Abstract
Central nervous system Infections (CNSIs) is a disease characterized by complex pathogens, rapid disease progression, high mortality rate and high disability rate. Here, we evaluated the clinical value of metagenomic next generation sequencing (mNGS) in the diagnosis of central nervous system infections and explored the factors affecting the results of mNGS. We conducted a retrospective study to compare mNGS with conventional methods including culture, smear and etc. 111 suspected CNS infectious patients were enrolled in this study, and clinical data were recorded. Chi-square test were used to evaluate independent binomial variables, taking p < 0.05 as statistically significant threshold. Of the 111 enrolled cases, 57.7% (64/111) were diagnosed with central nervous system infections. From these cases, mNGS identified 39.6% (44/111) true-positive cases, 7.2% (8/111) false-positive case, 35.1% (39/111) true-negative cases, and 18.0% (20/111) false-negative cases. The sensitivity and specificity of mNGS were 68.7% (44/64) and 82.9% (39/47), respectively. Compared with culture, mNGS provided a higher pathogen detection rate in CNSIs patients (68.7% (44/64) vs. 26.5% (17/64), p < 0.0001). Compared to conventional methods, positive percent agreement and negative percent agreement was 84.60% (44/52) and 66.1% (39/59) separately. At a species-specific read number (SSRN) ≥ 2, mNGS performance in the diagnosis of definite viral encephalitis and/or meningitis was optimal (area under the curve [AUC] 0.758, 95% confidence interval [CI] 0.663–0.854). In bacterial CNSIs patients with significant CSF abnormalities (CSF WBC > 300*106/L), the positive rate of CSF mNGS is higher. To sum up, conventional microbiologic testing is insufficient to detect all neuroinvasive pathogens, and mNGS exhibited satisfactory diagnostic performance in CNSIs and with an overall detection rate higher than culture (p < 0.0001).
Similar content being viewed by others
Introduction
Central nervous system infections (CNSIs) are usually classified as meningitis, encephalitis (or meningoencephalitis), or myelitis (or encephalomyelitis) based on the predominant anatomic site of infection. Infections of the CNS—associated with bacteria, viruses, fungi, and mycobacteria—compose a considerable portion of human morbidity and mortality1. Timely identification of the pathogen is essential for the process of antimicrobial therapy and its impact on clinical outcomes2. However, it is often difficult to determine the etiology of CNSIs. More than 100 pathogens have been reported as causative agents of CNSIs, most studies only identified the etiology in about 5.4–32.3% of cases3,4,5. The traditional detection methods for CNSIs include pathogen culture, serological detection, and molecular biology detection, among others. Culture is still the gold standard for the diagnosis of CNSIs, but the diagnosis and treatment of CNSIs will be delayed due to lack of sensitivity and long culture time. Besides, serological detection techniques such as immunofluorescence and complement binding assay have drawbacks such as long detection cycles, insufficient specificity or sensitivity6. Of course, the etiology in any individual case may be influenced by many factors including test availability, geographic region, host, and many others–the main point is that the exact etiology is not always uncovered7. With the development of detection technology, while ensuring specificity and sensitivity, researchers have further increased their requirements for detection speed. Molecular biology detection technology stands out because it is more suitable for laboratory testing and epidemiological investigations.
Recently, metagenomic next-generation sequencing (mNGS) has emerged as a novel and promising method for the detection of infectious agents. Based on non-preference and high sensitivity, mNGS provides more sensitive pathogen detection results than conventional culture8. To date, several studies have addressed value of mNGS in finding out pathogens and minimizing time of empirical treatment without pathogenic evidence9,10,11. In 2020, mNGS was recommended in ‘Clinical practice expert consensus for the application of metagenomic next generation sequencing’ as Level A evidence to assist clinical decision in CNS infections12. However, most previous studies have only focused on clinical cases caused by novel or rare pathogens. Thus, the performance of mNGS in diagnosis of CNSIs should be studied using a larger sample size. Therefore, we conducted a retrospective study involving hospitalized patients with suspected CNSIs. We describe the sensitivity and specificity of CSF mNGS assays to detect pathogens in patients with CNSIs. The aim of our study was to determine pathogens and the incremental yield of mNGS that could not be detected by conventional methods. Furthermore, we also explored potential factors that affect the mNGS outcomes of patients.
