Abstract
The gut microbiome is a significant factor in the pathophysiology of ulcerative colitis (UC), prompting investigations into the use of probiotic therapies to counter gastrointestinal inflammation. However, while much attention has been given to the therapeutic potential of microbes at the species and strain level, the discovery and application of their metabolic products may offer more precise and controlled solutions in battling disease. In this work, we examined the therapeutic potential of indole lactic acid (ILA) to alleviate inflammation in a murine model of colitis. A previously constructed ILA-producing Escherichia coli Nissle 1917 strain (EcN aldh) and its isogenic non-ILA producing counterpart (EcN) were studied in a murine model of Dextran Sodium Sulfate (DSS) induced colitis. The colitic animals suffered from severe colitic symptoms, with no differentiation between the groups in body weight loss and disease activity index. However, three days after cessation of DSS treatment the EcN aldh–treated mice showed signs of reduced intestinal inflammation, as manifested by lower concentrations of fecal lipocalin-2. Additionally, expression analysis of the inflamed tissue revealed distinct effects of the EcN aldh strain on proteins associated with intestinal health, such as TFF3, occludin and IL-1β expression. These results show no impact of EcN or EcN aldh on acute DSS-induced colitis, but suggest that in particular EcN aldh may assist recovery from intestinal inflammation.
Similar content being viewed by others
Introduction
Ulcerative colitis (UC) is one of the principal types of inflammatory bowel disease (IBD), a chronic condition that manifests as recurrent mucosal inflammation in the colon and rectum1. Its etiology is not clearly determined. Genetic background and mucosal immune dysregulation play central roles, in concert with a number of environmental factors, with the gut microbiota being a key factor2. Current first-line therapies involve anti-inflammatory drugs, such as corticosteroids, and in cases progressing from mild-to-severe disease, biological agents that intricately target immunological actors such as TNF or integrin α4β7 are commonly used3. However, a clinical response is not achieved in all patients and their prolonged use may cause significant adverse effects3,4,5. Hence, there is a need for alternative modalities with high clinical efficiency, fewer side effects and prolonged remission periods.
Gut bacterial species and their metabolites have been associated with disease outcomes6,7. Thus, alternative therapies, based on gut microbiome manipulations, are also being explored. Microbiota therapies against UC have shown promise towards disease cessation and remission8, although clinical studies on probiotics and/or fecal microbiome transplantation (FMT) treatments exhibit high variability in remission rates9,10. Since the exact molecular mechanisms of action of probiotics and FMTs are still unknown, it is difficult to standardize the therapies and develop reproducible interventions.
One solution is to combine knowledge on gut metabolites and synthetic biology to engineer advanced microbial therapeutics (AMTs) that can colonize the gastrointestinal tract and express specific bacterial metabolites in situ11. Localized delivery minimizes pharmacological loss of the active compound and reduces systemic exposure. Furthermore, engineered probiotic strains, such as Escherichia coli Nissle 1917 (EcN), allow for the development of complex genetic circuits that control strain colonization12, regulate the expression of specific metabolites13 and enable biocontainment of the strains14. This makes AMTs promising alternatives to both conventional therapies and traditional probiotic approaches. Especially since natural producers are typically not efficient colonizers of the colon15,16 and the production of the respective metabolites cannot be tightly controlled and, subsequently, lead to standardized therapies, as it is dependent on the composition of the microbiome17.
Several studies have used AMTs to alleviate colitis symptoms18,19,20, but so far none has employed microbial metabolites. Since such metabolites inherently interact with the intestine, their use as therapeutic agents may improve therapeutic specificity and potency with marginal side effects21. One promising candidate is indole lactic acid (ILA). Cumulative evidence suggests that ILA and ILA-producing species protect the intestinal barrier and decrease inflammation22,23,24,25. In addition, EcN has been reported to be a potent anti-colitic agent, whose properties are comparable to the golden standard of UC treatment, mesalazine26,27. EcN is known to interact with both the gut microbiota and the host, leading to its intestinal inflammation amelioration properties. Mechanisms of actions include direct and indirect antagonistic effects against enteroinvasive pathogens, reinforcement of the gut barrier, and immune-modulatory and anti-inflammatory effects28,29.
Previously, we developed an engineered EcN, namely EcN aldh, which boosts ILA production in the gut microbiome30. In the present study, we investigated the therapeutic potential of EcN aldh against DSS-induced colitis and epithelial recovery in a preclinical mouse model. We present an AMT that, upon colonization, increases systemic ILA, but does not alter acute disease progression. However, three days after cessation of DSS-treatment, EcN aldh mice showed lower intestinal inflammation levels and improvement in gene expression markers of intestinal epithelial health suggested an enhanced healing process in the presence of the AMT.
Results
Colitic symptoms of the DSS-treated mice
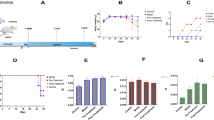
The study comprised seven days of DSS treatment with orally administered AMTs on Day 4 and a post-treatment period of three days (Fig. 1A). All DSS-treated animals exhibited similar symptoms, with drastic weight loss occurring from Day 5 onwards (Fig. 1B). The severity of the colitis symptoms was further displayed by the Disease Activity Index calculations, with no significant differences between the DSS treated groups (Supplementary Fig. S1). Colon length measurements indicated similar disease severity among the colitic mice groups, with the non-colitic colons being significantly longer (Fig. 1C). All colitic mice had slightly enlarged spleens, compared to the untreated group, only reaching statistical significance in the EcN aldh group (Fig. 1D).
DSS-induced colitis: macroscopic observations and markers of disease. (A) Experimental setup. (B) Body weight loss calculated relatively to the AMT administration day (Day 4). Statistical analysis performed with the mixed effects model (REML), uncorrected Fisher’s LSD. (C) Colon length measurements and (D) spleen weight measurements on termination day. Statistical significance was calculated by Kruskal Wallis, uncorrected Dunn test, and by ordinary one-way ANOVA, Fisher’s LSD test, respectively. (E) Fecal lcn-2 measurements on selective days of the experiment. Statistical analysis was performed with a two-way ANOVA, Tukey correction (adjusted p-values are depicted on the illustration). All data is shown as the mean ± SEM of the biological replicates from each group (n = 10, 10, 9, 8 in the non-colitic, DSS, EcN and EcN aldh groups respectively). *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Lipocalin-2 (lcn-2), an acute-phase protein of the innate immune response, exhibits bacteriostatic activity31 and is also involved in the regulation of systemic inflammation32,33 and mucosal repair34. Its expression is increased distinctly in inflammatory diseases35,36,37, and it is considered a great biomarker for disease progression. Fecal lcn-2, in particular, is an excellent real-time biomarker of intestinal inflammation considering its stability, sensitivity, and broad dynamic range to detect low-grade but also severe colitic symptoms38,39. Fecal lcn-2 was increased in all colitic groups by Day 10 as compared to the Day 0 fecal concentration (Fig. 1E). While no significant differences were detected on Day 0 among the groups (Kruskal Wallis, uncorrected Dunn test), on Day 10 administration of EcN aldh resulted in significantly lower levels of fecal lcn-2 compared to the other colitic groups (Fig. 1E), suggesting a beneficial effect of the strain on intestinal inflammation. The DSS and EcN groups displayed approximately a 40-fold increase compared to the control group (p < 0.00001, both comparisons), while EcN aldh exhibited a less pronounced tenfold increase in fecal lcn-2, which did not reach statistical significance (p = 0.118). The relatively high fecal lcn-2 levels observed in the control group is likely attributed to the mild inflammation resulting from streptomycin administration40. Serum lcn-2 as a general marker of pathophysiological conditions41, did not differ among the groups, although all groups experienced an increase by Day 9 (Supplementary Fig. S1B).
