Abstract
The efficacy of immunotherapy for clear cell renal cell carcinoma (ccRCC), especially advanced ccRCC, is limited, presenting a clinical challenge. This limitation is closely tied to the immune regulation network. Understanding the heterogeneity of the tumour microenvironment (TME) is crucial for developing advanced ccRCC therapies. Using publicly available ccRCC data (scRNA-seq, bulk RNA-seq, and somatic mutation data), a multiomics study was performed to explore TME heterogeneity. Three distinct ccRCC immune subtypes were identified through combined scRNA-seq and bulk RNA-seq analysis. A prognostic model based on unique cell signalling molecules in immunosuppressive tumour subtype was validated in the TCGA and CheckMate cohorts. MDK emerged as a critical regulatory gene in the immunosuppressive subtype, predicting a poor ccRCC prognosis and a poor immunotherapy response. MDK promotes M2 macrophage polarization via the MDK-LRP1 interaction, and the inhibition of MDK suppressed M2 polarization. This study revealed the heterogeneity of the ccRCC TME and a reliable prognostic model, shedding light on the vital role of MDK in the immunosuppressive TME and paving the way for optimized ccRCC immunotherapy.
Similar content being viewed by others
Introduction
Clear cell renal cell carcinoma (ccRCC) represents 75% of kidney cancers1 and is challenging to treat due to its aggressive nature. Surgical resection is the primary treatment for localized cases, but 20% of patients experience recurrence2. Diagnosis often occurs late, with 17% of patients already having metastatic disease at diagnosis, resulting in a 10% 5-year survival rate3,4. Patients with advanced disease are resistant to surgery, radiotherapy, and chemotherapy, which poses therapeutic challenges.
ccRCC is a highly vascularized and immunogenic tumour characterized by dense immune cell infiltration5. Immune checkpoint inhibitors (ICIs), which target CTLA-4, PD-1, and PD-L1, have transformed the treatment of advanced ccRCC. Despite advancements in immunotherapy, challenges persist, including limited patient responsiveness, rapid development of resistance, and immune-related toxicity6,7,8,9. In clinical practice, most ICI-treated patients show minimal sensitivity. Effective prognostic biomarkers are lacking for advanced ccRCC patients receiving ICI treatment10,11.
In recent years, the tumour microenvironment (TME) has become a focal point in tumour immunotherapy research. The TME is a complex network comprising diverse cellular and extracellular matrix elements, primarily cancer cells, immune cells, endothelial cells, and cancer-associated fibroblasts12. Key immune cells in the TME include CD8 + T cells, tumour-associated macrophages (TAMs), and regulatory T cells (Tregs), among others, which play pivotal roles in cancer progression13,14. Signals from this intricate microenvironment significantly influence tumour occurrence and development. Variability in TME components contributes to individual differences in immunotherapy response, thus impacting clinical efficacy15,16. Consequently, understanding TME heterogeneity and devising novel therapeutic strategies targeting diverse TME components and immunoregulatory mechanisms are crucial for optimizing immunotherapy and advancing precision medicine.
By integrating single-cell and bulk sequencing data, we conducted a comprehensive analysis of ccRCC TME characteristics and revealed three distinct immune subtypes with markedly different prognoses. Through detailed exploration of subtype heterogeneity, we identified crucial immunosuppression biomarkers by analysing cell communication. Using RNA-seq profiling data from The Cancer Genome Atlas Kidney Clear Cell Carcinoma (TCGA-KIRC) cohort and corresponding clinical data, we identified correlations between TME biomarkers and patient outcomes, revealing three specific prognostic markers. In particular, MDK, which is uniquely expressed in tumour cells, fibroblasts, and endothelial cells, plays a pivotal role in forming an immunosuppressive TME, promoting M2 macrophage polarization. MDK has significant prognostic value and therapeutic implications for ccRCC patients.
Materials and methods
Clinical specimens
Five pairs of ccRCC and adjacent normal kidney tissues, sourced from Zhongda Hospital Affiliated with Southeast University (Nanjing, China), were promptly frozen in liquid nitrogen for subsequent analysis. Detailed clinicopathological information for these patients is provided in Table S1.
Data source and preprocessing
Bulk RNA sequencing (RNA-seq) transcriptomic data, clinical information, and somatic mutation data for ccRCC patients were retrieved from The Cancer Genome Atlas (TCGA) database (https://www.cancer.gov/tcga). Single-cell RNA-seq data for ccRCC were obtained from the National Center for Biotechnology Information Sequence Read Archive under accession number PRJNA768891 and processed using Cell Ranger software. Clinical and transcriptomic data for the CheckMate cohort, comprising ccRCC patients receiving anti-PD1 medication, were extracted from a previously published study17.
Estimation of immune cell infiltration
We employed CIBERSORT (https://cibersort.stanford.edu/) to deconvolute immune cell subsets and determine the relative and total abundances of 22 immune cell types. Fibroblast infiltration was quantified using the Microenvironment Cell Populations-counter (MCP-counter) algorithm through the immunedeconv R package18. Furthermore, stromal and immune scores were calculated using the estimate R package19.
Bulk RNA-seq analysis
Initially, differential expression analysis was conducted to identify differentially expressed genes (DEGs) between tumour and normal samples in the TCGA cohort utilizing the limma package20, with |\({log}_{2}FC\)|> 1.5 and a false discovery rate (FDR) < 0.05 as cut-offs. Next, weighted gene correlation network analysis (WGCNA) was employed to filter hub genes among the DEGs using the WGCNA package21. The soft threshold power β was selected as the lowest power where the scale-free topology fit index reached 0.85. The resulting modules were presented in a dendrogram following the clustering process, and a module-trait correlation heatmap was generated to highlight the most significant modules involved in ccRCC development.