Materials and methods
Study design
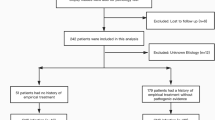
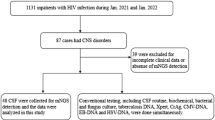
This study was a retrospective analysis of mNGS in CSF samples obtained from patients with suspected CNSIs. We retrospectively reviewed 131 patients with suspected CNSIs at the First Affiliated Hospital of Nanchang University from 15 February 2018 to 5 February 2023. A total of 111 samples were included in the current study, according to our inclusion/exclusion criteria (Fig. 1). The inclusion criteria of patients were as follows: (a) patients over 18 years of age; (b) high-level clinical suspicion of CNS infectious diseases; (c) CSF sample undergone mNGS. Patients were excluded from the study based on the following criteria: (a) patients with an uncertain final diagnosis; (b) patients with incomplete data. CSF samples were sent for: routine (quantity and classification of CSF cell) and biochemical tests (protein, glucose, chloride, and adenosine deaminase content of CSF); culturing of bacteria, fungi and tuberculosis; specific antibody tests; serologic tests and CSF smear. Cell biopsy and nucleic acid amplification testing (traditional PCR, Xpert MTB (Mycobacterium tuberculosis)) were conducted according to clinically assessed necessity. All conventional tests were done and collective conclusion made which is blinded to those doing and analysis and mNGS data. The final clinical diagnosis of the patient was independently determined by 3 physicians with expertise in CNSIs. Criteria for CNSIs refers to Chinese Expert Consensus on the Diagnosis and Treatment of Central Nervous System Infection in Neurosurgery (2021) (Supplementary Table S1)13. Then, based on final clinical diagnosis, all samples were categorized into 2 groups defined as CNS infection and non-CNS infection.
Metagenomic next-generation sequencing and analysis
Sample collection and nucleic acid extraction
CSF samples were collected according to the standard procedures and stored at − 20 °C. 1ml cerebrospinal fluids (CSF) were used for mNGS testing. Nucleic acid from all samples including the negative controls (NCs) was obtained using the TIANamp Micro DNA Kit (TIANGEN Biotech, Beijing, China) according to the manufacturer’s manual. Using Qubit dsDNA HS Assay Kit (ThermoFisher Scientific) for nucleic acid concentration determination following the manufacturer’s manual. Negative control (non-template control, NTC (sterile deionized water)) were also set for each batch of experiments using the same wet lab procedures from DNA extraction to end of sequencing and bioinformatics analysis.
Library construction and sequencing
Libraries were constructed by using the Nextera XT kit (Illumina) following the manufacturer manual. Illumina NextSeq-550Dx sequencer was employed for the sequencing reaction to gain the sample sequence information in the sample. The sequencing depth of each sample ≥ 20 million reads was sequenced using a 75-cycle single-end sequencing strategy. Raw sequencing data were filtered by the index reads removing any reads associated with indexes that had a quality score of less than Q30. This process was implemented to enhance the overall quality and reliability of the data.
Bioinformatics analysis
Bcl2fastq v2.20.0.422(Illumina, https://support.illumina.com/sequencing/sequencing_software/bcl2fastq-conversion-software.html) with default parameters was applied to demultiplex the primary sequencing output. For quality control adapter contamination and low-quality and low-complexity reads, raw reads and adapter removal were filtered by fastp (v0.19.5)14 and Komplexity v0.3.615. Samples with < 10 million reads after QC were excluded from this study, the QC information of samples is shown in online Supplementary Table S2. Reads that were mapped to the human reference assembly GRCh38 were removed with Bowtie2 v2.3.4.316. Then, reads were aligned to the microorganism database consisting of approximately 12,000 genomes with SNAP v1.0 beta.1817 as previously described18. The mapped reads were classified based on the NCBI RefSeq genome database, or the NCBI GenBank genome database (ftp://ftp.ncbi.nlm.nih.gov/genomes/)was selected for each species. After filtering low-complexity reads19, we counted the species or genus abundance with Perl scripts.The software, source code and proprietary binaries is available for download on GitHub (https://github.com/OpenGene/fastp).