Histological analysis of colonic tissue
The histopathological damage scores suggest no effect of the engineered strains on the inflammatory cell infiltrate compared to the DSS group, and a significant increase of the infiltrate in the DSS-treated mice compared to the non-colitic mice (Fig. 2A, left panel). DSS and EcN groups showed significantly higher scores in epithelial and mucosal damages compared to the non-colitic group, while this was not the case for the EcN aldh group, which scored lower than DSS and EcN (Fig. 2A, right panel), suggesting a slight impact of the strain. It should be noted that the non-colitic mice showed signs of mild inflammation, a known effect of streptomycin administration40, but no signs of architectural damage were observed in this group (Fig. 2A,B).
Histopathological analysis: influence of EcN aldh on inflammatory cell infiltrate and intestinal architecture. (A) Histological scores of colon sections. Data was generated by the histopathological analysis of 7 mice from each colitic group and 4 from the non-colitic group. The plots depict the mean ± SEM in each group. Statistical significance was calculated by Kruskal–Wallis, uncorrected Dunn’s test (inflammatory cell filtrate scores) and one-way ANOVA, uncorrected Dunn’s test (intestinal architecture scores). *p < 0.05 (B) Representative images of H&E-stained colon sections. Histological features: a: mucosal immune cell infiltration, a1: mucosal and submucosal immune cell infiltration, b: goblet cells loss, c: focal hyperplasia, d: ulcer formation.
No major differentiation in the type of artifacts was observed among the groups (Fig. 2B). We detected inactive ulcers, focal hyperplasia, depletion of goblet cells and the presence of immune cell infiltrates that occasionally reached the submucosa in all colitic tissues processed, confirming that the mice suffered from moderate to severe symptoms. The presence of granulation tissue and process of restitution of cell lining suggest that the intestinal epithelium of the three colitic groups were in process of healing by Day 10. Overall, mice treated with EcN aldh showed a lower percentage of tissue architectural damages per histological cut (data not shown).
AMTs and their impact on the gut metabolome
AMT colonization was similar for both strains with approx. 108 and 109 CFUs/g of gut content in the cecum and colon, respectively (Supplementary Fig. S2A). Antibiotic plates that selected only for the EcN and not the EcN aldh strain, confirmed that no substantial plasmid loss took place during the experiment (Supplementary Fig. S2B).
As EcN aldh influences the concentration of all three aromatic lactic acids (aLAs)30, besides ILA, we measured the concentration of phenyllactic acid (PLA) and 4-hydroxy-phenyllactic acid (4HPLA) in fecal samples from the mice. Before treatment, the average concentration of ILA was 1 ± 0.5 nmol/g, PLA 124.7 ± 30.2 nmol/g and 4HPLA 5 ± 0.9 nmol/g in the fecal samples. Introduction of streptomycin and DSS caused an increased ILA and 4HPLA and a decreased PLA concentration (REML model, Tukey test correction analysis, data not shown). When compared to the non-colitic and DSS animals, the AMTs appeared to only slightly affect the absolute values of the aLAs and no significant differences were observed between EcN and EcN aldh groups (Supplementary Fig. S3A-C).
To account for any individualized differences that may influence aLA production, we normalized the daily measurements for each mouse by their Day 0 concentration. AMTs increased ILA and 4HPLA concentrations, with higher ILA increase in EcN aldh and higher 4HPLA increase in EcN (Fig. 3A,C). However, none of these comparisons reached statistical significance. PLA concentrations did not increase by AMT administration (Fig. 3B). These results should however be interpreted with caution, since the fecal samples that were collected on the last day were too few to be considered a representative sample (Supplementary Table S1). The aAAs concentration decreased during DSS administration and increased during recovery (Supplementary Fig. S4A-C).
Impact of the AMT strains on fecal and serum aLAs. Fold change of (A) ILA, (B) PLA and (C) 4HPLA concentration in the fecal samples from D0 to D10. Values were calculated for each mouse individually. The bars depict the mean ± SEM of each group (n = 7, 9, 6, 4 in the non-colitic, DSS, EcN and EcN aldh groups respectively). Statistical significance was calculated with the Kruskal–Wallis method, uncorrected Dunn’s test. (D) ILA, (E) PLA and (F) 4HPLA concentrations in serum samples on day 9. The graphs illustrate the mean ± SD of the biological replicates (n = 8, 9, 7, 6 in the non-colitic, DSS, EcN and EcN aldh groups respectively). Each symbol represents an individual mouse. Statistical significance was calculated with the Kruskal–Wallis method, uncorrected Dunn’s test. *p < 0.05, **p < 0.01, ***p < 0.001.
EcN aldh affects the serum ILA levels
Higher serum ILA concentrations were observed in the EcN aldh treated mice on Day 9, its concentration was 1.6- and 1.7-fold higher in the EcN aldh than in the DSS and EcN group respectively (Fig. 3D). Although serum PLA was only detected in traces, the EcN aldh mice trended towards higher serum concentrations in comparison to the EcN mice (p = 0.08) (Fig. 3E). Similarly, serum 4HPLA exhibited a trend towards higher concentration in the EcN aldh mice when compared to the EcN (p = 0.08) (Fig. 3F). Analysis of serum samples on day 3 showed no previous differences among the groups (Supplementary Fig. S5A). Serum concentrations of tryptophan and phenylalanine on Day 9 appears elevated in the EcN aldh mice, although the effect seems to be driven by a single mouse (Supplementary Fig. S5B). No differences in serum tyrosine concentration were observed among the colitic animals (Supplementary Fig. S5B).
AMTs influences expression of intestinal epithelial health markers
The impact of AMTs was further examined by relative gene expression analysis on colonic tissue from the mice from the last day of the experiment and several effects on gene expression were observed (Fig. 4). Compared to the DSS and EcN, EcN aldh markedly upregulated Trefoil factor 3 (TFF3), a peptide with anti-apoptotic properties that is essential for epithelial repair42, suggesting a direct effect of ILA on TFF3 expression (Fig. 4A). Occludin (OCL), a protein integral to tight junction stability43 and gut barrier function44 was upregulated in both groups that received the AMTs, compared to DSS, although the effect was larger in the EcN aldh group (Fig. 4B). Expression of mucin 3 (Muc3), a glycoprotein responsible for epithelial restitution45, was mainly upregulated in EcN (Fig. 4C).
Impact of EcN aldh on molecular markers of intestinal epithelial recovery and inflammation. Relative expression of proteins related to (A–C) intestinal epithelial health (TFF3, OCL, Muc3), and (D–H) immunological markers (IL-1β, iCAM, iNOS, RIPK1, Foxp3, AhR, TRL9, TGF-β) in colonic tissue. The RT-qPCR results were normalized to the expression of NONO. The gene expression is shown relative to the non-colitic animals. Bars represent the mean ± SEM of the biological replicates (n = 9, 7, 7, 7; non-colitic, DSS, EcN and EcN aldh groups respectively). Statistical analysis was performed with one-way ANOVA, Fisher’s LSD test, except for IL-1β, iCAM-1, iNOS and TLR9 data that was Kruskal–Wallis, uncorrected Dunn’s test was used. Asterisks (*) on top of the bars note the p values in comparison to the non-colitic mice. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Analysis of mRNA expression of markers of intestinal inflammation revealed an effect of EcN aldh on the inflammatory response as well. IL-1β, a pro-inflammatory cytokine with essential roles in colonic inflammation46, was significantly upregulated in the DSS and EcN groups, with mean fold change of 1041 and 540 respectively, while EcN aldh mice exhibited a mean fold change of 63 when compared to the non-colitic group (Fig. 4D). Similarly, the intercellular adhesion molecule 1 (ICAM-1) expression, a molecule whose upregulation strongly correlates with increased DSS-colitis symptoms47,48, was significantly increased in the DSS and EcN groups, compared to the EcN aldh group (Fig. 4E). The expression of the inducible nitric oxide synthase (iNOS), an enzyme linked to aggravated inflammation due to excessive production of NO49 was also significantly upregulated only in the DSS and EcN groups (Fig. 4F). A similar trend was observed in Foxp3 expression, an anti-inflammatory protein when located in the colonic epithelium50, although the results were not statistically significant (Fig. 4G).