Single-cell RNA-seq analysis
Processed scRNA-seq data were transformed into Seurat files using the Seurat R package22. Cells that failed to meet the criteria (cells with > 300 genes and < 15% mitochondrial genes) were excluded. Gene expression normalization and identification of the top 2000 highly variable genes were performed. Principal component analysis (PCA) was conducted using the RunPCA function, and batch effects were removed using the harmony method. The top 15 harmony embeddings were chosen based on the ElbowPlot function for clustering and visualization utilizing the FindNeighbors, FindClusters, and RunUMAP functions. The cell types were preliminarily annotated using the SingleR R package23. To ensure accuracy, the recognized cell markers were cross-referenced with the SingleR results, and the final cell annotations were determined. Marker genes were identified across cell types using a threshold (adjusted P value < 0.05 and \({log}_{2}FC\) > 0.5) and the FindAllMarkers function. Intersecting genes between marker genes and DEGs identified by WGCNA were selected for further analysis.
Identification of subtypes and correlation analysis
Nonnegative matrix factorization (NMF)24 was utilized to categorize patients in the TCGA cohort into distinct groups based on intersecting genes. The appropriate number of clusters was determined using cophenetic, dispersion, and silhouette methods. A heatmap depicting gene expression patterns was generated to visualize transcriptional characteristics within subtypes. Prognostic differences among subtypes were assessed. Simultaneously, a pairwise correlation analysis of the intersecting genes was conducted to identify genes with significant coexpression relationships. Samples were scored based on these coexpression profiles to explore their the associations with tumour subtype and prognosis. Additionally, the composition of the tumour microenvironment in different subtypes was evaluated.
Functional enrichment analysis
We utilized Kyoto Encyclopedia of Genes and Genomes (KEGG)25,26 and Gene Ontology (GO) analyses to explore significantly enriched pathways of intersecting genes by employing the clusterProfiler R package27. Single-sample gene set enrichment analysis (ssGSEA) was conducted using the GSVA R package28. All analysed gene sets were obtained from the Molecular Signatures Database (MSigDB) and previously published studies29,30.
Cell‒cell communication analysis
Cell communication networks were analysed using the CellChat R package, which focuses on ligand‒receptor interactions31. The statistical significance of the communication probabilities was determined via a permutation test. Differential expression analysis was performed to compare the subtype of interest with others, revealing significantly upregulated genes. These genes were overlapped with ligand‒receptor molecules to identify specific cell communication patterns in the subtypes for further analysis.
Construction and validation of a prognostic model
Patients from the TCGA cohort were divided into two groups based on ligand‒receptor molecule medians. Prognosis-related genes were identified through Kaplan‒Meier survival analysis. These genes were subjected to multivariate Cox regression to identify independent prognostic factors to construct a prognostic model. Patients in both the TCGA and CheckMate cohorts were classified into high-risk or low-risk groups using median risk scores. Kaplan–Meier survival curves were generated to compare clinical outcomes. ROC curves for 1-, 3-, and 5-year overall survival (OS) were plotted, and the area under the curve (AUC) values were calculated using the timeROC R package.
Immunotherapy responses in different subgroups
To assess the associations among the identified key molecules, risk scores, and immunotherapy efficacy, patients in the CheckMate cohort were stratified based on the median values of these molecules and risk scores. Subsequently, the correlation of these variables with immunotherapy outcomes was analysed.
Gene mutation analysis
Somatic mutation data were visualized using the maftools package, and gene mutation details are presented in a waterfall plot. Different mutation types are represented by distinct colours at the plot's base.
Cell lines and cell culture
The THP-1 cell line was cultured in RPMI 1640 medium (Gibco, Thermo Fisher Scientific) supplemented with 10% foetal bovine serum (Gibco, Thermo Fisher Scientific) and 1% penicillin G and streptomycin sodium (Gibco, Thermo Fisher Scientific). Cell lines were obtained from the Shanghai Institutes for Biological Sciences and maintained in a 95% humidified atmosphere at 37 °C with 5% CO2. THP-1 cells were differentiated into M0 macrophages by incubating them with 100 ng/ml PMA (Sigma‒Aldrich) for 36 h.
Small interfering RNA
Small interfering RNAs (siRNAs) targeting MDK were custom-designed and synthesized by GenePharma Co. (Shanghai, China). The sense sequences for the siRNAs were 5′-AAGGCCAAAGCCAAGAAAGTT-3′ (si-MDK-1) and 5′-AACUGGAAGAAGGAGUUUGTT-3′ (si-MDK-2). Transfections of si-MDK or the negative control were carried out using a transfection reagent (Vazyme Biotech, Nanjing, China) according to the manufacturer's instructions.
Preparation of conditioned medium
si-MDK-Caki-1 or si-NC-Caki-1 cells were incubated at 37 °C for 2 days, followed by centrifugation at 800 rpm for 5 min to collect the supernatant. The supernatant was then mixed with freshly prepared 10% FBS at a ratio of 1:2 to obtain the final conditioned medium.
Flow cytometry
M0 macrophages were incubated with the specified conditioned medium for 48 h. For surface marker analysis, cells were suspended in 0.1% BSA/1 × PBS and stained with anti-human CD163 (PE, Bio-Legend) at 4 °C for 20 min. Data were acquired using LSRFortessa (BD Bioscience) and analysed with FACS Diva and FlowJo software.
