Abstract
The bidirectional effect of hyperuricemia on chronic kidney disease (CKD) underscores the importance of hyperuricemia as a risk factor for CKD. We evaluated the effect of hyperuricemia on the presence and development of CKD after considering genetic background by calculating polygenic risk scores (PRSs). We employed genome-wide association study summary statistics—excluding the United Kingdom Biobank (UKB) datasets among published CKD Gen Consortium papers—to calculate the PRSs for CKD in white background subjects. To validate PRS performance, we divided the UKB into two datasets to validate and test the data. We used logistic regression analysis to evaluate the association between hyperuricemia and CKD, and performed Kaplan–Meier survival analysis exclusively for subjects with available follow-up data. In total, 438,253 clinical data and 4,307,940 single nucleotide polymorphisms from 459,155 samples were included. We observed a significant positive association between PRS and CKD and the presence and development of CKD. Hyperuricemia significantly increased CKD risk (adjusted odds ratio 1.55, 95% confidence interval 1.48–1.61). The impact of hyperuricemia on CKD was maintained irrespective of PRS range. In addition, negative interaction between hyperuricemia and PRS for CKD was found. Survival analysis indicates that the presence of hyperuricemia significantly increased the risk of CKD development. The PRS for CKD thoroughly reflects the risk of CKD development. Hyperuricemia is a significant indicator of CKD risk, even after incorporating the genetic risk score for CKD. Irrespective of genetic risk, patients with a prospective risk of developing CKD require uric acid monitoring and management.
Similar content being viewed by others
Introduction
Chronic kidney disease (CKD) is a significant public health issue with an increasing prevalence1,2. As a progressive condition, identifying risk factors and prognostic factors, particularly those that can be modified, could significantly reduce the disease burden. Hyperuricemia is an emerging non-traditional risk factor for the development and progression of CKD3,4. Although epidemiological studies have shown that hyperuricemia increases the risk of kidney dysfunction,5,6 the role of hyperuricemia in CKD is not yet fully established.
Uric acid functions as an antioxidant in the extracellular environment, and plays a protective role in neurological diseases such as multiple sclerosis7,8. However, intracellular uric acid acts as a pro-oxidant, reducing endothelial nitric oxide activity and inhibiting cellular proliferation and migration9,10. Hyperuricemia can trigger hypertension and accelerate the progression of CKD as a result of long-term activation of the renin–angiotensin–aldosterone system, oxidative stress, and the loss of endothelial nitric oxide11,12,13. In addition, a high level of serum uric acid has been found to be independently associated with CKD, metabolic syndrome, and cardiovascular disease14,15.
Since the development of uric acid-lowering agents, may experimental and clinical studies have been conducted to demonstrate their disease-preventive effects16,17. Randomized trials have shown that allopurinol or febuxostat, which reduce uric acid levels, have a positive impact on systolic blood pressure, estimated glomerular filtration, and albuminuria18,19,20. Based on these results, several studies have attempted to identify genetic polymorphisms that affect serum uric acid levels and increase the risk of CKD21,22. However, the association between a genetic urate score and CKD has not yet been confirmed.
Recently, it was shown that polygenic risk scores (PRSs) calculated based on large-scale genome-wide association studies (GWAS) can estimate the effect of genetic variants on the risk of the disease of interest. Considering the complexity of the risk factors contributing to CKD development, the PRS for CKD was calculated to evaluate the genetic effects on CKD and their interactions with other risk factors of interest. We aimed to evaluate whether hyperuricemia has an additional effect on the risk of CKD, in addition to the genetic risk assessed by the PRS, given its nature as a modifiable risk factor.
Methods
Study populations and data acquisition
We utilized the United Kingdom Biobank (UKB) prospective cohort data (https://www.ukbiobank.ac.uk)23. The UKB is a large-scale database which comprises the data of 502,413 individuals aged between 40 and 69 years. For the phenotype dataset, we excluded subjects with non-white background, without data for serum creatinine, cystatin C, without applicable data for CKD or ESKD based on the ICD-10 code. We focused on non-Hispanic white subjects, who contributed to 94% of the UKB cohort, resulting in sample of 438,253 subjects for our final analysis. For the sensitivity analysis with longitudinal data set, we collected the second time repeated measurements of date of attending assessment center, which includes kidney-related phenotypes such as serum creatinine, cystatin C, ICD-10 code for CKD or ESKD.
