Abstract
Biofilm formation and toxin production are some of the virulence factors of Clostridioides difficile (C. difficile), which causes hospital-acquired C. difficile infection (HA-CDI). This work investigated the prevalence and distribution of different strains recovered from HA-CDI patients hospitalized in 4 medical centres across Israel, and characterized strains' virulence factors and antibiotic susceptibility. One-hundred and eighty-eight faecal samples were collected. C. difficile 's toxins were detected by the CerTest Clostridium difficile GDH + Toxin A + B combo card test kit. Toxin loci PaLoc and PaCdt were detected by whole-genome sequencing (WGS). Multi-locus sequence typing (MLST) was performed to classify strains. Biofilm production was assessed by crystal violet. Antibiotic susceptibility was determined using Etest. Fidaxomicin susceptibility was tested via agar dilution. Sequence type (ST) 42 was the most (13.8%) common strain. All strains harboured the 2 toxins genes; 6.9% had the binary toxin. Most isolates were susceptible to metronidazole (98.9%) and vancomycin (99.5%). Eleven (5.85%) isolates were fidaxomicin-resistant. Biofilm production capacity was associated with ST (p < 0.001). In conclusion, a broad variety of C. difficile strains circulate in Israel's medical centres. Further studies are needed to explore the differences and their contribution to HA-CDI epidemiology.
Similar content being viewed by others
Introduction
Clostridioides difficile (C. difficile) causes gastrointestinal infections (CDI), which can manifest by various symptoms, ranging from diarrhoea and fever to life-threatening colitis. Most cases are acquired within hospitals (HA-CDIs) and are associated with substantial mortality and contribute to rising healthcare expenditures1.
Elderly subjects are notably at risk, particularly those in healthcare institutions2. Antibiotics administration prior to infection is another major risk factor for CDI3.
The mechanism behind antibiotic-induced HA-CDI has been ascribed to disruption of the balanced gut microbiota, resulting in an environment conducive to C. difficile survival and proliferation4. Certain antibiotics, such as cephalosporins, have been identified as major CDI risk factors5. Nevertheless, the recommended CDI management continues to be antimicrobial therapy, primarily fidaxomicin and vancomycin; metronidazole may be given when the previous two antibiotics are not available6.
The pathogenicity of C. difficile is mediated by several virulence factors, including toxin A (TcdA) and toxin B (TcdB)7. Both toxins inactivate Rho GTPases, which trigger a series of events including inflammation marked by neutrophil presence in the colon, deterioration of the protective barrier of the epithelial gut and the eventual apoptosis of colon cells, resulting in diarrhoea8. Cytolethal distending toxin (CDT)/binary toxin, enhances CDI severity, and is found in some epidemic strains such as ribotype 027 and 0789.
Another key feature of C. difficile virulence is its ability to form biofilms, which are organized bacterial communities shielded by a self-generated matrix consisting of extracellular polymeric compounds. Inside these protective structures, the bacteria are safeguarded from outside stresses, such as the host immune response and antibiotics, and resists extreme conditions, which may contribute to infection duration and recurrence10.
While identification of the specific strain responsible for the infection is valuable for epidemiological surveillance, it is not a routine procedure. Generally, CDI diagnosis is based on algorithms that constitute molecular biology methods to identify toxin genes11, as this method delivers results quickly, whereas culturing C. difficile can take up to two days. PCR ribotyping is one of the commonly used methods for C. difficile molecular typing. This technique includes amplification of the 16-23S rDNA intergenic spacer region. Each ribotype (RT) has different size and pattern of amplified DNA fragments; thus, classification of strains is performed based on these differences12.
Multi-locus sequence typing (MLST) is another genotyping technique designed to distinguish between bacterial species by examining the nucleotide sequences from fragments of at least seven housekeeping genes. Each unique combination of alleles is given a sequence type (ST) number13. The technique highlights epidemiological associations between C. difficile isolates and addresses challenges in comparing typing results across laboratories14.
Data on the common strains and their characteristics in Israel is limited. Only 2 previous studies were reported, with one published in 2015, during an outbreak of the Nap1 strain15, and the second study, performed by our group in 2017, which focused on strains in one medical center in north Israel16. This encouraged us to perform the current study, aiming to explore the prevalence and distribution of strain types collected from HA-CDI patients across different regions in Israel during the years 2020–2022. It also endeavoured to profile various virulence attributes of C. difficile isolates, including toxins, biofilm-production capacity and antibiotics resistance. The findings offer an in-depth view of the HA-CDI epidemiological landscape in Israel.