Western blot analysis
Total protein was extracted using RIPA-Cocktail (1:1000) cell lysate (KeyGene Biotech, Nanjing, China), and the supernatant was collected. Protein quantification was conducted using the bicinchoninic acid assay (KeyGene Biotech). The extracted total protein was separated by 10% sodium dodecyl sulfate‒polyacrylamide gel electrophoresis and transferred to a polyvinylidene difluoride membrane (Merck Millipore, Burlington, Massachusetts, USA). The membrane was blocked for 1 h in Tris-buffered saline with Tween 20 containing 5% nonfat milk. Subsequently, the membrane was incubated overnight at 4 °C with the following primary antibodies: anti-MDK (1/2000, 13 kDa, Santa Cruz Biotechnology, Inc.) and anti-β-actin (1/3000, 43 kDa, Santa Cruz Biotechnology, Inc.) as an internal control. After incubation with horseradish peroxidase–conjugated secondary antibodies (Santa Cruz Biotechnology), protein bands were visualized through chemiluminescence. Densitometric analysis was performed using ImageJ software.
Quantitative real-time PCR assay
Total RNA was extracted following the manufacturer's protocol using TRIzol (Invitrogen, CA, USA) and then reverse transcribed into cDNA (complementary DNA) utilizing a reverse transcription kit and a SYBR Green Master Mix kit (Takara, Otsu, Japan). The sequences of primers used were as follows: for the MDK gene, 5′- CACCCCTAAGTGCCCAAA -3′ (F) and 5′- TGGGGAAGAACAAAAGCA -3′ (R); for the IL-10 gene, 5′- TTTAAGGGTTACCTGGGTTGC -3′ (F) and 5′- TTGATGTCTGGGTCTTGGTTC -3′ (R); and for the ARG1 gene, 5′- CACTCCCCTGACAACCAGC -3′ (F) and 5′- AGGACACAGGTTGCCCATG -3′ (R).
Cell migration and transwell wound healing assays
Caki-1 cells were inoculated into 6-well plates and cocultured with si-MDK or si-NC CM-induced polarized macrophages. A scratch was made across a monolayer of Caki-1 cells at a density of approximately 70% on a plate using a conventional pipette tip (10 µL tip). The cell wound edge was marked and photographed at the starting time point under a microscope. After 12 h, we measured the cell migration distance and analysed the wound closure percentage.
The transwell chamber (Corning, USA) contained 24 wells with a membrane (8.0 μm). We conducted a transwell assay to assess cell invasion and migration by using chemotactic chambers with and without a layer of Matrigel diluted with RPMI-1640 (BD Pharmingen Inc., San Diego, CA, USA) at 1:8 for the invasion assay and migration assay, respectively . Cells in serum-free medium (100 μL) were added to the upper chamber, and complete medium (600 μL) was added to the lower chamber as a chemoattractant. After incubation for 12 h, 4% PFA was used to fix the cells that passed through the membrane and then 1% crystal violet staining was performed for quantification.
Cell proliferation analysis
Colony formation and Cell Counting Kit-8 (CCK-8) assays were performed to compare cell proliferation under different conditions. Pretreated cells were cultured in 6-well plates at a density of 1000 cells/well. For the colony formation assay, cells were fixed with 4% methanol for 20 min after incubation at 37 °C for 2 weeks. We stained the cells using a 0.1% crystal violet dye solution and then manually counted the colonies under a light microscope. For the cell proliferation assay, CCK-8 solution from a kit (KeyGene Biotech) was added every 24 h. The absorbance was measured at 450 nm using an automatic microplate reader.
Mouse tumorigenicity assay
C57BL/6 male mice aged 4–6 weeks were purchased from the Comparative Medical Center of Yangzhou University and raised under standard conditions at the Animal Center of Southeast University for the mouse tumorigenicity assay. Prior to the experiments, the mice were acclimated in a laboratory environment for 2–3 weeks until they reached 6–8 weeks of age. The use of laboratory mice was approved by the Ethics Committee of the Affiliated Zhongda Hospital of Southeast University. Caki-1 cells (1 × 106) cocultured with CM-induced polarized macrophages were injected through the tail vein. After 3 weeks, the mice were euthanized (Cervical Dislocation Following Pentobarbital Anesthesia), and their lungs were removed.
Statistical analysis
All bioinformatics analyses were performed using R software (version 4.2.1). Differences in continuous variables between two groups were evaluated using the nonparametric Wilcoxon rank-sum test. Multigroup comparisons were conducted using the Kruskal‒Wallis test. Categorical variables were assessed using the chi-squared test or Fisher's exact test. Survival analysis was carried out utilizing the Kaplan‒Meier method and log-rank test. Correlation analysis was performed using the Spearman test. All experiments were independently repeated three times, and Student’s t test or one-way ANOVA was used to compare the studied groups. A two-sided P value < 0.05 was considered to indicate statistical significance.
Ethics statement
The methodology of this study followed the criteria outlined in the Helsinki Declaration. The study protocol was approved by the Ethics Committee and Institutional Review Board for Clinical Research of Zhongda Hospital (ZDKYSB077). All human tumor tissue samples were collected in accordance with the national and institutional ethical guidelines. All participants signed informed consent prior to their inclusion in the study. All animal experiments reported in this study were conducted in accordance with the ARRIVE guidelines (https://arriveguidelines.org).
