Abstract
The epithelial–mesenchymal transition (EMT) is a genetic reprogramming that tumor cells utilize for metastasis. Epsin-3 (EPN3) is an endocytic adapter protein involved in clathrin-mediated endocytosis and had been previously linked to EMT in breast cancer and glioma metastasis. In this study, identified the role of epsin-3 in lung adenocarcinoma and metastasis and epsin-3 levels identified using an expression profile analysis of patient data indicated the protein was abnormally overexpressed in lung adenocarcinoma patients and this was directly linked to disease severity. Gene knockdowns of EPN3 in human adenocarcinoma cell line A549 and the non-small cell lung carcinoma cell line H1299 decreased the levels of mesenchymal markers, including vimentin (VIM), N-cadherin (NCAD) and embryonic transcription factors like zinc finger E-box binding homeobox 1(ZEB1), snail, and the key molecules of Wnt pathway such as β-catenin and resulted in increased expression of the epithelial marker E-cadherin (ECAD). Our data links EPN3 to the EMT process in lung cancer and inhibition of its expression reduced the metastatic and invasive ability of lung adenocarcinoma cells by inhibiting the EMT process.
Similar content being viewed by others
Introduction
Lung cancer is one of the most common malignancies and the second most prevalent cancer with 2.2 million new cases in 20201. Pathological classifications divide lung cancer into the most prevalent non-small cell and small cell lung cancers. The non-small cell cancer can be further divided into lung adenocarcinoma and lung squamous cell carcinomas and the adenocarcinoma (LUAC) is the primary pathological type2. Lung cancer development occurs via tumor invasion and metastasis from in situ to distant sites and is often the primary reason for poor treatment effect and poor prognosis. In addition, cancer heterogeneity is also a primary reason for treatment failure and this is due to both genetic diversity and phenotypic plasticity3,4,5. For example, phenotypic transformations of tumors from proliferative to invasive are regulated by a genetic transformation termed the epithelial–mesenchymal transition (EMT) where tumor cells lose their epithelial characteristics and intercellular connections weaken. These cells acquire mesenchymal cell characteristics and the ability of independent migration and invasion6,7. EMT is regulated by many factors but the most prominent are the abnormal expression of embryonic transcription factors such as the Zn finger E-box binding homeobox 1 (ZEB1) and Zn finger E-box binding homeobox 2 (ZEB2) as well as other developmental regulators including snail (Snail1 and Snail2) and twist (Twist1 and Twist2). These gene families play a central role in the EMT process in cancer and a major outcome is the suppression of E-cadherin (ECAD) and other epithelial adhesion-related genes and cell polarity and cytoskeleton structure are altered via these and other transcriptional co-repressor and histone modifying enzyme complexes8,9.
The epsins encoded by EPN1-3 are a protein family of endocytic adapter proteins involved in clathrin-mediated endocytosis10,11. EPN1 and EPN2 are generally expressed throughout the body12 while EPN3 is restricted to gastric parietal cells under normal conditions13. EPN3 expression is significantly elevated in damaged epithelial tissues and numerous tumor tissues14. EPN3 overexpression enhances breast and glioma tumor cell metastasis and invasion by altering the EMT process. EPN3 enhances the migration and invasion of glioblastoma cells by activating the transcription factors Slug, Twist and ZEB1, but not Snail 1 or ZEB2, to induce EMT in glioma cells. And EPN3-induced partial EMT is instrumental for the transition from in situ to invasive breast carcinoma, the mechanism may be that EPN3 regulates and accelerates ECAD endocytosis on the cell membrane, activating and activating it β-catenin dependent EMT program, and, accordingly, high EPN3 levels are detected at the invasive front of human breast cancers and independently predict metastatic rather than loco-regional recurrence15,16, but the role of EPN3 in lung cancer is unclear.
In the current study, lung adenocarcinoma (LUAD) expression profile data from the TCGA (The Cancer Genome Atlas) database and immunohistochemistry were used to examine whether EPN3 expression differed between normal and lung adenocarcinoma tissues. Their findings were used to verify whether EPN3 expression differed between normal lung bronchial cell and lung cancer cell and then determine the expression relationship between EPN3 expression and EMT in lung cancer. Based on these results, to determine whether the expression of EPN3 in Adenocarcinoma of the lung cells enhances their invasive ability.