Methods
Sample collection
Stool samples of hospitalized patients who were diagnosed with HA-CDI between 2020 and 2022 at 4 medical centres in Israel, were included in this analysis. HA-CDI was defined as CDI that was developed > 48 h after hospitalization. All cases were confirmed at the hospital primary laboratories using the GeneXpert C. difficile PCR assay (Cepheid, Sunnyvale, CA, USA) which identifies presence of the genes encoding toxin B and the binary toxin, and identifies tcdC deletion. The following clinical and demographic data were collected from medical records: age, gender and mortality during the 30 days following hospitalization.
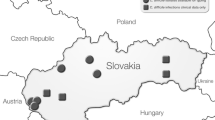
The participating medical centres are located in different geographic regions of Israel: Tzafon Medical Center, Poriya and W. Hirsch Regional Microbiology Laboratory Clalit Health Services, Haifa (north Israel), Edith Wolfson Medical Center, Holon (Israel's centre) and Soroka University Medical Center, Be'er Sheva (southern Israel).
All methods were carried out in accordance with relevant guidelines and regulations. All experimental protocols were approved by the Soroka medical center's Ethics (Helsinki) committee (approval no. SOR-0307-20), by the Tzafon medical center's Ethics (Helsinki) committee (Approval no. POR-0085-15) and by the Wolfson medical center's Ethics (Helsinki) committee (Approval no. WOMC-0115-20).
The need to obtain informed consent was waived by the Soroka medical center's Ethics (Helsinki), by the Tzafon medical center's Ethics (Helsinki) committee and by the Wolfson medical center's Ethics (Helsinki) committee.
Bacterial isolation and identification
Each stool sample was inoculated on a selective growth agar medium, chromID™ C. difficile (CDIF) (BioMérieux, Durham, NC), for 48 h, at 37 °C, under anaerobic conditions (10% CO2, 10% H2; and 80% N2), in a Bactron EZ 300 anaerobic chamber (Sheldon manufacturing, Cornelius, USA). C. difficile colonies were identified by their asymmetrical shape and black colour. Definitive identification was performed by matrix-assisted laser desorption ionization-time of flight (MALDI-TOF) mass spectrometry using the Bruker Biotyper system (Bruker Daltonics, Bremen, Germany)17. All isolates were stored as beads at −80 °C until further analysis.
Toxin detection
The CerTest Clostridium difficile GDH + Toxin A + B combo card test kit (Certest Biotec, S.L, Zaragoza, Spain) was used to detect C. difficile toxins in faecal samples, in accordance with the manufacturer's protocol. This assay simultaneously detects C. difficile antigens toxin A, toxin B, and glutamate dehydrogenase (GDH), with over 99% sensitivity and specificity18,19.
WGS and MLST of bacterial isolates
DNA extraction
The MagCore® Genomic DNA Bacterial Kit (ATRIDAB.V, Amersfoort, Nederland) was used with the MagCore® automated extraction instrument (RBCBioscience, New Taipei, Taiwan), to extract total genomic DNA from bacterial isolates according to the manufacturer’s instructions.
Library preparation and whole-genome sequencing
Following DNA clean-up, whole-genome sequencing was carried out using the Illumina DNA Prep kit (Illumina, Inc., San Diego, CA, USA), according to the manufacturer’s protocol. Samples were processed for tagmentation and adapter ligation using IDT for Illumina Nextera UD Indexes Set A, B, C, D (384 indexes, 384 samples). Further, enrichment and clean-up were performed according to the manufacturer’s instructions.
Libraries were pooled into 8 pools of 40 samples each. Pooled samples were quantified using the Qubit 4.0 fluorimeter using HS DS DNA kit (Thermo Fisher Scientific, Waltham, USA) and fragment sizes were determined using the TapeStation 4150 via DNA HS D1000 kit (Agilent Technologies, Santa Clara, CA, USA). The pooled libraries were normalized to 4 nM and 5μl of each normalized pool were combined in a fresh microcentrifuge tube. For sequencing, pooled libraries were denatured and neutralized with 0.2N NaOH and 400mM Tris–HCL (pH-8). Dual indexed paired-end sequencing with a 149-bp read length was performed with the NovaSeq6000 platform (Illumina, Inc.). Resulting FastQ files were analysed (alignment and QA/QC) by Partek flow, using the reference sequence of C. difficile strain 630 (NC_009089). To visually observe the depth and quality of the missing genes, the BAM files (created in PartekFlow) of the failed samples were analysed using the Genius program.