The receptor-interacting serine/threonine-protein kinase 1 (RIPK1), a protein integral to cell death and survival51,52, was significantly upregulated in both AMT-receiving groups compared to both non-colitic and DSS groups (Fig. 4H). The Aryl hydrocarbon Receptor (AhR), toll-like receptor 9 (TLR-9) and Tumor Growth Factor-beta (TGF-b) mRNAs had downregulation trends compared to the non-colitic group (Fig. 4I, J, K). All are implicated in immunological protection pathways in the gastrointestinal lumen53,54,55, and only TLR9 was significantly downregulated in the EcN aldh group, compared to the EcN group.
Effect of the AMTs on bacterial diversity and community composition
To evaluate the impact of the strains on the gut microbiota composition, we performed 16S rRNA gene analysis on fecal samples from pre-, during and post-treatment days. Before treatment, an average of 228 ± 15 ASVs was detected per mouse and no differences in alpha diversity were observed among the groups. The introduction of streptomycin and DSS decreased the observed richness by 75%, in non-colitic mice, and 55% in all DSS-treated mice from Day 2. All colitic mice exhibited a significantly higher observed richness than the non-colitic and no significant differences were observed between the colitic groups (Fig. 5A). Two days after DSS cessation (Day 10) all groups had similar observed richness (mean = 87 ± 6 ASVs). In terms of beta diversity, group as a variable explains most of the variation observed among all the groups. Depending on the day it explains 18.4 to 30.9% and 11.2 to 39.2% of the variation based on weighted UniFrac distances and Bray–Curtis dissimilarities respectively (Supplementary Table S2). However, the group variable did not significantly explain the variation observed among the colitic groups (Supplementary Table S3, Supplementary Fig. S6) and thus the differences observed across groups were largely explained by the DSS-treatment. The microbiota genus composition of all experimental groups on selected days are depicted in Fig. 5(B).
Compositional changes resulting from the AMT administration. (A) Observed richness of the four experimental groups. Box plots depict the median, minimum, and maximum values from the independent biological replicates on the selected days. (B) The microbiota composition of the experimental groups based on the detected genera on selected days of the experiment. (Relative abundance cutoff 0.1%) *q < 0.05, **q < 0.01, ***q < 0.001, ****q < 0.0001.
To further assess the AMT’s impact, we used LIMMA to determine differentially abundant phyla and genera among the colitic groups, including information about sampling day and individual animals. In the phylum level, Proteobacteria (q = 0.002) and Verrucomicrobia (q = 0.002) abundance was significantly elevated on days 8 and 10, in the AMT-receiving groups (data not shown). From the 129 genera detected in total, six were significantly different in their relative abundance, namely Escherichia/Shigella (q < 0.0001), Adlercreutzia (q = 0.006), Enteroscipio (q = 0.006), unclassified Eggerthellaceae (q = 0.006), Enterococcus (q = 0.01) and Acinetobacter (q = 0.02). From these genera, Escherichia/Shigella, Adlercreutzia, Enteroscipio and unclassified Eggerthellaceae were consistently more abundant in AMT treated animals (Supplementary Table S4).
Discussion
Probiotics and/or FMT are agents capable of managing symptoms of UC and induce disease remission in both rodents56,57 and humans58,59. However, studies with special focus on metabolic products that drive these effects and their mechanisms only now start to rise60,61,62. In this study, we opted to examine EcN aldh, an AMT that produces ILA, a natural gut microbial metabolite, on its potential in mitigating symptoms of intestinal inflammation. Overall, the data indicate no or minor differences in acute disease severity (body weight, colon length, histology) among DSS, EcN and EcN aldh mice. However, the EcN aldh mice showed signs of better initial recovery in terms of lower fecal lcn-2 and altered colon tissue gene expression already three days after DSS cessation. Analysis of expression levels of intestinal epithelial health markers suggested a putative positive effect of the AMT in the initial recovery phase.
DSS is widely used in experimental colitis models and a concentration of 4.5% falls within the high range of doses63,64, thus leading to the consistent colitic symptoms that we observed in the animals (Fig. 1B–D). Our results are however somewhat contrasting recent studies, which demonstrate that ILA and ILA-producing strains significantly attenuate weight loss, colon shortening, spleen enlargement and histopathological damage in colitis models65,66. Also, the anticipated healing effect of EcN in murine DSS-colitis67,68,69 was not reproduced in the present work. Since a DSS concentration of 3% w/v was used in all the aforementioned studies, we speculate that the high DSS concentration might have increased the severity of the symptoms, possibly masking the full therapeutic potential of the tested strains. Additionally, the efficacy of the ILA-producing strains against colitis is dependent on administration time65, thus strain administration before the DSS treatment may have resulted in stronger anti-colitic effect of the AMTs. The streptomycin-induced dysbiosis may have further contributed to symptom severity40,70 and the incapability of the interventions to exert their full effect.
Fecal lcn-2 measurements provided evidence that the ILA-producing EcN aldh might alleviate the inflamed gut (Fig. 1E), as the animals receiving the strain differed distinctively in their lcn-2 levels. Histopathological damage analysis revealed a weak positive trend of EcN aldh on the intestinal epithelium (Fig. 2A), although the disease features were similar among the colitic mice (Fig. 1B–E).
Both resistant strains colonized successfully in the streptomycin-treated mouse model. Their high presence in the ileum may be an indicator of the severity of the colitic symptoms, as degeneration of the ileocecal valve can lead to overgrowth of bacteria in the ileum71 and EcN and EcN aldh has not been previously observed in the ileum of healthy streptomycin-treated mice30. Despite the colonization level, EcN aldh did not result in differences between absolute levels of ILA in the feces. That is in contrast to our previous results, where comparable colonization levels in healthy mice led to a significant increase of ILA in fecal matter30. It is probable that the concomitant occurrence of streptomycin40, DSS72, the AMTs, an active inflammation73,74, and naturally occurring individualization of the mice microbiota due to cage-effects75, increased the inter-individual variability among the mice and, consequently, obscured the effect of EcN aldh regarding the amount of fecal ILA. Nevertheless, the ILA was significantly increased in the serum of EcN aldh colonized animals as compared to the EcN group (Fig. 3D) indicating that the produced ILA was readily absorbed.
Our RT-qPCR results suggest a better recovery of the EcN aldh mice and indicate potential routes through which ILA may function. Specifically, EcN adlh colonization led to upregulated the expression of TFF3 (Fig. 4A), a peptide capable of colitis symptoms amelioration19,76. It is specifically its luminal administration and not its systemic administration, that has been shown to significantly improve colitic symptoms in murine models76, thus rendering EcN aldh and its localized effect on TFF3 a promising result that warrants further investigation. Occludin and Muc3 expression was upregulated as well (Fig. 4B and C), although the lack of significant differences between EcN and EcN aldh groups suggests the presence of EcN as the effector of that upregulation. It is likely, that EcN aldh provides a combinatorial effect of ILA’s and EcN properties.
EcN aldh reduced the expression of the pro-inflammatory IL-1β in the colitic mice. That is in accordance with previously reported effects25,77. Likewise, lower levels of intestinal tissue expression of Foxp3 were observed in the EcN aldh mice (Fig. 4G), an effect that have been reported as a remission response to anti-colitic drug in UC patients50. Those results are in line with the observed reduced fecal lcn-2 in the EcN aldh mice on day 10 (Fig. 1E).