Materials and methods
EPN3 gene expression profiling and functional analysis
Gene expression and clinical data of LUAD patients were obtained from the University of Southern California (UCSC) https://xenabrowser.net/datapages/ using the Htseq-FPKM data set in TCGA-LUAD that were transformed into TPM data using R software. The Limma package in R 4.10 was used to identify differential gene expression levels between tumor and normal tissues using a setting of p < 0.05 and |log2 FC|≥ 1. LUAD patient survival data was then correlated with EPN3 expression and patient prognosis using the Surviviner and Survminer package in R that was then visualized using Kaplan Meier curves. Differentially expressed genes (DEG) were also correlated to EPN3 expression using Limma. Expression levels that grouped by expression level of EPN3 in LUAD with |log2 FC|≥ 0.585 and adjusted p values < 0.05 were used for Gene Ontology (GO) and KEGG library classifications utilizing tools from the David database https://david.ncifcrf.gov/. And KEGG data comes from www.kegg.jp/kegg/kegg1.html.
Immunohistochemistry
EPN3 protein levels in paraffin embedded sections of LUAD and normal lung tissues were examined after dewaxing and rehydrating. Antigen repair and blocking of endogenous peroxidase were carried out using hydrogen peroxide (3%) under a condition of avoiding light at room temperature. The slides were blocked with goat serum (AIFang Biologic, Hunan, China) and stained using a horseradish-peroxidase linked rabbit anti-EPN3 antibody that dilution ratio is 1:1000 (Cohesion Biosciences, Suzhou, China). The stained tissue sections were photographed by a microscope imaging system (Nikon, Japan).
Cell lines and construction of EPN3-knock down stable cell lines
The human normal bronchial cell line Beas-2b, human tumor adenocarcinoma cell line A549 and human non-small cell lung carcinoma cell line H1299 were purchased from ATCC (Manassas, VA, USA). They were cultured in DEME, Ham's F-12 K and RPMI 1640 (Gibco, Grand Island, NY, USA) with 10% fetal bovine serum (PAN, Germany) and maintained at 37 °C in a 5% CO2 atmosphere. A stable knock-down of EPN3 was constructed using the lentivirus strain Gv493 expressing a short hairpin using the Hu6-MCS-CBH-gcGFP-RES-puromycin vector (Genechem, Shanghai, China). The targeting sequence was constructed using the following oligonucleotides: 5′-AGTGGCCTTCACCGAAGTCAT-3′. Control (CTR) sequences were 5′-TTCTCCGAACGTGTCACGT-3′. According to the lentivirus user manual, stable EPN3 knock down cell lines were obtained by co-culturing cells at a concentration of 5 × 104/ml with a lentivirus titer of 1 × 108TU/ml for 24 h, followed by independent cultivation of the cell line for 48 h.
The lentivirus user manual comes from the official website of Genechem.
Cell migration and invasion assays
The wound healing assays were examined in 6 well plates (Corning, Ithaca, NY, USA) by allowing the cells to grow to confluence and were then removed in a line using a pipette tip and photographed. The cells were then incubated for 24 h and the monolayers were re-photographed17 and the extent of cell migration was assessed by comparison using Image J (https://imagej.nih.gov/ij/download.html, accessed as of April of 2022).
Transwell assays of A549 and H1299 cells was also conducted using 24-well Transwell plates (Corning) that were pre-coated with (for the purpose of testing invasion) or without (for the purpose of testing migration) matrigel-coated membrane (Corning, U.S.) and contained 3 × 105 cells/well in a serum-free basal medium in the upper chambers. Medium containing 10% serum was added to the lower chambers as chemotactic agent. The cells were allowed to migrate for a duration of 24 h. The chambers were fixed with paraformaldehyde for 30 min, followed by staining with crystal violet for 20 min18 before observing and photographing in a microscope. Cells were quantified using Image J (https://imagej.nih.gov/ij/download.html, accessed as of April of 2022).
Western blotting
RIPA lysis buffer, protease inhibitor mixture and BCA protein kits were obtained from (Beyotime, Shanghai, China). Total protein lysates were prepared using standard techniques and lysates were separated using SDS-PAGE (Epizyme Biotech, Shanghai, China). Primary antibodies used for Western blots were EPN3 (Cohesion) and Vimentin (5741), E-cadherin (3195), N-cadherin (13116), snail (3879), ZEB1 (3396) and β-Catenin (8480) antibodies were all obtained from Cell Signaling Technology (Beverly, MA, USA). Antibodies to β-actin (GB11001) and GAPDH (GB15004) were obtained from Service Bio (Wuhan, China). Secondary antibodies used were anti rabbit IgG HRP (Cell Signaling, 7074). And we used a total protein content of 50 μg in each well. The dilution ratio of the primary antibody included EPN3, Vimentin, E-cadherin, N-cadherin, snail, ZEB1and β-Catenin used in our experiment is 1:1000. The dilution ratio of the primary antibody included β-actin and GAPDH used in our experiment is 1:2000. And the dilution ratio of the secondary antibodies used in our experiment is 1:10,000.
