Abstract
Thyrinteina arnobia (Lepidoptera: Geometridae) is a native American species. Despite its historical importance as an insect pest in Eucalyptus plantations, more information is needed regarding the population diversity, demography, and climatic variables associated with its distribution in different regions of Brazil. We used a phylogeographic approach to infer the genetic diversity, genetic structure, and demographic parameters of T. arnobia. We also conducted an ecological niche modeling (ENM) to predict suitable areas for T. arnobia occurrence in Brazil and other countries worldwide. Although T. arnobia populations have low genetic diversity in Brazil, we identified mitochondrial haplogroups predominating in different Brazilian regions and high ФST and ФCT values in AMOVA, suggesting a low frequency of insect movement among these regions. These results indicate that outbreaks of T. arnobia in Eucalyptus areas in different regions of Brazil are associated with local or regional populations, with no significant contribution from long-distance dispersal from different regions or biomes, suggesting that pest management strategies would be implemented on a regional scale. In Brazil, the demographic and spatial expansion signals of T. arnobia seem to be associated with the history of geographical expansion of Eucalyptus plantations, a new sustainable host for this species. ENM indicated that isothermality and annual rainfall are critical climatic factors for the occurrence of T. arnobia in tropical and subtropical areas in the Americas. ENM also suggested that T. arnobia is a potential pest in Eucalyptus areas in all Brazilian territory and in regions from Africa, Asia, and Oceania.
Similar content being viewed by others
Introduction
The Brazilian trade in forest products has grown significantly in the last 100 years, with an inevitable demand for increased production in the forestry sector. Trees of the genus Eucalyptus (Myrtaceae), native to Australia, were introduced into Brazil due to their high wood and cellulose production potential for use in industrial sectors of energy production, textile sector, furniture manufacturing, and pharmacies1. Eucalyptus was first planted in Brazil in the southeast region (mainly in São Paulo and Minas Gerais states) in 19052. After 1950, Eucalyptus plantations were expanded to neighboring states and into northern Brazil around the 2000s, and by 2021, they had reached a total of 7.5 million hectares3.
Efforts to improve Eucalyptus productivity in Brazil are negatively affected by damage from native and exotic pests and pathogens4,5. Thyrinteina arnobia (Stoll, 1782) (Lepidoptera: Geometridae), known as the eucalyptus brown looper, is a species native to the Americas that occurs in nearly all South America and parts of Central and North America6. This species feeds on several native hosts, with a preference for members of the family Myrtaceae6,7. Thyrinteina arnobia has found a vulnerable host in Eucalyptus species and has become the primary defoliator of this crop, mainly in subtropical and tropical Brazil8,9. Eucalyptus plantations infested by T. arnobia can reduce their annual biomass production by up to 40%10.
Despite the historical importance of T. arnobia as an insect pest in Eucalyptus plantations in Brazil11, more information is needed regarding its population diversity and demography in different regions of the country. Studies phylogeography aims to investigate local patterns of historical dispersal, revealing the presence of genetic strains, population structure, and demographic expansion of a species. This approach increases understanding of historical and contemporaneous processes associated with the distribution of strains and can provide new prospects for the management of agricultural pests12,13,14,15. Divergent strains or populations of a species could respond differently to control strategies commonly used for pest control in farming areas, such as plant resistance (including transgenic crops), insecticides, pheromones, and natural enemies16,17,18.
Phylogeography uses molecular markers, mainly DNA sequencing, to provide insights into the history of the genetic diversity and distribution of species in both space and time19,20. Additionally, ecological niche models (ENMs) combine collection localities and Geographic Information System (GIS) maps of environmental data to create spatial predictions concerning a species’ historical and current ranges21,22,23. ENM is widely used to estimate the suitability of environments for organisms, helping to build ecological models for prediction in specific geographic areas. This way, we can examine species distribution patterns using ecological niche models, combining species presence records with environmental variables24,25,26. Despite T. arnobia being a native pest, reports of its occurrence in Brazil are frequently associated with Eucalyptus areas due to a lack of information on its presence and distribution in native environments. Thus, ENMs could bring new insights into the distribution of this species in Brazilian territory, including new Eucalyptus areas, and its potential as an invasive pest in Eucalyptus plantations worldwide. Primarily because invasions of agricultural and forest pests have become increasingly frequent over the past two decades27,28.
