cm top diameter (5-L pot) filled with a sterile mixture of topsoil and well-decomposed goat manure at a ratio of 5:1. The planting media were previously soil-drenched with 10 mL of FOL inoculum at a concentration of 1 × 108 conidia mL−1 on the same day. The experiment consisted of eight treatments with endophytically colonized tomato seedlings and FOL inoculum as follows: (i) T. asperellum M2RT4 and FOL; (ii) T. harzianum KF2R41 and FOL; (iii) H. lixii F3ST1 and FOL; (iv) T. asperellum M2RT4 alone; (v) T. harzianum KF2R41 alone; (vi) H. lixii F3ST1 alone; (vii) endophyte-free seedlings (negative controls) and with FOL; and (viii) endophyte-free seedlings alone without FOL (positive control). Each treatment consisted of a 5 L pot with two seedlings in each pot. A completely randomized design was used with three replications over time. The seedlings were kept inside a greenhouse and maintained as described above.
Fusarium wilt disease assessment
Fusarium wilt disease incidence and severity data were collected at 15-, 30-, 45-, and 60-days post inoculation (dpi) on FOL-inoculated tomato plant in reference to leaf wilting and vascular discolouration. The incidence was determined using the following formula:
\(\text{Disease incidence }\left(\text{\%}\right)\text{ = }\left(\text{Number of infected plants}/\text{Total number of inoculated sampled plants}\right)\times {{100}}\) adopted from55, while the severity was assessed using a 0–4 ranking scale, as described by Amini89 and Horinouchi et al.90.
The Area Under Disease Progress Curve (AUPDC) was determined as explained by Batista91 using a formula:
where: Yi = Severity of fusarium wilt at the ith observation, Ti = time (days after inoculation) at the ith observation and n = total number of evaluations.
Plant growth and yield parameters
Tomato growth parameters such as plant height, number of completely developed leaves, leaf width, and leaf length were assessed as defined by92. The dry weights of roots and shoots were obtained by placing them in an oven at 60 °C for 48 h and then weighed using Endeavour electronic weighing balance with a precision of ± 0.001 g93.
Three yield parameters such as number of fruits, fruit weight, and yield were collected in the manner outlined by94, where the total number of fruits per plant and the individual fruit weight were obtained by counting and weighing each marketable fruit. The yield (kg plant−1) was obtained by totalling the weight of all fruits harvested from each plant. The yield estimation in tons per ha was calculated using the below formula described by Ali et al.95:
Tomato defence gene expression
Based on the in planta experiment results above, the endophyte T. asperellum M2RT4, which displayed a high colonization rate, with complete suppression/inhibition of fusarium wilt pathogen, disease incidence and severity was selected and assessed for its ability to trigger the defence gene in the endophytically-colonized tomato crop. Five-liter pots were filled with a sterile mixture of topsoil and goat manure at a 5:1 ratio. The pots were soil-drenched with 10 mL of FOL inoculum at a concentration of 1 × 108 conidia mL−1. Afterwards, Trichoderma asperellum M2RT4 inoculated tomato seeds were sown into the planting medium on the same day. The experiment consisted of four treatments: (i) FOL-infected tomato plants, (ii) T. asperellum-endophytically colonized tomato plants, (iii) T. asperellum M2RT4 endophytically-colonized and FOL-infected tomato plants, and (iv) un-inoculated tomato plants (control). Each treatment had six replicates and was organized in a completely randomized design.
One (1) g of roots, stems, and leaves of each treatment sample were cut and placed into 2 mL Eppendorf tubes and stored at − 80 °C for RNA extraction. RNA extraction was done using the ISOLATE II RNA Mini Kit, (Bioline, London, UK) following the manufacturer’s guidelines. The RNA samples were reverse-transcribed into cDNA using SensiFAST cDNA Synthesis Kit (Meridian Bioscience, London, UK) following the manufacturer’s guidelines. The target primers were designed using primer 3.0 software, available at NCBI (https://www.ncbi.nlm.nih.gov/tools/primer-blast/) (Table 7). The qPCR reaction was performed using QuantStudio 5 Real-Time PCR system (Thermo Fisher, Scientific, Waltham, USA). PCR mixture constituted of PCR water (1.5 μL), SensiFAST SYBR® Hi-ROX Kit (5 μL), Forward primer (0.5 μL) and Reverse primer (0.5 μL) and RNA template (2.5 μL). Each qPCR reaction consisted of both positive (known sample) and negative (non-template control) samples. GAPDH1 was used as a housekeeping gene to normalize gene expression. Relative expression of tomato defence-related genes was analyzed using the delta-delta Ct method (2−ΔΔCt)45.