Specific steps:
-
(1)
Total cell protein extraction
-
➀
Remove the cells to be used for protein extraction from the cell culture incubator, discard the remaining culture medium, wash the cells twice with 4 °C PBS buffer, and discard the PBS buffer.
-
➁
Add RIPA lysis solution 100 containing 1% protease inhibitor mixture (universal, 100×) to a 6 cm culture dish containing cells after cleaning μl. Lyse on ice for 30 min and gently scrape the cells on the bottom of the culture dish with a cell scraper during this period to ensure even cell lysis.
-
➂
After 30 min of lysis, take a clean 1.5 ml centrifuge tube and suck the cell lysis solution into the centrifuge tube. Use ultrasound to shake on ice at 30% power for 5 s, with an interval of 10 s, repeating 3 times.
-
➃
Centrifuge at 4 °C for 10 min at a rate of 12,000 rpm/min. After completion, take 100% of the supernatant μ Add a new pre labeled 1.5 ml centrifuge tube and add SDS-PAGE protein loading buffer to it at a volume of 1:4. Then place the centrifuge tube in a 100 °C constant temperature heater and heat for 10 min. After heating, the total protein sample is prepared and stored in a − 20 °C refrigerator.
-
➄
Take the supernatant after centrifugation in ④ and measure the protein concentration according to the instructions of BCA reagent.
-
➀
-
(2)
Immunoblotting
-
➀
Configure SDS-PAGE gel according to the instructions of PAGE gel rapid preparation kit.
-
➁
After the gel in ① is prepared, assemble it into the electrophoresis instrument and add the electrophoresis solution. According to the measured total protein concentration, add the total protein sample to the upper gel. According to 80 V constant pressure, after the protein sample runs to the lower layer of gel, change to 120 V constant pressure, and then until the protein runs to the bottom of gel.
-
➂
When the protein in ② runs to the bottom of the gel, take out the gel, cut the band with the required molecular weight according to the molecular weight shown in the protein marker, and conduct electroporation. When the electroporation is set to 250 mA constant current, and electroporate each molecular weight of the protein according to its own protein for 1 min, and transfer the protein from the SDS-PAGE gel to the PVDF membrane.
-
➃
After the electrical conversion is completed, take out the PVDF membrane with the converted protein and place it in a serum-free protein blocking solution for 30 min to block.
-
➄
After the closure is completed, incubate the corresponding protein primary antibody overnight at 4 ℃ based on the band of the target protein.
-
➅
Take out the PVDF membrane that has been incubated with primary antibody, place it in TBST, wash for 10 min, and repeat 3 times; Then, incubate the cleaned PVDF membrane strips in the secondary antibody for 1 h.
-
➆
Wash the PVDF membrane strips that have completed secondary antibody incubation with TBST again for 10 min each time, repeating 3 times.
-
➇
Use ECL developer to complete imaging detection of the cleaned strips under the gel imager.
-
➀
Western blotting grayscale values were measured by imagelab that downloaded from BIO-RED official website https://www.bio-rad.com/zh-cn/product/image-lab-software?ID=KRE6P5E8Z#fragment-6. By comparing the grayscale values of the target gene and the house-keeping gene, the expression level of the target gene in cells can be measured.
Immune cell analysis
The TIMER (Tumor IMmune Estimation Resource) database (https://cistrome.shinyapps.io/timer/) was used to analyze the correlation between EPN3 expression and expression of immune cells in patients with LUAD. CIBESORT was used to analyze the effect of EPN3 expression on immune cell infiltration in LUAD on R software 4.1 by LUAD expression profile.
Statistics
Cell migration experiments were analyzed using Image J for area calculations and cell counting. Prism 9.0 was used for t-tests.
Western blotting results were measured by imagelab and analyzed using t-tests between two groups and one-way ANOVA between ≥ 3 groups.
All results were expressed as mean ± standard deviation and all experimental data were repeated 3 times.
Ethics statement
This study does not involve animal experiments and does not require ethical review related to animal experiments. And Ethical review and approval was not required for the study on human participants in accordance with the local legislation and institutional requirements. Written informed consent for participation was not required for this study in accordance with the national legislation and the institutional requirements.