Given the ongoing expansion of Eucalyptus plantations from southeastern to central and northeastern Brazil and the reports of T. arnobia infestation in old and new cultivation areas, it is imperative to comprehend the genetic relationships among its populations in Eucalyptus areas to know if the populations are connected among them and in demographic expansion in Brazil. Furthermore, estimating the suitable areas for this species in Brazil could help predict the potential occurrence of T. arnobia in new Eucalyptus areas in Brazil and worldwide. Therefore, integrating phylogeography with ENMs may increase understanding of the distribution and evolution of T. arnobia, offering critical data to enhance pest management strategies29,30. Specifically, our objectives were twofold: (i) to infer the genetic diversity, genetic structure, and demographic parameters of T. arnobia, aiming to estimate the historical dynamics of its colonization of the main Eucalyptus production areas in Brazil, and (ii) to conduct species modeling to forecast suitable areas for the occurrence of T. arnobia and its haplogroups in Brazil and other countries worldwide.
Diversity statics and genealogical inferences
We sequenced 139 individuals for the mitochondrial genes COI and COII. One hundred thirty-two T. arnobia from 16 Eucalyptus plantations across three geographical regions in Brazil: Atlantic Forest, the transition between Atlantic Forest and Southern Cerrado (AF/CE), and Northern Cerrado (Table S1 and Table 1). The other seven individuals from T. leucoceraea species were used to confirm the species identification (Table S1 and Table 1). We obtained a total of 1303 bp for the concatenated COI and COII gene sequences (642 and 660 bp, respectively). Twenty-two haplotypes were found for T. arnobia, with total haplotype diversity Hd = 0.858, nucleotide diversity π = 0.0028, and a mean number of nucleotide differences k = 3.72924 (Table 1). Sequence analysis of the EF-1α region identified 14 haplotypes with a haplotype diversity of 0.598, nucleotide diversity of 0.0013, and a mean of 0.749 nucleotide differences (Table 1).
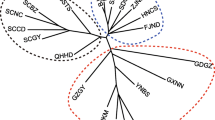
Genealogical relationships among 22 mitochondrial haplotypes of T. arnobia indicated three haplogroups distributed in Brazil (Fig. 1A, B). Haplogroup_G1 is more abundant in the Atlantic Forest biome; haplogroup_G2 is more abundant in the transition between Atlantic Forest and South Cerrado (AF/CE); and haplogroup_G3 is more abundant in the North Cerrado biome (Fig. 1C). Haplotypes H4, H9, and H16 are associated with haplogroups G1, G3, and G2, respectively, but do not show the same geographic distributions as their haplogroups.
(A) Bayesian phylogeny tree showing posterior probability values (> 90), and (B) haplotype network of Thyrinteina arnobia and T. leucoceraea (Lepidoptera: Geometridae) based on a 1303 bp fragment of the concatenated mitochondrial genes COI and COII. (C) Geographic distribution and frequency of the mitochondrial haplotypes in different regions of Brazil. Circles represent a single haplotype, and its colors are a haplogroup.
Analyses of the EF-1α region of T. arnobia revealed a single nucleotide polymorphism variation separating the haplotypes. The 14 haplotypes are separated by a one-step mutation, except for haplotype H14, which has two mutational steps (Fig. 2). The EF-1α network had lower haplotype diversity than the mitochondrial network. Haplotype H1 was the most frequent (58.6%) and was widely distributed across all regions (Fig. 2 and Table 1).
Haplotype network of Thyrinteina arnobia (Lepidoptera: Geometridae) based on a 532 bp fragment of the nuclear gene EF-1α. The colors represent mitochondrial haplogroups in Fig. 1.
Population structure
The analysis of molecular variance (AMOVA) of the mtDNA haplotypes found for T. arnobia, considering two hierarchical levels, revealed high variation among groups (65.17%, ФST = 0.652; p < 0.001) and consequently low variation within groups (34.83%) (Table 2a). When the AMOVA was divided into three hierarchical groups (Atlantic Forest, AF/CE, and North Cerrado biomes), we found a significantly high value, confirming our hypothesis that the genetic variation among T. arnobia populations is influenced by geographical regions where they were collected (ФCT = 0.578, p < 0.001). Furthermore, the value of genetic variation within populations (28.8%, ФST = 0.712) indicates an overall genetic differentiation in these populations into groups (Table 2a).