Data analyses
Before data analysis, the normality test was conducted to confirm if the data were normally distributed using the Shapiro–Wilk test96. Data on colonization rate, % inhibition of radial growth, and incidence and severity of fusarium wilt disease were analyzed using the generalized linear model (GLM) package of R statistical software97, assuming a binomial distribution with the log link function. The number of leaves and fruits per plant data were analyzed using GLM quasi-Poisson. Data on leaf width and length, plant height, fresh and dry weight of roots and shoots were analyzed using analysis of variance. The relative expression of tomato defence genes was analyzed using the delta-delta Ct method (2−ΔΔCt)45. All statistical analyses were done using R version 4.3.197. The treatment means were separated using the Student Newman Keuls (SNK) test. A p-value of less than 0.05 was considered for statistical significance.
Ethics approval
The experimental research and field studies on plants, including the collection of plant material, complied with relevant institutional, national, and international guidelines and legislation. The appropriate permissions and/or licenses for collection of plant or seed specimens were obtained for the study as approved by the National Commission of Science, Technology and Innovations, Kenya (License No: NACOSTI/P/23/23608). This article does not contain any studies with human participants performed by any of the authors.
Data availability
All relevant data are within the paper and supplementary materials.
References
Ahmed, A. A. & Reddy, G. H. A mobile-based system for detecting plant leaf diseases using deep learning. AgriEngineering 3, 478–493 (2021).
Cameron, K. H., Somachandra, K. P., Curry, C. N., Jenner, W. H. & Hobbs, S. L. A. Delivering actionable plant health knowledge to smallholder farmers through the Plantwise program. J. Agric. Food Inf. 17, 212–229 (2016).
Mahapatra, S. et al. Plant disease forecasting in the era of climate change : Trends and applications. 1–26 (2018) https://www.researchgate.net/publication/323725918.
Fenu, G. & Malloci, F. M. Review forecasting plant and crop disease: An explorative study on current algorithms. Big Data Cogn. Comput. 5, 1–24 (2021).
Sundararaman, B., Jagdev, S. & Khatri, N. Transformative role of artificial intelligence in advancing sustainable tomato (Solanum lycopersicum) disease management for global food security: A comprehensive review. Sustainability 15, 11681 (2023).
Ochilo, W. N. Characteristics and production constraints of smallholder tomato production in Kenya. Sci. Afr. 2, e00014 (2019).
Mwangi, M. W., Muiru, W. M. & Kimenju, J. W. Characterisation of Fusarium species infecting tomato in Mwea West Sub-county, Kirinyaga County, Kenya. Can. J. Plant Pathol 43, 1–6 (2020).
Waceke, W. J., Wafula, G. O. & Macharia, C. M. Integrated management of fusarium wilt-root knot nematode complex on tomato in Central Highlands of Kenya. Sustain. Agric. Res 7, 8–18 (2018).
Karuri, H. Nematode community response to intensive tomato production in the tropics, Rhizosphere. Rhizosphere 25, 100681 (2023).
Thomas, M. M. Technical efficiency in tomato production among smallholder farmers in Kirinyaga County, Kenya. Afr. J. Agric. Res 16, 667–677 (2020).
Ochieng, V. Tomato production manual. Kenya Agricultural and Livestock Institute (KALRO) Preprint at (2016).
Singh, V. K., Singh, A. K. & Kumar, A. Disease management of tomato through PGPB: Current trends and future perspective. 3 Biotech 7, 1–10 (2017).
Ma, M., Taylor, P. W. J., Chen, D., Vaghefi, N. & He, J. Z. Major soilborne pathogens of field processing tomatoes and management strategies. Microorganisms 11, 263 (2023).
