Abstract
Transitional waters are important habitats both for biodiversity and ecological functions, providing valuable natural resources and relevant ecosystem services. However, they are highly susceptible to climate changes and anthropogenic pressures responsible for biodiversity losses and require specific biomonitoring programs. Benthic macroinvertebrates are suitable as ecological indicators of transitional waters, being affected by biological, chemical, and physical conditions of the ecosystems about their life cycles and space-use behaviour. The advent of high-throughput sequencing technologies has allowed biodiversity investigations, at the molecular level, in multiple ecosystems and for different ecological guilds. Benthic macroinvertebrate communities’ composition has been investigated, at the molecular level, mainly through DNA extracted from sediments in marine and riverine ecosystems. In this work, benthic macroinvertebrate communities are explored through eDNA metabarcoding from water samples in a Mediterranean transitional water ecosystem. This research highlighted the validity of eDNA metabarcoding as an efficient tool for the assessment of benthic macroinvertebrate community structure in transitional waters, unveiling the spatial heterogeneity of benthic macroinvertebrate communities correlated to the measured environmental gradients. The results suggest that peculiar features of transitional water ecosystems, such as shallow waters and limited currents, facilitate the assessment of benthic macroinvertebrate communities through environmental DNA analysis from surface water samples, opening for more rapid and accurate monitoring programs for these valuable ecosystems.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
Transitional waters are coastal ecosystems originating from both marine and freshwater sources, representing a continuum between continental and marine ecosystems. The peculiar localization of these ecosystems makes them influenced by seasonal, tidal, and freshwater regimes, modifying the ecological niches in terms of salinity gradients, water depth, light availability, and nutrient regimes. This supports complex food webs which contribute to the high biological value of these ecosystems 1,2,3. The existence of such intense gradients makes transitional waters some of the most productive ecosystems, leading to the development of diverse communities and complex self-regulation mechanisms 4,5. Transitional waters provide relevant ecosystem services and societal goods and benefits, some of the most important represented by local mitigation of climate change as well as biodiversity and biomass reservoirs 5,6,7. The important ecological functions of transitional waters depend on the detrital food webs that support carbon and nutrient cycling; for these basic functions, the sediment-associated organisms play a key role 8,9,10.
Key functional groups within sediments belong to benthic macroinvertebrates 11,12,13. Benthic invertebrates are invertebrate animals inhabiting the bottom of water bodies 14. They can inhabit the bedforms surface or live within the sediments, and display different feeding strategies, e.g., deposit-feeding, filter-feeding, grazing, and predation 15. They have a role as suspension feeders by capturing organic particulate from the water column and introducing it to the bottom. Particulate organic material also reaches the sediments by physical deposition and chemical flocculation 16,17,18. Other benthic macroinvertebrates act in the sediment to accelerate the decomposition processes of organic matter 19,20. Benthic macroinvertebrates are suitable as ecological indicators of transitional water ecosystems for their critical role in the food web as well as being sensitive to the biological, chemical and physical conditions of the ecosystems due to their relatively sedentary life, their long life cycles, and their space-use behaviour 21,22,23.
Moreover, the biodiversity reduction of benthic macroinvertebrates can impact the ecosystem functioning as demonstrated for different freshwater ecosystems 24,25. Traditional assessment of benthic macroinvertebrate biodiversity is done through the identification of morphological traits and counting of sampled living individuals. This method requires intense screening work for taxa identification and its accuracy could be undetermined by difficulties in distinguishing phenotypically similar organisms, juvenile stages, and rare species.
In recent years, the application of molecular methods, such as environmental DNA (eDNA) metabarcoding, is improving the analysis of benthic macroinvertebrate biodiversity 26. eDNA metabarcoding refers to the analysis of DNA molecules present in an environmental sample, such as water or sediment, through PCR amplification of specific gene markers, high-throughput sequencing of amplicons, and taxonomic classification of DNA sequences comparing them with DNA barcodes deposited in the reference databases 27,28. DNA metabarcoding can also be applied starting from DNA extracted from a bulk sample of organisms 29,30. The application of eDNA metabarcoding for the biodiversity assessment of benthic macroinvertebrates has been performed in riverine and marine ecosystems starting from sediments 31,32,33,34 and in a few studies from water samples 35,36. In marine ecosystems, the assessment of benthic macroinvertebrate communities through water eDNA metabarcoding has highlighted the need to sample in close vicinity to the bottom and resulted in a poor proxy 37,38.