Most compellingly, the administration of EcN aldh led to the downregulation of ICAM1 and iNOS expression (Fig. 4E and F). ICAM1 is thought to have a central role in IBD conditions and treatment with anti-ICAM1 antibodies reduces colitis symptoms in rodents78,79. iNOS is negatively associated with intestinal inflammation, and its suppression leads to better outcomes in murine colitis as well. Hence, both proteins are candidate targets for the development of biologic agents80,81,82,83. Consequently, besides its potential assistance in epithelial recovery (Fig. 4A–C), our results indicate that EcN aldh is an interesting candidate to investigate further, as its colonization in the colitic gut delivers results that resemble candidate biologics’ effects.
The AMTs did not influence the alpha diversity, but elicited compositional changes in the gut microbiome, in agreement with previous reports on EcN impact on the microbiota84. Streptomycin and DSS affected the observed richness greatly. While their effect was anticipated40,85, the groups that were treated with both streptomycin and DSS showed an increase in their alpha diversity when compared to the streptomycin-treated group (Fig. 5A). As DSS is not known to affect intestinal microbiota in vitro86 and its suspected mechanism in vivo does not involve a direct impact on the microbiota63, this effect could be attributed to the host’s immune response, that was triggered by DSS, shaping the microbiome differently.
Even though ILA is known to influence the microbiome composition66, we found no major differences between the EcN and the EcN aldh groups (Fig. 5). Among the positively influenced genera by both AMT strains, Eggerthelaceae has been proposed as a potentially new source of probiotic species87. Members of this genus produce ellagic acid metabolites88 that can ameliorate colitis symptoms through downregulation of iNOS89. Genus Adlercrutzia has been also associated with better intestinal health90,91, overall suggesting a putative positive influence of EcN on the microbiome. However, the associations among EcN and the aforementioned genera should be treated with care, as the presence of streptomycin may have caused alterations and perturbations that influenced EcN’s effect.
EcN and EcN aldh did not affect hallmark measures of acute colitis, i.e. body weight, colon length, and tissue histology compared to DSS animals. Cui and colleagues revealed that strain administration several days before the DSS treatment is more effective against the colitic symptoms compared to concurrent strain and DSS administration65. Consequently, we speculate that a lower DSS dose or a chronic colitis model and an earlier administration of the strains might improve their impact on intestinal health. Studies on cell lines and/or organoids could more closely explore ILA’s effect on the intestine, and studies with animal models should be conducted to further investigate its therapeutic effect. Regarding the safety of the increased ILA systemic distribution, although a previous study of the strain in a conventional mouse model showed no major toxicity signs, such as weight loss, lethargy, changes in fur, eyes, behaviour patterns, diarrhea, etc.30, thorough in vitro and in vivo examinations must be conducted to assess the strain’s safety.
Altogether, this study presents an ILA-producing AMT that, when administered in DSS-treated rodents, shows limited therapeutic effect on chemically induced acute colitis, but beneficially affected gene expression markers of epithelial recovery and inflammation in colonic tissue in the initial recovery phase 3 days after treatment cessation. This finding suggests that EcN aldh and ILA may assist recovery from conditions that involve intestinal inflammation, however further studies are needed to confirm this.
Materials and methods
Strains and bacterial cultures
The engineered probiotic strains used in this study, namely EcN and EcN aldh, are modified versions of the wild type E. coli Nissle 1917 (tradename Mutaflor, Ardeypharm, Germany) that contain a vector with (EcN aldh) or without (EcN) the aromatic lactic acid dehydrogenase (aldh, GenBank accession no. WP_014484799.1) from Bifidobacterium longum subsp. infantis30. Both strains grew in LB medium supplemented with either 50 µg/ml streptomycin (EcN), or 50 µg/ml streptomycin and 30 µg/ml kanamycin (EcN aldh), and incubated at 37 °C at 200 rpm.
Bacterial suspensions of the strains were prepared from overnight cultures, harvested by centrifugation (4000 g for 10 min), washed twice with, and re-suspended in sterile PBS. The concentrations were quantified by flow cytometric detection of GFP on a flow cytometer (MACSQuant® VYB, Miltenyi Biotec) and adjusted to 108 cells/ml. The solutions were kept at room temperature until they were orally administered to the mice.
Statements of animal care, use and reporting
All methods were performed in accordance with the relevant guidelines and regulations. The Danish Animal Experiments Inspectorate provided ethical approval on the experimental design (Fig. 1A) and protocol, with authorization number: 2020-15-0201-00484 C1. The experiment was carried out at the DTU National Food Institute facilities and the in-house Animal Welfare Committee for animal care and use oversaw the experiments. The animal study was reported in accordance with the ARRIVE guidelines (https://arriveguidelines.org/).
In vivo experiment
Seven-week-old C57BL/6NTac mice (female, n = 40, Taconic Biosciences, Lille Skensved, Denmark) acclimatized in groups of 10 (Type III cages) for 4 days before the start of the experiment. On day -5, the mice were split into four groups by stratified randomization based on weight, moved to Type II cages and single-housed for the rest of the experiment. No a priori sample size calculations were performed. The groups were: (1) the control group of healthy mice (non-colitic), (2) the disease control mice (DSS), (3) the colitic mice that received EcN (EcN), and (4) the colitic mice that received EcN aldh (EcN aldh). After 5 days of acclimatization in their single-housing conditions, on Day 0, the appropriate treatment was administered to each group. As both EcN and EcN aldh encode for streptomycin resistance, the chronic colonization model40 was used in order to ensure the AMTs’ steady colonization. Accordingly, streptomycin was continuously provided as a drinking solution of streptomycin sulfate (5 g/l) in all groups. Groups DSS, EcN and EcN aldh received drinking solutions that additionally contained 4.5% Dextran sulfate sodium (DSS) 40 kDa (Sigma-Aldrich, USA) for the UC-introduction phase from Day 1 to Day 7. All solutions were sterile filtered (Bottle Top Vacuum Filter with 0.22 μm pores). The mice were monitored daily. Each animal was weighted individually, food and fluid intake were registered, and the feces were inspected for signs of disease (stool consistency and blood occurrence). Disease activity index (DAI) was calculated from the registered data according to Mu and colleagues92.
Throughout the whole experiment, environmental conditions were kept at 12-h light/ dark cycles, at 22 °C, humidity of 55%, and 50 air changes per hour. The mice had ab libitum access to water and feed (SAFE Scientific Diet A30, SAFE, Augy, France). The cages contained aspen bedding (Tapvei Aspen Bedding 4HBB), nesting material (EnviroDri), a rodent stick (Tapvei S-Bricks Aspen) and a hide (Mouse House, red polycarbonate).
On Day 4, all groups were orally gavaged with 100 µl of either bacterial solution (108 cells/ml) of EcN or EcN aldh (EcN and EcN aldh groups), or with sterile PBS (non-colitic and DSS groups). On Day 7, DSS treatment was ceased, followed 3 days post-treatment. On Day 7, three animals needed to be sacrificed due to the established human-endpoint criteria, which correspond to weight loss below the 20%. Data from those mice, one from the EcN and two from the EcN aldh group, were excluded from downstream analysis. On Day 10, the remaining mice were sedated by hypnorm/midazolam (0.1 ml/10 g SC) and euthanized by cervical dislocation.
Collection of in vivo samples
Two fecal pellets from each mouse were collected daily and stored at -80 °C until further processing. Peripheral blood samples were collected on Day -1, Day 3, and Day 9. All blood samples were processed the day of collection; they were incubated at room temperature for 30 to 60 min, to allow coagulation, and then centrifuged for 10 min at 2000 rpm followed by supernatant aspiration to obtain the serum.
