Abstract
Clear cell renal cell carcinoma (ccRCC) is characterized by a high incidence and mortality rate. Despite advancements in therapeutic interventions, the prognosis for renal cancer patients remains suboptimal. Of late, methylation modifications have emerged as promising molecular targets for tumor assessment and treatment, yet their potential has not been fully investigated in the context of ccRCC. Transcriptomic and clinical data were extracted from The Cancer Genome Atlas, Gene Expression Omnibus, and ArrayExpress databases, leading to the identification of 57 methylation-related genes (MRGs). Utilizing DESeq2 analysis, Cox regression analysis, and the LASSO regression algorithm, a Methylation-Related Risk Score (MARS) was constructed. Cluster analysis, Gene Ontology (GO) analysis, clinical feature analysis, immune infiltration analysis, and mutation analysis were further employed to evaluate the model. Our investigation identified six pivotal prognostic MRGs and established a risk score predicated on m6A/m5C/m1A/m7G regulatory factors. This score was validated across two external cohorts and can be utilized to assess individual immune infiltration statuses and predict responses to immunotherapy. Moreover, cluster analysis delineated two distinct m6A/m5C/m1A/m7G gene clusters. We have developed and validated a robust prognostic signature based on genes associated with m6A, m5C, m1A, and m7G modifications. This gene signature demonstrates significant prognostic value in assessing survival outcomes, clinical characteristics, immune infiltration, and responses to immunotherapy in ccRCC patients. This finding provides valuable insights for refining precision treatment strategies.
Similar content being viewed by others
Introduction
Annually, there are still in excess of 400,000 individuals diagnosed with kidney cancer on a global scale1. Patients with distant metastases have a 5-year relative survival rate of 15.3%. Clear cell renal cell carcinoma (ccRCC) is the predominant subtype of kidney cancer, making up approximately 70% of all instances2. The therapeutic landscape for metastatic ccRCC is continually evolving, yet it remains encumbered by heterogeneity in treatment responses, the emergence of resistance to therapeutic agents, and the attendant toxicities associated with treatment regimens3. In light of the significant incidence and mortality associated with ccRCC, the exploration of novel biomarkers is essential to enhance early detection, assess risk, and predict the effectiveness of therapeutic interventions.
Post-transcriptional modifications of RNA represent a pivotal component in the field of epigenetics, having garnered considerable attention as a focal point of research in recent years4. Mounting evidence suggests that the dynamic pathways of RNA modification constitute a sophisticated regulatory network implicated in a plethora of cellular processes, including stem cell differentiation, cellular motility, immune responses, and stress reactions5,6. These pathways are increasingly recognized as potential therapeutic targets in the battle against cancer. The cellular RNA landscape is adorned with over 170 distinct types of modifications, with methylation being the most prevalent7. This includes N6-methyladenosine (m6A), 5-methylcytosine (m5C), N1-methyladenosine (m1A), and 7-methylguanosine (m7G). These modifications are orchestrated by a suite of regulators comprising writers, readers, and erasers. Among these, N6-methyladenosine (m6A) is distinguished as one of the most ubiquitous, abundant, and evolutionarily conserved internal modifications within eukaryotic RNA, and it has been the subject of extensive research scrutiny8,9. The m6A modification is typically installed by a complex ensemble of 'writer' enzymes, such as the METTL3/METTL14 complex, recognized by 'reader' proteins like the YTHDF family, and can be removed by 'eraser' enzymes, including FTO and ALKBH55. The m5C modification, while prevalent in tRNA and rRNA, has also been identified in mRNA and long non-coding RNAs (lncRNAs)5. The m1A modification has been noted to influence the structure and function of tRNA. In contrast, the m7G modification forms part of the cap structure at the 5ʹ end of mRNA, playing a vital role in safeguarding RNA from exogenous nuclease degradation, facilitating the export of mature mRNA from the nucleus, enhancing translation efficiency, and regulating mRNA splicing10. An RNA molecule may harbor multiple methylation modifications simultaneously, which collectively constitute a complex regulatory network that impacts the biological function of RNA molecules and the progression of tumors11,12. The intricate interplay of these modifications and their specific roles in cancer remain a vibrant area of research. To date, comprehensive studies on the plethora of methylation modifications in ccRCC are lacking, with the specific mechanisms still necessitating more profound investigation to elucidate their intricacies.
We have engineered an innovative prognostic signature predicated upon methylation-related genes (MRGs) expression levels to ascertain the survival outcomes in patients afflicted with ccRCC. The robustness of this prognostic tool was corroborated through validation in independent patient cohorts. Furthermore, the signature demonstrates predictive prowess in assessing clinical and pathological variables, in addition to quantifying the degree of immune cell infiltration. Complementary functional analyses were executed to shed light on the underlying biological mechanisms that may dictate the prognostic landscape of ccRCC. These insights hold promise for enhancing prognostic accuracy concerning long-term survival in ccRCC, thereby propelling the field towards more tailored therapeutic strategies.