Criteria for positive mNGS detection20,21
-
(1)
For the detected bacteria (Mycobacterium excluded), fungi (Cryptococcus excluded), and parasites, the positive criteria for the mNGS result were set as follows: (a) genome coverage of the unique reads mapped to this microorganism ranked top10 of the same kind of microbes and the microorganism was not detected in the NTC; or (b) RPMsample/RPMNTC was > 10 (RPMNTC ≠ 0, Reads per million:RPM).
-
(2)
For viruses, Mycobacterium tuberculosis, and Cryptococcus, a positive mNGS result was considered when it was not detected in NTC and at least 1 unique read was mapped to species or when RPMsample/RPMNTC was > 5 (RPMNTC ≠ 0).
Diagnostic assessment of mNGS
Each mNGS result was identified as a true positive (TP), false positive (FP), true negative (TN), or false negative (FN) result relative to the clinical diagnosis. If the pathogen was found to be consistent with the clinical diagnosis, it was considered as true positive. A false positive was defined as a case in which the detected pathogen was inconsistent with the clinical diagnosis. A true negative case was defined if no pathogen was detected by mNGS and a clinical diagnosis of non-CNSIs was ultimately determined. Those whose CSF was detected no pathogen by mNGS but were clinically diagnosed as CNSIs were all false-negative cases.
Statistical analysis
All statistical analyses were conducted using SPSS ver. 26.0 (SPSS, Chicago, IL, USA; https://www.ibm.com/products/spss-statistics), GraphPad Prism (ver. 9.0; GraphPad Software, San Diego, CA; www.graphpad.com), with counting data described in terms of frequency and component ratio. Continuous variables of normal distribution are expressed as means ± SD(standard), or as medians (25th, 75th percentiles) for non-normal distribution. Continuous variables were compared by Mann–Whitney U-test or t-test, and categorical variables were analyzed using the Fisher’s exact test. p < 0.05 was considered statistically significant for the difference.
Ethical approval and consent to participate
According to the review and comments made by the Ethics Committee of the First Affiliated Hospital of Nanchang University, our research is in line with the exemption type of informed consent and ethics approval that “Using identifiable human body materials or data for research, it is no longer possible to locate the subject, and the research project does not involve personal privacy disclosure or commercial interests”. As this is a retrospective Cohort study based on previous clinical diagnosis and treatment results, the Ethics Committee of the First Affiliated Hospital of Nanchang University granted the study exemption status (AF-SQ-10-2.1). In addition, we declare that this study is in line with the ethical guidelines of the Declaration of Helsinki, and the patient related data is strictly confidential. All methods were carried out in accordance with relevant guidelines and regulations in the manuscript.
Results
General characteristics of the enrolled cohort
As displayed in Fig. 1, a total of 131 patients were enrolled, 1 patient was lost to follow-up and 17 patients were not included due to unclear final diagnosis. Among 111 participants, 64 patients were diagnosed with CNS infections, including 23 cases (35.9%) of viral infection, 19 (29.7%) of bacterial infection, 10 (15.6%) of Mycobacterium tuberculosis, 7 (10.9%) of fungal infection, and 5 (7.8%) of mixed infection(3 of bacterial and viral coinfection, 2 of tuberculosis and fungal coinfection). 47 patients were finally diagnosed with non-CNS infections, including 10 cases (21.3%, 10/47) of meningeal carcinoma, 12 cases (25.5%,12/47) of encephalopathy associated with autoimmune disease, 4 cases (8.5%, 4/47) of metabolic encephalopathy and 21 cases (44.7%, 21/47) of other non-infectious diseases of CNS. The underlying medical comorbidities, clinical characteristics, and laboratory examination are reported in Table 1. The mean length of stay in the hospital for patients with CNS infections was longer than that of patients diagnosed with non-CNS infections (p = 0.0309). 68.5% (76/111) patients had a history of empirical treatment before CSF sampling. There was a difference in empirical treatment history between the two groups (p = 0.0397). Compared with patients diagnosed with non-CNS infections, those with CNS infections exhibited significantly lower chloridion in peripheral blood (p = 0.0108), CSF chloridion (p = 0.0211) and glucose ratio of CSF to serum (p = 0.0484). We did not find any other significant differences between the two groups in terms of clinical presentation or other laboratory parameters.