Dita, M., Barquero, M., Heck, D., Mizubuti, E. S. G. & Staver, C. P. Fusarium wilt of banana: Current knowledge on epidemiology and research needs toward sustainable disease management. Front. Plant Sci. https://doi.org/10.3389/fpls.2018.01468 (2018).
Srinivas, C. et al. Fusarium oxysporum f. sp. lycopersici causal agent of vascular wilt disease of tomato: Biology to diversity—A review. Saudi J. Biol. Sci. 26, 1315–1324 (2019).
Maurya, S., Dubey, S., Kumari, R. & Verma, R. Management tactics for fusarium wilt of tomato caused by Fusarium oxysporum f. sp. lycopersici (Sacc.): A review. Int. J. Res. Pharm. Pharm. Sci. 4, 1–7 (2019).
Ajilogba, C. F. & Babalola, O. O. Integrated management strategies for tomato fusarium wilt. Biocontrol Sci. 18, 117–127 (2013).
Ivanov, A. A., Ukladov, E. O. & Golubeva, T. S. Phytophthora infestans: An overview of methods and attempts to combat late blight. J. Fungi 7, 1071 (2021).
de Lamo, F. J. & Takken, F. L. W. Biocontrol by Fusarium oxysporum using endophyte-mediated resistance. Front. Plant Sci. 11, 1–15 (2020).
Bastakoti, S., Belbase, S., Manandhar, S. & Arjyal, C. Trichoderma species as biocontrol agent against soil borne fungal pathogens identification of Trichoderma isolates. Nepal J. Biotechnol. 5, 39–45 (2017).
Akutse, K. S., Subramanian, S., Maniania, N. K., Dubois, T. & Ekesi, S. Biopesticide research and product development in Africa for sustainable agriculture and food security—experiences from the International Centre of Insect Physiology and Ecology (icipe). Front. Sustain. Food Syst. 4, 1–14 (2020).
Majumder, D., Deb, L. & Malakar, P. Entomopathogenic fungi: Potential tool for biological control of plant diseases. ResearchGate 549–580 (2021).
Agbessenou, A., Akutse, K. S., Yusuf, A. A. & Khamis, F. M. The endophyte Trichoderma asperellum M2RT4 induces the systemic release of methyl salicylate and (Z)-jasmone in tomato plant affecting host location and herbivory of Tuta absoluta. Front. Plant Sci. 13, 1–16 (2022).
El Komy, M. H., Saleh, A. A., Ibrahim, Y. E., Hamad, Y. K. & Molan, Y. Y. Trichoderma asperellum strains confer tomato protection and induce its defense-related genes against the Fusarium wilt pathogen. Trop. Plant Pathol. 41, 277–287 (2016).
Halifu, S., Deng, X., Song, X. & Song, R. Effects of two Trichoderma strains on plant growth, rhizosphere soil nutrients, and fungal community of Pinus sylvestris var. mongolica annual seedlings. Forests 10, 1–17 (2019).
Sinno, M. et al. Selection of endophytic Beauveria bassiana as a dual biocontrol agent of tomato pathogens and pests. Pathogens 10, 1242 (2021).
de Vries, S. et al. Broad-spectrum inhibition of Phytophthora infestans by fungal endophytes. FEMS Microbiol. Ecol. 94, 1–15 (2018).
Paradza, V. M., Khamis, F. M., Yusuf, A. A. & Subramanian, S. Endophytic colonisation of Solanum lycopersicum and Phaseolus vulgaris by fungal endophytes promotes seedlings growth and hampers the reproductive traits, development, and survival of the greenhouse whitefly, Trialeurodes vaporariorum. Front. Microbiol. 12, 1–17 (2021).
Alfano, G. et al. Systemic modulation of gene expression in tomato by Trichoderma hamatum 382. Phytopathology 97, 429–437 (2007).
Upadhyay, R. K. & Autar, M. K. Genome-wide identification of tomato (Solanum lycopersicum L.) lipoxygenases coupled with expression profiles during plant development and in response to methyl-jasmonate and wounding. J. Plant Physiol. 231, 1–45 (2018).