MLST analysis
The PubMLST C. difficile database (http://pubmlst.org/cdifficile/) was used to determine the strain type and clade of each isolate, based on the sequences of 7 housekeeping genes (adk, atpA, dxr, glyA, recA, sodA, tpi), as previously described13. A neighbour-joining tree was generated with MUSCLE-aligned concatenated allele sequences (created in Geneious Prime), to display the genetic diversity of the different strains. MUSCLE-aligned concatenated allele sequences were used as input for IQ-TREE v2.2.2.6 [https://doi.org/10.1093/molbev/msaa015], along with ModelFinder [https://doi.org/10.1038/nmeth.4285] to generate the maximum-likelihood phylogenetic tree. The tree was visualized with an online tool for phylogenetic tree display and annotation iTOL [https://doi.org/10.1093/nar/gkab301].
Toxin gene analysis
The toxin loci PaLoc and PaCdt were detected via the whole-genome sequencing data. A sequencing depth cutoff of 25 or higher was used to ensure the quality of each sample and to verify results' reliability. Additionally, this cutoff was employed to determine the presence of the tcdA, tcdB, cdtA, and cdtB genes. The presence of the full PaCDT locus, which results in the binary toxin formation, was determined by comparing the sequencing depth of the cdtA and cdtB genes by aligning each sample with C. difficile strain 630 (A + B + CDT−) and C. difficile strain 196 (A + B + CDT +). A CDT + isolate was expected to have the same average coverage when aligned to both reference strains. Conversely, in CDT- isolates, we anticipated seeing partial average coverage when aligned to C. difficile strain 196 compared to C. difficile strain 630.
Microtitre plate assay for the assessment of biofilm production
Biofilm production was assessed using a microtitre plate, as previously described20. Following incubation of C. difficile isolates in brain heart infusion broth supplemented with yeast extract + 0.1% L-cysteine (BHIS), at 37 °C for 24 h under anaerobic conditions, the suspension was diluted to an optical density (OD600nm) of 0.8 and then diluted 1:100 in BHIS. The diluted inoculum (200 μl) was transferred to a 96-well plate. The biofilm-forming ATCC® BAA-1382 C. difficile strain (strain 630) and BHIS served as positive and negative controls, respectively. Plates were incubated under anaerobic conditions for 24 h incubation, after which, spent medium was aspirated, and plates were washed twice with phosphate-buffered saline (PBS) (Oxoid, Cambridge, UK). An aqueous crystal violet solution (0.25% (w/v)) was then added to each well (200 μl) for 5 min. Then, the plates were rinsed eight times with PBS and left to air-dry. To dissolve the dye, 200 μl of an ethanol:acetone (1:1) solution was added to each well. Absorbance was measured within 5 min at a wavelength of 570 nm using an ELISA reader (Multiskan Go, Fisher-Scientific Ltd., Vantaa, Finland). The cut-off OD570nm (ODc) was defined as three standard deviations above the mean OD of the negative control. Biofilm formation capacity was classified as previously described21: non-producers—isolates with OD < ODC, weak producers—isolates with ODc < OD ≤ 2 × ODc, moderate producers—2 × ODc < OD ≤ 4 × ODc and strong producers—OD > 4 × ODc. The experiment was repeated three times, with triplicates of each strain tested in each experiment.
Antimicrobial susceptibility testing
The Etest technique – a gradient-diffusion method used to determine the lowest concentration of an antibiotic that inhibits the growth of a microorganism, was used to determine the minimum inhibitory concentration (MIC) of ampicillin, amoxicillin/clavulanate, ceftriaxone, cefuroxime, ceftazidime, chloramphenicol, ciprofloxacin, clindamycin, doxycycline, erythromycin, gentamycin, moxifloxacin, metronidazole, tigecycline, and vancomycin. C. difficile colonies were inoculated in thioglycollate broth medium (Becton Dickinson, Heidelberg, Germany) until 1.0 McFarland standard's turbidity was achieved. The bacterial suspension was inoculated on Brucella blood agar supplemeted with hemin and vitamin K1 (Hy Laboratories, Rehovot, Israel) and a gradient Etest strip (bioMérieux, Durham, NC) of each antibiotic was added. Plates were incubated under anaerobic conditions at 37 °C for 48 h, after which, MIC of each antibiotic was visually determined. Strains were classified as susceptible or resistant in accordance with the European Committee on Antimicrobial Susceptibility Testing (EUCAST) guidelines, versions 10.0–13.022.