Results
Identification of tumour microenvironment-related genes
We utilized the CIBERSORT algorithm to assess the proportions of 22 immune cell types in the TCGA-KIRC cohort. As expected, tumour samples exhibited significantly greater infiltration of immune cells, particularly CD8 + T cells, macrophages, and regulatory T (Treg) cells, than normal samples (Fig. 1A). Subsequently, we conducted differential gene expression analysis between tumour and normal samples and identified 1372 differentially expressed genes (DEGs), 570 of which were upregulated and 802 of which were downregulated in tumours (Fig. 1B,C). These DEGs were subjected to WGCNA, which revealed four modules, among which the magenta and grey modules were strongly associated with tumour phenotype and immune cell infiltration (Supplementary Fig. S1A–C).
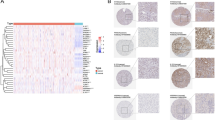
Immune infiltration and differential expression analysis in the TCGA cohort. (A) Abundance of 22 immune cells in the TCGA-KIRC cohort derived from the CIBERSORT algorithm. (B) Heatmap of the top 40 DEGs between tumour samples and normal samples. (C) Volcano plot displaying all 40 DEGs between tumour samples and normal samples (adjusted P value < 0.05 and |\({log}_{2}FC\)|> 1.5).
We concurrently analysed single-cell RNA-seq data from four ccRCC patients. Leveraging the SingleR package and established cell markers, we successfully identified 13 cell types (Fig. 2A–C) and identified marker genes for each type (Fig. 2D).
Considering the immune cell infiltration and WGCNA results based on the bulk RNA-seq data, we focused on marker genes of tumour cells, CD8 + T cells, proliferative CD8 + T cells, CD4 + T cells, and macrophages for further analysis. These immune cells demonstrated significant infiltration in the TME of ccRCC and were closely correlated with tumour-related modules. Intersection of these markers with module genes revealed 191 tumour microenvironment-related genes (Fig. 3A,B). KEGG and GO analyses of these genes revealed significant enrichment in immune activation-related pathways and some metabolism-related pathways, such as "central carbon metabolism in cancer" and the "HIF − 1 signalling pathway". Notably, an immune checkpoint-related pathway, "PD − L1 expression and PD − 1 checkpoint pathway in cancer," was also enriched (Fig. 3C,D).
Identification of tumour microenvironment-related genes. (A) Venn diagram illustrating the overlap between significant modules from WGCNA and markers identified from the scRNA-seq data. (B) UpSet plot detailing the relationships among gene sets. (C,D) Enrichment analysis of KEGG pathways and GO terms for identified tumour microenvironment-related genes.
Identification of different immune subtypes
To further explore the heterogeneity of the tumour microenvironment in ccRCC, we applied NMF dimension reduction and clustering to 191 tumour microenvironment-related genes within the TCGA cohort. By evaluating the relevant characteristics, we identified three clusters as the optimal number (Fig. 4A). The three-dimensional principal component analysis (PCA) plot verified the efficacy of the clustering method (Fig. 4B). Gene expression heatmaps unequivocally demonstrated distinct differences among the three subtypes (Fig. 4C). Notably, Kaplan‒Meier survival analysis revealed significant discrepancies in prognosis among the subtypes, with C1 displaying the worst prognosis and C3 exhibiting the most favourable outcomes (Fig. 4D). An examination of the clinical traits of patients with each subtype highlighted a higher incidence of advanced ccRCC in patients with the C1 subtype (Table 1).
Identification of immune subtypes. (A) Line plots depicting NMF-related parameter variations with changing K values from 2 to 6. (B) Three-dimensional principal component analysis illustrating the subtypes. (C) Heatmap displaying the expression patterns of TME-related genes across the three subtypes. (D) Kaplan‒Meier overall survival curves for patients with the three subtypes. (E) Pairwise correlation clustering of TME-related genes. (F) Kaplan‒Meier overall survival curves of TCGA-KIRC patients grouped by coexpression program score (threshold set at the median value). (G) Boxplot showing the coexpression program score for each immune subtype. Wilcoxon test. (H) Expression pattern of the coexpression program across cell types. ns not significant; *P < 0.05; **P < 0.01; ***P < 0.001.
Additionally, we conducted pairwise correlation analysis on these 191 genes. The resulting correlation coefficient heatmap revealed a substantial coexpression program (Fig. 4E). The ssGSEA algorithm was used to score each sample based on this program, and it was found that patients with high program scores experienced poorer overall survival than those with low scores (Fig. 4F). Among the three subtypes, C2 displayed the highest program score, followed by C1, while C3 had the lowest score (Fig. 4G). Furthermore, we scored this expression program in individual cells in the scRNA-seq data, which indicated significant expression in CD8 + T cells, macrophages, and monocytes, underscoring its association with immune cell infiltration (Fig. 4H).
TME heterogeneity in ccRCC
We conducted a comprehensive analysis using the CIBERSORT and MCP-counter algorithms. Our results revealed that the C2 subtype, followed by the C1 subtype, exhibited significant immune cell infiltration, particularly CD8 + T cell infiltration, while the C3 subtype was immune cell deficient. Notably, the C1 and C2 subtypes showed reduced M1 macrophage infiltration, with C1 displaying the greatest infiltration of cancer-associated fibroblasts (Fig. 5A,B). To provide a more nuanced assessment of immune activity within the tumour microenvironment, we calculated four immune signature scores for each sample. The analysis indicated that the C2 subtype had the highest scores for CD8 + T-cell-related functions and phagocytosis. In contrast, the C1 subtype exhibited the highest angiogenesis score (Fig. 5C). Additionally, the results of the ESTIMATE algorithm indicated that the C2 subtype had the highest immune score, followed by the C1 subtype, and the C3 subtype had the lowest score. Furthermore, the C1 and C2 subtypes displayed higher stromal scores than did the C3 subtype (Fig. 5D). We also explored differences in functional pathway enrichment among the subtypes. Our findings revealed distinct pathway enrichment patterns across the three subtypes (Fig. 5E,F).