The AMOVA results for the EF-1α gene showed low variation among populations at two hierarchical levels (6.21%, ФST = 0.062; p > 0.01) and, consequently, high variation within groups. When the AMOVA was divided into three hierarchical levels, the variation among groups remained low and non-significant, and the variation within populations remained high (Table 2b).
Pairwise FST values calculated with mtDNA data varied from 0.00 to 0.971 (Table S2). The Mantel test, which examines the relationship between Slatkin’s linearized FST and linear geographic distances, revealed a low but significant correlation coefficient among the variables (r2 = 0.069; p = 0.0214) (Fig. S1).
Demographic analysis using mtDNA genes
The neutrality tests indicated that the populations of T. arnobia are not recent population expansion or contraction (Tajima’s D = − 0.8; p = 0.22; Fu’s Fs = − 4.524; p = 0.099) (Table 3). These results suggest that the polymorphisms found conform to the neutrality model proposed by TAJIMA31, and in general, the populations of T. arnobia are not expanding32. This holds true when considering the AMOVA groups separately, except for the Atlantic Forest population, which shows evidence of demographic expansion as seen in the results for Tajima’s D and Fu’s Fs (p-value), respectively: D = − 1.756 (p < 0.02), Fs = − 3.742 (p < 0.01) (Table 3).
The results of mismatch distribution analyses for observed and simulated values for spatial expansion were not significant for T. arnobia individuals when evaluated as a whole (SSD = 0.01718, p > 0.23; Raggedness = 0.049, p > 0.16). When divided into groups, the SSD and Raggedness index values were also not significant, indicating a spatial expansion for T. arnobia in the locations sampled. This is evidenced by the rejection of the null hypothesis of unimodal curve patterns and non-significance (p > 0.05) (Table 3).
According to the Extended Bayesian Skyline Plot (EBSP) analysis, the population expansion of T. arnobia in Brazil occurred within the last 100 years (Fig. 3). In the Atlantic Forest region, the expansion also occurred in the last 100 years (Fig. 3A), and in AF/CE and North Cerrado in the last 10 years (Fig. 3B, C).
Extended Bayesian Skyline Plot (EBSP) showing population size dynamics throughout time for Thyrinteina arnobia (Lepidoptera: Geometridae) in Brazil based on a 1303 bp fragment of the concatenated mitochondrial genes COI and COII. The y-axis indicates effective population size (Ne) scaled by mutation rate (µ) as a function of time. The black horizontal line shows the median EBSP estimate, and the gray area shows the upper and lower 95% highest posterior density limits. (A) Atlantic Forest (B) North Cerrado (C) transition between Atlantic Forest and South Cerrado biomes/region (AF/CE).
Modeling performance
The test AUC values for the initial and final models were 0.970 and 0.952, respectively, with standard deviations of 0.005 and 0.011. These results attest to the models’ high performance33.
Contribution of environmental variables
The Pearson correlation coefficients between the 20 environmental variables are presented in Table S3, while the percentage contributions of the variables in the initial model are shown in Table S4. Following the screening procedure, Isothermality (Bio3), Mean temperature in the coldest quarter of the year (Bio11), Annual rainfall precipitation (Bio12), and Precipitation in the hottest quarter of the year (Bio18) were selected for the final model. Isothermality and annual precipitation collectively contributed 78.4% of the model output (Table S5), and their response curves are shown in Fig. 4A and B, respectively.
As seen in Fig. 4A, the highest probability of T. arnobia occurrence will be in regions with isothermality, approximately 77%. Likewise, Fig. 4B shows that the peak probability of occurrence is expected in areas with annual precipitation close to 3500 mm; beyond this threshold, the standard deviation increases substantially. Each of these response curves represents a model created by the MaxEnt using only the corresponding variable. They reflect the dependence of the predicted suitability on the selected variable. According to the Jackknife test of variable importance, Bio3 emerged as the environmental variable with the highest gain when used alone. Hence, it provides the most helpful information on its own (Fig. S2). The environmental variable that showed the most significant decrease in gain when omitted is Bio3, suggesting that it contains information not present in the other variables. These values were averaged over replicate runs.