Results
Expression and prognosis of EPN3 in lung adenocarcinoma
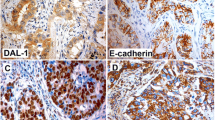
In order to verify the expression of EPN3 in lung adenocarcinoma, we compared the expression level of EPN3 in cancerous and normal tissues through the expression profile data of LUAD in the TCGA database. EPN3 levels in lung adenocarcinoma was significantly higher than that in normal tissues (Fig. 1A). The expression profile data in TCGA is also linked to patient prognosis data and we found that patients with high expression of EPN3 in lung adenocarcinoma had a worse prognosis (Fig. 1B). We also confirmed in Western blotting experiments that our cell lines A549 and H1299 expressed significantly greater levels of the protein than did the control normal bronchial cell line Beas-2b derived from a non-cancerous patient (Fig. 1C). These results suggested that EPN3 may play an important role in the occurrence and development of lung adenocarcinoma. The presence of EPN3 protein in lung adenocarcinoma was also apparent and was elevated in cancer versus normal control tissues using immunohistochemical staining (Fig. 2).
(A) EPN3 expression level between normal tissue and lung Adenocarcinoma. (B) Prognosis of LUAD patients between high EPN3 expression level group and low EPN3 expression level group. (C) EPN3 expression level between normal lung cell line (Beas-2B) and lung cancer cell line(A549 and H1299) by western blot. n = 3, p < 0.05. Quantification of bands relative to β-actin using Graphpad prism (*p < 0.05; **p < 0.005).
EPN3 is an EMT regulator
In order to verify the role of EPN3 expression in lung adenocarcinoma, we used LUAD expression profile data in the TCGA database to locate DEGs associated with epn3 expression and then analyzed them using GO and KEGG. EPN3 expression was linked to cell connections, adhesion and immune functions in patients with lung adenocarcinoma and this correlated with EMT processes involving the decline of cell connection and adhesion functions (Fig. 3). To verify a role for EPN3 in lung adenocarcinoma, we constructed EPN3 gene knockdowns in A549 and H1299 cells using shRNAs and examined EPN3 expression using western blotting (Fig. 4A,B). We compared Epn3 patterns to those of other markers known to be involved in EMT. E-cadherin (ECAD) expression declines with EMT onset while N-cadherin (NCAD) is increased19,20,21. We found that E-cadherin was increased in A549 EPN3-knockdown (KD) cells while N-cadherin and vimentin (VIM) levels decreased (Fig. 4C,E,G). In H1299 cells, E-cadherin was absent (Fig. 4F) but the mesenchymal markers N-cadherin and vimentin decreased significantly (Fig. 4D,H). These data indicated that EPN3 knockdown inhibits the expression of known EMT markers suggesting EPN3 expression may be required for EMT.
Analysis of DEGs from LUAD patient data relative to EPN3 grouping by expression level. (A) Analyzing the cellular functions involved in EPN3 in lung adenocarcinoma cells by GO enrichment analysis. (B) Analyzing the signaling pathways involved in the regulation of EPN3 in lung adenocarcinoma cells by KEGG pathway analysis.
Western blot analysis of EPN3-knockdown (KD) cell lines and expression of EMT markers. The differences in expression level of EPN3 between normal cell, control group and EPN3-KD group in A549 and H1299 (A,B). The differences in expression level of NCAD, ECAD and VIM between control group and EPN3-KD group in A549 (C,E,G). The differences in expression level of NCAD, ECAD and VIM between control group and EPN3-KD group in H1299 (D,F,H). NCAD, N-cadherin; ECAD, E-cadherin; VIM, vimentin. n = 3, *p < 0.05, **p < 0.01, ***p < 0.001.
EPN3-KD decreased the ability of invasion and metastasis of lung adenocarcinoma cells
We further examined whether EPN3 expression was also reflected in its ability to direct cellular migration, a key component of EMT and metastasis. In EPN3-KD cells, migration and invasion ability for both A549 and H1299 cells was decreased using in vitro assays (Figs. 5 and 6).
Mechanism of EMT mediated by EPN3 expression
We further explored the mechanism of the effects of EPN3 on EMT by examining the expression of EMT-associated transcription factors. ZEB1 and Snail protein expression were decreased in EPN3-KD cells and this was consistent with decreased levels of β-catenin which is a primary regulator of the Wnt signaling pathway (Fig. 7).