Definition of clinical outcome and exposures
CKD was considered the main outcome, and we defined CKD as the presence of one or more of the following conditions in the baseline or second stage cohort of the UKB database: 1) estimated glomerular filtration rate (eGFR) < 60 mL/min/1.73 m2 by creatinine (field 30,700) or cystatin C (field 30,720) based on the CKD Epidemiology Collaboration (CKD-EPI) equation; 2) a diagnostic code for CKD stage 3–5 (N18.3–N18.5) or end-stage kidney disease (ESKD) (N18.6, Z94.0) based on the International Classification of Disease 10th revision (ICD-10) code (field 41,270); or 3) a record of ESKD diagnosis (field 42,026 or 42,027).
As the main exposure, uric acid was used as a continuous and categorical variable, defining hyperuricemia as urate levels > 420 μmol/L in males and > 360 μmol/L in females.
Genotyping, quality control, and imputation
Among the 502,413 subjects in the UKB, 50,000 were genotyped using the UK Biobank Lung Exome Variant Evaluation (UK BiLEVE) Axiom Array by Affymetrix and the remaining subjects were genotyped using the UK Biobank Axiom Array. Pre-phasing was carried out with SHAPEIT324 with 1000 Genome phase 3,25 and imputation of untyped single nucleotide polymorphisms (SNPs) was performed with IMPUTE4 (https://jmarchini.org/software/)26 using UK10K, 1000 Genome phase 3, and the HRC panel,27 resulting in 93,095,623 autosomal SNPs. The imputed dataset was downloaded and quality control was performed. SNPs were filtered out if the missing genotype rates were ≥ 0.01, Hardy–Weinberg Equilibrium P values were < 10−6, or minor allele frequencies (MAF) were < 0.01. Subjects were excluded if the they were not from a White background. The detailed procedure is illustrated in Fig. 1. All data management was performed using PLINK,28 PLINK2,29 GCTA,30 and ONETOOL31.
GWAS and heritability analyses
We estimated SNP heritability and genetic correlations using linkage disequilibrium (LD) score regression32. LD score regression requires summary statistics from GWASs, and GWASs were conducted with the UKB dataset using logistic regressions for CKD and hyperuricemia after adjusting for the effects of baseline age, sex, and ten principal components (PC) corresponding to the top 10 largest eigenvalues.
PRS calculation and its evaluation
PRS derivation requires GWAS summary statistics and we considered the following GWAS summary data which did not include UKB dataset among the published CKD Gen Consortium papers (https://ckdgen.imbi.uni-freiburg.de/)33. Results from SNPs with MAF ≥ 0.005 were utilized for PRS. PRSs were calculated with clumping + thresholding (CT),34 LDpred with infinitesimal, grid and auto models,35,36 lassosum,37 and PRS continuous shrinkage (CS)38. For CT, we set the clumping at P = 10–5 and pruning at r2 = 0.2. For the LDpred grid model, the proportions of causal variants \(\rho\) were set at 0.03, 0.01, 0.3, 0.1, and 1. A 1000 Genomes Phase 3 European subpopulation was used as the linkage disequilibrium (LD) reference panel. For the other hyperparameters, we used the default values. To examine the prevalence of CKD as an increase in PRS, the calculated PRSs for CKD were categorized into deciles. In addition, for logistic regression, we re-defined PRS groups as tertile ranges, and 2nd tertile range was regarded as a reference in the logistic regression model for evaluating the effect of PRS on prevalent CKD.
To validate the PRS performance, we separated the UKB into two datasets of a white background group to validate and test the data. For validation and testing, we used 47,611 subjects genotyped using the UK BiLEVE array chip, and 390,642 subjects genotyped using the Axiom chip. Validation data was used to fit the logistic regression of PRS on CKD after adjusting for baseline age, sex, PRS, and PC 1–10 on CKD outcome. Akaike information criterion (AIC) was used to choose the best PRS method.
The PRS significance was evaluated by applying logistic regression to the test data. We included baseline age, sex, eGFR, body mass index (BMI), smoking status, physical activity, comorbidities (type 2 diabetes mellitus [T2DM], hypertension [HTN], cardiovascular diseases [CVD]), hemoglobin, glucose, albumin, calcium, cholesterol and PC 1–10 as covariates. We also conducted survival analyses using the test data. The age at CKD onset was considered the main outcome, and the same covariates as in the logistic regression were used. Survival analyses were conducted using the survival package39 (version 3.4.0) and ggsurvfit package40 (version 0.2.0) of R (version 4.5.0).