Heterogeneity of the three immune subtypes. (A) Boxplot showing differences in immune cell infiltration among the three subtypes. Kruskal‒Wallis test. (B) Heatmap displaying immune cell infiltration according to the MCP algorithm. (C) Boxplot showing the signature scores of four immune gene sets for each subtype. Wilcoxon test. (D) Boxplot representing the ESTIMATE algorithm results: stromal score and immune score for each subtype. Wilcoxon test. (E,F) Heatmap of ssGSEA enrichment results for hallmark pathways and KEGG pathways. ns not significant; *P < 0.05; **P < 0.01; ***P < 0.001.
Identification of key communication molecules
According to our previous findings, the C1 subtype exhibited high infiltration of cancer-associated fibroblasts and a lack of CD8 + T cells and had the poorest prognosis. We hypothesized that the C1 subtype exhibits an immunosuppressive microenvironment. To further substantiate this hypothesis and explore the mechanisms underlying this immunosuppressive microenvironment, we conducted a differential gene expression analysis between the C1 subtype and other subtypes, identifying 965 upregulated genes specific to the C1 subtype. KEGG and GO enrichment analyses of these genes highlighted their significant enrichment in pathways related to the extracellular matrix (Fig. 6A,B), consistent with our earlier observation of increased cancer-associated fibroblast infiltration in the C1 subtype.
Analysis of the immunosuppressive tumour microenvironment. (A,B) KEGG pathways and GO terms enriched in the upregulated genes in the C1 subtype. (C) Dot plot showing all significant ligand‒receptor interactions in the C1 subtype. (D) Boxplot showing the expression of all ligands and receptors among the three subtypes. Wilcoxon test. (E) Kaplan‒Meier analyses of overall survival in TCGA-KIRC patients, categorized by individual ligand and receptor expression levels (median value as the threshold). (F) Multivariate Cox regression analysis of prognostic factors. (G,H) Kaplan‒Meier analyses of overall survival between the low-risk and high-risk groups in the TCGA and CheckMate cohorts. ns not significant; *P < 0.05; **P < 0.01; ***P < 0.001.
Furthermore, we performed cell communication analysis using scRNA-seq data and identified all ligand‒receptor pairs involved in cellular communication within the tumour microenvironment. By intersecting these ligand-receptors with the upregulated genes in the C1 subtype, we identified specific cell communication patterns that were significantly enriched in the C1 subtype. These patterns shed light on the mechanisms shaping the immunosuppressive microenvironment in the C1 subtype. Specifically, we identified seven ligand‒receptor pairs, comprising a total of nine signalling molecules (Fig. 6C,D).
Prognostic model construction and validation
We conducted Kaplan‒Meier survival analysis for the nine essential signalling molecules and identified six prognosis-related genes (Fig. 6E). Subsequently, via multivariate Cox regression analysis, we identified three independent prognostic factors: MDK, CD44, and COL6A2 (Fig. 6F). Utilizing these factors, we constructed a prognostic model that stratified the TCGA cohort into high-risk and low-risk groups that exhibited significant differences in their prognoses (Fig. 6G). The validity of the model was further confirmed in the CheckMate cohort, and this analysis yielded consistent results (Fig. 6H). Multivariate Cox regression analysis of the prognostic model, in combination with clinical characteristics, further verified that the risk score is an independent prognostic factor for patients, substantiating the model's reliability (Supplementary Fig. S2A). The risk score exhibited robust performance in predicting overall survival in the TCGA cohort (AUC for 1-, 3-, and 5-year OS: 0.696, 0.644, and 0.637; Supplementary Fig. S2B). These findings were consistent with those in the CheckMate cohort (Supplementary Fig. S2C).
MDK predicts the response to immunotherapy
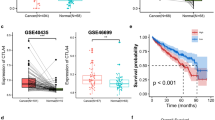
Three immunosuppressive microenvironment-related genes, which are also independent prognostic factors for ccRCC, were identified. However, their connection with immunotherapy efficacy remains unclear. To investigate this connection, we explored their relationships in the CheckMate cohort. The findings suggested that MDK might be a gene associated with the immunotherapy response and that patients with low MDK expression are more likely to benefit from immunotherapy than those with high MDK expression (Table 2).
In terms of cell communication, tumour cells, fibroblasts, and endothelial cells communicate with macrophages through MDK-LRP1 interactions (Fig. 7A,B). This led us to hypothesize that MDK-LRP1 influences macrophage polarization. To confirm this hypothesis, we examined the correlation between MDK-LRP1 expression and macrophage infiltration. The results revealed a significant positive correlation between MDK-LRP1 expression and M2 macrophage infiltration, while a negative correlation was observed with M1 macrophages (Fig. 7C,D). Additionally, MDK-LRP1 expression was significantly correlated with Treg and M0 macrophage infiltration (Fig. 7E,F) but inversely correlated with resting dendritic cells and mast cells (Fig. 7G,H). These findings collectively suggest that MDK-LRP1 expression promotes M2 macrophage polarization, contributing to the establishment of an immunosuppressive microenvironment.
Relationship between MDK-LRP1 and macrophage polarization. (A,B) Expression profiles of MDK and LRP1 across cell types. (C,D) Scatterplots revealing a significant positive correlation between MDK-LRP1 expression and M2 macrophage infiltration, while a significant negative correlation was observed with M1 macrophage infiltration. Spearman test. (E–H) Scatterplots demonstrating the significant positive correlation between MDK-LRP1 expression and Treg and M0 macrophage infiltration and the significant negative correlation between MDK-LRP1 expression and resting dendritic cell and mast cell infiltration. Spearman test.