Western blot analysis of protein expression of key EMT-related factors in EPN3-KD cell line. The differences in expression level of ZEB1, β-catenin and Snail between control group and EPN3-KD group in A549 (A). The differences in expression level of ZEB1, β-catenin and Snail between control group and EPN3-KD group in H1299 (B). n = 3, *p < 0.05, **p < 0.01.
Effects of EPN3 expression on immune infiltration in lung adenocarcinoma
We further explored the effect of EPN3 expression on immune function in lung adenocarcinoma using TIMER database analysis of immune cells expression in lung adenocarcinoma patients with EPN3 expression. EPN3 expression was positively correlated with tumor purity, i.e., the percentage of cancer cells in a tumor tissue. In contrast, the presence of immune cells including B and CD8-positive T cells, neutrophils and dendritic cells was negatively correlated with EPN3 expression. Infiltration of B and T cells was more obvious in the low-risk group while regulatory T cells and M2 macrophages were elevated in the high-risk groups (Fig. 8A,B). These data suggested that EPN3 expression in lung adenocarcinoma may also be related to the immune escape of tumor cells and the deconstruction of the immune microenvironment during metastasis development.
Discussion
For this study, we explored the high expression of EPN3 in lung adenocarcinoma, and analyzed its possible negative impact on the prognosis of lung adenocarcinoma patients. We also explored that EPN3 may affect the development of lung adenocarcinoma by mediating and regulating the occurrence of cell EMT in lung adenocarcinoma. We utilized the expression profile data of lung adenocarcinoma patients in the TCGA database and found that EPN3 expression in lung adenocarcinoma patients increased significantly compared with matched normal tissues. These data allowed the prognosis of patients with lung adenocarcinoma to be predicted by EPN3 gene expression. We found a worse prognosis for patients with high expression of EPN3. These patients also presented higher levels of EPN3 protein in lung adenocarcinoma tissues. EPN3 gene knockdowns reduced invasion and migration of A549 and H1299 cells in vitro. These results implicate EPN3 as an oncogene in lung adenocarcinoma. Inhibiting the expression of EPN3 can significantly reduce the spread to surrounding tissues and migration of lung adenocarcinoma cells.
We further found a tight linkage of EPN3 gene expression to expression of cell adhesion molecules. EMT results in the loss of epithelial characteristics and cell–cell connections and adhesion6,7 and our data suggested that EPN3 is linked to EMT. In glioma and breast cancer, increasing EPN3 expression enhanced metastasis and invasion ability of tumor cells by inducing EMT15,16. Expression of the EPN3 gene in lung adenocarcinoma may also be related to the EMT of lung adenocarcinoma cells that thereby enhance tumor cell invasion and migration. EPN3 knockdowns in A549 and H1299 cells resulted in decreased expression of E-cadherin, a primary mediator of cell connection and adhesion and is an inhibitory protein for the invasion and growth of a variety of epithelial tumors19,20,21. In A549 cells, expression of E-cadherin increased with the inhibition of EPN3 expression although in H1299 cells, E-cadherin was not expressed. In contrast, elevation of EPN3 was positively correlated with N-cadherin expression which is abnormally up-regulated in metastatic tumor cells. Interestingly, N-cadherin synergizes with the FGF receptor that results in MMP-9 over-expression and tumor cell invasion that are all related to the EMT21. We found that in A549 and H1299 cells N-cadherin decreased significantly with the inhibition of EPN3 expression. Another EMT related protein is vimentin that is a major component of intermediate filaments and an EMT marker22. Vimentin is overexpressed in lung cancer23,24 and its overexpression is closely related to tumor growth and metastasis25,26,27. We found that vimentin protein expression decreased significantly in EPN3-KD cells consistent with the link to EMT.
We also found that ZEB1 protein expression was decreased significantly with the inhibition of EPN3 expression in A549 and H1299 cells. ZEB family proteins inhibit E-cadherin expression28 and these functions were consistent with our results. Snail is a zinc finger transcription factor that can also inhibit E-cadherin transcription and acts to mediate the occurrence of EMT resulting in local tumor cell infiltration29,30,31,32,33. We found that Snail protein expression was decreased significantly in EPN3-KD cells. β-Catenin is a key effector molecule in Wnt signaling pathway34 that activates ZEB1 and induces the EMT to enhance the migration and invasion of tumor cells35. EPN3 belongs to the epsin family of adaptors that link PM proteins with the endocytic machinery. EPN3 also possess lipid-binding and membrane-bending ability. At present, studies have shown that EPN3 in breast cancer may regulate key EMT proteins such as ECAD, β-Catenin, to induce the occurrence of cell EMT process16. This may suggest that EPN3 mediated regulation β-Catenin induced EMT may occur on the cell membrane in lung adenocarcinoma cells. Our results using lung adenocarcinoma cell lines A549 and H1299 indicated β-catenin protein expression decreased significantly after the expression of EPN3 was inhibited. Therefore, when EPN3 expression is inhibited, the activation of WNT signal pathway mediating EMT is weakened, and the expression of transcription factors ZEB1 and snail that activate EMT process is significantly decreased. These indicated that EPN3 may induce EMT process of tumor cells through Wnt pathway regulation and regulation of transcription factors that mediate EMT activation in lung adenocarcinoma cells.