Materials and methods
Data acquisition
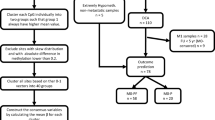
We conducted a meticulous interrogation of transcriptomic landscapes and correlative clinical metrics harvested from The Cancer Genome Atlas (TCGA; https://portal.gdc.cancer.gov/projects/TCGA-KIRC/) and ArrayExpress repositories (https://www.ebi.ac.uk/arrayexpress/) to substantiate our hypotheses. The Gene Expression Omnibus (GEO) dataset GSE40435 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=gse40435) encompassing 101 paired specimens of ccRCC and non-cancerous tissue counterparts, was employed to corroborate the differential expression of genes.To maintain uniformity in data interpretation, RNA-Seq TPM outputs were subjected to a binary logarithmic conversion. Post elimination of specimens devoid of expression profiles or survival indices, our study retained a cohort comprising 530 ccRCC specimens juxtaposed with 72 non-cancerous tissue counterparts from TCGA, in addition to 101 ccRCC specimens from the E-MTAB-1980 collective. TCGA and E-MATB-1980 dataset collects rapidly frozen untreated primary tumor samples, and all tumors must have matched normal samples from the same patient. For the TCGA-KIRC dataset, of the 530 patient samples included in the study, 175 experienced outcome events (patient death), and 355 were censored, with a median follow-up time of 39.85 months, a median follow-up time of 27.3 months for event occurrence, and a median follow-up time of 48.3 months for censoring. The E-MATB-1980 dataset collects untreated primary tumor samples, with 38 experiencing outcome events (patient death) and 202 censored, with a median follow-up time of 55.5 months, a median follow-up time of 26.5 months for event occurrence, and a median follow-up time of 60 months for censoring. Genes bereft of expression data or those registering TPM values below unity in excess of a quarter of the sample population were excluded to preclude the incidence of spurious positive detections.
Identification of prognostic MRGs
Through an exhaustive literature review, we identified genes involved in RNA methylation processes such as m6A, m5C, m1A, and m7G. Recent studies have revealed that certain DNA methylation genes (e.g., the DNMT and TET families) possess molecular functions that extend beyond gene silencing, also participating in RNA methylation-mediated post-transcriptional regulation. Ultimately, we identified 57 m6A/m5C/m1A/m7G regulators as MRGs for further analysis11,12,13,14,15. Information for each of these 57 MRGs is provided in the Supporting information (Supplementary Table 1). The "DESeq2" R package was employed to calculate the fold changes in mRNA expression levels between ccRCC tumor specimens and adjacent normal tissue samples16. We established a threshold for selecting differentially expressed genes (DEGs) as an absolute log2FoldChange greater than 1, coupled with an adjusted p-value of less than 0.05. Cox regression analyses were conducted using the "ezcox" package, selecting genes with an p-value of less than 0.05 as prognostically significant17. Volcano plots were crafted utilizing the "ggrepel" package18. Venn diagrams were generated with the "VennDiagram" package, while forest plots were created using the "forestplot" package19. The protein–protein interaction (PPI) network of selected MRGs was constructed using the STRING database (https://string-db.org/) version 11.0, selecting interacting proteins with a combined score of 0.4 or higher for graphical representation, and setting the k-means clustering value to three20. Visualization of the PPI network was achieved through Cytoscape software version 3.7.221. Bar and scatter plots featured in this article were also produced using the "ggplot2" package22.
Identification of functional pathways associated with MRGs
The "CancerSubtypes" package harnesses a multitude of integrative learning algorithms to consolidate heterogeneous biological markers, thereby identifying and validating molecular cancer clusters23. We utilized the "CancerSubtypes" package to cluster 530 samples from the TCGA-KIRC cohort based on the expression patterns of prognostic MRGs. The silhouette width serves as a metric to assess the consistency of clustering outcomes, with values ranging from − 1 to 1, indicating that a larger coefficient signifies less variation within clusters and greater dissimilarity between them. The silhouette width was employed to evaluate the consistency of samples within clusters24. Principal component analysis was conducted using the "FactoMineR" package, concurrently corroborating the analytical results of the "CancerSubtypes" package. GO analysis was performed with "clusterProfiler v4.4.4"25. Genes between two clusters with |log2FC|> 1 and p < 0.05 were selected for GO analysis, and the top three enriched pathways with the smallest adjusted p-values from the "BP," "CC," and "MF" categories were chosen for visualization26. The reference gene set was "c2.all.v2022.1.Hs.entrez.gmt".
Construction and validation of a methylation-associated risk scoring model
LASSO regression analysis is conducted via the "glmnet" package, which incorporates a penalty term by adding the sum of the absolute values of the regression coefficients to the loss function, thereby constraining or compressing the model to enhance its generalizability. The regularization parameter, λ (lambda), governs the intensity of the penalty and is determined through cross-validation, with the optimal λ employed to construct the scoring formula. The risk scoring formula is as follows:
Patients are assigned a Methylation-associated Risk Scoring (MARS) score based on the expression levels of prognostic MRGs and are stratified into high and low MARS groups according to the median MARS score.We utilize the "caret" package to randomly divide TCGA-KIRC into a training set and a validation set in a 7:3 ratio. The efficacy of the risk scoring model is assessed using Kaplan–Meier survival curves and ROC curves. KM survival analysis is performed using the "survival" and "survminer" packages, with survival curves plotted accordingly. The Receiver Operating Characteristic Curve is depicted utilizing the "pROC" package. Sankey diagrams are crafted with "ggplot2" to illustrate the distribution of clustering, risk grouping, and survival status.