Further exploration of the relationship between MDK-LRP1 and clinicopathological characteristics revealed a significant association between MDK expression and ccRCC pathological stage and grade (Supplementary Fig. S3A). Analysis of somatic mutation data indicated a substantial correlation between MDK expression and PBRM1 and BAP1 mutations. PBRM1 mutations were more prevalent in the MDK-low-expression group, whereas BAP1 mutations were more common in the MDK-high-expression group (Supplementary Fig. S3B–D). These findings highlight MDK as a pivotal molecule in the tumour microenvironment that promotes ccRCC progression.
MDK promotes M2 macrophage polarization to accelerate the migration, invasion and proliferation of ccRCC cells
We examined MDK levels in five pairs of peritumour and tumour tissues. The results demonstrated the upregulation of MDK in tumour tissues (Fig. 8A). Additionally, we assessed MDK expression in several RCC cell lines compared with that in normal renal tubular epithelial cells (HK2 cell lines) and observed MDK upregulation in the Caki-1 cell line (Fig. 8B,C). Using two independent small interfering RNAs (siRNAs), we successfully knocked down MDK in the ccRCC cell line Caki-1, as confirmed by Western blot and qPCR analyses (Fig. 8D,E).
MDK promotes immunosuppressive (M2) macrophage polarization. (A) Western blot analysis comparing MDK levels in five pairs of peritumoral and tumour tissues. (B,C) Western blotting and qPCR results showing MDK levels in four renal carcinoma cell lines. (D,E) Western blotting and qPCR results illustrating MDK expression in cells after MDK knockdown. (F) Representative images demonstrating morphological changes in M0 macrophages (from oval cells to antennal, spindle-shaped adherent cells) in the different treatment groups. (G) Quantitative data of M2 markers in the different treatment groups. (H) Relative percentage of CD163-positive cells in the different treatment groups.
To investigate the role of MDK in M2 macrophage polarization, a crucial regulator linking inflammation and cancer, differentially treated Caki-1 cells (ccRCC cells) were used to culture M0 macrophages. Our experiments revealed that treatment with conditioned medium from Caki-1 cells treated with si-MDK-2 resulted in reduced M0 macrophage polarization, as evidenced by fewer M0 macrophages in the CM-treated group than in the negative control group (Fig. 8F). Moreover, the relative RNA expression of M2 marker genes, namely, ARG1 and IL-10, indicated that MDK knockout led to M2 macrophage depolarization (Fig. 8G). Furthermore, flow cytometry analysis of the surface molecule CD163 in M2 macrophages revealed a 2.5-fold decrease (from 83.16% to 33.52%) in the proportion of CD163-positive cells when MDK expression was inhibited (Fig. 8H). Taken together, these results suggest that MDK promotes M2 macrophage polarization in ccRCC tissues.
M0 macrophages were treated with conditioned medium from si-MDK or si-NC Caki-1 cells, after which the conditioned medium-induced polarized macrophages were cocultured with Caki-1 cells. The invasion and migration of Caki-1 cells were significantly lower in the si-MDK group than in the si-NC group (Fig. 9A–C). Similarly, the proliferation ability of Caki-1 cells was also diminished in the si-MDK group (Fig. 9D,E). In vivo experiments involved injecting cocultured cells through the tail vein of C57BL/6J mice and monitoring lung metastatic nodules (Fig. 9F). The results demonstrated that macrophage polarization induced by cells transfected with si-MDK significantly suppressed the metastasis of Caki-1 cells.
Effect of MDK on ccRCC cell growth and aggressive behaviour. (A) Wound healing assay for Caki-1 cells cocultured with CM-induced polarized macrophages. (B,C) Representative images of migration and invasion assays for the indicated cells. Scale bars, 100 μm. (D,E) Colony formation and proliferation assays of Caki-1 cells cocultured with CM-induced polarized macrophages. (F) Representative images of lung metastasis specimens derived from the tail vein injected with the indicated cells.
Discussion
Clear cell renal cell carcinoma (ccRCC) is a prevalent primary renal cancer, and advanced ccRCC is very difficult to treat32. The effectiveness of immunotherapy remains suboptimal, largely because of the intricacies of the tumour microenvironment (TME)33. Elucidating the heterogeneity of the TME in ccRCC patients can guide the development of innovative therapeutic strategies. Leveraging recent advancements in single-cell sequencing technology and analytical methodologies, this study employed a comprehensive multiomics approach to dissect cell‒cell interactions34. This facilitated a deeper understanding of the tumour microenvironment (TME), facilitated prognosis prediction, and enabled the exploration of novel therapeutic strategies for clear cell renal cell carcinoma (ccRCC).
Previous ccRCC studies have focused mainly on specific cell types, so the applicability of the results has been limited35,36. In contrast, our strategy, which involved the integration of single-cell RNA (scRNA) and bulk RNA data, meticulously revealed genes associated with tumour progression, ensuring precision across all cell types. Notably, we discerned that elevated CD8 + T-cell infiltration did not consistently indicate poor prognosis. We revealed a more detrimental immunosuppressive subtype (C1) characterized by cancer-associated fibroblast infiltration, myeloid inflammation, and angiogenesis but lacking CD8 + T-cell infiltration.
C1 represents the immune-excluded subtype, C2 represents the immune-infiltrated subtype, and C3 represents the immune-desert subtype37. Although the C2 subtype was highly infiltrated with immune cells, it also displayed characteristics of high exhaustion. In contrast, the C3 subtype was devoid of CD8 + T-cell infiltration and showed enhanced fatty acid metabolism and Wnt/β-catenin signalling, as well as neuroendocrine features such as pancreatic β cells and steroid hormone biosynthesis.