Our GO enrichment analysis indicated that EPN3 may alter tumor immune responses in lung cancer. According to TIMER assays, EPN3 expression was positively correlated with the purity of the tumor; i.e., higher EPN3 levels indicated a greater proportion of tumor cells versus stromal cells. In contrast, B and CD8-positive T cells, neutrophils and dendritic cells involved in anti-tumor effect were negatively correlated indicating that EPN3 can inhibit the killing effect of immune cells on tumors in lung adenocarcinoma and thereby promote tumor growth. According to the CIBESORT results also suggested that the high-risk patient group displayed decreased humoral immune response (B cell infiltration) often represented a worse prognosis36,37. However, the decrease of B cell content indicated that EPN3 can inhibit B cell infiltration and this is more conducive to tumor growth. Additionally, EPN3 regulated T cell content in the high-risk group and a decreased T cell recruitment promotes tumor growth. The M0 macrophage content increased in the high-risk group; however, it shows more M0 macrophages to be polarized to M2 subtype. M2 macrophages are immunosuppressive and allows tumor growth38,39,40,41. EPN3 upregulation may inhibit the immune response to lung adenocarcinoma cells to promote proliferation and metastasis.
EPN3 expression in patients with lung adenocarcinoma occurred most likely via activation of the WNT/β-Catenin signaling pathway regulates ZEB1 and Snail transcription factors that promote EMT. Our bioinformatics analyses indicated that in lung adenocarcinoma, EPN3 expression is linked to the immune response in tumor tissues and elevation of its expression may be immunosuppressive.
However, our study still has significant limitations regarding the role of EPN3 in lung adenocarcinoma. In this study, the impact of EPN3 expression on the clinical survival and prognosis of lung adenocarcinoma patients lacks systematic clinical prospective research support. Further research is needed on the molecular mechanism of EPN3 mediated EMT in lung adenocarcinoma and the expression of EPN3 may also regulate the growth, invasion, immune regulation and other processes of lung adenocarcinoma cells through other pathways, and further research is needed. These effects require further experimental verification and is planned for the near future in our laboratory.
Data availability
The original contributions presented in the study are included in the article. Gene expression and clinical data of LUAD patients were obtained from the University of Southern California https://xenabrowser.net/datapages/ using the Htseq-FPKM data set in TCGA-LUAD that were transformed into TPM data using R software. Gene Ontology (GO) and KEGG library classifications utilizing tools from the David database https://david.ncifcrf.gov/. And we have obtained KEGG usage authorization. And KEGG data comes from www.kegg.jp/kegg/kegg1.html. The TIMER database (https://cistrome.shinyapps.io/timer/) was used to analyze the correlation between EPN3 expression and expression of immune cells in patients with LUAD. Further inquiries can be directed to the corresponding author.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71(3), 209–249. https://doi.org/10.3322/caac.21660 (2021).
Travis, W. D., Brambilla, E., Burke, A. P., Marx, A. & Nicholson, A. G. Introduction to the 2015 world health organization classification of tumors of the lung, pleura, thymus, and heart. J. Thorac. Oncol. 10(9), 1240–1242. https://doi.org/10.1097/JTO.0000000000000663 (2015).
Mehlen, P. & Puisieux, A. Metastasis: A question of life or death. Nat. Rev. Cancer 6(6), 449–458 (2006).
Ling, S. et al. Extremely high genetic diversity in a single tumor points to prevalence of non-Darwinian cell evolution. Proc. Natl. Acad. Sci. U.S.A. 112(47), E6496–E6505. https://doi.org/10.1073/pnas.1519556112 (2015).
Williams, M. J., Werner, B., Barnes, C. P., Graham, T. A. & Sottoriva, A. Identification of neutral tumor evolution across cancer types. Nat. Genet. 48(3), 238–244. https://doi.org/10.1038/ng.3489 (2016).