Global and local predicted suitable areas for Thyrinteina arnobia
Our ENM analysis revealed suitable areas for the species in diverse regions over Brazil and in Central America (Fig. 5). As a potential invasive pest, we found suitable areas in Africa, such as the sub-Saharan region, on the east coast, and in Madagascar. Suitable areas are also present in Southeast Asia, in some parts of India, Myanmar, Thailand, Laos, Cambodia, Vietnam, and Malaysia; the Philippines; and Indonesia. In Oceania, Australia’s east coast and parts of New Zealand have suitable areas.
We also modeled the suitability for each T. arnobia haplogroup within Brazil. Haplogroup_G1 and Haplogroup_G2 have similar potential suitability in Brazil, while Haplogroup_G3 is restricted more to North Brazil (Fig. 6). The future T. arnobia haplogroups suitability in Brazil is little changed until 2050 (Fig. S3).
Discussion
This study is the first to employ molecular tools to gain insights into the diversity and history of dispersal of T. arnobia populations collected in Eucalyptus areas in Brazil. In general, mitochondrial (mtDNA) and nuclear markers (ncDNA) revealed a moderate haplotype and low nucleotide diversity across the country. However, the mtDNA genes proved to be more valuable markers for phylogeographic inferences of T. arnobia, generating a larger number of haplotypes and, notably, greater nucleotide diversity compared to the ncDNA marker. The higher diversity of mtDNA genes than single-copy nuclear genes aligns with typical findings in insect phylogeographic and phylogenetic studies34,35,36,37. This phenomenon is likely associated with differences in evolution between mitochondrial and nuclear genomes38,39,40.
The mitochondrial and nuclear haplotype networks revealed a close genealogical relationship among T. arnobia haplotypes. Despite being native to the Americas, our results suggest that T. arnobia populations in Brazil have recently undergone demographic changes, possibly linked to recent genetic differentiation or dispersal events. This is supported by the descriptions of five T. arnobia subspecies, only one of which is found in Brazil6,41. Our study confirmed the presence of one subspecies based on the low genetic diversity in Brazil. This implies that the diversification center of T. arnobia is not in Brazil but probably in Central America. Similar findings of low genetic diversity associated with recent dispersal to South America have been reported for other agricultural insect pests native to the Americas42,43,44,45.
Despite the low nucleotide diversity, the mitochondrial haplotype network revealed the presence of three haplogroups, which were further supported by a Bayesian phylogeny. A clear relationship exists between the mtDNA network architecture and the geographic origin of individuals across different regions in Brazil. Haplogroup_G1, the most ancestral among the haplogroups, is more abundant in the Atlantic Forest region. Haplogroup_G2 is more abundant in the northern Cerrado, and Haplogroup_G3 is more abundant in the AF/CE. Three haplotypes are not geographically located together with their respective haplogroups, but each of these haplotypes occurs in only one region (see Fig. 1). This suggests the possibility of insect movement among Eucalyptus areas in different Brazilian regions, but not in high frequency, at least in the last few years. Specifically, these long-distance dispersal events can be associated with natural events or mediated by human activity related to the spatial expansion of Eucalyptus plantations, forestry industries, or pest-management activities in the past.
The AMOVA confirmed a high population structure among populations, particularly when considering geographical regions. This supports the inference of a low movement frequency of T. arnobia individuals among Eucalyptus areas in different Brazilian regions. Thus, considering the distribution pattern of the mtDNA haplotypes and the AMOVA index, we can confirm that the insects associated with population outbreaks of T. arnobia in Eucalyptus areas originated from local or regional populations, with no significant contribution via long-distance dispersal of insects from different regions. However, we cannot infer insect movement and gene flow among T. arnobia populations within the same geographical regions using these markers alone. This regional genetic structure, observed in T. arnobia populations using mtDNA markers, was also observed in populations of Diatraea saccharalis (Fabricius, 1794) (Lepidoptera: Crambidae) in Brazil. Some hypotheses for this genetic structure are associated with isolation by distance, demographic expansion, and genetic drift promoted by agricultural practices46,47. However, the high genetic structure is not standard for all lepidopteran pests in Brazil, e.g., Noctuidae moths, associated with annual crops (e.g., maize, soybean, and cotton), show low genetic structure in agricultural landscapes48,49,50, probably associated with a high dispersion capacity of this group51,52,53.