Correlative analysis of clinical characteristics in MARS
Utilizing the "rms" package, a nomogram is constructed to intuitively align risk factors with their respective scores, subsequently transforming the aggregate total into a predictive probability. The calibration plot, a graphical method to evaluate the precision of predictive models, is rendered via the "rms" package to quantitatively assess the accuracy of the scoring model's predictions.
Analysis of immune infiltration in MARS
The single-sample Gene Set Enrichment Analysis (ssGSEA) is an analytical method employed to quantify gene set enrichment within an individual sample, assessing the infiltration levels of 23 immune cell types at the single-sample scale. The implementation of ssGSEA is adeptly facilitated by the Gene Set Variation Analysis (GSVA) package27. The ESTIMATE algorithm leverages specific gene expression signatures to quantify the non-cancerous cellular components within tumor tissues. The Immune Score reflects the level of immune cell infiltration, whereas the Stromal Score indicates the abundance of stromal cells in the tumor matrix, with the ESTIMATE Score integrating both to estimate tumor purity28. Tumor Immune Dysfunction and Exclusion (TIDE) analysis, a computational method, predicts a cancer patient's response to immune checkpoint blockade therapy. The TIDE analysis is conducted through the Response Prediction module accessible at http://tide.dfci.harvard.edu/. The Dysfunction value and IFNG value are computed by evaluating the expression levels of biomarkers within the tumor29.
Mutation and TMB analysis
The oncoplot delineates the genomic alteration frequency and mutation types within the TCGA-KIRC cohort. Tumor Mutational Burden (TMB) enumerates the quantum of mutations harbored within the genomic landscape of neoplastic tissues, typically encompassing non-synonymous alterations, insertions, or deletions that have the potential to alter the proteomic script. This metric is quantified as mutations per megabase (mut/Mb) of the genomic sequence30. The generation of oncoplots and the computation of TMB are facilitated by the R package "maftools"31.
Statistical methodology
We used R 4.2.1 software to perform statistical analysis32. The Pearson correlation algorithm was used to examine the relationship between numerical variables. To assess the difference between two independent samples, we employed the Wilcoxon rank-sum test.This study employs the Benjamini–Hochberg method for correction.Throughout the study, a default significance level of p-value less than 0.05 was considered as statistically significant, unless stated otherwise.
Results
Identification of prognostic MRGs in ccRCC patients
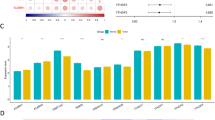
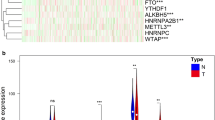
Transcriptional data of fifty-seven MRGs procured from TCGA repository were harnessed for differential expression analysis. MRGs exhibiting an absolute log2 fold change (log2FC) greater than 1 and a p-value of less than 0.05 were designated as DEGs, with the identification results depicted in Fig. 1A. Cox regression analysis was utilized to further appraise the prognostic significance of DEGs on survival outcomes, identifying nine MRGs as OS-related genes—comprising five risk factors and four protective factors—as illustrated in Fig. 1B. The expression variance of MRGs was validated using a cohort of 202 samples from the external dataset GSE40435 (Fig. 1C). Subsequently, eight MRGs were discerned as prognostic genes for ensuing analysis (Fig. 1D). The PPI networks and co-expression relationships of the eight prognostic genes were further elucidated (Fig. 1E,F).
(A) The DESeq2 analysis delineated nine upregulated MRGs alongside five downregulated MRGs. (B) Prognostic significance of DEGs on survival outcomes assessed by Cox regression analysis. (C) Validation of MRG expression variance using external dataset GSE40435. (D) Venn diagram illustrating the overlap in the identification of prognostic MRGs. (E) Protein–protein interaction networks of prognostic MRGs. (F) Co-expression analysis elucidates relationships among the eight prognostic MRGs. ns p > 0.05; *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
The clustering analysis based on MRGs
Building upon the previously identified prognosis-related MRGs, we conducted a Non-negative Matrix Factorization clustering analysis on 530 cases from the TCGA-KIRC dataset, as depicted in Fig. 2A. The silhouette width, an index assessing the consistency of clustering results with values ranging from -1 to 1, suggests that a greater coefficient corresponds to diminished intra-cluster variation and enhanced inter-cluster divergence. We compared the silhouette widths for different numbers of clusters (k = 2 to k = 9), finding that the silhouette width peaked at k = 2, indicating an optimal cluster number as shown in Fig. 2B. As depicted in Fig. 2C, a pronounced divergence in survival probabilities is evident between patient clusters 1 and 2, with the survival outcome of cluster 1 being inferior to that of cluster 2. Principal Component Analysis (PCA) corroborated the clustering consistency internally, as shown in Fig. 2D. Further investigation into the potential mechanisms of MRGs in ccRCC is warranted. The Gene Ontology (GO) analysis entries with the smallest adjusted p-value, primarily associated with complement activation signaling pathways and humoral immunity, are presented in Fig. 2E. This may offer a novel perspective for exploring the underlying mechanisms of methylation in the pathogenesis and progression of ccRCC.
(A) Non-negative Matrix Factorization clustering of prognosis-related MRGs in TCGA-KIRC dataset. (B) Silhouette width analysis for optimal cluster determination. (C) Kaplan–Meier survival curves demonstrating significant differences in survival outcomes between clusters C1 and C2. (D) Principal Component Analysis corroborats the internal consistency of the identified clusters. (E) Gene Ontology analysis revealing MRG-associated biological pathways.