Performance of mNGS and conventional test to clinical diagnosis
Among 64 CNSIs patients, 44 were confirmed by mNGS and 17 were confirmed by conventional methods. Using clinical diagnosis as the gold standard, positive predictive value (PPV), negative predictive value (NPV), sensitivity, and specificity were calculated to evaluate the diagnostic value of mNGS and conventional test. The sensitivity and specificity of diagnosing CNSIs by mNGS were 68.7% (95% confidence interval (CI) 56.6–78.7%) and 82.9% (95% CI 56.6–78.7%) respectively, with NPV and PPV being 66.1% (95% CI 53.4–76.8%) and 84.6% (95% CI 72.5–92.0%) (Fig. 2a). The sensitivity and specificity of diagnosing CNSIs by conventional diagnostic testing were 26.5% (95% CI 17.3–38.4%) and 95.7% (95% CI 85.8–99.2%) respectively, with NPV and PPV being 48.9% (95% CI 38.9–58.9%) and 89.4% (95% CI 68.6–98.1%) (Fig. 2b). Furthermore, we compared the detected and clinically approved pathogens by mNGS and conventional test (Fig. 2c). The etiological diagnosis showed that mNGS reported 44 positive cases. mNGS detection revealed that bacteria (n = 17) was the most common identified potential pathogens. The top one causative pathogens identified were Klebsiella pneumoniae. The most detected fungal infection is Cryptococcal meningitis and Aspergillus meningitis. The most detected viral infection is Epstein-Barr virus. Overall, 10.9% (7/64) were diagnosed by both conventional testing and mNGS. mNGS detected an additional 27 types of bacteria and fungi in both culture negative patients and conventional methods-negative tests (excluding culture) (Table 2). Infections that were diagnosed solely by mNGS included Leptospira renal, Varicella-zoster virus, Mycobacterium frothy, Nocardia farcinica, Listeria monocytogenes, these pathogens had not been considered by the treating clinicians for the patients. mNGS also identified pathogens for which there was some degree of clinical suspicion, although conventional testing had returned negative(Acinetobacter baumannii, Klebsiella pneumoniae, Pseudomonas aeruginosa). More detailed information on results of mNGS and conventional methods was showed in Supplementary Table S3.
Influence of antimicrobial treatment on diagnostic detection rate of mNGS
Among CNS infected patients who underwent or did not undergo empirical treatment before CSF sampling, the detection rates of mNGS for potential pathogens were 70.73% and 65.22%, respectively. The detection rate of mNGS was significantly higher than conventional methods, in regardless of whether it is in the experience treatment group or the non experience treatment group (70.73% vs. 36.59%, p = 0.0016; 65.22% vs. 8.70%, p = 0.000 respectively). In bacterial CNS infections, detection rate of mNGS was significantly higher than that of conventional methods, both empirically treated and non-empirically treated groups (76.92% vs. 23.08%, p = 0.0045; 77.78% vs. 11.11%, p = 0.0023, respectively). In fungal CNS infections, detection rate of mNGS was significantly higher than that of conventional methods in empirically treated groups (85.71% vs. 28.57%, p = 0.0306). In viral CNS infections, detection rate of mNGS was also significantly higher than that of conventional methods in empirically treated groups (60.0% vs. 13.3%, p = 0.0067). No other significant difference was observed in fungal and viral infections in non-empirically treated groups. The detection rate of mNGS compared to final diagnosis were not statistically different, between non-empirically treated groups and treated groups. In bacterial CNS infections, the detection rate of conventional methods compared to final diagnosis in empirically treated was significantly higher than non-empirically treated groups (36.59% vs. 8.70%, p = 0.0187) (Table 3).