Multiple databases related to cross-disciplinary research, which allow an in-depth exploration of specialised subfields within a certain academic or scientific discipline, such as Web of Science (http://www.webofknowledge.com/), Science Direct (http://www.sciencedirect.com) and Google Scholar (http://scholar.google.com) have been searched using different combinations of the keywords “environmental DNA”, “metabarcoding”, “benthic invertebrates”, “Mediterranean transitional waters”. Up to date, no works exploring macroinvertebrate communities using eDNA metabarcoding from surface water samples in Mediterranean transitional water ecosystems have been found. Accordingly, for the first time, in this study, benthic macroinvertebrate communities are explored through eDNA metabarcoding from surface water samples in a Mediterranean transitional water ecosystem. The objective of the work is to verify whether the shallow water and limited currents in such ecosystems facilitate the assessment of benthic macroinvertebrate communities through eDNA metabarcoding from surface water samples. Furthermore, we investigated the spatial heterogeneity of benthic macroinvertebrate communities and correlated it to the measured environmental gradients. Finally, we investigated whether the use of biological replicates supports the validity of eDNA metabarcoding as an efficient tool for the assessment of benthic macroinvertebrate community structure in transitional water ecosystems.
Materials and methods
Study area
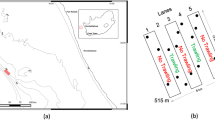
The study area is represented by the coastal lagoon included within the NATURA 2000 Site “Aquatina di Frigole” (IT50003) located in southeastern Italy along the Salento peninsula in the Apulia Region (Fig. 1), which also hosts the Research Centre for Fisheries and Aquaculture of Aquatina di Frigole of the University of Salento. The Aquatina di Frigole lagoon has a surface area of about 42 hectares and a length of about 2 km. The maximum registered depth is about 1.5 m, and the maximum registered tidal excursion is about 34 cm, annually 39.
Map of the Aquatina di Frigole Lagoon showing the localisation of the seven sampling sites. The map was generated using QGIS v3.32.0 (https://qgis.org).
The lagoon is directly communicating with the Adriatic Sea through a 400 m long and 15 m wide channel on the southern boundary. On the northern boundary, the Aquatina di Frigole lagoon receives a freshwater input from a lateral ramification of the Giammatteo canal 39.
Sampling and DNA extraction
During the spring of 2021, three replicates of 1 L of surface water were collected in each of the seven sampling sites shown in Fig. 1. The samples were immediately transferred to the Research Centre for Fisheries and Aquaculture of Acquatina di Frigole—University of Salento, Italy, located within the NATURA 2000 site and processed.
To avoid filter clogging and reduce Polymerase Chain Reaction (PCR) inhibitors, a prefiltration process was applied before filtration 40. Specifically, each 1-L replicate has been pre-filtered through a 0.5 mm conical sieve mesh to retain suspended particulate material. Each 1-L replicate was then filtered through a 0.45 μm cellulose filter of 47 mm diameter (Advantec® Mixed Cellulose Ester filters). Following, each filter was singularly used in the DNA extraction process using the DNeasy PowerWater kit (QIAGEN, Germany) following the manufacturer’s protocol. The extracted DNA was immediately stored at − 20 °C.
Abiotic parameters (Salinity, Temperature, pH, and Dissolved Oxygen) are monthly surveyed in the Aquatina Lagoon in correspondence with the 7 sampling sites using the multiparametric probe YSI ProQuatro® (xylem INC, USA).
DNA amplification, sequencing, and bioinformatic analysis
A region of approximately 350 bp of the mitochondrial cytochrome c oxidase subunit I (COI) marker gene was amplified using the degenerated primers BF2 [5′-GCHCCHGAYATRGCHTTYCC-3′] and BR2 [5′-TCDGGRTGNCCRAARAAYCA-3′] 31. The PCR reaction was performed in a volume of 50 μL composed as follows: 5 μL 10X reaction buffer; 1 μL MgCl2 (50 mM); 1 μL dNTP mix (10 mM); 1 μL forward primer (10 mM); 1 μL reverse primer (10 mM); 10 ng DNA; 0.2 μL Platinum Taq (5 U/μL; Life Technologies, USA); sterile water to reach a volume of 50 μL. The amplification program included the following step: denaturation at 95 °C for 5 min, followed by 30 cycles of denaturation (95 °C for 30 s), annealing (50 °C for 30 s), extension (72 °C for 1 min), and a final extension at 72 °C for 10 min.