For the C1 subtype, extensive extracellular matrix pathway activation hindered immune cell infiltration, shaping the immunosuppressive microenvironment. Unique intercellular communication patterns in C1 led to the development of a prognostic model comprising three genes that effectively predicted ccRCC patient outcomes. Notably, MDK exhibited strong correlations with the immunotherapy response, suggesting its potential as a therapeutic target.
MDK was predominantly expressed in tumour cells, fibroblasts, and endothelial cells, while LRP1 was highly expressed in macrophages. MDK-LRP1, a pivotal ligand‒receptor pair, is significantly correlated with M2 macrophage infiltration and plays a key role in immunosuppression by promoting M2 polarization. Clinical and in vitro validation experiments confirmed these findings. We found that MDK is highly expressed in renal tumours, and coculture of si-MDK-transfected ccRCC cells with macrophages significantly reduced macrophage M2 polarization. Furthermore, we cocultured macrophages induced by different conditions with tumour cells and found that such an immunosuppressive environment promoted tumour progression.
In addition, MDK was found to promote immunosuppression via coordinated activation of nuclear factor-κB and downregulation of interferon-associated pathways38. ErbB pathway-mutated gallbladder cancers upregulate MDK expression in epithelial cells via the PI3K/AKT pathway, potentiate the MDK-LRP1 interaction between malignant epithelial cells and macrophages, and promote the differentiation of macrophages into immunosuppressive M2 macrophages39. Our study also revealed the same effect in ccRCC and revealed that the PI3K/AKT pathway was enriched in genes upregulated in the C1 subtype. We also found that the expression of MDK was significantly related to PBRM1 and BAP1 mutations. Previous research has linked PBRM1 mutations, resulting in loss of protein function, to sensitivity to PD-1 inhibitors in advanced ccRCC40. The protein corresponding to PBRM1 is BAF180, which is involved in making up the SWI/SNF chromosome remodelling complex. Loss of function of the protein or complex activates an inflammatory response that releases cytokines such as IL-2 and IL-12 and the chemokine CCL21. Interestingly, BAP1 also appears to be involved in regulating immune responses via the SWI/SNF complex41. Loss of BAP1 expression is also associated with an immunosuppressive microenvironment42. Although the associations among PBRM1, BAP1, and immune responses are not well understood, MDK undoubtedly plays an essential role in immune remodelling.
Moreover, a study revealed that p53-mediated upregulation of MDK reshaped the immunosuppressive microenvironment43. Another investigation demonstrated that IFN-γ induces MDK expression through STAT1 activation in cancer44. Previous research revealed functional binding sites for retinoic acid receptor and hypoxia response elements in the promoter region of MDK that may facilitate MDK overexpression in diverse tumours45,46. Thus, MDK can be used as both a diagnostic marker and a promising emerging target in renal cancer. In fact, there is already a large amount of evidence proving the value of MDK as a therapeutic target47, and a series of MDK inhibitors, including antibodies, aptamers, GAGs, peptides, and low molecular weight compounds, are under preclinical development. It is believed that novel therapies targeting MDK and its associated pathways could improve the prognosis of ccRCC in the near future.
In summary, our study precisely classified ccRCC immunophenotypes, offering valuable insights for precise patient treatment. MDK, a crucial player in the immunosuppressive microenvironment, has emerged as a potential therapeutic target. Deeper investigations are warranted to elucidate the transcriptional activation mechanisms of MDK in advanced renal cancer.
Data availability
The datasets presented in this study are available in online repositories. Please refer to the “Materials and methods” section for the name of the repositories and accession numbers.
Abbreviations
- ccRCC:
-
Clear cell renal cell carcinoma
- TME:
-
Tumour microenvironment
- TAMs:
-
Tumour-associated macrophages
References
Hsieh, J. J. et al. Renal cell carcinoma. Nat. Rev. Dis. Primers 3, 17009 (2017).
Capitanio, U. & Montorsi, F. Renal cancer. Lancet 387, 894–906 (2016).
Kotecha, R. R., Motzer, R. J. & Voss, M. H. Towards individualized therapy for metastatic renal cell carcinoma. Nat. Rev. Clin. Oncol. 16, 621–633 (2019).
Capitanio, U. et al. Epidemiology of renal cell carcinoma. Eur. Urol. 75, 74–84 (2019).
Krishna, C. et al. Single-cell sequencing links multiregional immune landscapes and tissue-resident T cells in ccRCC to tumor topology and therapy efficacy. Cancer Cell 39, 662-677.e666 (2021).
Motzer, R. J. et al. Nivolumab plus ipilimumab versus sunitinib in advanced renal-cell carcinoma. N. Engl. J. Med. 378, 1277–1290 (2018).
de Miguel, M. & Calvo, E. Clinical Challenges of Immune Checkpoint Inhibitors. Cancer Cell 38, 326–333 (2020).
Rizzo, A. et al. Hypertransaminasemia in cancer patients receiving immunotherapy and immune-based combinations: The MOUSEION-05 study. Cancer Immunol. Immunother. 72, 1381–1394 (2023).
Dall’Olio, F. G. et al. Immortal time bias in the association between toxicity and response for immune checkpoint inhibitors: A meta-analysis. Immunotherapy 13, 257–270 (2021).
Guven, D. C. et al. The association between albumin levels and survival in patients treated with immune checkpoint inhibitors: A systematic review and meta-analysis. Front. Mol. Biosci. 9, 1039121 (2022).