Use of mNGS to diagnose viral encephalitis and/or meningitis
mNGS detected an additional 8 cases of viruses in patients diagnosed with non-CNS infections. Therefore, we evaluated the ability of mNGS in identifying viruses. 26 patients with definite viral encephalitis and/or meningitis were enrolled. An mNGS result was considered positive if the CSF sample exhibited species-specific read number (SSRNs) ≥ 1, 2, 5, or 10; the diagnostic sensitivity for viral encephalitis and/or meningitis were 53.8%, 57.7%, 30.8% and 19.2%, respectively. The specificity were 91.7%, 88.2%, 92.9% and 94.1%, respectively. The area under curve (AUC) for the four positive mNGS criteria were 0.727, 0.758, 0.617 and 0.539, respectively. When an SSRN ≥ 2 was considered positive, the corresponding AUC (0.758) was relatively larger than when other SSRNs were considered positive (AUC 0.758, 95% CI 0.663–0.854) (Fig. 3).
Correlative analysis between mNGS and CSF laboratory
To provide additional insights for the clinical indication of CNSIs, we investigated the clinical features between the CNS infection groups and its subgroups. The CSF chloridion in the mNGS positive/case consistent group were significantly lower than those in the mNGS negative/case consistent group (Fig. 4a). We further evaluated the influence of CSF laboratory examinations on mNGS detection rate (Fig. 4b). The detection rate of mNGS in CNSIs patients is higher in patients with CSF WBC > 300*106/L, CSF protein > 500mg/L, glucose ratio > 5, or CSF chloridion ≤ 110 (mmol/L), but there was no statistically significant difference between the two groups. In bacterial subgroup, mNGS detection rate was signifcantly higher in patients with CSF WBC > 300*106/L (p = 0.0274).
Discussion
In our study, we assessed diagnostic performance of mNGS and conventional methods compared with clinical final diagnosis. Overall, mNGS detected extra 27 bacteria and fungi in culture negative patients and conventional methods-negative tests excluding culture. mNGS detected a wide range of pathogens, including those causative agents commonly and infrequently reported in CNS infections. Compared to conventional methods, we further studied diagnostic performance of mNGS in each type of CNS infections. In bacterial, fungal and viral CNS infections, mNGS was superior to culture and conventional methods regardless of administration of empirical treatment. Wilson et al. also observed similar finding that mNGS detected significantly more pathogens than traditional methods in patients receiving empirical treatment with antibiotics22. Besides, our research also revealed that the detection rate of mNGS compared to final diagnosis were not statistically different between non-empirically treated groups and treated groups, which is contrary to other studies. Zhang et al.’s research supports the advantages of mNGS, especially in patients who have previously received antibiotic treatment23. Further analysis showed that if delayed-sampling after effective antimicrobial therapy is more than 4 days, it could decrease the mNGS detection rate significantly24. The reason behind this phenomenon may be due to different diagnostic mechanisms behind it. And the sensitivity of culture is limited by the use of antibiotics and antifungals25. Previous researches have shown that the detection rate of CSF culture in meningitis could be reduced to 9–11% after effective treatment26,27. Due to the fact that culture requires the existence of livable pathogens and therefore is easily influenced by the administration of antimicrobial treatment.
Pneumoniae and Neisseria meningitidis are the most common pathogens causing bacterial meningitis, However, in our study, K. pneumoniae and A. baumannii were the main pathogens. Listeria monocytogenes has become one of the common pathogens causing bacterial meningitis, this may be due to the differences in the types of pathogens in different areas, which is consistent with many cases reported in China28. Viruses are also a common cause of CNS infections, the most common viral infection are herpes simplex virus29. Among the 15 patients with viral infections, except for one case of dengue virus, all pathogens identified via mNGS were DNA viruses. RNA viruses, such as enteroviruses and Japanese encephalitis virus, are also common causes of CNSIs. Thus, DNA/RNA co-extraction methods must be improved and DNA and RNA sequenced simultaneously to improve virus detection rates30. In addition, identification of specific viral encephalitis/meningitis pathogens is difficult, some viruses are very common pathogen that may cause a false-positive result if present in CSF. In our research, when SSRNs ≥ 2 were considered positive, mNGS performance was optimal in terms of viral meningitis diagnosis (AUC = 0.758); mNGS may be very useful in this situation.