PRS analysis results for different models are shown in Supplementary Table S1. Among SNPs with GWAS results from Wuttke et al.33, genotypes for 3,898,527 SNPs were observed in UKB, and they were used for PRS analyses. LDpred grid1 (P = 0.03) had the best performance (P = 1.18 × 10–43 and of model AIC = 25,199.84), and was used for subsequent analysis.
Statistical analysis
The baseline characteristics of subjects with and without hyperuricemia were compared using the Student’s t-test and the chi-square test. Logistic regression analysis was used to evaluate the association between hyperuricemia and CKD, adjusting for covariates such as age, sex, BMI, smoking status, comorbidities (HTN, T2DM, and CVD), physical activity, blood laboratory findings (hemoglobin, glucose, albumin, calcium, and cholesterol), PRS, and PC 1–10. Adjusted odds ratios (ORs) and corresponding 95% confidence intervals (CIs) were estimated for incident CKD outcomes based on the PRSs. To rigorously investigate the interaction between hyperuricemia and PRS for CKD, we incorporated an interaction term as a covariate in the logistic regression analysis. In addition, we categorized the participants into six groups for analysis based on the tertile range of PRS and hyperuricemia status: PRS for CKD 1st tertile & hyperuricemia (−), PRS for CKD 1st tertile & hyperuricemia (+), PRS for CKD 2nd tertile & hyperuricemia (−), PRS for CKD 2nd tertile & hyperuricemia (+), PRS for CKD 3rd tertile & hyperuricemia (−), and PRS for CKD 3rd tertile & hyperuricemia (+). The group PRS for CKD 1st tertile & hyperuricemia (−) was used as the reference. Furthermore, we assessed the impact of hyperuricemia on CKD development based on sex (male and female), baseline age (age < 60 and age ≥ 60), and T2DM (control and T2DM).
We performed Kaplan–Meier survival analysis to access the risk of CKD development among subjects without underlying CKD who were sequentially followed up in the UKB database. We used hyperuricemia status and PRS grade as independent variables, and the newly diagnosed CKD as the outcome. All analyses were performed using R software, version 3.6.3 (R Foundation) and Rex. All Two-sided P values were two-sided reported, and P < 0.05 was considered to indicate statistical significance41.
Ethical considerations
This study was performed in accordance with the principles of the Declaration of Helsinki and approved by the Institutional Review Board of Seoul National University Boramae Medical Center (IRB No. 07-2022-45). The usage of the UK Biobank data was approved by the UK Biobank consortium (application No. 53799). Acquisition of informed consent was not required, as the study investigated anonymous public databases and genetic summary statistics.
Results
Study populations
A total of 438,253 subjects with available clinical data were included in the study, of which 28,970 (6.6%) had CKD and 57,656 (13.2%) had hyperuricemia. Subjects with hyperuricemia were more likely to be male, older, have a higher BMI, and more likely to have HTN, T2DM, and CVD (Table 1). In addition, subjects with hyperuricemia showed a lower eGFR and a higher incidence of CKD, greater albuminuria, higher serum glucose, lower high-density lipoprotein cholesterol, higher low-density lipoprotein cholesterol, and higher triglyceride levels (Table 1).
Heritability analyses and genetic correlation
The estimated heritability from the summary statistics of GWAS on CKD and hyperuricemia were 0.0292 (SE = 0.0021, P = 5.93 × 10–44) and 0.0709 (SE = 0.009, P = 3.33 × 10–15), respectively. Their genetic correlation was 0.5076 (SE = 0.0505, P = 9.55 × 10–24).
Association between PRSs and the risk of CKD
The prevalence of CKD increased from 1645 (4.23%) to 3624 (9.28%) between the 1st decile and the 10th decile (Supplementary Table S2). Different PRS densities in subjects with and without CKD are shown in Supplementary Figure S1.
According to the PRSs, subjects within the 1st tertile of PRSs showed a significantly decreased risk of the development of CKD (OR 0.78), and it was maintained after adjustment of covariates (aOR 0.93, 95% CI 0.89–0.97). On the contrary, subjects within the 3rd tertile showed a significantly increased risk for the development of CKD (OR 1.31), and it was also maintained after adjustment of covariates (aOR 1.12, 95% CI 1.08–1.17) (Table 2).
In the stratified analysis using the tertile ranges of PRS, there was a decreased risk of CKD in the 1st tertile of PRS and an increased risk of CKD in the 3rd tertile of PRS in females and in subjects without T2DM. The impact of PRS was significant irrespective of age groups stratified by a cut-off age of 60 years (Supplementary Table S3).