Post-termination, the colon was removed, and its length was measured. Gut content was collected from ileum, cecum and colon for plating and CFU counting. Two colonic tissue samples (approx. 0.5 cm each) were collected and either stored in RNA later (Invitrogen, USA) in -20 °C, for subsequent RNA extraction, or fixed and embedded in paraffin for histological analysis.
Lipocalin-2 detection assays
Frozen serum samples were thawed at 4 °C for 30 min and then allowed to equilibrate to room temperature before processing. Frozen fecal samples were thawed at room temperature and reconstituted in PBS. Lcn-2 was quantified using Lipocalin-2 (NGAL) Mouse ELISA Kit (Abcam, UK) according to the manufacturer’s instructions. The broad and narrow spectrum versions of the kit were used for the fecal and serum samples respectively. Measurements were conducted with the BioTek ELx800 Absorbance Microplate Reader (BioTek, USA).
Histology analysis
Five-µm sections were cut (Epredia HM 450 Sliding Microtome, Thermo Fisher Scientific, US) from the paraffin-embedded tissue and stained with hematoxylin (Avantor, USA) and eosin (0.5% solution, Sigma-Aldrich, USA). Histopathological damage was evaluated by two investigators in a blinded manner (Supplementary Table S5), by light microscopy regarding (1) the severity and extent of the inflammatory cell infiltration, and (2) the changes of epithelial and mucosal architecture, using the criteria of Erben and colleagues93.
Gut content samples—Plate CFU counting
The gut content samples were placed at 4 °C, until processing. On the same day, the samples were diluted 1:5 w/v in sterile MilliQ (MQ) water and vortexed in 30-s intervals, until fully dissolved. Ten-fold dilutions were prepared and plated, for plate counting in triplicates94, on LB plates with either 50 µg/ml streptomycin (selects for both AMTs), or 30 µg/ml kanamycin and 50 µg/ml streptomycin (EcN aldh selection).
Fecal DNA isolation
The fecal samples were thawed at 4 °C for approximately 30 min. DNA was extracted using the DNeasy PowerSoil Kit (12,855-100, Qiagen, Netherlands) modified on step 5 to include a two-step centrifugation: first 3 min at 13,000 rpm, and, after removing most of the supernatant, for 1 min at 13,000 rpm. DNA concentrations were measured with the Qubit dsDNA HS and Qubit dsDNA BS (when appropriate) assay kits (Thermo Fisher Scientific, USA). The DNA was stored at -20 °C until further use.
LC-HRMS analysis—aromatic amino acids and metabolites profiling
The fecal samples were thawed at 4 °C for approximately 30 min. The thawed pellets were diluted 1:5 w/v in sterile MQ water, and the samples were vortexed in 30-s intervals, until fully dissolved, and then processed as previously described30. Briefly, a liquid–liquid extraction was employed, acetonitrile was used as a protein precipitation agent, and the samples underwent a series of centrifugations to extract the analytes, then they were concentrated to dryness, and diluted before injection in the column. Internal standard (IS) solutions were prepared for three stable isotopes, namely L-tryptophan (indole-d5), L-phenylalanine (ring-d5) and L-tyrosine (ring-d4), in MQ water (16 µg/ml). Standard solutions (SDs) were prepared for the analytes (the three aromatic amino acids and their respective lactic acid and pyruvic acid derivatives) and the three stable isotopes in their respective diluents to a final concentration of 1 mg/ml (Supplementary Table S6). The SDs were further diluted to 0.1 µg/ml, 0.5 µg/ml, 1 µg/ml, 2 µg/ml, 4 µg/ml, 10 µg/ml and 20 µg/ml and the IS diluted to 4 µg/ml. A calibration curve was calculated by the analysis of the SDs. All serum samples were processed according to Zhu and colleagues95.
RNA extraction and RT-qPCR
RNA was extracted from colonic tissue samples, stored in RNAlater, using the RNAqueous™-4PCR Total RNA Isolation Kit (Invitrogen, USA), according to manufacturer’s instructions. The RNA yield was measured with the Qubit HS RNA assay (Invitrogen, USA) and the samples were stored in − 80 °C. The RNA was reverse transcribed using the SuperScript™ IV VILO™ Master Mix (Invitrogen, USA). An ezDNase (Invitrogen, USA) treatment step preceded the reverse transcription reaction. Both steps were performed according to the kit protocols, with the reverse transcription reactions scaled up to 40 µl.
Relative quantification analysis of the genes-of-interest expression levels was performed with mRNA-specific primers (Supplementary Table S7) using the cDNA. The LightCycler 480 SYBR Green I Master Mix (Roche, Switzerland) was used for the assay according to its protocol, with a final volume of 10 µl. All samples were run in triplicates on the LightCycler 480 Instrument II (Roche, Switzerland). Single outliers were excluded from average Ct-value calculation in case the triplicates showed SDCt values > 0.5. Non-POU Domain Containing Octamer Binding protein (NONO) served as a reference gene96. Data processing was performed with the LinRegPCR software97. The results were analyzed with the ΔΔCT method98 using the healthy control group as reference.
16S rRNA amplicon sequencing
16S rRNA gene sequencing was performed on the Ion Torrent platform on the fecal samples from Days -1, 6, 8, and 10. The 16S rRNA gene V3-region was PCR amplified using the Phusion High-Fidelity DNA Polymerase (Thermo Fisher Scientific, USA), according to the manufacturer’s instructions, with a 20 µl total reaction volume. We used forward primer: PBU; 5ʹ-A-adapter-TCAG-barcode-CCTACGGGAGGCAGCAG-3ʹ and reverse primer PBR_P1: 5ʹ-trP1-adapter-ATTACCGCGGCTGCTGG-3ʹ. PBUs contained a unique barcode of 10–12 bp (Ion Xpress™ Barcode Adapters) for each sample and PBR_P1 the universal trP1 adapter, both needed for Ion Torrent sequencing. The PCR program comprised denaturation (98 °C for 30 secs), 24 cycles amplification (98 °C for 15 s, 72 °C for 30 s), and finally 72 °C for 5 min. PCR products were purified with the HighPrep PCR clean-up system (MagBIO), according to the manufacturer. DNA concentrations were measured with the Qubit dsDNA HS assay (Invitrogen, USA) and samples were pooled to obtain equimolar libraries. Sequencing was performed on the Ion S5™ sequencing system, on two separate Ion 520™ chips prepared with the Ion OneTouch™ 2 system.
Microbial community analysis—bioinformatic and statistical analysis
16S rRNA gene amplicon reads were processed using an in-house pipeline99. Raw amplicon sequences were demultiplexed by cutadapt (v. 4.1)100 and denoised using DADA2 (v. 1.22)101. ASVs were classified against rdp_train_set_18102. All downstream processing was done by Phyloseq (v.1.42.0) running in R (v. 4.2)103 and ASV decontamination was conducted with the decontam package104.
Statistical analysis
Except for the 16S rRNA analysis, all statistical analysis was performed in GraphPad Prism, version 9 (GraphPad Software Inc., USA). Normal distribution of the variables was assessed with the Kolmogorov–Smirnov test (p > 0.05). Outliers were identified and removed using the ROUT method (Q = 1%). For LC-HRMS data analysis, values below the limit of quantification (LOQ) and above the limit of detection were replaced with the LOQ/2. A p value of less than 0.05 was considered significant. The three prematurely terminated mice were removed from the analysis, hence, data presented in this study comprises results from n = 10 (Non-colitic group), n = 10 (DSS group), n = 9 (EcN group), n = 8 (EcN aldh group) mice. However, due to missing fecal samples fewer samples were processed on LC-HRMS, for 16S rRNA gene amplicon sequencing and for fecal lcn-2 measurements (Supplementary Table S1).