All PCR products were purified with a PureLink PCR purification kit (Invitrogen, Carlsbad, CA, USA). The Celero™ DNA-Seq Library Preparation Kit cod. 0360A (Tecan, Männedorf, CH) was used for library preparation following the manufacturer’s instructions, without fragmenting the amplicons. Both input and final libraries were quantified by a Qubit 2.0 Fluorometer (Invitrogen, Carlsbad, CA) and quality tested by an Agilent 2100 Bioanalyzer High Sensitivity DNA assay (Agilent Technologies, Santa Clara, CA). Libraries were then sequenced on the MiSeq instrument (Illumina, San Diego, CA) in paired-end 300 bp mode by IGATech, Udine, Italy. High throughput sequencing of the COI amplicons generated a total of 8,654,348 raw reads, with an average of about 412,111 raw reads per sample. An internally developed pipeline and the USEARCH algorithm were used for chimaera filtering, grouping of replicate sequences, and OTU clustering. An internally developed classifier and the IGATech COI reference database were used for taxonomic classification. Specifically, the reference database has been generated by downloading the COI sequences from the NCBI Nucleotide section using the keywords “COI[all fields] OR Cytochrome c oxidase subunit I [all fields]) AND biomol_genomic[PROP] AND animal[All Fields] AND 200:1000[SLEN]”. Steps for the analysis pipeline have been developed as follows: (1) reads were trimmed to eliminate primer sequences by cutadapt1 41, (2) low-quality bases were removed from 3’ by erne-filter2 42 by applying default parameters, excluding reads < 60 bp and (3) reads with an error rate > 1% from further analysis; (4) chimeric sequences were removed with uchime_denovo3 43 implemented in usearch; (5) reads were clustered to a minimum identity of 97% generating representative sequences with cluster_fast 44 implemented in usearch; (6) clusters were blasted against the COI database, without a minimum identity filter; (7) the classifier analysed all the possible blast hits and reported the lowest unambiguous taxonomy possible. The resulting Operational Taxonomic Unit (OTU) annotated list was filtered to contain only macrozoobenthic annotated OTUs.
Statistical OTU diversity estimation
To quantify the macrozoobenthic diversity in terms of OTU richness and the number of reads per OTU in each sampling site, the Shannon–Wiener OTUs Diversity Index has been calculated as a comparative information statistic index. One-way ANOVA was calculated to investigate differences between replicates and between sampling sites. All the above calculations have been done through the software Microsoft Excel. The index results obtained through Microsoft Excel calculations have been uploaded in R and are visualised as a boxplot obtained using the R package ggplot2 v3.5.0 45. To quantify the compositional dissimilarity between sampling sites in terms of the number of reads per OTU, a pairwise Bray–Curtis dissimilarity matrix was calculated for the Wisconsin double standardised data using the R package vegan v2.6-446. The results are visualized as a non-metric multidimensional scaling (nMDS) plot obtained using the R package ggplot2 v3.5.0. Multiple regressions were also calculated to investigate the OTUs which significantly drive the nMDS distribution of the sampling sites and replicates, referred to as intrinsic variables, using the R package vegan v2.6-4. Significance was tested by permutation test (n = 999 permutations). To better understand the differences in terms of OTU composition of the samples and replicates and correlate it to the abiotic characteristics of the study area, a Bray–Curtis distance-based Redundancy Analysis (db-RDA) was performed on the Wisconsin double standardised OTUs data and standardised environmental data using the R package vegan v2.6-4. Response variables are represented by the number of reads per OTU retrieved in each replicate, while explanatory variables are represented by the mean abiotic parameter values registered in the Aquatina Lagoon during the survey held in the same sampling month (Salinity, Temperature, Dissolved Oxygen, and pH). Two-way ANOVA was calculated for significance values. The results are visualised as a db-RDA plot obtained using the R package ggplot2 v3.5.0.
Results
eDNA metabarcoding for the assessment of benthic macroinvertebrate communities composition in the lagoon ecosystem
High throughput sequencing of the COI amplicons generated a total of 8,654,348 raw reads, with an average of about 412,111 raw reads per sample. The observed features and Shannon index rarefaction curve of filtered sequences indicated that the samples reached a plateau (Figs. S1; Fig. S2), highlighting that sequencing depth was appropriate in all the sample replicates. OTUs annotated as macrozoobenthic taxa were selected for further analyses and reported in Table S1.
The total and relative number of OTUs annotated in the seven sampling sites were considered as the sum of the three replicates (Fig. 2). The analysis highlighted a specific and different macrozoobenthic community composition throughout the seven selected sampling sites of the Aquatina Lagoon (Fig. 2, Table S2). Overall, 49% of the OTUs belong to the Mollusca phylum, 16% to the Annelida phylum, 13% to the Gastrotricha phylum, 8% to the Chordata phylum, 7% to the Arthropoda phylum, 3% to the Cnidaria phylum, 2% to the Porifera phylum, and the remaining 2% to the Echinodermata and Nemertea phyla.