Rizzo, A. et al. Quality of life assessment in renal cell carcinoma Phase II and III clinical trials published between 2010 and 2020: A systematic review. Future Oncol. 17, 2671–2681 (2021).
Wu, T. & Dai, Y. Tumor microenvironment and therapeutic response. Cancer Lett. 387, 61–68 (2017).
Hanahan, D. & Coussens, L. M. Accessories to the crime: Functions of cells recruited to the tumor microenvironment. Cancer Cell 21, 309–322 (2012).
Turley, S. J., Cremasco, V. & Astarita, J. L. Immunological hallmarks of stromal cells in the tumour microenvironment. Nat. Rev. Immunol. 15, 669–682 (2015).
Munn, D. H. & Bronte, V. Immune suppressive mechanisms in the tumor microenvironment. Curr. Opin. Immunol. 39, 1–6 (2016).
Galluzzi, L., Chan, T. A., Kroemer, G., Wolchok, J. D. & López-Soto, A. The hallmarks of successful anticancer immunotherapy. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aat7807 (2018).
Braun, D. A. et al. Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nat. Med. 26, 909–918 (2020).
Becht, E. et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 17, 218 (2016).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Langfelder, P. & Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Brunet, J. P., Tamayo, P., Golub, T. R. & Mesirov, J. P. Metagenes and molecular pattern discovery using matrix factorization. Proc. Natl. Acad. Sci. U. S. A. 101, 4164–4169 (2004).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951 (2019).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: An R package for comparing biological themes among gene clusters. Omics 16, 284–287 (2012).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7 (2013).
Long, Z. et al. Single-cell multiomics analysis reveals regulatory programs in clear cell renal cell carcinoma. Cell Discov. 8, 68 (2022).
Liberzon, A. et al. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Jin, S. et al. Inference and analysis of cell-cell communication using Cell Chat. Nat. Commun. 12, 1088 (2021).
Barata, P. C. & Rini, B. I. Treatment of renal cell carcinoma: Current status and future directions. CA Cancer J. Clin. 67, 507–524 (2017).
Riaz, N. et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171, 934-949.e916 (2017).
Song, Q. et al. Single-cell sequencing reveals the landscape of the human brain metastatic microenvironment. Commun. Biol. 6, 760 (2023).
Liu, B., Chen, X., Zhan, Y., Wu, B. & Pan, S. Identification of a gene signature for renal cell carcinoma-associated fibroblasts mediating cancer progression and affecting prognosis. Front. Cell Dev. Biol. 8, 604627 (2020).
Wu, X. et al. CD8(+) T cell-based molecular classification with heterogeneous immunogenomic landscapes and clinical significance of clear cell renal cell carcinoma. Front. Immunol. 12, 745945 (2021).
Hegde, P. S. & Chen, D. S. Top 10 challenges in cancer immunotherapy. Immunity 52, 17–35 (2020).
Cerezo-Wallis, D. et al. Midkine rewires the melanoma microenvironment toward a tolerogenic and immune-resistant state. Nat. Med. 26, 1865–1877 (2020).
Zhang, Y. et al. Single-cell RNA-sequencing atlas reveals an MDK-dependent immunosuppressive environment in ErbB pathway-mutated gallbladder cancer. J. Hepatol. 75, 1128–1141 (2021).
Miao, D. et al. Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science 359, 801–806 (2018).
Louie, B. H. & Kurzrock, R. BAP1: Not just a BRCA1-associated protein. Cancer Treat. Rev. 90, 102091 (2020).
Figueiredo, C. R. et al. Loss of BAP1 expression is associated with an immunosuppressive microenvironment in uveal melanoma, with implications for immunotherapy development. J. Pathol. 250, 420–439 (2020).
Meng, X. et al. DNA damage repair alterations modulate M2 polarization of microglia to remodel the tumor microenvironment via the p53-mediated MDK expression in glioma. EBioMedicine 41, 185–199 (2019).
Zheng, L. et al. Targeting MDK abrogates IFN-γ-elicited metastasis in cancers of various origins. Front. Oncol. 12, 885656 (2022).
Pedraza, C., Matsubara, S. & Muramatsu, T. A retinoic acid-responsive element in human midkine gene. J. Biochem. 117, 845–849 (1995).
Reynolds, P. R., Mucenski, M. L., Le Cras, T. D., Nichols, W. C. & Whitsett, J. A. Midkine is regulated by hypoxia and causes pulmonary vascular remodeling. J. Biol. Chem. 279, 37124–37132 (2004).
Filippou, P. S., Karagiannis, G. S. & Constantinidou, A. Midkine (MDK) growth factor: A key player in cancer progression and a promising therapeutic target. Oncogene 39, 2040–2054 (2020).
Acknowledgements
This work was supported by Biobank, Zhongda Hospital, Southeast University.
Funding
This work was supported by the National Natural Science Foundation of China (grant numbers 823B2067), Medical Research Project of Jiangsu Provincial Health Commission (ZD2022010).
Author information
Authors and Affiliations
Contributions
Ding X.F. and Chen M. designed and supervised the study. Shi N.P. and Wang D. collected and analysed data and wrote the original draft. Wu T.G. revised the paper. Chen S.S. and Zhang N.K. conducted experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shi, N., Chen, S., Wang, D. et al. MDK promotes M2 macrophage polarization to remodel the tumour microenvironment in clear cell renal cell carcinoma. Sci Rep 14, 18254 (2024). https://doi.org/10.1038/s41598-024-69183-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-69183-z
- Springer Nature Limited