CSF chloridion showed significant differences between mNGS positive/Case consistent group and mNGS negative/Case consistent group, and this can be easily explained by that decreased CSF chloridion are associated with higher chance of CNS infections. Cut-off value analysis found that patients with CSF WBC > 300*106/L, CSF protein > 500 mg/L, glucose ratio > 4.5, or CSF chloridion ≤ 110(mmol/L) have a higher mNGS detection rates, but there was no statistically significant difference between two groups. Further research is needed to determine whether these indications hint that these patients may be more likely to benefit from mNGS. In bacterial subgroup, CSF WBC > 300*106/L with higher mNGS detection rate (p = 0.0274), implying that these patients with bacterial CNSIs may be more likely to benefit from mNGS. Another study found that CSF WBC > 300 * 106/L were related with higher mNGS detection rate, indicating that higher CSF WBC count might reflect a more active diseases status and higher pathogen loads, leading to higher detection rate23.
mNGS reported 8 failed false-positive cases. Non-pathogenic microbes’sequences detected in our study may come from three possible sources: environmental contamination; skin flora; reagent pollution; errors occurred in sequencing and mapping. We used negative controls in each sequencing to control contamination, and all samples in this study were processed in the same laboratory according to the same protocol. Incorporation of quality controls and positivity-thresholds chosen to avoid major errors is a crucial step in ensuring the results of mNGS30. How to determine the cutoff value of a positive result from mNGS test is always an issue for laboratories. So this metagenomic approach need to prompt reassessment of their significance when identified above reporting thresholds31. More studies should include comparisons with usual practice and application of mNGS to infectious and non-infectious patient, to better inform distinction between colonisation and infection and quantify clinical impacts32,33. mNGS also missed pathogen testing for 20 infected patients, which has many reasons behind it. For example, the abundance of pathogens in CSF is very low34; the fungal cell wall is thick and hard to break, and DNA fragments are difficult to obtain, which may affect the detection rate of fungi28. Qian et al.'s study pointed out that pre-use of antibiotics can also cause false negative mNGS test results35. It can be seen that multiple stages can affect the mNGS detection results. When CNS infections are more complex, we recommend a combination of mNGS and traditional testing methods.
Nevertheless, there were some deficiencies in our study. Firstly, the limited sample size may affect the accuracy of the study. Secondly, although each subject underwent CSF culture, the choice to use serological or molecular-based testing was entirely at the discretion of the clinical team. Therefore, most of the results of mNGS were not further verified by a standard reference, such as PCR and serologic confirmation. Thirdly, for practical applications of mNGS, the subjective judgment of clinicians is still needed, which is highly dependent on clinical experience. Although consensus was reached in this study by three experienced senior physicians and was based on the clinical manifestations of patients combined with other laboratory results, subjective bias is still possible.
Conclusion
Conventional microbiologic methods are often insufficient to detect all neuroinvasive pathogens. In this study, the detection rate of mNGS for potential pathogens is generally superior to traditional methods. Our findings also discovered that elevated CSF WBC and protein level or decreased glucose ratio is correlated with higher mNGS positivity in CNS infections, the analysis of clinical features associated with positive mNGS results further supports the clinical feasibility and utility of this technique. The combination of higher specificity from culture tests and higher sensitivity from mNGS may be more useful for physicians to determine the causative pathogen for a given case. We also suggest that the positive cut off should be more flexible for mNGS in the current stage of clinical applications.
Data availability
The datasets generated and/or analysed during the current study are available in the National Center Biotechnology Information BioProject database repository [ID 1,018,096—BioProject—NCBI (nih.gov)], accession number PRJNA1018096.
References
Bahr, N. C. & Boulware, D. R. Methods of rapid diagnosis for the etiology of meningitis in adults. Biomark. Med. 8, 1085–1103. https://doi.org/10.2217/bmm.14.67 (2014).
Hong, N. T. T. et al. Cerebrospinal fluid MinION sequencing of 16S rRNA gene for rapid and accurate diagnosis of bacterial meningitis. J. Infect. 80(4), 469–496. https://doi.org/10.1016/j.jinf.2019.12.011 (2020).
Sulaiman, T., Salazar, L. & Hasbun, R. Acute versus subacute community-acquired meningitis: Analysis of 611 patients. Medicine 96, e7984. https://doi.org/10.1097/MD.0000000000007984 (2017).
McGill, F. et al. The UK joint specialist societies guideline on the diagnosis and management of acute meningitis and meningococcal sepsis in immunocompetent adults. J. Infect. 72, 405–438. https://doi.org/10.1016/j.jinf.2016.01.007 (2016).