Analysis of the diversity of the benthic macroinvertebrate communities among lagoon sites
The Shannon–Wiener diversity index (Fig. 3) shows that the sampling sites closest to the seawater inlet (namely AF, A, and B) host a higher macrozoobenthic OTUs diversity with respect to the others. The diversity differences are significant among the sampling sites (one-way ANOVA, p-value < 0.01; df = 6 between groups, 14 within groups; F = 13.96), whereas they are not significant among replicates (one-way ANOVA; p-value > 0.5; df = 2 between groups, 18 within groups; F = 0.01).
The pairwise Bray Curtis dissimilarity matrix (Fig. S3) and the nMDS ordination (Fig. 4; Fig. S4) clearly suggest a distribution of the benthic macroinvertebrate diversity into three main communities: one localised in correspondence of sampling site AF; one shared between sampling sites A, B, and C; and one shared between sampling sites D, E, and F.
In Table S3, OTUs significantly driving the nMDS ordination (p-values < 0.05) have been marked with an asterisk. In particular, Acroloxus lacustris, Ascidiella aspersa, Botrylloides leachii, Bythinella charpentieri, Bythinella sp., Chaetonotus sp., Distomus variolosus, Euglesa casertana, Gammarus pulex, Gyraulus rossmaessleri, Gyraulus sp., Ichthydium sp., Idiopoma sp., Nais sp., Pseudamnicola sp., Radix sp., Semibalanus balanoides, Squilla mantis, and Theodoxus meridionalis are the species significantly correlated with the nMDS ordination (p-values < 0.05).
Benthic macroinvertebrate communities structure is correlated to the temperature and salinity gradient of the lagoon
To correlate the dissimilarities between the sampling sites and the main abiotic drivers that characterize the environment under study, a constrained ordination has been performed. In particular, a Bray–Curtis distance-based Redundancy Analysis (db-RDA) has been calculated. Four environmental parameters have been used as explanatory variables: Temperature (T), Salinity (PSU), Dissolved Oxygen (DO) and pH (Fig. 5; Table S4). All loading and score values are summarised in Table S5. According to the two-way ANOVA p-values calculated for each of the abiotic parameters, salinity (p-value < 0.01) and temperature (p-value < 0.01) are the main explanatory variables for the OTUs distribution across the sampling sites. pH (p-value < 0.05) and DO (p-value < 0.1) do not affect it.
Bray–Curtis distance-based Redundancy Analysis (db-RDA), calculated to explain the differences in the macrozoobenthic taxonomic composition of the Aquatina di Frigole lagoon. PSU = Salinity (Practical Salinity Units, PSU); T = Temperature (°C); DO = Dissolved oxygen (mg/L). Black dots and letters indicate the seven sampling sites and replicates.
The db-RDA ordination suggests a clear distinction of the diversity described in the sampling sites related to abiotic factors in different macrozoobenthic communities, showing a distribution similar to that shown in the nMDS ordination.
Discussion
This study represents the first description of Mediterranean transitional waters macroinvertebrate communities' composition and spatial variation using eDNA metabarcoding from surface water samples, replicated three times in each sampling site. Since the lagoon is characterized by shallow water, only surface water was sampled. In addition, sampling surface water avoids using the bottom sediments that are characterised, in these peculiar coastal ecosystems, by a high concentration of humic substances. These substances are natural organic compounds derived from the decomposition of plant and animal material and are important components of lagoon ecosystem sediments47,48,49. The presence of humic substances in natural samples negatively interferes with the extraction of high-quality environmental DNA and with the PCR amplification process 50,51,52. Through the use of surface water samples, we overcame these issues, and we were able to obtain results coherent with the environmental features of transitional water ecosystems. Indeed, we analysed the benthic communities’ spatial variation by collecting surface water samples along the natural gradients of the lagoon, from the connection with the sea to the freshwater input. Specifically, salinity and temperature are two key environmental factors that play a crucial role in shaping the structure and function of lagoon ecosystems 53,54,55. Salinity and temperature in lagoon ecosystems can have profound effects on the distribution and abundance of species 56, as well as the overall productivity and biodiversity of the ecosystem2,23,57,58. High throughput sequencing results from this study demonstrated their reliability in defining the spatial heterogeneity of the study site, with species occurrences distributed according to the monitored salinity and temperature gradients. For instance, despite the high connection and circulation rates generally characterising lagoon ecosystems 59,60,61,62,63, in this eDNA metabarcoding assessment we identified freshwater species like Bythinella charpentieri, Euglesa casertana, Idiopoma sp. and Radix sp. in the lagoon sites closer to the freshwater input. At the same time, we identified marine species like Botrylloides leachii, Distomus variolosus, Semibalanus balanoides and Squilla mantis in the lagoon sites closer to the sea connection. Finally, we identified more broadly distributed species, like the brackish Nais sp. along the entire lagoon.