Susceptibility to fidaxomicin was determined by agar dilution in accordance with the procedures of the Clinical and Laboratory Standards Institute (CLSI-M11-9th)23. Fidaxomicin (Sigma-Aldrich, Missouri, US) was first dissolved in dimethyl sulfoxide (DMSO) and then diluted with distilled water. The antibiotic was mixed with Brucella agar + 5% defibrinated sheep blood (Hy Laboratories), to final concentrations of 0.03–32 µg/mL. C. difficile colonies were suspended in thioglycollate broth medium (Becton Dickinson) to 0.5 McFarland turbidity, and then placed as spots on the fidaxomicin-supplemented agar plates. Following incubation at 35 °C, for 48 h, under anaerobic conditions, plates were screened for bacterial growth. MIC was defined as the lowest concentration of fidaxomicin that inhibited bacterial growth. Strains were classified as susceptible or resistant in accordance with the European Committee on Antimicrobial Susceptibility Testing (EUCAST) guidelines22.
Statistical analysis
Descriptive statistics are presented as follows: numbers and percentages are presented for categorical variables and mean, standard deviation and median are presented for continuous variables. Differences between probabilities of categorical variables were analysed using the Fisher exact test and the Pearson's Chi-squared test. Differences between two independent groups of continuous parameters were analysed by the independent samples Wilcoxon rank sum test. The independent samples Kruskal–Wallis rank sum test was performed to compare the median of more than two independent groups. Statistical significance was defined as p value < 0.05. Data were analysed using the RStudio® version 2021.09.0 Build 351.
Results
Demographic characteristics of patients
One hundred and eighty-eight CDI patients were enrolled in this study, 46.8% (88/188) of whom were females. The average age of the study population was 71.06 ± 17.22 years. The samples were collected from the northern (25.5% (48/188)), central (41.5% (78/188)) and southern region (33% (62/188)) regions of the country. The 30-day mortality rate was 21.8% (Supplementary Table 1).
Distribution of STs
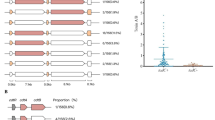
MLST analysis results were available for 89.4% (168/188) of the samples, while results were inconclusive for the rest (Fig. 1). The isolates were categorized into four major groups; each group contained at least eight isolates and an additional group called "others" included 89 different STs (with fewer than 8 isolates per ST). The most frequent STs were ST42 (13.8%, 26/188), ST2 (10.6%, 20/188), ST104 (6.9%, 13/188), ST11 (6.4%, 12/188) and ST3 (4.3%, 8/188), which all belonged to Clade 1, with the exception of ST11, which belonged to Clade 5.. Most of the "Others" group's isolates belonged to Clade 1, expect for isolates ST39 and ST37 which belonged to Clade and ST5 which belonged to Clade 3 (Fig. 1).
Phylogenetic tree of C. difficile strains, based on whole-genome sequencing. The sequence type (ST) is indicated after the strain label. MLST clades are marked with colored squares: Clade 1 (red), Clade 3 (blue), Clade 4 (purple) and Clade 5 (green). NC_009089 was used as a reference genome for alignment. Tree scale: 0.001. The Phylogenetic tree was created in iTol (https://itol.embl.de/), version 6.9.1.
Demographic characteristics of CDI patients versus ST
The mean age of patients in the different ST groups was not significantly different (p = 0.362). Similarly, no significant associations was found between ST and gender, ST and geographic location and ST and 30-day mortality (Table 1).
C. difficile’s virulence factors
All isolates harboured the genes for both toxins A + B. However, the toxin proteins were not detected in all stool samples; 5.3% (10/188) of the samples were positive for toxin A only, 21.8% (41/188) samples were positive for toxin B only, and 72.9% (137/188) were positive for both toxins. In 13 (6.9%) isolates, the binary toxin gene was detected. Of these 13 isolates, 11 (84.6%) belonged to ST11 and 2 (15.4%) belonged to ST5.