Hasbun, R. et al. Epidemiology of meningitis and encephalitis in the United States, 2011–2014. Clin. Infect. Dis. 65, 359–363. https://doi.org/10.1093/cid/cix319 (2017).
Tang, X. et al. Psittacosis caused severe community-acquired pneumonia accompanied by acute hypoxic respiratory failure: A multicenter retrospective cohort study from China. BMC Infect. Dis. 23, 532. https://doi.org/10.1186/s12879-023-08283-z (2023).
Matta, S. K., Rinkenberger, N., Dunay, I. R. & Sibley, L. D. Toxoplasma gondii infection and its implications within the central nervous system. Nat. Rev. Microbiol. 19, 467–480. https://doi.org/10.1038/s41579-021-00518-7 (2021).
Xing, X. W. et al. Metagenomic next-generation sequencing for diagnosis of infectious encephalitis and meningitis: A large, prospective case series of 213 patients. Front Cell Infect. Microbiol. 10, 88. https://doi.org/10.3389/fcimb.2020.00088 (2020).
Wilson, M. R. et al. Actionable diagnosis of neuroleptospirosis by next-generation sequencing. N. Engl. J. Med. 370, 2408–2417. https://doi.org/10.1056/NEJMoa1401268 (2014).
Wylie, K. M. et al. High-throughput sequencing of cerebrospinal fluid for diagnosis of chronic Propionibacterium acnes meningitis in an allogeneic stem cell transplant recipient. Transpl. Infect. Dis. 18, 227–233. https://doi.org/10.1111/tid.12512 (2016).
Wilson, M. R. et al. Clinical metagenomic sequencing for diagnosis of meningitis and encephalitis. N. Engl. J. Med. 380, 2327–2340. https://doi.org/10.1056/NEJMoa1803396 (2019).
Editorial Board of Chinese Journal of Infectious Diseases. Clinical practice expert consensus for the application of metagenomic next generation sequencing. Chin. J. Infect. Dis. 38, 681–689. https://doi.org/10.3760/cma.j.cn311365-20200731-00732 (2020).
The Neurocritical Expert Committee of the Neurosurgery Branch of the Chinese Medical Association, and the Neurosurgery Critical Care Group of the Neurosurgery Branch of the Beijing Medical Association. Chinese expert consensus on the diagnosis and treatment of central nervous system infection in neurosurgery. Chin. J. Neurosurg. 37(14), 2021. https://doi.org/10.3760/cma.j.cn112050-2020831-00480 (2021).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890. https://doi.org/10.1093/bioinformatics/bty560 (2018).
Clarke, E. L. et al. Sunbeam: An extensible pipeline for analyzing metagenomic sequencing experiments. Microbiome 7, 46. https://doi.org/10.1186/s40168-019-0658-x (2019).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359. https://doi.org/10.1038/nmeth.1923 (2012).
Naccache, S. N. et al. A cloud-compatible bioinformatics pipeline for ultrarapid pathogen identification from next-generation sequencing of clinical samples. Genome Res. 24, 1180–1192. https://doi.org/10.1101/gr.171934.113 (2014).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864. https://doi.org/10.1093/bioinformatics/btr026 (2011).
Zaragoza, R., Ramírez, P. & López-Pueyo, M. J. Infección nosocomial en las unidades de cuidados intensivos [Nosocomial infections in intensive care units]. Enferm Infecc Microbiol Clin 32, 320–327. https://doi.org/10.1016/j.eimc.2014.02.006 (2014).
Miller, S. et al. Laboratory validation of a clinical metagenomic sequencing assay for pathogen detection in cerebrospinal fluid. Genome Res. 29(5), 831–842. https://doi.org/10.1101/gr.238170.118 (2019).
Zhang, H. et al. Metagenomic next-generation sequencing of plasma cell-free DNA improves the early diagnosis of suspected infections. BMC Infect. Dis. 24, 187. https://doi.org/10.1186/s12879-024-09043-3 (2024).
Zhang, S. et al. Understanding etiology of community-acquired central nervous system infections using metagenomic next-generation sequencing. Front. Cell Infect. Microbiol. 12, 979086. https://doi.org/10.3389/fcimb.2022.979086 (2022).