In this study, eDNA samples were amplified using universal primers targeting a fragment of the mitochondrial COI gene. This region of the mitochondrial DNA has been chosen for its broad records in the reference libraries databases for animal species 64 and for benthic macroinvertebrates in understudied regions 26. In particular, the BF2/BR2 COI primer pair has been selected due to its higher detection and precision rates, compared with other combinations, for benthic macroinvertebrates 65.
The obtained results confirm the validity of this primer pair in detecting a wide range of benthic macroinvertebrate taxa. Considering the overall biodiversity of benthic macroinvertebrates detected via eDNA metabarcoding, we have identified families such as Gammaridae, Hydrobiidae, Muricidae, Palaemonidae, Penaeidae, and Spionidae, common in this ecosystem as reported in the most comprehensive morphological inventory of species of Aquatina di Frigole lagoon66. Not all families reported in the morphological inventory have been detected. Probably, this is due to the absence, concerning less-studied environments, of the publicly available molecular sequences of several species, hindering the detection of their presence 67. The OTUs obtained in this study and unassigned at the species level represent about 40% of total OTUs. This data agrees with the previous gap analysis of COI DNA barcodes in the international databases for transitional waters benthic macroinvertebrate species of the region in which Aquatina di Frigole lagoon is located 67. In this previous study, we reported the absence of a DNA barcode for 36% of benthic macroinvertebrate species. In more detail, both gap analysis and eDNA metabarcoding assessment showed that Mollusca represents the Phylum with the highest number of OTUs unassigned to the species level (Table S1) 67. This suggests the ability of eDNA metabarcoding assessments to recover high biodiversity rates in agreement with the observed gaps in the international and publicly available DNA barcode reference libraries.
Furthermore, the high throughput sequencing data identified 48 families (Table S1) not reported in the morphological inventory66 , including easily misidentified families such as Naididae and Talitridae 68,69. This suggests the role of molecular surveys in increasing the resolution of biodiversity assessments by unveiling the presence of hard-to-identify taxonomic groups.
In conclusion, the results from this study appear robust to state that the eDNA metabarcoding high-throughput data obtained from surface water samples allow the description of the benthic communities’ composition, as well as their spatial variation, in shallow coastal lagoon ecosystems, opening for more rapid and reliable monitoring programs of such valuable ecosystems.
Data availability
The FASTQ files are deposited in GenBank (SRA) of NCBI with identification number PRJNA849247.
References
Newton, A. et al. An overview of ecological status, vulnerability and future perspectives of European large shallow, semi-enclosed coastal systems, lagoons and transitional waters. Estuar. Coast. Shelf Sci. 140, 95–122 (2014).
Pérez-Ruzafa, A., Marcos, C., Pérez-Ruzafa, I. M. & Pérez-Marcos, M. Coastal lagoons: “Transitional ecosystems” between transitional and coastal waters. J. Coast. Conserv. 15, 369–392 (2011).
Underwood, G. J., Dumbrell, A. J., McGenity, T. J., McKew, B. A. & Whitby, C. The microbiome of coastal sediments. Marine Microbiome 479–534 (2022).
Berthold, M. & Paar, M. Dynamics of primary productivity in relation to submerged vegetation of a shallow, eutrophic lagoon: a field and mesocosm study. PLoS One 16(5), e0247696 (2021).
Rodrigues-Filho, J. L., Macêdo, R. L., Sarmento, H., Pimenta, V. R., Alonso, C., Teixeira, C. R., et al. From ecological functions to ecosystem services: Linking coastal lagoons biodiversity with human well-being. Hydrobiologia 1–43 (2023).
Maes, J. et al. An indicator framework for assessing ecosystem services in support of the EU Biodiversity Strategy to 2020. Ecosyst. Serv. 17, 14–23 (2016).
Newton, A. et al. Assessing, quantifying and valuing the ecosystem services of coastal lagoons. J. Nat. Conserv. 44, 50–65 (2018).