The nuclear haplotype network showed a star-like shape, with the central haplotype (H1) distributed in all populations of T. arnobia and connected to other low-frequency haplotypes. These low-frequency haplotypes are separated by a single mutation step, with their distributions each limited to a single biome. The absence of haplogroups supported by nuclear markers in our analysis is probably associated with a low genetic diversity of this marker compared to mtDNA markers. It is reinforced by the limitation of each low-frequency haplotype to one region or biome. However, the nuclear data confirmed the recent genealogical relationship among T. arnobia populations and the possibility of gene flow among populations located in different geographical areas in Brazil.
The neutrality tests Fu’s Fs and Tajima’s D assessed demographic aspects for T. arnobia populations. These tests are typically based on the distribution of pair-wise differences among sequences within populations. The neutrality tests gave significantly negative values for the Atlantic Forest biome, indicating that populations in this region are undergoing demographic expansion. Although most populations are not under the effect of demographic expansion, all are experiencing spatial expansion, as revealed by the Mismatch Distribution test. The skyline analysis confirmed a recent demographic expansion of T. arnobia in Brazil, agreeing with the above discussion. Furthermore, the expansion of T. arnobia occurred within the last 100 years, suggesting a strong association with the history of the expansion of Eucalyptus plantations in Brazil, which reached 7.6 million hectares in 20223,54. Demographic and spatial expansion associated with the increase of cultivated areas in Brazil is reported to other agricultural pests such as moths, stink bugs, and leaf miners14,55,56. This fact is frequently justified by the rise of resources provided for the agricultural areas for these species14,45.
With the continuous territorial expansion of Eucalyptus areas in Brazil, it is essential to understand which areas are most suitable for T. arnobia and its haplogroups in the country, mainly due to occurrence reports of this species being more frequently associated with Eucalyptus areas. Our analysis produced robust models with excellent performance, considering the high AUC values (AUC value > 0.90)41,54. Within Brazil, the suitable area for T. arnobia, as revealed by our analyses, aligns with the current distribution in the tropical and subtropical regions of the Americas. It confirms the potential of T. arnobia as a pest in new Eucalyptus areas in Brazil and as an invasive pest in parts of Africa, Asia, and Oceania.
According to the model, isothermality and annual precipitation are the critical climatic factors for T. arnobia occurrence. They could be future indicators of population outbreaks of this pest in Eucalyptus plantation areas. These variables are shown to influence the global distribution of T. arnobia, impacting the spread of the pest. Temperature contributes significantly more than precipitation conditions57, indicating a critical role in the spread of T. arnobia. Isothermality, i.e., the day-to-night temperature oscillations relative to the summer-to-winter (annual) oscillations (Bio2/Bio7)58.
Our results indicated little probability of T. arnobia movement among Brazilian regions. However, it is essential to understand the potential distribution of T. arnobia haplogroups within the country. Therefore, we also used the niche model approach to estimate the potential geographical distribution for each T. arnobia haplogroup. Although we lack information about the biotic potential of the different T. arnobia populations associated with each haplogroup, hybridization can increase the genetic diversity and the potential success of this species as an Eucalyptus pest in Brazil59,60. Our results showed that considering the haplogroups’ potential distribution, the present T. arnobia populations are not restricted by environmental factors in dispersing and colonizing other areas in Brazil, with the possibility of gene flow among populations from different haplogroups. For instance, haplogroups_G1 and G2 may eventually occupy nearly all of Brazil. Haplogroup_G3 may occupy a significant portion of the central and northern regions. In projected future climate scenarios from 2030 to 2050, our models do not indicate any significant change in the size of areas favorable to this insect pest in Brazil.
In conclusion, T. arnobia populations have low/moderate genetic diversity in Brazil. However, we identified the presence of mitochondrial haplogroups that predominate in different Brazilian regions, suggesting a low frequency of insect movement among these regions. Therefore, population outbreaks of T. arnobia in Brazil consist of local or regional insects, with no significant contribution through long-distance dispersal from different regions or biomes. It suggests that, when possible, the management strategies for T. arnobia populations would be implemented on a regional scale. The demographic and spatial expansion signals of T. arnobia are associated with the history of expanding Eucalyptus plantations, confirming it as the new sustainable host for T. arnobia. According to the ENM, isothermality, and annual precipitation are the critical climatic factors for T. arnobia occurrence in Brazil, with potential for distribution in Eucalyptus areas worldwide. Finally, a genomic approach is necessary to measure T. arnobia gene flow among populations located within and among geographical regions. Furthermore, genomic data can provide new insight into this species’ adaptation to Eucalyptus plantations and landscape variables in Brazil, providing crucial information for this pest management.