Alpha diversity was analysed by paired mixed effects model (REML), followed by Tukey multiple comparison test to correct for multiple testing. Beta-diversity contributing factors were analysed by permutational analysis of variance (PERMANOVA)105. Differential abundance analysis was conducted with the Limma model106. A Spearman’s correlation analysis was conducted to study the relationship of the differentially abundant genera, as detected in each group, with collection day, fecal lcn-2 levels and the aromatic lactic acids (aLAs) and aromatic amino acids (AAs) concentration in the fecal samples. To correct for multiple testing, the Benjamini–Hochberg method was applied107 and all q values presented stand for the Benjamini–Hochberg adjusted p-value. All analyses were conducted in R103 and bar and box plots were created in GraphPad Prism, version 9 (GraphPad Software Inc., USA).
Data availability
Data available from the corresponding author (M.F.L.) upon request.
References
Baumgart, D. C. & Sandborn, W. J. Inflammatory bowel disease: clinical aspects and established and evolving therapies. Lancet 369, 1641–1657 (2007).
Kaser, A., Zeissig, S. & Blumberg, R. S. Inflammatory bowel disease. Annu. Rev. Immunol. 28, 573–621 (2010).
Kobayashi, T. et al. Ulcerative colitis. Nat. Rev. Dis. Primers 6, 74 (2020).
Waljee, A. K. et al. Corticosteroid use and complications in a US inflammatory bowel disease cohort. PLoS One 11, e0158017 (2016).
Seo, G. S. & Chae, S.-C. Biological therapy for ulcerative colitis: an update. World J. Gastroenterol. 20, 13234–13238 (2014).
Chang, C.-S. et al. Identification of a gut microbiota member that ameliorates DSS-induced colitis in intestinal barrier enhanced Dusp6-deficient mice. Cell Rep. 37, 110016 (2021).
Mo, Q. et al. Novel gut microbiota patterns involved in the attenuation of dextran sodium sulfate-induced mouse colitis mediated by glycerol monolaurate via inducing anti-inflammatory responses. MBio 12, e0214821 (2021).
Chibbar, R. & Dieleman, L. A. Probiotics in the management of ulcerative colitis. J. Clin. Gastroenterol. 49(Suppl 1), S50–S55 (2015).
Pandey, K. R., Naik, S. R. & Vakil, B. V. Probiotics, prebiotics and synbiotics- a review. J. Food Sci. Technol. 52, 7577–7587 (2015).
Cammarota, G. & Ianiro, G. FMT for ulcerative colitis: closer to the turning point. Nat. Rev. Gastroenterol. Hepatol. 16, 266–268 (2019).
Charbonneau, M. R., Isabella, V. M., Li, N. & Kurtz, C. B. Developing a new class of engineered live bacterial therapeutics to treat human diseases. Nat. Commun. 11, 1738 (2020).
Kurtz, C. B. et al. An engineered E. coli Nissle improves hyperammonemia and survival in mice and shows dose-dependent exposure in healthy humans. Sci. Transl. Med. Soi https://doi.org/10.1126/scitranslmed.aau7975 (2019).
Rondon, R. E., Groseclose, T. M., Short, A. E. & Wilson, C. J. Transcriptional programming using engineered systems of transcription factors and genetic architectures. Nat. Commun. 10, 4784 (2019).
Mandell, D. J. et al. Biocontainment of genetically modified organisms by synthetic protein design. Nature 518, 55–60 (2015).
Han, S. et al. Probiotic gastrointestinal transit and colonization after oral administration: A long journey. Front. Cell Infect. Microbiol. 11, 609722 (2021).
Zmora, N. et al. Personalized gut mucosal colonization resistance to empiric probiotics is associated with unique host and microbiome features. Cell 174, 1388-1405.e21 (2018).
Chevrette, M. G. et al. Microbiome composition modulates secondary metabolism in a multispecies bacterial community. Proc. Natl. Acad. Sci. USA 119, e2212930119 (2022).
Yan, X. et al. Construction of a sustainable 3-hydroxybutyrate-producing probiotic Escherichia coli for treatment of colitis. Cell Mol. Immunol. 18, 2344–2357 (2021).
Praveschotinunt, P. et al. Engineered E. coli Nissle 1917 for the delivery of matrix-tethered therapeutic domains to the gut. Nat. Commun. 10, 5580 (2019).
Hanson, M. L. et al. Oral delivery of IL-27 recombinant bacteria attenuates immune colitis in mice. Gastroenterology 146, 210-221.e13 (2014).
Liu, J. et al. Functions of gut microbiota metabolites, current status and future perspectives. Aging Dis. 13, 1106–1126 (2022).
Ehrlich, A. M. et al. Indole-3-lactic acid associated with Bifidobacterium-dominated microbiota significantly decreases inflammation in intestinal epithelial cells. BMC Microbiol. 20, 357 (2020).
Wilck, N. et al. Salt-responsive gut commensal modulates TH17 axis and disease. Nature 551, 585–589 (2017).
Cervantes-Barragan, L. et al. Lactobacillus reuteri induces gut intraepithelial CD4+CD8αα+ T cells. Science 357, 806–810 (2017).
Meng, D. et al. Indole-3-lactic acid, a metabolite of tryptophan, secreted by Bifidobacterium longum subspecies infantis is anti-inflammatory in the immature intestine. Pediatr. Res. 88, 209–217 (2020).
Rembacken, B. J., Snelling, A. M., Hawkey, P. M., Chalmers, D. M. & Axon, A. T. Non-pathogenic Escherichia coli versus mesalazine for the treatment of ulcerative colitis: A randomised trial. Lancet 354, 635–639 (1999).
Kruis, W. et al. Maintaining remission of ulcerative colitis with the probiotic Escherichia coli Nissle 1917 is as effective as with standard mesalazine. Gut 53, 1617–1623 (2004).
Sonnenborn, U. Escherichia coli strain Nissle 1917-from bench to bedside and back: History of a special Escherichia coli strain with probiotic properties. FEMS Microbiol. Lett. 363(19), 212 (2016).
Scaldaferri, F. et al. Role and mechanisms of action of Escherichia coli Nissle 1917 in the maintenance of remission in ulcerative colitis patients: An update. World J. Gastroenterol. 22, 5505–5511 (2016).
Dimopoulou, C. et al. An engineered Escherichia coli Nissle 1917 increase the production of indole lactic acid in the gut. FEMS Microbiol. Lett. https://doi.org/10.1093/femsle/fnad027 (2023).
Singh, V. et al. Microbiota-inducible innate immune, siderophore binding protein lipocalin 2 is critical for intestinal homeostasis. Cell. Mol. Gastroenterol. Hepatol. 2, 482-498.e6 (2016).
Schroll, A. et al. Lipocalin-2 ameliorates granulocyte functionality. Eur. J. Immunol. 42, 3346–3357 (2012).
Warszawska, J. M. et al. Lipocalin 2 deactivates macrophages and worsens pneumococcal pneumonia outcomes. J. Clin. Invest. 123, 3363–3372 (2013).
Playford, R. J. et al. Effects of mouse and human lipocalin homologues 24p3/lcn2 and neutrophil gelatinase-associated lipocalin on gastrointestinal mucosal integrity and repair. Gastroenterology 131, 809–817 (2006).
Jaberi, S. A. et al. Lipocalin-2: Structure, function, distribution and role in metabolic disorders. Biomed. Pharmacother. 142, 112002 (2021).
Zollner, A. et al. Faecal biomarkers in inflammatory bowel diseases: calprotectin versus lipocalin-2-a comparative study. J. Crohns Colitis 15, 43–54 (2021).