Lloret, J. & Marín, A. The role of benthic macrophytes and their associated macroinvertebrate community in coastal lagoon resistance to eutrophication. Mar. Pollut. Bull. 58(12), 1827–1834 (2009).
Prado, P., Ibanez, C., Caiola, N. & Reyes, E. Evaluation of seasonal variability in the food-web properties of coastal lagoons subjected to contrasting salinity gradients using network analyses. Ecol. Modell. 265, 180–193 (2013).
Schmitz, O. J. & Leroux, S. J. Food webs and ecosystems: Linking species interactions to the carbon cycle. Annu. Rev. Ecol. Evol. Syst. 51, 271–295 (2020).
Edegbene, A. O., Odume, O. N., Arimoro, F. O. & Keke, U. N. Identifying and classifying macroinvertebrate indicator signature traits and ecological preferences along urban pollution gradient in the Niger Delta. Environ. Pollut. 281, 117076 (2021).
He, S., Soininen, J., Chen, K. & Wang, B. Environmental factors override dispersal-related factors in shaping diatom and macroinvertebrate communities within stream networks in China. Front. Ecol. Environ. 8, 141 (2020).
Tampo, L. et al. Benthic macroinvertebrates as ecological indicators: their sensitivity to the water quality and human disturbances in a tropical river. Front. Water. 3, 662765 (2021).
Heip, C. Eutrophication and zoobenthos dynamics. Ophelia 41(1), 113–136 (1995).
Grebmeier, J. M., McRoy, C. P. & Feder, H. M. Pelagic-benthic coupling on the shelf of the northern Bering and Chukchi Seas. I. Food supply source and benthic biomass. Mar. Ecol. 48(1), 57–67 (1988).
Chakraborty, A., Saha, G. K. & Aditya, G. Macroinvertebrates as engineers for bioturbation in freshwater ecosystem. Environ. Sci. Pollut. Res. 29(43), 64447–64468 (2022).
Kim, C., Kang, H. Y., Lee, Y. J., Yun, S. G. & Kang, C. K. Isotopic variation of macroinvertebrates and their sources of organic matter along an estuarine gradient. Estuaries Coast 43, 496–511 (2020).
Lloret, J. & Marín, A. The contribution of benthic macrofauna to the nutrient filter in coastal lagoons. Mar. Pollut. Bull. 62(12), 2732–2740 (2011).
Alam, M. K., Negishi, J. N., Pongsivapai, P., Yamashita, S. & Nakagawa, T. Additive effects of sediment and nutrient on leaf litter decomposition and macroinvertebrates in hyporheic zone. Water 13(10), 1340 (2021).
van der Meer, T. V., van der Lee, G. H., Verdonschot, R. C. & Verdonschot, P. F. Macroinvertebrate interactions stimulate decomposition in WWTP effluent-impacted aquatic ecosystems. Aquat. Sci. 83(4), 65 (2021).
Hering, D. et al. Assessment of European streams with diatoms, macrophytes, macroinvertebrates and fish: A comparative metric-based analysis of organism response to stress. Freshw. Biol. 51(9), 1757–1785 (2006).
Pinna, M., Janzen, S., Franco, A., Specchia, V. & Marini, G. Role of habitats and sampling techniques on macroinvertebrate descriptors and ecological indicators: An experiment in a protected Mediterranean lagoon. Ecol. Indic. 83, 495–503 (2017).
Pinna, M. et al. How do sampling methods affect the body size-abundance relationships of benthic macroinvertebrates in freshwater ecosystems? A comparative analysis. Water 15(3), 386 (2023).
Cao, X. et al. Loss of biodiversity alters ecosystem function in freshwater streams: Potential evidence from benthic macroinvertebrates. Ecosphere 9(10), e02445 (2018).
Johnson, C. N. et al. Biodiversity losses and conservation responses in the Anthropocene. Science 356(6335), 270–275 (2017).
Tzafesta, E., Zangaro, F., Specchia, V. & Pinna, M. An overview of DNA-based applications for the assessment of benthic macroinvertebrates biodiversity in Mediterranean aquatic ecosystems. Diversity 13(3), 112 (2021).
Pawlowski, J. et al. The future of biotic indices in the ecogenomic era: Integrating (e) DNA metabarcoding in biological assessment of aquatic ecosystems. Sci. Total Environ. 637, 1295–1310 (2018).
Taberlet, P., Coissac, E., Pompanon, F., Brochmann, C. & Willerslev, E. Towards next-generation biodiversity assessment using DNA metabarcoding. Mol. Ecol. 21(8), 2045–2050 (2012).