Methods
Insect collection and DNA extraction
Adults of T. arnobia were collected from 16 Eucalyptus plantations across three geographical regions in Brazil: Atlantic Forest, the transition between Atlantic Forest and Southern Cerrado (AF/CE), and Northern Cerrado (Table S1). For DNA extraction, we macerated the thoraxes of individual adult moths in liquid nitrogen and followed the standard genomic DNA extraction method, using CTAB (Cetyltrimethylammonium bromide) buffer with proteinase K for protein/enzyme digestion, followed by washing with a chloroform (CIA) solution and total DNA precipitation with isopropanol61. The DNA obtained at the end of the process was resuspended in 50 µL of ultrapure water and stored in a freezer at − 20 °C.
Mitochondrial and nuclear gene amplification and sequencing
Two mitochondrial gene fragments, COI (cytochrome c oxidase subunit I) and COII (cytochrome c oxidase subunit II), and a fragment of the nuclear gene EF-1α (eukaryotic translation elongation factor 1 alpha 1), were sequenced in our study. The primers used to amplify the COI gene were designed by Folmer et al.62, LCO1490 (Forward) (5′-GGTCAACAAATCATAAAGATATTGG- 3′) and HCO2198 (Reverse) (5′-TAAACTTCAGGGTGACCAAAAAATCA-3′). For the COII gene, the primers PATRICK (Forward) (5′-CTAATATGGCAGATTATATGTAATGGA-3′) and EVA (Reverse) (5′- GAGACCATTACTTGCTTTCGATCATCT-3′) were applied for amplification63. For the nuclear EF-1α gene, the primers used were EF1A-2 (Forward) (5′-GGCATCGAGGGCTTCAATAA- 3′) and EF1A-3R (Reverse) (5′-GTCCCATCCCTCAGCRTTAC-3′). The PCR reactions followed the pattern: 2.0 µL dNTP (2.5 mM) (Sinapse Inc.), 2.0 uL Primer F (5 µM), 2.0 uL Primer R (5 µM), 2.5 uL MgCl2 (25 mM), 2.5 uL buffer (Mg2+ Free; Sinapse Inc.), 10.7 uL MilliQ water, 0.3 uL Taq Polymerase (1 U) (Sinapse Inc.), and 3 uL of the individual’s DNA, for a total volume of 25u L per reaction. The PCR cycles consisted of an initial denaturation at 95 °C for 3 min; 35 cycles of denaturation for 30 s at 95 °C, annealing for 30 s, and extension for 2 min at 72 °C; and a final extension at 72 °C for 10 min. The annealing temperatures were 53, 62, and 59 °C for the COI, COII, and EF-1α genes, respectively. PCR amplicons were visualized under ultraviolet light after electrophoresis using 3 uL of the amplicon in a 1.5% v/v agarose gel. All PCR amplicons were purified using 1.0 µL of EXO + SAP enzymes (Cellco©) for every 10 µL of amplicon and sent for bidirectional Sanger sequencing at the Laboratório de Genômica e Biologia Molecular de Plantas (CEBTEC) at ESALQ/USP.
Assembly of sequencing datasets
All sequences were aligned and edited manually using the software BioEdit64. To eliminate missing data, sequences were interrupted at 643 bp for the COI gene, 660 bp for the COII gene, and 532 bp for the EF-1α gene. No insertions or deletions were found in the sequences obtained. We examined the mitochondrial gene fragments for the presence of nuclear paralogs of mitochondrial origin, known as numts65, using MEGA v.5.2 software66, after we concatenated the COI and COII genes for subsequent analysis. We created two alternative EF-1α sequences for all individuals that showed ambiguity in the polymorphic site. When two or more polymorphic sites were observed, we used the criteria of the haplotype frequency in a population to define the haplotypes. All datasets generated in this study have been submitted to NCBI-GenBank with the accession numbers PP723897-PP724035 (COI), PP750573–PP750711 (COII), and PP750741–PP750786 (EF-1α).