Building and verifying prognostic risk scoring models in ccRCC patients
To facilitate a quantitative prognostic evaluation of ccRCC patients, a risk scoring model predicated on MRGs was constructed. The TCGA-KIRC dataset was partitioned into a training cohort and a validation cohort in a 7:3 ratio. To mitigate the potential for overfitting, LASSO regression was employed in the development of the risk scoring model (Fig. 3A,B). At a lambda value of six, the model's predictive inaccuracy is minimized. Subsequently, six MRGs were utilized to compute the Methylation-associated Risk Scoring (MARS), stratifying patients into high and low-risk categories in equal proportions (Fig. 3C). The prognostic value of MARS was examined, revealing significant survival discrepancies between different risk groups, with MARS demonstrating commendable predictive accuracy for patient survival rates at 1, 3, and 5-year intervals (Fig. 3D,E). The correlation between patient clustering, risk group allocation, and prognostic outcomes is depicted in Fig. 3F.
(A,B) Optimization of lambda in the LASSO regression. (C) Distribution of Methylation-associated Risk Scoring (MARS) in ccRCC patients. (D) Kaplan–Meier survival analysis for MARS stratified risk groups. (E) Predictive accuracy of MARS for survival at 1, 3, and 5-year intervals. (F) Correlation of patient clustering, MARS risk groups, and outcomes.
The external validation of the MARS
To further substantiate the prognostic value of MARS for patients with ccRCC, we incorporated the TCGA validation cohort and the E-MATE-1980 cohort. The MARS was calculated for each patient within the two cohorts based on the expression levels of MRGs, stratifying them into high and low-risk groups accordingly. MARS proficiently delineated patients with divergent survival outcomes (Fig. 4A,B). Concurrently, Kaplan–Meier survival curves demonstrated that patients in the high-risk category exhibited inferior long-term survival rates across both validation cohorts (Fig. 4C,D). The Receiver Operating Characteristic (ROC) curves underscored MARS's robust predictive capability for 1-year, 3-year, and 5-year survival rates within both cohorts (Fig. 4E,F). The aforementioned findings suggest that the prognostic predictive value of MARS may possess a degree of universality among patients with ccRCC.
(A,B) MARS stratification of the TCGA-test cohort and E-MATE-1980 cohort. (C,D) Kaplan–Meier survival analysis for TCGA-test cohort and E-MATE-1980 cohort. (E,F) ROC curves evaluating the predictive accuracy of MARS for 1-year, 3-year, and 5-year survival rates in the TCGA-test cohort and E-MATE-1980 cohort.
The correlation of MARS with clinical characteristics
A cohort of 247 clinical-feature-complete TCGA-KIRC cases was incorporated into the analysis of this section. Bar graphs indicate that patients with higher Tumor, Nude, Metastasis stages, more advanced stages, and poorer nuclear grading have higher MARS (Supplementary Fig. 1A–E). No significant difference in MARS was observed between genders (Supplementary Fig. 1F). To integrate clinical features into a comprehensive analysis for assessing patient prognosis, we conducted a Cox regression analysis, identifying Age, M, and MARS as independent risk factors impacting the survival outcome of ccRCC patients (Fig. 5A,B). Each independent risk factor's level corresponded to a point, with the nomogram intuitively displaying the long-term survival probability associated with the total points (Fig. 5C). The calibration plot is a graphical tool used to demonstrate the accuracy of model-predicted probabilities, with actual points adhering closer to the ideal line (45-degree line) indicating higher prediction reliability. The prognostic prediction model demonstrated good reliability in predicting 1-year, 3-year, and 5-year patient survival outcomes (Fig. 5D–F).
(A,B) Cox regression analysis plot identifying Age, Metastasis (M), and MARS as independent risk factors affecting the survival outcome in ccRCC patients. (C) Nomogram representing the correlation of each independent risk factor's level with points. (D–F) Calibration plot for the prognostic prediction model displaying the concordance between predicted probabilities and actual outcomes for 1-year, 3-year, and 5-year survival.
Correlative analysis of MARS with the tumor immune microenvironment
In the preceding discourse, GO analysis indicated that MRGs may influence patient prognosis in ccRCC by engaging in immune-related pathways (Fig. 2E). Consequently, we employed single-sample Gene Set Enrichment Analysis (ssGSEA) to elucidate the relationship between MARS and the tumor immune microenvironment (TIME). A significant elevation in the majority of immune cell types was observed in patients with high MARS, notably marked by CD8+ T cells, B cells, and various differentiation clusters of CD4+ T cells (Fig. 6A). Concurrently, ESTIMATE analysis revealed that patients with elevated MARS possessed higher estimate scores and immune scores, both demonstrating a significant positive correlation (Fig. 6B–E). TIDE analysis delineates a correlation wherein an escalation in MRGS is attendant with a decrement in interferon levels and an amplification in functional dysregulation scores (Fig. 6F,G). These findings suggest that patients with elevated MRGS exhibit higher levels of immune cell infiltration and diminished immune cell functional capacity, which may be linked to mechanisms in ccRCC that suppress T cell activation and promote tumor immune evasion33.