Nieto, M. A., Huang, R.Y.-J., Jackson, R. A. & Thiery, J. P. EMT: 2016. Cell 166(1), 21–45. https://doi.org/10.1016/j.cell.2016.06.028 (2016).
Lambert, A. W., Pattabiraman, D. R. & Weinberg, R. A. Emerging biological principles of metastasis. Cell 168(4), 670–691. https://doi.org/10.1016/j.cell.2016.11.037 (2017).
Thiery, J. P., Acloque, H., Huang, R. Y. J. & Nieto, M. A. Epithelial–mesenchymal transitions in development and disease. Cell 139(5), 871–890. https://doi.org/10.1016/j.cell.2009.11.007 (2009).
Skrypek, N., Goossens, S., De Smedt, E., Vandamme, N. & Berx, G. Epithelial-to-mesenchymal transition: Epigenetic reprogramming driving cellular plasticity. Trends Genet. TIG 33(12), 943–959. https://doi.org/10.1016/j.tig.2017.08.004 (2017).
Xie, X., Cho, B. & Fischer, J. A. Drosophila Epsin’s role in Notch ligand cells requires three Epsin protein functions: The lipid binding function of the ENTH domain, a single Ubiquitin interaction motif, and a subset of the C-terminal protein binding modules. Dev. Biol. 363(2), 399–412. https://doi.org/10.1016/j.ydbio.2012.01.004 (2012).
Chen, H. et al. Epsin is an EH-domain-binding protein implicated in clathrin-mediated endocytosis. Nature 394(6695), 793–797 (1998).
Rosenthal, J. A. et al. The epsins define a family of proteins that interact with components of the clathrin coat and contain a new protein module. J. Biol. Chem. 274(48), 33959–33965 (1999).
Ko, G. et al. Selective high-level expression of epsin 3 in gastric parietal cells, where it is localized at endocytic sites of apical canaliculi. Proc. Natl. Acad. Sci. U. S. A. 107(50), 21511–21516. https://doi.org/10.1073/pnas.1016390107 (2010).
Spradling, K. D., McDaniel, A. E., Lohi, J. & Pilcher, B. K. Epsin 3 is a novel extracellular matrix-induced transcript specific to wounded epithelia. J. Biol. Chem. 276(31), 29257–29267 (2001).
Wang, Y. et al. Overexpression of Epsin 3 enhances migration and invasion of glioma cells by inducing epithelial–mesenchymal transition. Oncol. Rep. 40(5), 3049–3059. https://doi.org/10.3892/or.2018.6691 (2018).
Schiano Lomoriello, I. et al. A self-sustaining endocytic-based loop promotes breast cancer plasticity leading to aggressiveness and pro-metastatic behavior. Nat. Commun. 11(1), 3020. https://doi.org/10.1038/s41467-020-16836-y (2020).
Zhong, Y. et al. Long non-coding RNA AFAP1-AS1 accelerates lung cancer cells migration and invasion by interacting with SNIP1 to upregulate c-Myc. Signal Transduct. Target. Ther. 6(1), 240. https://doi.org/10.1038/s41392-021-00562-y (2021).
Zan, X. et al. COL8A1 promotes NSCLC progression through IFIT1/IFIT3-mediated EGFR activation. Front. Oncol. 12, 707525. https://doi.org/10.3389/fonc.2022.707525 (2022).
Wheelock, M. J. & Johnson, K. R. Cadherins as modulators of cellular phenotype. Annu. Rev. Cell Dev. Biol. 19, 207–235 (2003).
Christofori, G. Changing neighbours, changing behaviour: Cell adhesion molecule-mediated signalling during tumour progression. EMBO J. 22(10), 2318–2323 (2003).
Hazan, R. B., Qiao, R., Keren, R., Badano, I. & Suyama, K. Cadherin switch in tumor progression. Ann. N. Y. Acad. Sci. 1014, 155–163 (2004).
Gilles, C. et al. Vimentin contributes to human mammary epithelial cell migration. J. Cell Sci. 112(Pt 24), 4615–4625 (1999).
Al-Saad, S. et al. The prognostic impact of NF-kappaB p105, vimentin, E-cadherin and Par6 expression in epithelial and stromal compartment in non-small-cell lung cancer. Br. J. Cancer 99(9), 1476–1483. https://doi.org/10.1038/sj.bjc.6604713 (2008).
Upton, M. P. et al. Expression of vimentin in surgically resected adenocarcinomas and large cell carcinomas of lung. Am. J. Surg. Pathol. 10(8), 560–567 (1986).