Aamir, M. et al. Trichoderma erinaceum bio-priming modulates the WRKYs defense programming in tomato against the Fusarium oxysporum f. sp. lycopersici (Fol) challenged condition. Front. Plant Sci. 10, 911 (2019).
Morán-Diez, M. E., Tranque, E., Bettiol, W., Monte, E. & Hermosa, R. Differential response of tomato plants to the application of three Trichoderma species when evaluating the control of Pseudomonas syringae populations. Plants 9, 626 (2020).
Risoli, S. et al. Trichoderma-induced resistance to Botrytis cinerea in Solanum species: A meta-analysis. Plants 11, 180 (2022).
Prasetyawan, S., Sulistyowati, L. & Aulanni’Am. Glucanase and chitinase from some isolates of endophytic fungus Trichoderma spp. In IOP Conference Series: Materials Science and Engineering Vol. 299, (2018).
Ueki, A., Takehara, T., Ishioka, G., Kaku, N. & Ueki, K. β-1,3-Glucanase production as an anti-fungal enzyme by phylogenetically different strains of the genus Clostridium isolated from anoxic soil that underwent biological disinfestation. Appl. Microbiol. Biotechnol. 104, 5563–5578 (2020).
Yu, Z., Wang, Z., Zhang, Y., Wang, Y. & Liu, Z. Biocontrol and growth-promoting effect of Trichoderma asperellum TaspHu1 isolate from Juglans mandshurica rhizosphere soil. Microbiol. Res. 242, 126596 (2021).
Sinno, M., Ranesi, M., Gioia, L., D’errico, G. & Woo, S. L. Endophytic fungi of tomato and their potential applications for crop improvement. Agriculture 10, 1–22 (2020).
Strobel, G. Special issue: Fungal endophytes in plants. J. Fungi 4, 104 (2018).
Mwangi, M. W., Muiru, W. M., Narla, R. D., Kimenju, J. W. & Kariuki, G. M. Management of Fusarium oxysporum f. sp. lycopersici and root-knot nematode disease complex in tomato by use of antagonistic fungi, plant resistance and neem. Biocontrol Sci. Technol. 29, 207–216 (2019).
Dara, S. K. Non-entomopathogenic roles of entomopathogenic fungi in promoting plant health and growth. Insects 10, 277 (2019).
Wamani, A. O. Screening of Microbial Antagonists and Plant Extracts against Selected Tomato Pathogens and Their Potential in the Management of Bacterial Wilt (University of Nairobi repository, 2020).
Lu, H., Wei, T., Lou, H., Shu, X. & Chen, Q. A critical review on communication mechanism within plant-endophytic fungi interactions to cope with biotic and abiotic stresses. J. Fungi 7, 719 (2021).
Karim, H., Azis, A. A. & Jumadi, O. Antagonistic activity and characterization of indigenous soil isolates of bacteria and fungi against onion wilt incited by Fusarium sp. Arch. Microbiol. https://doi.org/10.1007/s00203-021-02663-2 (2022).
Leslie, J. F. & Summerell, B. A. The Fusarium Laboratory Manual (Blackwell Publishing Ltd, 2006).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25, 402–408 (2001).
Mwangi, M. T., Njiri Ndirangu, S. & Isaboke, N. H. Technical efficiency in tomato production among smallholder farmers in Kirinyaga County, Kenya. Afr. J. Agric. Res. 16, 667–677 (2020).
Nagaraj, G. et al. Morphological and molecular characterization of Fusarium spp. infecting tomato in Tamil Nadu. Source: Mycologia 21, 597–609 (2022).
Mwangi, M. W., Muiru, W. M. & Kimenju, J. W. Characterisation of Fusarium species infecting tomato in Mwea West Sub-county, Kirinyaga County, Kenya. Can. J. Plant Pathol. 43, 1–6 (2020).
Ragavendran, C., Dubey, N. K. & Natarajan, D. Beauveria bassiana (Clavicipitaceae): A potent fungal agent for controlling mosquito vectors of Anopheles stephensi, Culex quinquefasciatus and Aedes aegypti (Diptera: Culicidae). RSC Adv. 7, 3838–3851 (2017).
Shah, S., Nasreen, S. & Sheikh, P. A. Cultural and morphological characterization of Trichoderma spp. associated with green mold disease of Pleurotus spp. in Kashmir. Res. J. Microbiol. 7, 139–144 (2012).