(A) Bar graph illustrating the significant increase in various immune cell types in patients with high MARS. (B,C) Scatter plot displaying the positive correlation between MARS levels and ESTIMATE scores and Immune scores. (D,E) Box plot representing higher ESTIMATE scores and Immune scores in patients with increased MARS. (F,G) Bar graph demonstrating the association between higher MRGS and lower interferon levels and higher immune cell dysfunction scores. ns p > 0.05; *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
The assessment of the correlation between MARS and tumor mutational burden
Tumor Mutational Burden (TMB) is extensively characterized as the density of somatic mutations within each megabase of the genome examined, and its quantification serves as a prognostic instrument for anticipating the efficacy of immunotherapeutic regimens in a multitude of oncological contexts34. Given the significance of immunotherapy in ccRCC and the correlation between MARS and TIME, it is imperative to discuss the relationship between mutational burden and TMB. Oncoplots reveal that patients within the high mutational burden cohort exhibit an increased probability of VHL mutations (Fig. 7A,B). Concurrently, these patients demonstrate elevated levels of TMB, with a significant disparity between the two cohorts (Fig. 7C). Exploration of the differential mutagenic genes between these groups identified six genes—VHL, SETD2, BAP1, MTOR, SYNE1, and TP53—as having a significantly augmented mutation rate in the high-risk group, as determined by Fisher's exact test (Fig. 7D). Stratifying patients into four categories based on TMB and MARS levels, the cohort with both high TMB and high MARS exhibited the poorest long-term survival rates. Conversely, the group with low TMB and MARS demonstrated the most favorable long-term survival outcomes (Fig. 7E). This suggests that patients within the high MARS category tend to present with higher levels of TMB, correlating with lower long-term survival rates but potentially more likely to derive clinical benefit from immunotherapy35.
(A,B) The oncoplot delineates the prevalent mutational gene frequencies and modes of mutation between two MARS groups. (C) Comparison of TMB levels between the two MARS groups, showing a significant disparity. (D) Identification of six genes with significantly augmented mutation rates between two MARS groups. (E) Stratification of patients based on TMB and MARS levels, revealing the association with long-term survival rates.
Discussion
The therapeutic conquest of distant metastatic renal carcinoma remains an arduous quest. The introduction of targeted and immune therapies has markedly enhanced survival outcomes for patients with advanced RCC and those with a heightened risk of relapse2. Nonetheless, the individual variability in therapeutic efficacy, the side effects of treatments, and the emergence of drug resistance persist as substantial obstacles for patients36,37. The imperative development of innovative biomarkers and novel pharmacological entities is essential to refine the paradigms of clinical decision-making.
Owing to the reversibility of epigenetic modifications, a new strategic avenue has been unveiled for cancer therapy5. Recently, methylation modifications, including m6A, m5C, m1A, and m7G, as integral components of the epigenetic landscape, have been extensively and profoundly investigated for their roles in cancer diagnosis and targeted therapy. A plethora of studies have corroborated that dysregulation of methylation modifications or their associated enzymes precipitates the onset and progression of human cancers38,39,40,41,42,43,44,45. Currently, an array of pharmaceuticals targeting specific epigenetic alterations has been developed37,38. METTL3 (methyltransferase-like 3), a member of the METTL family, is responsible for catalyzing the transfer of methyl groups from donor molecules to specific target sites on RNA molecules46. Accumulating evidence suggests that METTL3 could serve as a potential therapeutic target. The development of METTL3 inhibitors such as adenosine analogs, UZH1a, and CDIBA is burgeoning46,47.
In previous studies, scholars have predominantly focused on individual RNA modifications, neglecting the synergistic effects of various RNA modifications within the tumor microenvironment and on immune infiltration. There is a dearth of research that considers m6A, m5C, m1A, and m7G modifications collectively in the context of ccRCC. Recent evidence has revealed the crosstalk between RNA m6A modifications mediated by m6A reader proteins FXR1 and TET1 and DNA cytosine 5mC modifications, which can regulate chromatin activation of transposable elements and cell fate determination in human pluripotent stem cells, demonstrating a significant regulatory potential in development, immune response, aging, and cancer progression48. Therefore, exploring these overlapping methylation modifications, including m6A, m5C, m1A, and m7G, may deepen our understanding of the TIME characteristics and promote the development of precise and convenient clinical scoring systems to aid in the diagnosis and personalized treatment of cancer patients.
In the realm of this investigation, we have adeptly delineated m6A/m5C/m1A/m7G-associated factors that manifest differential expression and prognostic significance within the cohort of ccRCC patients. Subsequently, we have formulated a prognostic risk assessment score for individuals afflicted with ccRCC, predicated upon these biomarkers. The prognostic validity of our scoring system was substantiated through Kaplan–Meier and Receiver Operating Characteristic analyses. The E-MTAB-1980 Cohort served as an external validation set, validating the robustness of our risk scoring model. In this study, we conducted an in-depth exploration into the correlation between MARS and clinical characteristics, the tumor immune microenvironment, and tumor mutational burden. Furthermore, we elucidated the utility of MARS as a prognostic indicator for patient outcomes and as a referential tool in the selection of therapeutic strategies.