Zhang, M. et al. Metagenomic next-generation sequencing for diagnosis of infectious encephalitis and meningitis: A retrospective study of 90 patients. Neurol. Res. 46, 187–194. https://doi.org/10.1080/01616412.2023.2265243 (2024).
Chen, C. et al. The incidence and risk factors of meningitis after major craniotomy in China: A retrospective cohort study. PloS one 9, e101961. https://doi.org/10.1371/journal.pone.0101961 (2014).
Wilson, M. R. et al. Acute West Nile virus meningoencephalitis diagnosed via metagenomic deep sequencing of cerebrospinal fluid in a renal transplant patient. Am. J. Transplant. 17, 803–808. https://doi.org/10.1111/ajt.14058 (2017).
Bronska, E. et al. Dynamics of PCR-based diagnosis in patients with invasive meningococcal disease. Clin. Microbiol. Infect. 12, 137–141. https://doi.org/10.1111/j.1469-0691.2005.01327.x (2006).
Nigrovic, L. E. et al. Effect of antibiotic pretreatment on cerebrospinal fluid profiles of children with bacterial meningitis. Pediatrics 122, 726–730. https://doi.org/10.1542/peds.2007-3275 (2008).
Liu, Y., Zhu, W., Jiao, M., Guo, W. & Luo, Y. Clinical application value of metagenomic next-generation sequencing in the diagnosis of central nervous system infections. Front. Bioeng. Biotechnol. 11, 885877. https://doi.org/10.3389/fbioe.2023.885877 (2023).
Gu, W., Miller, S. & Chiu, C. Y. Clinical metagenomic next-generation sequencing for pathogen detection. Annu. Rev. Pathol. 14, 319–338. https://doi.org/10.1146/annurev-pathmechdis-012418-012751 (2019).
Charalampous, T. et al. Evaluating the potential for respiratory metagenomics to improve treatment of secondary infection and detection of nosocomial transmission on expanded COVID-19 intensive care units. Genome Med. 13, 182. https://doi.org/10.1186/s13073-021-00991-y (2021).
Charalampous, T. et al. Nanopore metagenomics enables rapid clinical diagnosis of bacterial lower respiratory infection. Nat. Biotechnol. 37, 783–792. https://doi.org/10.1038/s41587-019-0156-5 (2019).
Chiu, C. Y. & Miller, S. A. Clinical metagenomics. Nat. Rev. Genet. 20, 341–355. https://doi.org/10.1038/s41576-019-0113-7 (2019).
Govender, K. N., Street, T. L., Sanderson, N. D. & Eyre, D. W. Metagenomic sequencing as a pathogen-agnostic clinical diagnostic tool for infectious diseases: A systematic review and meta-analysis of diagnostic test accuracy studies. J. Clin. Microbiol. 59, e0291620. https://doi.org/10.1128/JCM.02916-20 (2021).
Wang, S. et al. The feasibility of metagenomic next-generation sequencing to identify pathogens causing tuberculous meningitis in cerebrospinal fluid. Front. Microbiol. 10, 1993. https://doi.org/10.3389/fmicb.2019.01993 (2019).
Galardi, M. M. et al. Pathogen and antibody identification in children with encephalitis in Myanmar. Ann. Neurol. 93, 615–628. https://doi.org/10.1002/ana.26560 (2023).
Acknowledgements
All authors wish to thank the patient for participating in this study and all the staff members at our institution.
Funding
This research was supported by the Science and Technology Program of Jiangxi Traditional Chinese Medicine (grant number 2021B723), the National Natural Science Fund of China (grant number: 82260085) and the Natural Science Foundation of Jiangxi Province(grant number: 20242BAB217046). The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
All authors made a significant contribution to the work whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; agreed to submit to the current journal; and agreed to be accountable for all aspects of the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yuan, L., Zhu, X.Y., Lai, L.M. et al. Clinical application and evaluation of metagenomic next-generation sequencing in pathogen detection for suspected central nervous system infections. Sci Rep 14, 16961 (2024). https://doi.org/10.1038/s41598-024-68034-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-68034-1
- Springer Nature Limited