Elbrecht, V., Lindner, A., Manerus, L. & Steinke, D. A bright idea—Metabarcoding arthropods from light fixtures. PeerJ. 9, e11841 (2021).
Macher, J. N. et al. Comparison of environmental DNA and bulk-sample metabarcoding using highly degenerate cytochrome c oxidase I primers. Mol. Ecol. Resour. 18(6), 1456–1468 (2018).
Elbrecht, V. & Steinke, D. Scaling up DNA metabarcoding for freshwater macrozoobenthos monitoring. Freshw. Biol. 64(2), 380–387 (2019).
Hickling, S. et al. Benthic invertebrate biodiversity enhancement with reef cubes®, evidenced by environmental DNA analysis of sediment samples. Ecol. Eng. 195, 107064 (2023).
Ji, F. et al. Assessment of benthic invertebrate diversity and river ecological status along an urbanized gradient using environmental DNA metabarcoding and a traditional survey method. Sci. Total Environ. 806, 150587 (2022).
Sinniger, F. et al. Worldwide analysis of sedimentary DNA reveals major gaps in taxonomic knowledge of deep-sea benthos. Front. Mar. Sci. 3, 92 (2016).
Hupało, K., Schmidt, S., Macher, T. H., Weiss, M. & Leese, F. Fresh insights into Mediterranean biodiversity: Environmental DNA reveals spatio-temporal patterns of stream invertebrate communities on Sicily. Hydrobiologia 849(1), 155–173 (2022).
Reinholdt Jensen, M. et al. Seasonal turnover in community composition of stream-associated macroinvertebrates inferred from freshwater environmental DNA metabarcoding. Environ. DNA 3(4), 861–876 (2021).
Antich, A. et al. Marine biomonitoring with eDNA: Can metabarcoding of water samples cut it as a tool for surveying benthic communities?. Mol. Ecol. 30(13), 3175–3188 (2021).
Leduc, N. et al. Comparing eDNA metabarcoding and species collection for documenting Arctic metazoan biodiversity. Environ. DNA 1(4), 342–358 (2019).
Caroppo, C., Pinna, M. & Vadrucci, M. R. Phytoplankton size structure and diversity in the transitional system of the Aquatina Lagoon (Southern Adriatic Sea, Mediterranean). Microorganisms. 11(5), 1277 (2023).
Specchia, V. et al. Environmental DNA detects biodiversity and ecological features of phytoplankton communities in Mediterranean transitional waters. Sci. Rep. 13(1), 15192 (2023).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal. 17(1), 10–12 (2011).
Del Fabbro, C., Scalabrin, S., Morgante, M. & Giorgi, F. M. An extensive evaluation of read trimming effects on Illumina NGS data analysis. PLoS One. 8(12), e85024 (2013).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinform. 27(16), 2194–2200 (2011).
Müller, R. & Nebel, M. On the use of sequence-quality information in OTU clustering. PeerJ. 9, e11717 (2021).
Wickham, H. & Wickham, H. Data Analysis (Springer International Publishing, 2016).
Oksanen, J. vegan: Community Ecology Package-R package version 1.17-8. http://CRAN.R-project.org/package=vegan (2011).
Pinna, M. et al. Influence of aperiodic summer droughts on leaf litter breakdown and macroinvertebrate assemblages: Testing the drying memory in a Central Apennines River (Aterno River, Italy). Hydrobiologia 782, 111–126 (2016).
Santín, C., Yamashita, Y., Otero, X. L., Álvarez, M. Á. & Jaffé, R. Characterizing humic substances from estuarine soils and sediments by excitation-emission matrix spectroscopy and parallel factor analysis. Biogeochemistry 96, 131–147 (2009).
Viaroli, P., Bartoli, M., Giordani, G., Magni, P. & Welsh, D. T. Biogeochemical indicators as tools for assessing sediment quality/vulnerability in transitional aquatic ecosystems. Aquat. Conserv. Mar. Freshw. 14(S1), S19–S29 (2004).
Albers, C. N., Jensen, A., Bælum, J. & Jacobsen, C. S. Inhibition of DNA polymerases used in Q-PCR by structurally different soil-derived humic substances. Geomicrobiol. J. 30(8), 675–681 (2013).
Matheson, C. D., Gurney, C., Esau, N. & Lehto, R. Assessing PCR inhibition from humic substances. Open Enzyme Inhib. J. 3(1), 38–45 (2010).
Wnuk, E. et al. The effects of humic substances on DNA isolation from soils. PeerJ 8, e9378 (2020).