Gene Ontology (GO) analysis indicates a predominance of immune-related pathways in enriched pathways. In immune analyses, ssGSEA analysis suggests that patients with higher MARS scores exhibit a more significant degree of immune infiltration. Concurrently, TIDE analysis reveals that patients with higher MARS scores have elevated immune cell dysfunction scores. Recent studies have elucidated that epigenetic modifications are instrumental in the differentiation, maturation, and pathogen response of immune cells. This may correlate with the tumor cells' evasion of immune surveillance, achieved by methylation-induced silencing of antigen-presenting molecules or immune checkpoint molecules, thereby inhibiting T cell activation and proliferation. The regulatory role of DNA methylation in tumor immunity has been thoroughly investigated in recent years. Cutting-edge research has also substantiated the role of various RNA methylation mechanisms in the regulation of the TIME. Targeting methylation presents a promising strategy to overcome immune evasion in renal cancer. A small fraction of tumor mutations are translated, processed, and presented on MHC molecules, recognized by T cells as neoantigens; hence, an increase in TMB augments the likelihood of tumors presenting more neoantigens. TMB has been validated as an essential predictor of patient response to immunotherapy across multiple cancers. In our study, patients with higher MARS scores tend to have elevated TMB levels, suggesting that MARS could potentially serve as an indicator for predicting the efficacy of immunotherapy.
This investigation has identified DNMT3B, FTO, IGF2BP2, IGF2BP3, LRPPRC, and NSUN7 as pivotal prognostic MRGs and has incorporated them into a risk scoring model.
Functioning as a DNA (cytosine-5)-methyltransferase, DNA methyltranferase 3B (DNMT3B) plays a role by engaging in the epigenetic modification process via the de novo methylation pathway49. An increasing body of evidence substantiates the DNMT3B dysregulation in the oncogenesis and progression of cancers. The significance of DNMT3B in the progression and prognosis of hepatocellular carcinoma has been characterized50. Fat mass and obesity-associated protein (FTO), an m6A demethylase, modulates the progression of leukemia by regulating the m6A modification of downstream genes such as CEBPA51. The FTO inhibitor R-2HG, which impedes the FTO/MYC/CEBPA axis, has been sanctioned by the FDA for the treatment of refractory/relapsed acute myeloid leukemia51. All members of the insulin-like growth factor 2 mRNA-binding proteins (IGF2BP) family are overexpressed during early development and the oncogenic process, with their upregulation correlating with adverse outcomes in a spectrum of malignancies, including breast cancer, colorectal cancer, esophageal adenocarcinoma, glioma, hepatocellular carcinoma, pancreatic cancer, acute myeloid leukemia, Ewing's sarcoma, and head and neck squamous cell carcinoma52. This may be attributed to the critical roles that IGF2BP plays in cellular growth, the maintenance of stem cells, and differentiation during developmental processes53. Leucine-rich pentatricopeptide repeat containing (LRPPRC) exerts its oncogenic influence primarily by modulating mitochondrial homeostasis and eliciting oxidative stress54. The attenuation of LRPPRC expression can suppress the proliferation and invasion of tumor cells, induce apoptosis, and surmount drug resistance55. NOP2/Sun RNA methyltransferase family member 7 (NSUN7) is responsible for the m5C modification of RNA56. In hepatocarcinogenesis, the suppression of NSUN7 expression impedes proper mRNA methylation, playing a pivotal role in the process of cancer metastasis57.
Currently, clinicians may predict the outcomes for patients with ccRCC based on established prognostic assessment methodologies, including the Eastern Cooperative Oncology Group (ECOG) performance status scoring system, Fuhrman's histological grading, and the TNM staging system. However, these traditional approaches are challenged by the issue of interindividual heterogeneity. Our research has developed a methylation-based risk scoring model, MARS, which serves as a valuable augmentation in predicting outcomes for patients with ccRCC. Nevertheless, our investigation still encounters substantial limitations. On one hand, we have only preliminarily investigated the correlation between the expression levels of MRGs and clinical characteristics of patients, necessitating further in vivo and in vitro experiments to fully comprehend their biological mechanisms. On the other hand, our research is retrospective, based on databases such as TCGA, GEO, and ArrayExpress, and thus prospective studies across multiple centers are imperative to corroborate the efficacy of our scoring model.
Conclusions
We have developed and validated a robust prognostic signature based on genes associated with m6A, m5C, m1A, and m7G modifications. This gene signature consistently demonstrates significant prognostic utility for appraising survival outcomes, clinical attributes, immune infiltration, and the efficacy of immunotherapy in patients with ccRCC. These results highlight the potential of our gene signature to inform the refinement of precision treatment strategies for ccRCC patients.
Data availability
The transcriptomic data, clinical information, and somatic mutation data utilized in this study are freely available from the TCGA database (https://portal.gdc.cancer.gov/projects/TCGA-KIRC/). External validation data, E-MTAB-1980, is accessible from the ArrayExpress database (https://www.ebi.ac.uk/arrayexpress/). Access to the GSE40435 dataset is available via the following link: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE40435. The TIDE analysis was performed using publicly accessible data, which can be obtained from the TIDE website (http://tide.dfci.harvard.edu).
References
Bukavina, L. et al. Epidemiology of renal cell carcinoma: 2022 update. Eur. Urol. 82, 529–542 (2022).
Motzer, R. J. et al. Patient-reported outcome measurement and reporting for patients with advanced renal cell carcinoma: A systematic literature review. Eur. Urol. 84, 406–417 (2023).
Semenescu, L. E. et al. An overview of systemic targeted therapy in renal cell carcinoma, with a focus on metastatic renal cell carcinoma and brain metastases. CIMB 45, 7680–7704 (2023).
Song, P., Tayier, S., Cai, Z. & Jia, G. RNA methylation in mammalian development and cancer. Cell Biol. Toxicol. https://doi.org/10.1007/s10565-021-09627-8 (2021).