Overall, 99% (186/188) of isolates produced biofilm. Most of the biofilm-producing isolates were weak producers (57.4%, 108/188), while 19.1% (36/188) were moderate producers and 22.3% (42/188) were strong producers.
Associations between bacterial virulence factors and ST
No significant association was found between ST and toxin production (Table 2). In contrast, biofilm production capacity was associated with certain STs (p < 0.001). For example, all ST3 isolates (100%, 8/8) and most (60%, 12/60) isolates of the inconclusive group were weak biofilm producers. In contrast, most (27.7%, 15/26) isolates of ST42 and of ST11 (58.3%, 7/12) were strong producers. Among all isolates, only two isolates were non-producers, one isolate of ST104 (7.7%, 1/13) and one of the inconclusive group (5%, 1/20) (Table 2).
Antibiotic susceptibility patterns
Overall, two (1.1%, 2/188) isolates, belonging to ST34 (MIC = 3 µg/mL) and ST12 (MIC = 4 µg/mL), were resistant to vancomycin (Supplementary Table 2, Fig. 2). Two (1.1%, 2/188) isolates, belonging to ST139 and ST12, were resistant to metronidazole (both with MIC = 256 µg/mL. Eleven (5.85%, 11/188) isolates were resistant to fidaxomicin; 1 isolate (ST2) had an MIC of 1 µg/mL, 2 isolates, belonging to Inconclusive group, had an MIC of 2 µg/mL, 2 isolates of the Other group (ST12) and ST42 had an MIC of 4 µg/mL; MIC of 8 µg/mL was observed for 2 isolates of the Other (ST34) and the Inconclusive groups. MIC of 16 µg/mL was observed for ST2, 2 isolates of the Other group (ST13, ST48) and 1 isolate of the inconclusive group.
High geometric mean MICs (GM-MIC) were recorded for cephalosporins and gentamicin (Fig. 3). ST28 had the lowest GM-MIC (0.047 μg/mL) for metronidazole while ST12 and ST139 had the highest GM-MIC (256 μg/mL for both). For vancomycin, ST237 and ST604 had the lowest GM-MIC (0.38 μg/mL), while ST12 had the highest (4 μg/mL) GM-MIC. For moxifloxacin, ST604, ST160, ST139 and ST49 shared the lowest GM-MIC (0.5 μg/mL) while ST578 had the highest GM-MIC (32 μg/mL). ST17 had the lowest GM-MIC (0.06 μg/mL) while ST12 had the highest GM-MIC (4 μg/mL) for fidaxomicin (Fig. 3).
No statistically significant association was found between ST and susceptibility to metronidazole, vancomycin, or fidaxomicin (Supplementary Table 3).
Discussion
The current study characterized C. difficile isolates of patients with HA-CDI. Patient characteristics aligned with the typical patient profile reported in the literature. Specifically, mean patient age (71.06 y) corroborated reports of a tenfold higher risk for CDI development in individuals' ages 65 years and above, compared to younger people24. The 21.8% mortality rate was similar to the rate (24.6%) found in a study conducted in a Brazilian hospital25. These findings align with previous reports according to which patients with HA-CDI have increased risk for death during hospitalization, compared to other patients26,27.
The most prevalent strain in the current study was ST42 (13.8%). This was also the most common strain in the aforementioned study conducted in Brazil (43%). A study conducted in Canada during 2015–2019, found ribotype RT106 (equivalent to ST42) to be the second most common strain among HA-CDI patients, with similar prevalence (11.5%) as in our study28. This strain was also the second most prevalent strain in a 2020–2021 surveillance study in six US medical centres, where it was identified in 10.3% of patient samples29.
Previous study has demonstrated high spore production by RT10630. In addition, RT106 exhibited reduced susceptibility to antimicrobial peptides secreted by neutrophils and Paneth cells, compared to hypervirulent and epidemic ribotypes31. These characteristics that contribute to increased virulence may explain its high distribution of ST42/RT106 in healthcare facilities. RT106-induced CDI resulted in 100% mortality among infected hamsters32. In line with these findings, patients infected with RT106 had increased odds of poor clinical outcome, as compared to infection with other strains33.