Association between hyperuricemia and prevalent CKD
As a continuous variable, high uric acid levels increased the risk of CKD (Fig. 2). The presence of hyperuricemia (aOR 1.55, 95% CI 1.48–1.61) and higher PRS (aOR 1.12, 95% CI 1.09–1.14) increased risk of CKD, respectively. Additionally, there were a significant negative interaction between hyperuricemia and PRS for CKD (Table 3).
Spline curve for the risk of CKD based on the level of uric acid. Multivariable logistic regression included covariates including age, sex, body mass index, smoking status, comorbidities (hypertension, diabetes, cardiovascular disease), physical activity, laboratory findings (hemoglobin, serum glucose, albumin, calcium, cholesterol, estimated glomerular filtration rate), PC1 ~ 10, PRS for CKD, interaction between urate and PRS for CKD. CKD, chronic kidney disease.
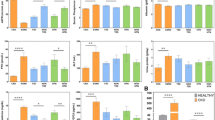
To enhance intuitive comprehension, we conducted an integrated grouping analysis based on the PRS for CKD and the presence of hyperuricemia. Compared to subjects within the 1st tertile of PRS without hyperuricemia, there was a trend of increasing risk of CKD in subjects within the 2nd and 3rd tertile of PRS; additionally, the risk of CKD was particularly prominent among the subjects with hyperuricemia (Fig. 3).
Effect of hyperuricemia on CKD according to the PRS group. PRS group was divided into tertile ranges, and subjects included 1st terile of PRS without hyperuricemia was regarded as a reference group. The X-axis represents the adjusted odds ratio, and the covariates are as follows. Multivariable logistic regression included covariates including age, sex, body mass index, smoking status, comorbidities (hypertension, diabetes, cardiovascular disease), physical activity, laboratory findings (hemoglobin, serum glucose, albumin, calcium, cholesterol, estimated glomerular filtration rate), PC1 ~ 10. CKD, chronic kidney disease; PRS, polygenic risk score.
In the stratified analysis based on sex, the significance of hyperuricemia was higher in females (aOR 1.88, 95% CI 1.69–2.08) than males (aOR 1.29, 95% CI 1.17–1.42) after adjustment for CKD. In addition, the significance of hyperuricemia in CKD was maintained irrespective of age and the presence of T2DM (Supplementary Table S4).
Risk of the development of CKD
A total of 14,699 subjects with follow-up data were included in the survival analysis. During the 51.9 ± 11.0 months of follow-up period, 114 patients (0.8%) were newly diagnosed with CKD. According to the PRS grade, 34 (0.7%), 27 (0.6%), and 53 (1.1%) patients had newly developed CKD in the 1st tertile, 2nd tertile, and 3rd tertile of the PRS grade, respectively. The different baseline characteristics according to PRS grades are shown in Supplementary Table S5. Except for kidney function-related indicators, there were no significant differences in characteristics according to PRS grade. In the survival analysis for CKD development according to the tertile grades of PRS, subjects within the 3rd tertile of PRS showed the highest risk of incident CKD (Supplementary Figure S2).
A total of 2382 (16.2%) patients had hyperuricemia. Participants with hyperuricemia were older, had a higher prevalence of obesity, and engaged in less physical activity. Although there were statistically significant differences in laboratory results between the two groups, these differences were found to be subtle (Supplementary Table S6). Hyperuricemia also significantly increased the risk of developing CKD (Supplementary Figure S3).
Discussion
As a significant modifiable risk factor, we investigated the effect of hyperuricemia on the development of CKD, adding to the genetic risk of CKD, which was estimated based on the PRS. PRSs for CKD were significantly associated with the presence and development of CKD. Considering these genetic risk factors for CKD with PRSs, hyperuricemia was found to play a major role in both the development and presence of CKD. Regardless of the genetic risk of CKD, hyperuricemia may be a significant factor that needs to be managed.
CKD involves complex disease entities including genetic and environmental factors. Significant progress has been made in human genetics in recent decades, leading to improvements in high-throughput genotyping, sequencing technologies, statistical genetics, and bioinformatics. Currently, 600 genes are implicated in monogenic kidney diseases, and GWASs have suggested that hundreds of genes contribute to complex kidney diseases33,42,43,44. The development of GWASs with large-scale population-based data significantly facilitates the risk stratification of CKD using PRS. In this study, we also found that the PRS for CKD showed a positive association with the risk of CKD, even after adjusting for well-known risk factors.