Kumar, G. et al. Morphomolecular identification of Trichoderma sp. and their mycoparasitic activity against soil borne pathogens. Int. J. Bio-resour. Stress Manag. 11, 613–627 (2020).
El-Komy, M. H., Saleh, A. A., Eranthodi, A. & Molan, Y. Y. Characterization of novel Trichoderma asperellum isolates to select effective biocontrol agents against tomato fusarium wilt. Plant Pathol. J. 31, 50–60 (2015).
Kariuki, W. G. et al. Antagonistic effects of biocontrol agents against Phytophthora infestans and growth stimulation in tomatoes. Angew. Chem. Int. Ed. 6(11), 951–95 28, 55–70 (2020).
Hewedy, O. A. et al. Phylogenetic diversity of Trichoderma strains and their antagonistic potential against soil-borne pathogens under stress conditions. Biology 9, 1–21 (2020).
Andrade-Hoyos, P., Silva-Rojas, H. V. & Romero-Arenas, O. Endophytic Trichoderma species isolated from Persea americana and Cinnamomum verum roots reduce symptoms caused by Phytophthora cinnamomi in avocado. Plants 9, 1–17 (2020).
Ayele, T. M., Gebremariam, G. D. & Patharajan, S. Isolation, identification and in vitro test for the biocontrol potential of Trichoderma viride on Fusarium oxysporum f. sp. lycopersici. Open Agric. J. 15, 10–20 (2021).
Alamri, S., Hashem, M. & Mostafa, Y. S. In vitro and in vivo biocontrol of soil-borne phytopathogenic fungi by certain bioagents and their possible mode of action. Biocontrol Sci. 17, 155–167 (2012).
Sood, M. et al. Trichoderma: The “secrets” of a multitalented biocontrol agent. Plants 9, 1–25 (2020).
Agbessenou, A., Akutse, K. S., Yusuf, A. A. & Ekesi, S. Endophytic fungi protect tomato and nightshade plants against Tuta absoluta (Lepidoptera : Gelechiidae ) through a hidden friendship and cryptic battle. Sci. Rep. 10, 1–13 (2020).
Baron, N. C. & Rigobelo, E. C. Endophytic fungi: A tool for plant growth promotion and sustainable agriculture. Mycology 13, 39–55 (2022).
Robert, B. & Brown, E. B. Ascending endophytic migration of locally isolated diazotroph Enterobacter sp. strain USML2in rice. Biotechnologhy 10, 521–527 (2011).
Tucci, M., Ruocco, M., De Masi, L., De Palma, M. & Lorito, M. The beneficial effect of Trichoderma spp. on tomato is modulated by the plant genotype. Mol. Plant Pathol. 12, 341–354 (2011).
Hermosa, R., Viterbo, A., Chet, I. & Monte, E. Plant-beneficial effects of Trichoderma and of its genes. Microbiology 158, 17–25 (2012).
Bamisile, B. S., Dash, C. K., Akutse, K. S., Keppanan, R. & Wang, L. Fungal endophytes: Beyond herbivore management. Front. Microbiol. 9, 1–11 (2018).
Idowu, O., Olawole, O., Idumu, O. & Salami, A. Bio-control effect of Trichoderma asperellum (Samuels) Lieckf. and Glomus intraradices Schenk on Okra seedlings infected with Pythium aphanidermatum (Edson) Fitzp and Erwinia carotovora (Jones). Am. J. Exp. Agric. 10, 1–12 (2016).
Tiru, Z., Sarkar, M., Pal, A., Chakraborty, A. P. & Mandal, P. Three dimensional plant growth promoting activity of Trichoderma asperellum in maize (Zea mays L.) against Fusarium moniliforme. Arch. Phytopathol. Plant Prot. 54, 764–781 (2021).
Aranega-Bou, P., de la O Leyva, M., Finiti, I., Garcfa-Agustfn, P. & Gonzalez-Bosch, C. Priming of plant resistance by natural compounds. Hexanoic acid as a model. Front. Plant Sci. 5, 1–12 (2014).