All isolates in the current study harboured both toxin A and B genes, similar to what was found in a study conducted in a Japanese university hospital34. Binary toxin was detected in 6.9% of the isolates. A study performed in Brazil detected the binary toxin in 12.3% of the isolates25, while a study conducted in East China reported on a 9% prevalence35. Most of the binary toxin- positive isolates belonged to ST11. This finding correlates with the literature according to which ST11/RT078 isolates harbour the binary toxin. ST11/RT078 isolates were associated with severe CDI in North America and Europe36.
Most (98.9%) isolates produced biofilm. As mentioned earlier, biofilm contributes to bacterial resistance against antibiotics and thus increase bacterial virulence. Therefore, it is reasonable that isolates recovered from hospitalized patients, which usually have a moderate-to-severe infection, would be highly virulent. Interestingly, biofilm-production capacity was associated with ST; for example, most ST42 and ST11 isolates were strong producers, while all ST3 isolates were weak producers. These results contradict our previous study in which 92.3% of ST42 and all ST11 isolates were weak biofilm producers37. However, it should be noted that the previous work analysed CA-CDI isolates only. This may suggest that ST characteristics can be influenced by origin/environment. In addition, this may indicate further evidence of the increased virulence of HA isolates, as manifested by higher biofilm-production capacity.
We found low resistance rates to both metronidazole and vancomycin. Several groups reported that none of their isolates was resistant to these antimicrobials23,38,39. The shift of treatment guidelines to use of fidaxomicin as a first-line treatment may have contributed to the low resistance to metronidazole and vancomycin, which were the recommended treatments in previous guidelines40. Reduced susceptibility to Fidaxomicin was observed in 5.85% of the isolates. No previous study has reported on similar rate, probably due to the fact that until 2024 there were no breakpoints for fidxomicin susceptibility in guidelines of CLSI and EUCAST. Thus, it is possible that fidaxomicin reduced susceptibility is underestimated.
No associations between ST and antibiotic susceptibility were found. However, the overall resistance rates were low and in some STs, we had small numbers of isolates. Further studies should be performed with larger numbers of isolates per each ST for definitive conclusion in this issue.
Most isolates showed high MICs for antibiotics from the cephalosporin family. This is not surprising, as these antibiotics are one of the most commonly used antibiotics for treatment of various infections. Furthermore, cephalosporin use was associated with increased risk for CDI41.
This is the first comprehensive study investigating the characteristics of HA-CDI isolates from patients hospitalized in 4 medical centres across Israel. A variety of strains was found circulating in these medical centres, with ST42 being the most common strain. The strains differed in some of their characteristics, such as biofilm production. Additional studies from other medical centeres in Israel should be conducted to further explore the epidemiology of HA-CDI in Israel.
Data availability
The data used and/or analyzed during the current study are available from the corresponding author on reasonable request. DNA sequences were deposited in NCBI repository, accession number PRJNA1044122 (accession numbers of isolates SRR26919831- SRR26920158).
References
Lee, H. S., Plechot, K., Gohil, S. & Le, J. Clostridium difficile: Diagnosis and the consequence of over diagnosis. Infect. Dis. Ther. 10, 687–697. https://doi.org/10.1007/s40121-021-00417-7 (2021).
Rea, M. C. et al. Clostridium difficile carriage in elderly subjects and associated changes in the intestinal microbiota. J. Clin. Microbiol. 50, 867–875 (2012).
Cho, H. J. et al. Epidemiology and clinical characteristics of Clostridium difficile -associated disease in children: Comparison between community- and hospital-acquired infections. Korean J. Pediatr. Gastroenterol. Nutr. 13, 146 (2010).
Calatayud, M. et al. Long-term lactulose administration improves dysbiosis induced by antibiotic and C. difficile in the PathoGutTM SHIME model. Antibiotics 11, 1464 (2022).
Sacco, M. D. et al. A unique class of Zn2+-binding serine-based PBPs underlies cephalosporin resistance and sporogenesis in Clostridioides difficile. Nat. Commun. 13, 4370 (2022).
van Prehn, J. et al. European Society of Clinical Microbiology and Infectious Diseases: 2021 update on the treatment guidance document for Clostridioides difficile infection in adults. Clin. Microbiol. Infect. 27, S1–S21. https://doi.org/10.1016/j.cmi.2021.09.038 (2021).