Uric acid is usually generated in the liver and intestine as a byproduct of the purine metabolic pathway, and the kidney has responsible for excreting two-thirds of the uric acid45. Although uric acids have a potent role as an antioxidant that can neutralize superoxide, hydroxyl radicals, and singlet oxygen,46 it has been widely accepted that hyperuricemia with hyperuricosuria causes kidney disease by depositing intraluminal crystal in the collecting duct of the nephron, similar to how gout does with the joints47,48. In addition, epidemiological studies have demonstrated that higher levels of uric acid are significantly related to an increased risk of CKD4,49,50. Nevertheless, the causal relationship between hyperuricemia and CKD is not clearly understood because of the bidirectional effects between uric acid and kidney function and the abundance of confounding factors.
Similar to attempts to determine the genetic effect on CKD, GWASs have uncovered roughly 30 loci that regulate serum uric acid, including uric acid transporters and regulatory transporter-associated proteins51,52. Moreover, among 50 loci associated with eGFR and CKD, nine were identified that are commonly associated with serum uric acid concentration53. Although a causal relationship between hyperuricemia and CKD has not been established yet, we found that there is a relative genetic correlation between CKD and hyperuricemia in this study. Nevertheless, the relationship between uric acid and CKD cannot be easily understood because of the uncontrollable and complex pathway that involves genetic factors related to the regulation of uric acid levels and such target diseases, as well as environmental and/or dietary factors. Therefore, using the PRS for CKD, we evaluated the additional impact of hyperuricemia on CKD. Finally, we found that hyperuricemia played a significant role in increasing the risk of CKD, regardless of the genetic risk score for CKD.
Our findings demonstrate that the effect of hyperuricemia on CKD supports those of previously reported randomized clinical trials demonstrating the efficacy of uric acid-lowering agents in preserving kidney function18,19,20. Recent research involving advanced CKD patients with a high risk of progression found no statistically significant differences between allopurinol and the control group regarding eGFR decline54. Another clinical trial using febuxostat also showed an insignificant effect on preserving kidney function in advanced CKD patients55. These findings may be related to an attenuation of the effectiveness of pharmacological management due to confounding factors that expand as CKD progresses. Despite the negative results of pharmacological interventions for uric acid control, it is undeniable that uric acid plays a crucial role as a marker, correlate, or representative of metabolic syndrome, cardiovascular disease, and CKD. Our findings, along with those of other studies, indicate that uric acid may be a useful prognostic marker in clinical practice.
This study demonstrates that hyperuricemia has a significant impact on CKD, even after considering the genetic risk score for CKD. We calculated the PRS based on large-population cohort data and validated these scores using an independent data set. However, this study has several limitations. Firstly, while our analysis demonstrates that PRS significantly impact CKD development, the ability to predict individual disease risk based on PRS alone is limited. This is because CKD is influenced by a multitude of factors beyond genetic predisposition. Although we adjusted for certain clinical variables to focus on the pure genetic effect, our study does not fully capture the complexity of CKD etiology, including various environmental and lifestyle factors, which are known to contribute to CKD risk. Ancestry is the parameter that varies the most when calculating the PRS; therefore, it should be a requirement that individuals of the same ancestry be treated as a single population group when calculating scores. In this regard, we included only non-Hispanic whites and the results could not be adjusted for overall ancestry.
Although we performed survival analysis in subpopulations with follow-up data, we could not conduct a Cox proportional hazard analysis that considers various covariates due to the insufficient number of subjects and the limited duration of follow-up. We used the UKB data, which consists of healthy general populations, therefore, healthy volunteer bias should be considered. Despite the genetic correlation between hyperuricemia and CKD, we could not consider share effects in the PRS analysis. Lastly, we could not evaluate medication history, including the use of uric acid lowering agents. To generalize the findings, further research that restricts the analysis to other ethnic populations and considers more comprehensive factors, including drug implications, is needed. Furthermore, given the significant interaction between hyperuricemia and PRS on CKD risk, identifying dietary, life-style, environmental, and genetic risks of hyperuricemia, which are not included in the CKD PRS, may be beneficial for CKD prevention and treatment.
This study demonstrates that hyperuricemia is a significant risk factor representing the risk of CKD, even after considering the genetic risk score for CKD. Monitoring and managing uric acid levels are warranted, especially in patients with a potential risk for CKD, regardless of genetic risk.