Diversity and genealogical analysis
Using DnaSP software v6.12.0167, we calculated the haplotype diversity indices (Hd), number of haplotypes (H), nucleotide diversity (π), and mean number of nucleotide differences (k) for the T. arnobia populations. We established genealogical relationships among T. arnobia sequences by constructing a Bayesian phylogeny68 and a Median Joining haplotype network in PopArt software v1.769. For the Bayesian analysis, we found the best nucleotide substitution model and parameters HKY + I + G for phylogenetic analysis, using the software MrModeltest v2.470 using the Akaike Information Criterion71. The tree was generated using two simultaneous runs of 100 million generations each in MrBayes software68. We obtained a 50%-majority-rule consensus tree of the two independent runs with posterior probabilities that were equal to bipartition frequencies.
Subsequently, we conducted a molecular variance analysis (AMOVA) using the Arlequin v3.1 program72, applying 5000 replicates for bootstrap parameters and a 5% significance level. We performed non-hierarchical analyses, which considered all sampling locations. Afterward, we examined the variance between populations based on each collection site and among geographically structured populations separated into groups that include the criteria of biome and geographical distance (Atlantic Forest, transition between Atlantic Forest and South Cerrado (AF/CE), and North Cerrado).
Using the program Arlequin, we calculated the pairwise FST matrix as described by Slatkin73. To determine if there is an inverse relationship between gene flow and geographic distance, we examined the correlation between the pairwise FST values and the linear geographic distances. The Mantel test74 was employed to test the hypothesis of genetic isolation by geographic distance (IBD), utilizing 10,000 permutations.
Demographic analysis using mtDNA genes
Tajima’s D and Fu’s Fs neutrality tests were computed using Arlequin v.3.1. Both tests involved 1000 permutations utilizing coalescing simulations. The significance of Fu’s Fs statistic was determined at the 95% confidence level, considering a p-value less than 0.02. Additionally, we calculated tau (Τ) with its 95% confidence intervals for each group, using a generalized least-squares approach and 1000 coalescent simulations in Arlequin v.3.1.
We conducted a mismatch distribution analysis, using a spatial expansion model to test for population expansion and relate the results to Tajima’s D and Fu’s Fs neutrality tests. The sum of squared differences (SSD), Raggedness index (R) statistics, and their associated p-values were also calculated using Arlequin v.3.1. A non-significant SSD value implies that the hypothesis of population expansion cannot be rejected, and a non-significant raggedness index suggests a good fit of the data to the spatial expansion model75.
Additionally, we employed an Extended Bayesian Skyline Plot (EBSP) in Beast to reconstruct the demographic history, utilizing TRACER v.1.6, based on the COI-COII data. The HKY substitution model was chosen because it works with fewer parameters. A strict clock model with a mutation rate based on similar organisms was employed, given the absence of a specific value for this species. We used the divergence rates of 3.54% My1 for COI76 and 2.76% My1 for COII77. We assumed that T. arnobia has 6 generations per year, as described by Pereira et al.78. The chain length was set at 100 million generations, sampled every 4000 states, and a 20% burn-in was applied in three independent runs. Results were examined in Tracer, and the posterior ESS (effective sample size) and convergence among the runs were assessed.
Thyrinteina arnobia occurrence dataset
To obtain the worldwide occurrence records of T. arnobia, we use the online database Global Biodiversity Information Facility79 and the occurrence records of the species collected by the company Suzano S.A. (Tables S2 and S6). To mitigate potential bias in the dataset, we implemented spatial filtering using spThin, an R package80. Following the filtering procedure, all 360 remaining occurrence points (Tables S2 and S6) were separated by a minimum of 10 km and were preserved for model construction. This ensured that each grid cell contained only one occurrence81,82. The total occurrence dataset was subdivided into three subgroups of occurrences according to T. arnobia haplogroups: haplogroup_G1 (more abundant in the Atlantic Forest), haplogroup_G2 (Transition Atlantic Forest/South Cerrado), and haplogroup_G3 (North Cerrado). In addition to the global model, a model was developed for each haplogroup with its occurrence points.
Environmental data layers
Initially, we opted for climatic and elevation factors as the initial environmental variables. Twenty variables sourced from WorldClim version 2.1 were initially employed as potential predictors (Table S7). These data layers had a 2.5-min spatial resolution (approximately 5 km), which proved adequate given their worldwide coverage of all land surfaces.