Chen, K. Y. et al. The transcriptional regulator Lrp contributes to toxin expression, sporulation, and swimming motility in Clostridium difficile. Front. Cell Infect. Microbiol. 9, 356 (2019).
Wedari, N. L. P. H., Budayanti, N. N. S. & Darwinata, A. E. Clostridium difficile virulence factors as the cause of antibiotic-associated diarrhea (AAD): A literature review. Bali Med. J. 11, 1277–1281. https://doi.org/10.15562/bmj.v11i3.3696 (2022).
Gerding, D. N. & Lessa, F. C. The epidemiology of Clostridium difficile infection inside and outside health care institutions. Infect. Dis. Clin. North Am. 29, 37–50. https://doi.org/10.1016/j.idc.2014.11.004 (2015).
Frostid, L. R., Chengid, J. K. J. & Unnikrishnanid, M. Clostridioides difficile biofilms: A mechanism of persistence in the gut?. PLoS Pathog. 17, e1009348 (2021).
Florea, D. et al. Clinical utility of the GeneXpert assay for the diagnosis of Clostridium difficile infections. BMC Infect. Dis. https://doi.org/10.1186/1471-2334-14-S7-O34 (2014).
Abad-Fau, A., Sevilla, E., Martín-Burriel, I., Moreno, B. & Bolea, R. Update on commonly used molecular typing methods for Clostridioides difficile. Microorganisms 11, 1752. https://doi.org/10.3390/microorganisms11071752 (2023).
Griffiths, D. et al. Multilocus sequence typing of Clostridium difficile. J. Clin. Microbiol. 48, 770–778 (2010).
Lemee, L., Dhalluin, A., Pestel-Caron, M., Lemeland, J. F. & Pons, J. L. Multilocus sequence typing analysis of human and animal Clostridium difficile isolates of various toxigenic types. J. Clin. Microbiol. 42, 2609–2617 (2004).
Adler, A. et al. A national survey of the molecular epidemiology of Clostridium difficile in Israel: The dissemination of the ribotype 027 strain with reduced susceptibility to vancomycin and metronidazole. Diagn. Microbiol. Infect. Dis. 83, 21–24 (2015).
Rohana, H. et al. Characterization of Clostridioides difficile strains, the disease severity, and the microbial changes they induce. J. Clin. Med. 9, 4099 (2020).
Singhal, N., Kumar, M., Kanaujia, P. K. & Virdi, J. S. MALDI-TOF mass spectrometry: An emerging technology for microbial identification and diagnosis. Front. Microbiol. https://doi.org/10.3389/fmicb.2015.00791 (2015).
Gateau, C., Couturier, J., Coia, J. & Barbut, F. How to: Diagnose infection caused by Clostridium difficile. Clin. Microbiol. Infect. 24, 463–468. https://doi.org/10.1016/j.cmi.2017.12.005 (2018).
Biotec SL. CERTEST Clostridium Difficile GDH+Toxin A+B Revision 00 3 CerTest. www.certest.es (2013).
Hammond, E. N., Donkor, E. S. & Brown, C. A. Biofilm formation of Clostridium difficile and susceptibility to Manuka Honey. BMC Complement. Altern. Med. https://doi.org/10.1186/1472-6882-14-329 (2014).
Stepanovic, S., Vukovic, D., Dakic, I., Savic, B. & Svabic-Vlahovic, M. A Modified Microtiter-Plate Test for Quantification of Staphylococcal Biofilm Formation. Journal of Methods Microbiological Journal of Microbiological Methods vol. 40 www.elsevier.com/locate/jmicmeth (2000).
The European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters. EUCAST.
Meng, X. et al. Antibiotic resistances and molecular characteristics of Clostridioides difficile in ICUs in a teaching hospital from Central South China. Front. Med. https://doi.org/10.3389/fmed.2021.745383 (2021).
Czepiel, J. et al. Clostridium difficile infection: Review. Eur. J. Clin. Microbiol. Infect. Dis. https://doi.org/10.1007/s10096-019-03539-6 (2019).
Carvalho, F. A. C., Silva, R. O. S., Dos Santos, B. M. R. T., Diniz, A. N. & Vilela, E. G. Clinical outcome and severity of Clostridioides (Clostridium) Difficile infection at a tertiary referral hospital in Brazil. Arq. Gastroenterol. 60, 330–338 (2023).