Data availability
All data used in the present study were provided by the UK biobank and could be downloaded at https://www.ukbiobank.ac.uk.
References
Lv, J. C. & Zhang, L. X. Prevalence and disease burden of chronic kidney disease. Adv. Exp. Med. Biol. 1165, 3–15. https://doi.org/10.1007/978-981-13-8871-2_1 (2019).
Jung, S. et al. Risk of mortality and cause of death according to kidney function parameters: A nationwide observational study in Korea. Kidney Res. Clin. Pract. 43, 202–215. https://doi.org/10.23876/j.krcp.22.088 (2024).
Srivastava, A., Kaze, A. D., McMullan, C. J., Isakova, T. & Waikar, S. S. Uric acid and the risks of kidney failure and death in individuals with CKD. Am. J. Kidney Dis. 71, 362–370. https://doi.org/10.1053/j.ajkd.2017.08.017 (2018).
Zhu, P., Liu, Y., Han, L., Xu, G. & Ran, J. M. Serum uric acid is associated with incident chronic kidney disease in middle-aged populations: A meta-analysis of 15 cohort studies. PLoS One 9, e100801. https://doi.org/10.1371/journal.pone.0100801 (2014).
Borghi, C. et al. Serum uric acid and the risk of cardiovascular and renal disease. J. Hypertens. 33, 1729–1741. https://doi.org/10.1097/hjh.0000000000000701 (2015) (discussion 1741).
Borghi, C. & Cicero, A. F. G. Serum uric acid and cardiometabolic disease: Another brick in the wall?. Hypertension 69, 1011–1013. https://doi.org/10.1161/hypertensionaha.117.09081 (2017).
Justicia, C. et al. Uric acid is protective after cerebral ischemia/reperfusion in hyperglycemic mice. Transl. Stroke Res. 8, 294–305. https://doi.org/10.1007/s12975-016-0515-1 (2017).
Hooper, D. C. et al. Uric acid, a natural scavenger of peroxynitrite, in experimental allergic encephalomyelitis and multiple sclerosis. Proc. Natl. Acad. Sci. USA 95, 675–680. https://doi.org/10.1073/pnas.95.2.675 (1998).
Sautin, Y. Y., Nakagawa, T., Zharikov, S. & Johnson, R. J. Adverse effects of the classic antioxidant uric acid in adipocytes: NADPH oxidase-mediated oxidative/nitrosative stress. Am. J. Physiol. Cell Physiol. 293, C584-596. https://doi.org/10.1152/ajpcell.00600.2006 (2007).
Kang, D. H., Park, S. K., Lee, I. K. & Johnson, R. J. Uric acid-induced C-reactive protein expression: implication on cell proliferation and nitric oxide production of human vascular cells. J. Am. Soc. Nephrol. 16, 3553–3562. https://doi.org/10.1681/asn.2005050572 (2005).
Kang, D. H. et al. A role for uric acid in the progression of renal disease. J. Am. Soc. Nephrol. 13, 2888–2897. https://doi.org/10.1097/01.asn.0000034910.58454.fd (2002).
Nakagawa, T. et al. Hyperuricemia causes glomerular hypertrophy in the rat. Am. J. Nephrol. 23, 2–7. https://doi.org/10.1159/000066303 (2003).
Mazzali, M. et al. Hyperuricemia induces a primary renal arteriolopathy in rats by a blood pressure-independent mechanism. Am. J. Physiol. Renal. Physiol. 282, F991-997. https://doi.org/10.1152/ajprenal.00283.2001 (2002).
Masuo, K., Kawaguchi, H., Mikami, H., Ogihara, T. & Tuck, M. L. Serum uric acid and plasma norepinephrine concentrations predict subsequent weight gain and blood pressure elevation. Hypertension 42, 474–480. https://doi.org/10.1161/01.Hyp.0000091371.53502.D3 (2003).
Kuwabara, M. et al. Asymptomatic hyperuricemia without comorbidities predicts cardiometabolic diseases: Five-year Japanese cohort study. Hypertension 69, 1036–1044. https://doi.org/10.1161/hypertensionaha.116.08998 (2017).
Sánchez-Lozada, L. G. et al. Treatment with the xanthine oxidase inhibitor febuxostat lowers uric acid and alleviates systemic and glomerular hypertension in experimental hyperuricaemia. Nephrol. Dial. Transpl. 23, 1179–1185. https://doi.org/10.1093/ndt/gfm783 (2008).