Nombela, P., Miguel-López, B. & Blanco, S. The role of m6A, m5C and Ψ RNA modifications in cancer: Novel therapeutic opportunities. Mol. Cancer https://doi.org/10.1186/s12943-020-01263-w (2021).
Shima, H. & Igarashi, K. N 1-methyladenosine (m1A) RNA modification: The key to ribosome control. J. Biochem. 167, 535–539 (2020).
Ma, S. et al. The interplay between m6A RNA methylation and noncoding RNA in cancer. J. Hematol. Oncol. https://doi.org/10.1186/s13045-019-0805-7 (2019).
Patrasso, E. A., Raikundalia, S. & Arango, D. Regulation of the epigenome through RNA modifications. Chromosoma 132, 231–246 (2023).
Shen, L. et al. N 6-Methyladenosine RNA modification regulates shoot stem cell fate in arabidopsis. Dev. Cell 38, 186–200 (2016).
Zhang, X., Zhu, W.-Y., Shen, S.-Y., Shen, J.-H. & Chen, X.-D. Biological roles of RNA m7G modification and its implications in cancer. Biol. Direct. https://doi.org/10.1186/s13062-023-00414-5 (2023).
Huang, F. et al. Using integrated analysis from multicentre studies to identify RNA methylation-related lncRNA risk stratification systems for glioma. Cancer Cell Int. https://doi.org/10.1186/s12935-023-03001-w (2023).
Pan, C. et al. An m1A/m6A/m5C-associated long non-coding RNA signature: Prognostic and immunotherapeutic insights into cervical cancer. J. Gene Med. https://doi.org/10.1002/jgm.3618 (2023).
Zhang, X., Zhang, Y., Wang, C. & Wang, X. TET (Ten-eleven translocation) family proteins: Structure, biological functions and applications. Sig. Transduct. Target Ther. 8, 297 (2023).
Lyko, F. The DNA methyltransferase family: A versatile toolkit for epigenetic regulation. Nat. Rev. Genet. 19, 81–92 (2017).
Liu, J., Dou, M., Liu, X., Lu, Y. & Lu, W. A novel m6A/m5C/m1A score signature to evaluate prognosis and its immunotherapy value in colon cancer patients. J. Cancer Res. Clin. Oncol. 149, 11995–12012 (2023).
Love, M., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol. 15, 550 (2014).
Wang, S. ezcox: Easily Process a Batch of Cox Models. R package version 1.0.0. https://github.com/ShixiangWang/ezcox. (2021).
K, S. ggrepel: Automatically Position Non-Overlapping Text Labels with ‘ggplot2’. https://ggrepel.slowkow.com/, https://github.com/slowkow/ggrepel. @Manual. (2024).
Chen, H. & Boutros, P. C. VennDiagram: A package for the generation of highly-customizable Venn and Euler diagrams in R. BMC Bioinform. https://doi.org/10.1186/1471-2105-12-35 (2011).
Szklarczyk, D. et al. The STRING database in 2021: Customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucl. Acids Res. 49, D605–D612 (2020).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
H, W. Ggplot2: Elegant Graphics for Data Analysis. (Springer, 2016).
Xu, T. et al. CancerSubtypes: an R/Bioconductor package for molecular cancer subtype identification, validation and visualization. Bioinformatics 33, 3131–3133 (2017).
Rousseeuw, P. J. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 20, 53–65 (1987).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation. 2, 100141 (2021).
Ashburner, M. et al. Gene Ontology: Tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-Seq data. BMC Bioinform. 14, 7 (2013).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. https://doi.org/10.1038/ncomms3612 (2013).
Jiang, P. et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 24, 1550–1558 (2018).
Goodman, A. M. et al. Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers. Mol. Cancer Ther. 16, 2598–2608 (2017).
Mayakonda, A., Lin, D.-C., Assenov, Y., Plass, C. & Koeffler, H. P. Maftools: Efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 28, 1747–1756 (2018).
Team, R. R: A Language and Environment for Statistical Computing. (2014).
Monjaras-Avila, C. U. et al. The tumor immune microenvironment in clear cell renal cell carcinoma. IJMS 24, 7946 (2023).
Strickler, J. H., Hanks, B. A. & Khasraw, M. Tumor mutational burden as a predictor of immunotherapy response: Is more always better?. Clin. Cancer Res. 27, 1236–1241 (2021).
Principe, D. R. Patients deriving long-term benefit from immune checkpoint inhibitors demonstrate conserved patterns of site-specific mutations. Sci. Rep. https://doi.org/10.1038/s41598-022-15714-5 (2022).
Blas, L. et al. Current status and future perspective of immunotherapy for renal cell carcinoma. Int. J. Clin. Oncol. https://doi.org/10.1007/s10147-023-02446-3 (2023).
Zhou, X. et al. The role of RNA modification in urological cancers: Mechanisms and clinical potential. Discov. Onc https://doi.org/10.1007/s12672-023-00843-8 (2023).
Qiu, L., Jing, Q., Li, Y. & Han, J. RNA modification: Mechanisms and therapeutic targets. Mol. Biomed. https://doi.org/10.1186/s43556-023-00139-x (2023).
Chen, B. et al. N7-methylguanosine tRNA modification promotes tumorigenesis and chemoresistance through WNT/β-catenin pathway in nasopharyngeal carcinoma. Oncogene 41, 2239–2253 (2022).