Contreras-Cornejo, H. A., Macías-Rodríguez, L., Beltrán-Peña, E., Herrera-Estrella, A. & López-Bucio, J. Trichoderma-induced plant immunity likely involves both hormonal- and camalexin dependent mechanisms in Arabidopsis thaliana and confers resistance against necrotrophic fungus Botrytis cinerea. Plant Signal. Behav. 6, 1554–1563 (2011).
Akbudak, M. A., Yildiz, S. & Filiz, E. Pathogenesis related protein-1 (PR-1) genes in tomato (Solanum lycopersicum L.): Bioinformatics analyses and expression profiles in response to drought stress. Genomics 112, 4089–4099 (2020).
Win, T. T., Bo, B., Malec, P., Khan, S. & Fu, P. Newly isolated strain of Trichoderma asperellum from disease suppressive soil is a potential bio-control agent to suppress Fusarium soil borne fungal phytopathogens. J. Plant Pathol. 103, 549–561 (2021).
Soliman, S. A. et al. Differences in pathogenesis-related protein expression and polyphenolic compound accumulation reveal insights into Tomato-Pythium aphanidermatum interaction. Sustainability 15, 1–16 (2023).
Yan, L. et al. Role of tomato lipoxygenase D in wound-induced jasmonate biosynthesis and plant immunity to insect herbivores. PLoS Genet. 9, e1003964 (2013).
Hernández-Aparicio, F., Lisón, P., Rodrigo, I., Bellés, J. M. & López-Gresa, M. P. Signaling in the tomato immunity against Fusarium oxysporum. Molecules 26, 1818 (2021).
Bittel, P. & Robatzek, S. Microbe-associated molecular patterns (MAMPs) probe plant immunity. Curr. Opin. Plant Biol. 10, 335–341 (2007).
Boller, T. & Felix, G. A renaissance of elicitors: Perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu. Rev. Plant Biol. 60, 379–406 (2009).
Mugao, L. G., Muturi, P. W., Gichimu, B. M. & Njoroge, E. K. In vitro control of Phytophthora infestans and Alternaria solani using crude extracts and essential oils from selected plants. Int. J. Agron. 2020, 1–10 (2020).
Hall, T. A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98 (1999).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Mount, D. W. Using the basic local alignment search tool (BLAST). Cold Spring Harb. Protocols 2007, pdb.top17 (2007).
Akutse, K. S., Maniania, N. K., Fiaboe, K. K. M., Van den Berg, J. & Ekesi, S. Endophytic colonization of Vicia faba and Phaseolus vulgaris (Fabaceae) by fungal pathogens and their effects on the life-history parameters of Liriomyza huidobrensis (Diptera: Agromyzidae). Fungal Ecol. 6, 293–301 (2013).
Akutse, K. S., Subramanian, S., Khamis, F. M., Ekesi, S. & Mohamed, S. A. Entomopathogenic fungus isolates for adult Tuta absoluta (Lepidopter: Gelechiidae) management and their compatibility with Tuta pheromone. J. Appl. Entomol. 144, 777–787 (2020).
Inglis, G. D., Enkerli, J., Goettel, M. S. & Tj, C. Laboratory techniques used for entomopathogenic fungi : hypocreales. Manual of Techniques in Invertebrate Pathology (Elsevier Ltd, 2012). https://doi.org/10.1016/B978-0-12-386899-2.00007-5.
Goettel, M. S. & Inglis, G. D. Fungi : Hyphomycetes. In Manual of Techniques in Insect Pathology 213–249 (ScienceDirect, 1997). https://doi.org/10.1016/B978-012432555-5/50013-0.
Myrchiang, P., Dkhar, M. S. & Devi, H. R. Studies on endophytic fungi associated with medicinally important aromatic plant Artemisia nilagirica (C.B. Clarke) Pamp. and their antagonistic activity against Phytophthora infestans. Adv. Lab. Res. Biol. 5, 112–119 (2014).
Karuppiah, V., Sun, J., Li, T., Vallikkannu, M. & Chen, J. Co-cultivation of Trichoderma asperellum GDFS1009 and Bacillus amyloliquefaciens 1841 causes differential gene expression and improvement in the wheat growth and biocontrol activity. Front. Microbiol. 10, 1068 (2019).