Abella, V. et al. The potential of lipocalin-2/NGAL as biomarker for inflammatory and metabolic diseases. Biomarkers 20, 565–571 (2015).
Chassaing, B. et al. Fecal lipocalin 2, a sensitive and broadly dynamic non-invasive biomarker for intestinal inflammation. PLoS One 7, e44328 (2012).
Yadav, S. K. et al. Fecal Lcn-2 level is a sensitive biological indicator for gut dysbiosis and intestinal inflammation in multiple sclerosis. Front. Immunol. 13, 1015372 (2022).
Spees, A. M. et al. Streptomycin-induced inflammation enhances Escherichia coli gut colonization through nitrate respiration. MBio 4(4), 10–128 (2013).
Asaf, S. et al. Lipocalin 2-not only a biomarker: a study of current literature and systematic findings of ongoing clinical trials. Immunol. Res. https://doi.org/10.1007/s12026-022-09352-2 (2022).
Podolsky, D. K., Gerken, G., Eyking, A. & Cario, E. Colitis-associated variant of TLR2 causes impaired mucosal repair because of TFF3 deficiency. Gastroenterology 137, 209–220 (2009).
Saitou, M. et al. Complex phenotype of mice lacking occludin, a component of tight junction strands. Mol. Biol. Cell 11, 4131–4142 (2000).
Chelakkot, C. et al. Intestinal epithelial cell-specific deletion of PLD2 alleviates DSS-induced colitis by regulating occludin. Sci. Rep. 7, 1573 (2017).
Ho, S. B. et al. Cysteine-rich domains of muc3 intestinal mucin promote cell migration, inhibit apoptosis, and accelerate wound healing. Gastroenterology 131, 1501–1517 (2006).
Bersudsky, M. et al. Non-redundant properties of IL-1α and IL-1β during acute colon inflammation in mice. Gut 63, 598–609 (2014).
Bendjelloul, F. et al. Intercellular adhesion molecule-1 (ICAM-1) deficiency protects mice against severe forms of experimentally induced colitis. Clin. Exp. Immunol. 119, 57–63 (2000).
Bennett, C. F. et al. An ICAM-1 antisense oligonucleotide prevents and reverses dextran sulfate sodium-induced colitis in mice. J. Pharmacol. Exp. Ther. 280, 988–1000 (1997).
Zamora, R., Vodovotz, Y. & Billiar, T. R. Inducible nitric oxide synthase and inflammatory diseases. Mol. Med. 6, 347–373 (2000).
Li, Z. et al. Reciprocal changes of Foxp3 expression in blood and intestinal mucosa in IBD patients responding to infliximab. Inflamm. Bowel Dis. 16, 1299–1310 (2010).
Ofengeim, D. & Yuan, J. Regulation of RIP1 kinase signalling at the crossroads of inflammation and cell death. Nat. Rev. Mol. Cell Biol. 14, 727–736 (2013).
Zhang, X. et al. Ubiquitination of RIPK1 suppresses programmed cell death by regulating RIPK1 kinase activation during embryogenesis. Nat. Commun. 10, 4158 (2019).
Schmitt, H. et al. The TLR9 agonist cobitolimod induces IL10-producing wound healing macrophages and regulatory T cells in ulcerative colitis. J. Crohns Colitis 14, 508–524 (2020).
Ihara, S., Hirata, Y. & Koike, K. TGF-β in inflammatory bowel disease: a key regulator of immune cells, epithelium, and the intestinal microbiota. J. Gastroenterol. 52, 777–787 (2017).
Lamas, B., Natividad, J. M. & Sokol, H. Aryl hydrocarbon receptor and intestinal immunity. Mucosal Immunol. 11, 1024–1038 (2018).
Xu, L. et al. Probiotic consortia and their metabolites ameliorate the symptoms of inflammatory bowel diseases in a colitis mouse model. Microbiol. Spectr. 10, e0065722 (2022).
Yang, Y. et al. Human fecal microbiota transplantation reduces the susceptibility to dextran sulfate sodium-induced germ-free mouse colitis. Front. Immunol. 13, 836542 (2022).
Lopez, J. & Grinspan, A. Fecal microbiota transplantation for inflammatory bowel disease. Gastroenterol. Hepatol. 12, 374–379 (2016).
Zigra, P. I., Maipa, V. E. & Alamanos, Y. P. Probiotics and remission of ulcerative colitis: a systematic review. Neth. J. Med. 65, 411–418 (2007).
Shen, Z.-H. et al. Relationship between intestinal microbiota and ulcerative colitis: Mechanisms and clinical application of probiotics and fecal microbiota transplantation. World J. Gastroenterol. 24, 5–14 (2018).
Wei, Y.-L. et al. Fecal microbiota transplantation ameliorates experimentally induced colitis in mice by upregulating ahr. Front. Microbiol. 9, 1921 (2018).
Ding, S. et al. Lactobacillus brevis alleviates DSS-induced colitis by reprograming intestinal microbiota and influencing serum metabolome in murine model. Front. Physiol. 10, 1152 (2019).
Chassaing, B., Aitken, J. D., Malleshappa, M. & Vijay-Kumar, M. Dextran sulfate sodium (DSS)-induced colitis in mice. Curr. Protoc. Immunol. 104(1), 15–25 (2014).
Wirtz, S. et al. Chemically induced mouse models of acute and chronic intestinal inflammation. Nat. Protoc. 12, 1295–1309 (2017).
Cui, Q. et al. Bifidobacterium bifidum ameliorates DSS-induced colitis in mice by regulating AHR/NRF2/NLRP3 inflammasome pathways through indole-3-lactic acid production. J. Agric. Food Chem. 71, 1970–1981 (2023).
Zhang, Q. et al. Lactobacillus plantarum-derived indole-3-lactic acid ameliorates colorectal tumorigenesis via epigenetic regulation of CD8+ T cell immunity. Cell Metab. 35, 943-960.e9 (2023).
Park, J. et al. Anti-inflammatory properties of Escherichia coli Nissle 1917 in a murine colitis model. Intest. Res. 19, 478–481 (2021).
Rodríguez-Nogales, A. et al. The administration of Escherichia coli nissle 1917 ameliorates development of dss-induced colitis in mice. Front. Pharmacol. 9, 468 (2018).
Fábrega, M.-J. et al. Intestinal anti-inflammatory effects of outer membrane vesicles from escherichia coli nissle 1917 in DSS-experimental colitis in mice. Front. Microbiol. 8, 1274 (2017).
Strati, F. et al. Antibiotic-associated dysbiosis affects the ability of the gut microbiota to control intestinal inflammation upon fecal microbiota transplantation in experimental colitis models. Microbiome 9, 39 (2021).
Miller, L. S. et al. Ileocecal valve dysfunction in small intestinal bacterial overgrowth: A pilot study. World J. Gastroenterol. 18, 6801–6808 (2012).
Nagalingam, N. A., Kao, J. Y. & Young, V. B. Microbial ecology of the murine gut associated with the development of dextran sodium sulfate-induced colitis. Inflamm. Bowel Dis. 17, 917–926 (2011).
Wang, J., Chen, W.-D. & Wang, Y.-D. The relationship between gut microbiota and inflammatory diseases: The role of macrophages. Front. Microbiol. 11, 1065 (2020).
Hansen, J., Gulati, A. & Sartor, R. B. The role of mucosal immunity and host genetics in defining intestinal commensal bacteria. Curr. Opin. Gastroenterol. 26, 564–571 (2010).
McCafferty, J. et al. Stochastic changes over time and not founder effects drive cage effects in microbial community assembly in a mouse model. ISME J. 7, 2116–2125 (2013).