Gao, T. et al. Association of Clostridium difficile infection in hospital mortality: A systematic review and meta-analysis. Am. J. Infect. Control 43, 1316–1320 (2015).
Wenisch, J. M. et al. A prospective cohort study on hospital mortality due to Clostridium difficile infection. Infection 40, 479–484 (2012).
Du, T. et al. Characterization of Healthcare-Associated and Community-Associated Clostridioides difficile Infections among Adults, Canada, 2015–2019. Emerg. Infect. Dis. https://doi.org/10.3201/eid2806.212262 (2022).
Snydman, D. R. et al. A US s of Clostridioides difficile associated diarrheal isolates with special reference to ridinilazole: 2020–2021. Open Forum Infect Dis 9, 1299 (2022).
Vohra, P. & Poxton, I. R. Comparison of toxin and spore production in clinically relevant strains of Clostridium difficile. Microbiology 157, 1343–1357 (2011).
Furci, L. et al. New role for human α-defensin 5 in the fight against hypervirulent Clostridium difficile strains. Infect. Immun. 83, 986–995 (2015).
Roxas, B. A. P. et al. Phylogenomic analysis of Clostridioides difficile ribotype 106 strains reveals novel genetic islands and emergent phenotypes. Sci. Rep. https://doi.org/10.1038/s41598-020-79123-2 (2020).
Almutairi, M. S. et al. Comparative clinical outcomes evaluation of hospitalized patients infected with Clostridioides difficile ribotype 106 vs. other toxigenic strains. Anaerobe 72, 102440 (2021).
Tokimatsu, I. et al. Molecular epidemiologic study of Clostridium difficile infections in university hospitals: Results of a nationwide study in Japan. J. Infect. Chemother. 24, 641–647 (2018).
Shu, C. et al. Genomic epidemiology and antimicrobial resistance profiles of Clostridioides difficile from multi-hospitals in a City in Eastern China. Infect Drug Resist 16, 3379–3388 (2023).
He, M. et al. Emergence and global spread of epidemic healthcare-associated Clostridium difficile. Nat. Genet. 45, 109–113 (2013).
Schwartz, O. et al. Characterization of community-acquired Clostridioides difficile strains in Israel, 2020–2022. Front. Microbiol. https://doi.org/10.3389/fmicb.2023.1323257 (2023).
Cui, Y. et al. An epidemiological surveillance study (2021–2022): Detection of a high diversity of Clostridioides difficile isolates in one tertiary hospital in Chongqing, Southwest China. BMC Infect. Dis. 23, 703 (2023).
Boudjelal, Y. et al. Molecular epidemiology and antimicrobial resistance patterns of Clostridioides difficile isolates in Algerian hospitals. J. Infect. Dev. Ctries. 16, 1055–1063 (2022).
Johnson, S. et al. Clinical Practice Guideline by the Infectious Diseases Society of America (IDSA) and Society for Healthcare Epidemiology of America (SHEA): 2021 Focused Update Guidelines on Management of Clostridioides difficile infection in adults. Clin. Infect. Dis. 73, 1029–1044 (2021).
Slimings, C. & Riley, T. V. Antibiotics and hospital-acquired Clostridium difficile infection: Update of systematic review and meta-analysis. J. Antimicrob. Chemother. 69, 881–891. https://doi.org/10.1093/jac/dkt477 (2014).
Funding
This research received no specific grant from any funding agency, commercial or not-for-profit sectors. *This paper is part of a research work performed in partial fulfilment of the requirements for a Ph.D. degree by Orna Schwartz, Bar Ilan University.
Author information
Authors and Affiliations
Contributions
O.SC., and A.P. conceptualized the research. O.SC., H.R., and M.A. curated the data. O.SC., H.R., M.A., A.S., N.R., and A.P. investigated the data. O.SC. performed the experiments and prepared the figures. H.R., A.S., N.R., Y.M., L.N., and O.SA assisted in some of the experiments. M.A. and A.P. supervised the project. O.SC., M.A., and A.P. wrote the original manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Schwartz, O., Rohana, H., Azrad, M. et al. Virulence factors, antibiotic susceptibility and sequence type distribution of hospital-associated Clostridioides difficile isolates in Israel, 2020–2022. Sci Rep 14, 20607 (2024). https://doi.org/10.1038/s41598-024-71492-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-71492-2
- Springer Nature Limited