Barra-Bucarei, L. et al. Antifungal activity of Beauveria bassiana endophyte against Botrytis cinerea in two Solanaceae crops. Microorganisms 8, 1–15 (2020).
Schulz, B., Guske, S., Dammann, U. & Boyle, C. Endophyte-host interactions. II. Defining symbiosis of the endophyte-host interaction. Symbiosis 25, 213–227 (1998).
Seo, Y. & Kim, Y. H. Potential reasons for prevalence of fusarium wilt in oriental melon in Korea. Plant Pathol. J. 33, 249–263 (2017).
Amini, J. Physiological race of Fusarium oxysporum f. sp. lycopersici in Kurdistan province of Iran and reaction of some tomato cultivars to race 1 of pathogen. Plant Pathol. J. 8, 68–73. https://doi.org/10.3923/ppj.2009.68.73 (2009).
Horinouchi, H., Muslim, A. & Hyakumachi, M. Biocontrol of Fusarium wilt of spinach by the plant growth promoting fungus Fusarium equiseti GF183. J. Plant Pathol. 92, 249–254 (2010).
Batista, R. O. et al. Resistance to Fusarium wilt in common bean. Crop Breed. Appl. Biotechnol. 16, 226–233 (2016).
Aydi-Ben-Abdallah, R., Jabnoun-Khiareddine, H. & Daami-Remadi, M. Fusarium wilt biocontrol and tomato growth stimulation, using endophytic bacteria naturally associated with Solanum sodomaeum and S. bonariense plants. Egypt. J. Biol. Pest Control 30, 1–13 (2020).
Tanveer, K. et al. Effect of salt stress on tomato plant and the role of calcium. J. Plant Nutr. 43, 28–35 (2020).
Liu, J., Hu, T., Feng, P., Wang, L. & Yang, S. Tomato yield and water use efficiency change with various soil moisture and potassium levels during different growth stages. PLoS ONE 14, 1–14 (2019).
Ali, A. et al. Evaluation of various tomato (Lycopersicon esculentum Mill.) cultivars for quality, yield and yield components under agroclimatic condition of Peshawar. ARPN J. Agric. Biol. Sci. 11, 59–62 (2016).
Shapiro, S. & Wilk, M. An analysis of variance test for normality (Complete samples). Biometrika 52, 591–611 (1965).
R Core Team. R: A language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2019)
Acknowledgements
This work received financial support from the UK’s Foreign, Commonwealth and Development Office (FCDO) (B2329A-FCDO-FAW and B2291A-FCDO-BIOPESTICIDE) through the International Centre of Insect Physiology and Ecology (icipe). The authors gratefully acknowledge the icipe core funding provided by the Swedish International Development Cooperation Agency (Sida); the Swiss Agency for Development and Cooperation (SDC); the Australian Centre for International Agricultural Research (ACIAR); the Norwegian Agency for Development Cooperation (Norad); the German Federal Ministry for Economic Cooperation and Development (BMZ); and the Government of the Republic of Kenya. The first author was also supported through the Inter-University Council for East Africa (IUCEA) of the East African Community (EAC) Scholarship Programme. We are also thankful to Wafula Sospeter, Jane Kimemia and Levi Ombura for their technical assistance. The views expressed herein do not necessarily reflect the official opinion of the donors.
Author information
Authors and Affiliations
Contributions
Conceptualization: KSA and FMK, Funding acquisition: KSA, Experimental design: MCM, KSA and FMK, Software: MCM and KSA, Investigation: MCM, Visualization: MCM, FMK and KSA, Resources: KSA and FMK; Supervision: ESN, WW and KSA, Project administration: KSA, writing—original draft preparation: MCM, Writing—review and editing: MCM, KSA, FMK, ESN and WW. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Muhorakeye, M.C., Namikoye, E.S., Khamis, F.M. et al. Biostimulant and antagonistic potential of endophytic fungi against fusarium wilt pathogen of tomato Fusarium oxysporum f. sp. lycopersici. Sci Rep 14, 15365 (2024). https://doi.org/10.1038/s41598-024-66101-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-66101-1
- Springer Nature Limited