Kim, H. S. et al. The protective effect of febuxostat on chronic tacrolimus-induced nephrotoxicity in rats. Nephron 135, 61–71. https://doi.org/10.1159/000449289 (2017).
Goicoechea, M. et al. Effect of allopurinol in chronic kidney disease progression and cardiovascular risk. Clin. J. Am. Soc. Nephrol. 5, 1388–1393. https://doi.org/10.2215/cjn.01580210 (2010).
Whelton, A., Macdonald, P. A., Zhao, L., Hunt, B. & Gunawardhana, L. Renal function in gout: Long-term treatment effects of febuxostat. J. Clin. Rheumatol. 17, 7–13. https://doi.org/10.1097/RHU.0b013e318204aab4 (2011).
Goicoechea, M. et al. Allopurinol and progression of CKD and cardiovascular events: Long-term follow-up of a randomized clinical trial. Am. J. Kidney Dis. 65, 543–549. https://doi.org/10.1053/j.ajkd.2014.11.016 (2015).
Hughes, K., Flynn, T., de Zoysa, J., Dalbeth, N. & Merriman, T. R. Mendelian randomization analysis associates increased serum urate, due to genetic variation in uric acid transporters, with improved renal function. Kidney Int. 85, 344–351. https://doi.org/10.1038/ki.2013.353 (2014).
Yang, Q. et al. Multiple genetic loci influence serum urate levels and their relationship with gout and cardiovascular disease risk factors. Circ. Cardiovasc. Genet. 3, 523–530. https://doi.org/10.1161/circgenetics.109.934455 (2010).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209. https://doi.org/10.1038/s41586-018-0579-z (2018).
O’Connell, J. et al. Haplotype estimation for biobank-scale data sets. Nat. Genet. 48, 817–820 (2016).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74. https://doi.org/10.1038/nature15393 (2015).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLOS Genetics 5, e1000529. https://doi.org/10.1371/journal.pgen.1000529 (2009).
Consortium, the Haplotype Reference Consortium. A reference panel of 64,976 haplotypes for genotype imputation. Nature genetics 48, 1279-1283 (2016).
Purcell, S. et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575. https://doi.org/10.1086/519795 (2007).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience https://doi.org/10.1186/s13742-015-0047-8 (2015).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: A tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82. https://doi.org/10.1016/j.ajhg.2010.11.011 (2011).
Song, Y. E. et al. ONETOOL for the analysis of family-based big data. Bioinformatics 34, 2851–2853 (2018).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295. https://doi.org/10.1038/ng.3211 (2015).
Wuttke, M. et al. A catalog of genetic loci associated with kidney function from analyses of a million individuals. Nat. Genet. 51, 957–972. https://doi.org/10.1038/s41588-019-0407-x (2019).
Euesden, J., Lewis, C. M. & O’Reilly, P. F. PRSice: Polygenic risk score software. Bioinformatics 31, 1466–1468. https://doi.org/10.1093/bioinformatics/btu848 (2014).
Vilhjálmsson, B. J. et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet. 97, 576–592. https://doi.org/10.1016/j.ajhg.2015.09.001 (2015).
Privé, F., Arbel, J. & Vilhjálmsson, B. J. LDpred2: Better, faster, stronger. Bioinformatics 36, 5424–5431. https://doi.org/10.1093/bioinformatics/btaa1029 (2020).
Mak, T. S. H., Porsch, R. M., Choi, S. W., Zhou, X. & Sham, P. C. Polygenic scores via penalized regression on summary statistics. Genet. Epidemiol. 41, 469–480. https://doi.org/10.1002/gepi.22050 (2017).
Ge, T., Chen, C.-Y., Ni, Y., Feng, Y.-C.A. & Smoller, J. W. Polygenic prediction via Bayesian regression and continuous shrinkage priors. Nat. Commun. 10, 1776. https://doi.org/10.1038/s41467-019-09718-5 (2019).
Therneau, T. M. (2022).
Sjoberg, D. D., Baillie, M., Fruechtenicht, C., Haesendonckx, S., & Treis, T. ggsurvfit: Flexible Time-to-Event Figures, https://github.com/ddsjoberg/ggsurvfit, http://www.danieldsjoberg.com/ggsurvfit/ (2022).
Lee, B., An, J., Lee, S. & Won, S. Rex: R-linked EXcel add-in for statistical analysis of medical and bioinformatics data. Genes Genomics 45, 295–305. https://doi.org/10.1007/s13258-022-01361-7 (2023).