Sethi, S., Macoska, J., Chen, W. & Sarkar, F. H. Molecular signature of epithelial–mesenchymal transition (EMT) in human prostate cancer bone metastasis. Am. J. Transl. Res. 3(1), 90–99 (2010).
Jin, H. et al. Vimentin expression of esophageal squamous cell carcinoma and its aggressive potential for lymph node metastasis. Biomed. Res. (Tokyo, Japan) 31(2), 105–112 (2010).
Hu, L. et al. Association of Vimentin overexpression and hepatocellular carcinoma metastasis. Oncogene 23(1), 298–302 (2004).
Vandewalle, C., Van Roy, F. & Berx, G. The role of the ZEB family of transcription factors in development and disease. Cell. Mol. Life Sci. CMLS 66(5), 773–787. https://doi.org/10.1007/s00018-008-8465-8 (2009).
Barrallo-Gimeno, A. & Nieto, M. A. The Snail genes as inducers of cell movement and survival: Implications in development and cancer. Development (Cambridge, England) 132(14), 3151–3161 (2005).
Cano, A. et al. The transcription factor snail controls epithelial–mesenchymal transitions by repressing E-cadherin expression. Nat. Cell Biol. 2(2), 76–83 (2000).
Batlle, E. et al. The transcription factor snail is a repressor of E-cadherin gene expression in epithelial tumour cells. Nat. Cell Biol. 2(2), 84–89 (2000).
Yokoyama, K. et al. Increased invasion and matrix metalloproteinase-2 expression by Snail-induced mesenchymal transition in squamous cell carcinomas. Int. J. Oncol. 22(4), 891–898 (2003).
Yokoyama, K. et al. Reverse correlation of E-cadherin and snail expression in oral squamous cell carcinoma cells in vitro. Oral Oncol. 37(1), 65–71 (2001).
Cadigan, K. M. & Nusse, R. Wnt signaling: A common theme in animal development. Genes Dev. 11(24), 3286–3305 (1997).
Kahlert, U. D. et al. Activation of canonical WNT/β-catenin signaling enhances in vitro motility of glioblastoma cells by activation of ZEB1 and other activators of epithelial-to-mesenchymal transition. Cancer Lett. 325(1), 42–53. https://doi.org/10.1016/j.canlet.2012.05.024 (2012).
Germain, C. et al. Presence of B cells in tertiary lymphoid structures is associated with a protective immunity in patients with lung cancer. Am. J. Respir. Crit. Care Med. 189(7), 832–844. https://doi.org/10.1164/rccm.201309-1611OC (2014).
Lohr, M. et al. The prognostic relevance of tumour-infiltrating plasma cells and immunoglobulin kappa C indicates an important role of the humoral immune response in non-small cell lung cancer. Cancer Lett. 333(2), 222–228. https://doi.org/10.1016/j.canlet.2013.01.036 (2013).
Biswas, S. K. & Mantovani, A. Macrophage plasticity and interaction with lymphocyte subsets: Cancer as a paradigm. Nat. Immunol. 11(10), 889–896. https://doi.org/10.1038/ni.1937 (2010).
Edin, S., Wikberg, M. L., Rutegård, J., Oldenborg, P.-A. & Palmqvist, R. Phenotypic skewing of macrophages in vitro by secreted factors from colorectal cancer cells. PLoS One 8(9), e74982. https://doi.org/10.1371/journal.pone.0074982 (2013).
Gordon, S. & Martinez, F. O. Alternative activation of macrophages: Mechanism and functions. Immunity 32(5), 593–604. https://doi.org/10.1016/j.immuni.2010.05.007 (2010).
Mantovani, A. et al. The chemokine system in diverse forms of macrophage activation and polarization. Trends Immunol. 25(12), 677–686 (2004).
Author information
Authors and Affiliations
Contributions
B.Z. and Y.L. initiated and designed this work. Y.L. undertook most of the experiments. P.Z., Y.L., Z.W., G.T., J.Z. analyzed the data and drew the chart and figures. Y.L. wrote the original manuscript.
Corresponding author
Ethics declarations
Competing interests
This study is supported by research funding from the corresponding author Bing Zhu. And the corresponding author Zhu Bing participated in guiding the design of the research. All authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Y., Zhang, P., Tang, G. et al. Lowering expression of Epsin-3 inhibits migration and invasion of lung adenocarcinoma cells by inhibiting the epithelial–mesenchymal transition. Sci Rep 14, 17069 (2024). https://doi.org/10.1038/s41598-024-68193-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-68193-1
- Springer Nature Limited