Wu, Y. et al. N1-Methyladenosine (m1A) regulation associated with the pathogenesis of abdominal aortic aneurysm through YTHDF3 modulating macrophage polarization. Front. Cardiovasc. Med. https://doi.org/10.3389/fcvm.2022.883155 (2022).
Cheng, J. X. et al. RNA cytosine methylation and methyltransferases mediate chromatin organization and 5-azacytidine response and resistance in leukaemia. Nat. Commun. https://doi.org/10.1038/s41467-018-04518-9 (2018).
Li, Y. et al. Novel long noncoding RNA NMR promotes tumor progression via NSUN2 and BPTF in esophageal squamous cell carcinoma. Cancer Lett. 430, 57–66 (2018).
Chen, L. et al. METTL3-mediated m6A modification stabilizes TERRA and maintains telomere stability. Nucl. Acids Res. https://doi.org/10.1093/nar/gkac1027 (2022).
Cui, Q. et al. m 6 A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep. 18, 2622–2634 (2017).
Liu, H.-T. et al. lncRNA THAP7-AS1, transcriptionally activated by SP1 and post-transcriptionally stabilized by METTL3-mediated m6A modification, exerts oncogenic properties by improving CUL4B entry into the nucleus. Cell Death Differ. https://doi.org/10.1038/s41418-021-00879-9 (2021).
Ma, C. et al. The molecular mechanism of METTL3 promoting the malignant progression of lung cancer. Cancer Cell Int. https://doi.org/10.1186/s12935-022-02539-5 (2022).
Fiorentino, F., Menna, M., Rotili, D., Valente, S. & Mai, A. METTL3 from target validation to the first small-molecule inhibitors: A medicinal chemistry journey. J. Med. Chem. 66, 1654–1677 (2023).
Sun, T., Xu, Y., Xiang, Y., Soderblom, E. J. & Diao, Y. Crosstalk between RNA m6A and DNA methylation regulates transposable element chromatin activation and cell fate in human pluripotent stem cells. bioRxiv (2022) https://doi.org/10.1101/2022.09.08.507172.
So, J. Y. et al. DNA methyltransferase 3B–mediated intratumoral heterogeneity and therapeutic targeting in breast cancer recurrence and metastasis. Mol. Cancer. Res. 20, 1674–1685 (2022).
Kim, J., Park, J.-H. & Lee, J.-H. Clinical and prognostic values of DNMT3B expression in hepatocellular carcinoma. Keimyung Med. J. 41, 13–16 (2022).
Su, R. et al. R-2HG exhibits anti-tumor activity by targeting FTO/m6A/MYC/CEBPA signaling. Cell 172, 90-105.e23 (2018).
Ramesh-Kumar, D. & Guil, S. The IGF2BP family of RNA binding proteins links epitranscriptomics to cancer. Semin. Cancer Biol. 86, 18–31 (2022).
Christiansen, J., Kolte, A. M., Hansen, T. V. O. & Nielsen, F. C. IGF2 mRNA-binding protein 2: Biological function and putative role in type 2 diabetes. J. Mol. Endocrinol. 43, 187–195 (2009).
Cui, J., Wang, L., Ren, X., Zhang, Y. & Zhang, H. LRPPRC: A multifunctional protein involved in energy metabolism and human disease. Front. Physiol. https://doi.org/10.3389/fphys.2019.00595 (2019).
Zhao, Z., Sun, Y., Tang, J., Yang, Y. & Xu, X. LRPPRC regulates malignant behaviors, protects mitochondrial homeostasis, mitochondrial function in osteosarcoma and derived cancer stem-like cells. BMC Cancer https://doi.org/10.1186/s12885-023-11443-8 (2023).
Zhang, Q. et al. The role of RNA m5C modification in cancer metastasis. Int. J. Biol. Sci. 17, 3369–3380 (2021).
Ortiz-Barahona, V. et al. Epigenetic inactivation of the 5-methylcytosine RNA methyltransferase NSUN7 is associated with clinical outcome and therapeutic vulnerability in liver cancer. Mol. Cancer https://doi.org/10.1186/s12943-023-01785-z (2023).
Acknowledgements
We gratefully acknowledge Horizontal Project of Provincial Hospital Affiliated to Shandong First Medical University. Moreover, our heartfelt appreciation goes out to the TCGA, GEO, and ArrayExpress databases for their indispensable resources. We also express our thanks to the developers of the R language packages.
Funding
Funding was provided by Shandong First Medical University (Shandong Academy of Medical Sciences) Youth Science Cultivation Funding Program (Grant number: 202201-044).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Conceptualization: Z.X.C, W.H, T.H.Y; Methodology: Z.X.C, Z.H.N, Z.S.Y, Y.M.Z; Writing—original draft preparation: Z.X.C, X.J, C.T.N, X.T.S; Writing—review and editing: Z.H.N, Q.F, W.H, F.J.C; Supervision: Z.H.N, Q.F. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
He, W., Cong, Z., Niu, C. et al. A prognostic signature based on genes associated with m6A/m5C/m1A/m7G modifications and its immunological characteristics in clear cell renal cell carcinoma. Sci Rep 14, 18708 (2024). https://doi.org/10.1038/s41598-024-69639-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-69639-2
- Springer Nature Limited