Rasouly, H. M. et al. The burden of candidate pathogenic variants for kidney and genitourinary disorders emerging from exome sequencing. Ann. Intern. Med. 170, 11–21. https://doi.org/10.7326/m18-1241 (2019).
Boyle, E. A., Li, Y. I. & Pritchard, J. K. An expanded view of complex traits: From polygenic to omnigenic. Cell 169, 1177–1186. https://doi.org/10.1016/j.cell.2017.05.038 (2017).
Park, S. et al. Genetic variations in HMGCR and PCSK9 and kidney function: A Mendelian randomization study. Kidney Res. Clin. Pract. 42, 460–472. https://doi.org/10.23876/j.krcp.22.237 (2023).
Gondouin, B. et al. Plasma xanthine oxidase activity is predictive of cardiovascular disease in patients with chronic kidney disease independently of uric acid levels. Nephron 131, 167–174. https://doi.org/10.1159/000441091 (2015).
Ames, B. N., Cathcart, R., Schwiers, E. & Hochstein, P. Uric acid provides an antioxidant defense in humans against oxidant- and radical-caused aging and cancer: A hypothesis. Proc. Natl. Acad. Sci. U S A 78, 6858–6862. https://doi.org/10.1073/pnas.78.11.6858 (1981).
Waisman, J., Mwasi, L. M., Bluestone, R. & Klinenberg, J. R. Acute hyperuricemic nephropathy in rats. An electron microscopic study. Am. J. Pathol. 81, 367–378 (1975).
Spencer, H. W., Yarger, W. E. & Robinson, R. R. Alterations of renal function during dietary-induced hyperuricemia in the rat. Kidney Int. 9, 489–500. https://doi.org/10.1038/ki.1976.63 (1976).
Johnson, R. J. et al. Uric acid and chronic kidney disease: Which is chasing which?. Nephrol. Dial. Transpl. 28, 2221–2228. https://doi.org/10.1093/ndt/gft029 (2013).
De Cosmo, S. et al. Serum uric acid and risk of CKD in type 2 diabetes. Clin. J. Am. Soc. Nephrol. 10, 1921–1929. https://doi.org/10.2215/cjn.03140315 (2015).
Köttgen, A. et al. Genome-wide association analyses identify 18 new loci associated with serum urate concentrations. Nat. Genet. 45, 145–154. https://doi.org/10.1038/ng.2500 (2013).
Okada, Y. et al. Meta-analysis identifies multiple loci associated with kidney function-related traits in east Asian populations. Nat. Genet. 44, 904–909. https://doi.org/10.1038/ng.2352 (2012).
Pattaro, C. et al. Genetic associations at 53 loci highlight cell types and biological pathways relevant for kidney function. Nat. Commun. 7, 10023. https://doi.org/10.1038/ncomms10023 (2016).
Badve, S. V. et al. Effects of allopurinol on the progression of chronic kidney disease. N. Engl. J. Med. 382, 2504–2513. https://doi.org/10.1056/NEJMoa1915833 (2020).
Kimura, K. et al. Febuxostat therapy for patients with stage 3 CKD and asymptomatic hyperuricemia: A randomized trial. Am. J. Kidney Dis. 72, 798–810. https://doi.org/10.1053/j.ajkd.2018.06.028 (2018).
Acknowledgements
This work was supported by a multidisciplinary research grant-in-aid from the Seoul Metropolitan Government Seoul National University (SMG-SNU) Boramae Medical Center (04-2023-0040).
Author information
Authors and Affiliations
Contributions
The principal investigator was JL. The study proposal and design were conducted by YK and JL. Acquisition, analysis, or interpretation of data were performed by JJ, YJ, EB, KL, and SW. The study protocol was reviewed by JHP, KJ, SH, JPL, DKK, CHSL, SW and JL. Material and technical supports were provided by YJ, KL, and SW. Each author contributed important intellectual content during manuscript drafting or revision and accepts accountability for the overall work by ensuring that questions pertaining to the accuracy or integrity of any portion of the work are appropriately investigated and resolved.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kim, Y., Jo, J., Ji, Y. et al. Impact of hyperuricemia on CKD risk beyond genetic predisposition in a population-based cohort study. Sci Rep 14, 18466 (2024). https://doi.org/10.1038/s41598-024-69420-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-69420-5
- Springer Nature Limited