García-Oliva, M. et al. Modelling the impact of dredging inlets on the salinity and temperature regimes in coastal lagoons. Ocean Coast. Manag. 180, 104913 (2019).
Harris, C. M., McClelland, J. W., Connelly, T. L., Crump, B. C. & Dunton, K. H. Salinity and temperature regimes in eastern Alaskan Beaufort Sea lagoons in relation to source water contributions. Estuaries Coasts 40, 50–62 (2017).
Newton, A. & Mudge, S. M. Temperature and salinity regimes in a shallow, mesotidal lagoon, the Ria Formosa, Portugal. Estuar. Coast. Shelf Sci. 57(1–2), 73–85 (2003).
Dhib, A., Frossard, V., Turki, S. & Aleya, L. Dynamics of harmful dinoflagellates driven by temperature and salinity in a northeastern Mediterranean lagoon. Environ. Monit. Assess. 185, 3369–3382 (2013).
Alongi, D. M. The ecology of tropical soft-bottom benthic ecosystems. Oceanogr. Mar. Biol. 28(3), 381–496 (1990).
Jeppesen, E. et al. Salinity induced regime shift in shallow brackish lagoons. Ecosystems 10(1), 48–58 (2007).
Duck, R. W. & da Silva, J. F. Coastal lagoons and their evolution: A hydromorphological perspective. Estuar. Coast. Shelf Sci. 110, 2–14 (2012).
Ghezzo, M. et al. Connectivity in three European coastal lagoons. Estuaries Coasts. 38, 1764–1781 (2015).
Hearn, C. J., Lukatelich, R. J. & McComb, A. J. Coastal Lagoon Ecosystem Modelling. Elsevier Oceanogr. Ser. 60, 471–506 (1994).
Kjerfve, B. & Magill, K. E. Geographic and hydrodynamic characteristics of shallow coastal lagoons. Mar. Geol. 88(3–4), 187–199 (1989).
Panda, U. S., Mahanty, M. M., Rao, V. R., Patra, S. & Mishra, P. Hydrodynamics and water quality in Chilika Lagoon—A modelling approach. Procedia Eng. 116, 639–646 (2015).
Ratnasingham, S. & Hebert, P. D. BOLD: The barcode of life data system. Mol. Ecol. Notes 7(3), 355–364 (2007).
Elbrecht, V. & Leese, F. Validation and development of COI metabarcoding primers for freshwater macroinvertebrate bioassessment. Front. Environ. Sci. 5, 11 (2017).
Zangaro, F. et al. Building a transnational biodiversity geo-database of the protected areas in the Adriatic-Ionian Macro-Region: Approaches and results from the IMPRECO Project. Biodivers. Data J. 9, e67169 (2021).
Pinna, M. et al. Testing the influence of incomplete DNA barcode libraries on ecological status assessment of Mediterranean Transitional Waters. Biology 10(11), 1092 (2021).
Chakma, S., Martinsson, S. & Naveed, M. I. DNA barcoding and genetic distances in three genera of Naididae (Annelida: Clitellata). Biologia 79(5), 1359–1366 (2024).
Lobo, J. et al. Contrasting morphological and DNA barcode-suggested species boundaries among shallow-water amphipod fauna from the southern European Atlantic coast. Genome 60(2), 147–157 (2017).
Funding
This research was supported by the RIPARTI project (Code a467414b) funded by Apulia Region, Italy awarded to V. Specchia and supporting F. Zangaro and by the PRO-COAST project funded by EU HORIZON-CL6-2022-BIODIV-01 to Maurizio Pinna.
Author information
Authors and Affiliations
Contributions
Conceptualization, Valeria Specchia and Maurizio Pinna; Data curation, Francesco Zangaro, Valeria Specchia and Maurizio Pinna; Formal analysis, Valeria Specchia, Francesco Zangaro, Maurizio Pinna; Funding acquisition, Valeria Specchia and Maurizio Pinna; Methodology, Francesco Zangaro Resources, Valeria Specchia, Maurizio Pinna; Supervision, Valeria Specchia and Maurizio Pinna; Writing – original draft, Valeria Specchia, Francesco Zangaro and Maurizio Pinna; Writing – review & editing, Valeria Specchia, Francesco Zangaro, Maurizio Pinna.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pinna, M., Zangaro, F. & Specchia, V. Assessing benthic macroinvertebrate communities’ spatial heterogeneity in Mediterranean transitional waters through eDNA metabarcoding. Sci Rep 14, 17890 (2024). https://doi.org/10.1038/s41598-024-69043-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-69043-w
- Springer Nature Limited