Poulsen, S. S., Kissow, H., Hare, K., Hartmann, B. & Thim, L. Luminal and parenteral TFF2 and TFF3 dimer and monomer in two models of experimental colitis in the rat. Regul. Pept. 126, 163–171 (2005).
Huang, W., Cho, K. Y., Meng, D. & Walker, W. A. The impact of indole-3-lactic acid on immature intestinal innate immunity and development: a transcriptomic analysis. Sci. Rep. 11, 8088 (2021).
Abdel-Wahab, B. A., Alkahtani, S. A., Alqahtani, A. A. & Hassanein, E. H. M. Umbelliferone ameliorates ulcerative colitis induced by acetic acid via modulation of TLR4/NF-κB-p65/iNOS and SIRT1/PPARγ signaling pathways in rats. Environ. Sci. Pollut. Res. Int. 29, 37644–37659 (2022).
Song, Y. et al. Caveolin-1 protects against DSS-induced colitis through inhibiting intestinal nitrosative stress and mucosal barrier damage in mice. Biochem. Pharmacol. 180, 114153 (2020).
Du Plessis, J. et al. Activated intestinal macrophages in patients with cirrhosis release NO and IL-6 that may disrupt intestinal barrier function. J. Hepatol. 58, 1125–1132 (2013).
Reinisch, W., Hung, K., Hassan-Zahraee, M. & Cataldi, F. Targeting endothelial ligands: ICAM-1/alicaforsen, MAdCAM-1. J. Crohns Colitis 12, S669–S677 (2018).
Panés, J. et al. Efficacy of an inhibitor of adhesion molecule expression (GI270384X) in the treatment of experimental colitis. Am. J. Physiol. Gastrointest. Liver Physiol. 293, G739–G748 (2007).
Warner, T. D. & McCartney, S. A. NOS inhibitors in colitis: a suitable case for treatment?. Gut 42, 152–153 (1998).
Geervliet, M. et al. Effects of Escherichia coli Nissle 1917 on the Porcine Gut Microbiota, Intestinal Epithelium and Immune System in Early Life. Front. Microbiol. 13, 842437 (2022).
Forster, S. C. et al. Identification of gut microbial species linked with disease variability in a widely used mouse model of colitis. Nat. Microbiol. 7, 590–599 (2022).
Krause, J. L. et al. DSS treatment does not affect murine colonic microbiota in absence of the host. Gut Microbes 16, 2297831 (2024).
Selma, M. V. et al. Isolation of human intestinal bacteria capable of producing the bioactive metabolite isourolithin a from ellagic acid. Front. Microbiol. 8, 1521 (2017).
Beltrán, D., Romo-Vaquero, M., Espín, J. C., Tomás-Barberán, F. A. & Selma, M. V. Ellagibacter isourolithinifaciens gen. nov., sp. nov. a new member of the family Eggerthellaceae, isolated from human gut. Int. J. Syst. Evol. Microbiol. 68, 1707–1712 (2018).
Bunt, D. V., Minnaard, A. J. & El Aidy, S. Potential modulatory microbiome therapies for prevention or treatment of inflammatory bowel diseases. Pharmaceuticals (Basel) 14(6), 506 (2021).
Mayo, B., Vázquez, L. & Flórez, A. B. Equol: A bacterial metabolite from the daidzein isoflavone and its presumed beneficial health effects. Nutrients 11(9), 2231 (2019).
Dong, Y. et al. Bifidobacterium BLa80 mitigates colitis by altering gut microbiota and alleviating inflammation. AMB Express 12, 67 (2022).
Mu, H.-X. et al. Anti-inflammatory Actions of (+)-3’α-Angeloxy-4’-keto-3’,4’-dihydroseselin (Pd-Ib) against Dextran Sulfate Sodium-Induced Colitis in C57BL/6 Mice. J. Nat. Prod. 79, 1056–1062 (2016).
Erben, U. et al. A guide to histomorphological evaluation of intestinal inflammation in mouse models. Int. J. Clin. Exp. Pathol. 7, 4557–4576 (2014).
Thomas, P., Sekhar, A. C., Upreti, R., Mujawar, M. M. & Pasha, S. S. Optimization of single plate-serial dilution spotting (SP-SDS) with sample anchoring as an assured method for bacterial and yeast cfu enumeration and single colony isolation from diverse samples. Biotechnol. Rep. (Amst) 8, 45–55 (2015).
Zhu, W. et al. Quantitative profiling of tryptophan metabolites in serum, urine, and cell culture supernatants by liquid chromatography-tandem mass spectrometry. Anal. Bioanal. Chem. 401, 3249–3261 (2011).
Eissa, N., Kermarrec, L., Hussein, H., Bernstein, C. N. & Ghia, J.-E. Appropriateness of reference genes for normalizing messenger RNA in mouse 2,4-dinitrobenzene sulfonic acid (DNBS)-induced colitis using quantitative real time PCR. Sci. Rep. 7, 42427 (2017).
Cikos, S., Bukovská, A. & Koppel, J. Relative quantification of mRNA: comparison of methods currently used for real-time PCR data analysis. BMC Mol. Biol. 8, 113 (2007).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Mortensen, M. S. Optimized DADA2 pipeline for 16S rRNA gene sequencing using IonTorrent. Tech. Univ. Denmark https://doi.org/10.11583/dtu.22657339.v1 (2023).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10 (2011).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Callahan, B. RDP taxonomic training data formatted for DADA2 (RDP trainset 18/release 11.5). Zenodo https://doi.org/10.5281/zenodo.4310151 (2020).
R Foundation for Statistical Computing, R. C. T. R Core Team (2023). R: A language and environment for statistical' ' computing. R Foundation for Statistical Computing, Vienna, Austria. at <https://www.R-project.org/.>
Davis, N. M., Proctor, D., Holmes, S. P., Relman, D. A. & Callahan, B. J. Simple statistical identification and removal of contaminant sequences in marker-gene and metagenomics data. BioRxiv https://doi.org/10.1101/221499 (2017).
Anderson, M. J. in Wiley statsref: statistics reference online (eds. Balakrishnan, N. et al.) 1–15 (John Wiley & Sons, Ltd, 2014). https://doi.org/10.1002/9781118445112.stat07841
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Benjamini, Y. & Hochberg, Y. On the adaptive control of the false discovery rate in multiple testing with independent statistics. J. Educ. Behav. Statist. 25, 60–83 (2000).
Acknowledgements
The authors thank Liljana Petrevska for technical assistance regarding LC-HRMS.
Funding
This work was supported by the Novo Nordisk Foundation under NFF grant number: NNF10CC1016517 and The Novo Nordisk Foundation, Challenge programme, CaMiT under grant agreement: NNF17CO0028232.
Author information
Authors and Affiliations
Contributions
C.D. was responsible for the design of this work, acquisition, analysis, and interpretation of data, drafted this paper, and created all illustrations. P.R.G. contributed in the design and conduction of the in vivo study and the histological analysis. M.S.M. developed the 16S rDNA gene pipeline and the subsequent statistical analysis, K.A.K. assisted in the in vivo experiment, samples processing, and performed the ELISA assays, M.P. oversaw all LC-HRMS analysis, M.I.B., M.A.O.S., T.R.L., and M.F.L. contributed in the study conception and provided substantial feedback on the manuscript. All authors reviewed the manuscript. The authors thank Liljana Petrevska for technical assistance regarding LC-HRMS.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dimopoulou, C., Guerra, P.R., Mortensen, M.S. et al. Potential of using an engineered indole lactic acid producing Escherichia coli Nissle 1917 in a murine model of colitis. Sci Rep 14, 17542 (2024). https://doi.org/10.1038/s41598-024-68412-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-68412-9
- Springer Nature Limited