Recognizing that the selection of environmental variables is pivotal in determining simulation accuracy and considering that MaxEnt calculates the contribution of each predictor variable to the model, we employed a method to screen the most suitable variables for our dataset82. Choosing the best variables for the final model included several steps. First, an initial model was created to assess the percentage contribution of each initial variable (as shown in Table S7). Then, ArcGIS 10.1 was used to extract in each of the 360 presence records the respective attribute values from each of the initial variables. After that, the Pearson correlation between the initial variables was calculated. If the correlation coefficient between any two variables was higher than 0.7, the most important variable based on its percentage contribution in the initial model was retained, while the other variable was excluded. Finally, the remaining variables were sorted by percentage contribution, and only those with more than 1% contribution were kept for the final model.
Future projections for Thyrinteina arnobia
Our study focused on two future periods (2030 and 2050) under the SSP5-8.5 scenarios outlined in the sixth assessment report of the Intergovernmental Panel on Climate Change (IPCC AR6). The SSP5-8.5 scenario defines the upper boundary among the range of scenarios described in the literature83. The Global Climate Model (GCM) utilized in our analysis is the Model for Interdisciplinary Research on Climate (MIROC-6). This model was developed through collaboration between the Atmosphere and Ocean Research Institute (University of Tokyo), the National Institute for Environmental Studies (Japan), and the Japan Agency for Marine-Earth Science and Technology. MIROC-6 is a newly developed climate model featuring updates to its physical parameterizations across all sub-modules84.
Model development and validation
The global potential distributions of T. arnobia were predicted using MaxEnt version 3.4.485. First, we created an initial model as part of the variable selection process for the final model, such as the descriptive Sect. “Environmental data layers”. Given that the data were sourced from external providers and we lacked control over the sampling process, we constructed a sampling bias surface, utilizing the kernel density estimate available in the SDMToolbox86. The final T. arnobia model was based on fine-tuning the default MaxEnt settings for specific combinations of resource types and the regularization multiplier (RM)57,87,88. Linear (L), quadratic (Q), product (P), and hinge (H) feature sets were employed, utilizing automatic feature selection alongside RM = 1 to regulate the complexity of the model for the species89. Additionally, regarding modeling accuracy, we evaluated the AUC (area under the receiver operating characteristic [ROC] curve) for the final model performance33.
We calculated the AUC by running a tenfold cross-validation in MaxEnt. An AUC value of 0.5 indicates that model predictions are no better than random; values < 0.5 are worse than random; values between 0.5 and 0.7 indicate poor performance; values between 0.7 and 0.9 indicate moderate performance; and values > 0.9 indicate high performance54. We used the Jackknife test to identify the environmental variables that most influenced the final model90,91,92. The final Maxent maps showed values ranging from 0 to 1, indicating T. arnobia occurrence probability. We selected the Maximum Test Sensitivity Plus Specificity threshold (MTSPS) to distinguish suitable and unsuitable areas for T. arnobia. MTSPS is considered simple and equal to or more effective than other, more intricate approaches93,94.
Data availability
The gene sequences produced in this work were submitted to GenBank: PP723897-PP724035 (COI), PP750573–PP750711 (COII), and PP750741–PP750786 (EF-1α). The dataset used in the Ecological Modelling Niche analysis was included in the Supplementary Material.
References
Queiroz, L. R., de S. & Barrichelo, L. E. G. O Eucalipto, Um Século No Brasil = The Eucalypt - a Century in Brazil. (Antonio Nellini Editora & Cultura, São Paulo, 2007).
Jacobs, M. R. Eucalypts for Planting. FAO Forestry Series. vol. 11 (Food and Agriculture Organization of the United Nations, Urbana, IL, 1981).
Rodrigues, G. S. de S. C., Ross, J. L. S., Teixeira, G., Santiago, O. R. P. L. & Franco, C. EUCALIPTO NO BRASIL - Expansão Geográfica e Impactos Ambientais. (Composer, Uberlândia, 2021).
Schühli, G. S., Penteado, S. C., Barbosa, L. R., Reis Filho, W. & Iede, E. T. A review of the introduced forest pests in Brazil. Pesqui. Agropecu. Bras. 51, 397–406 (2016).
Barbosa, L. R., Domingues, M. M., Wilcken, C. F. & Zanuncio, J. C. Eucalyptus pests. In Forest microbiology 227–249 (Elsevier, 2023). https://doi.org/10.1016/B978-0-443-18694-3.00020-1.
Scoble, M. J. Geometrid Moths of the World a Catalogue: (Lepidoptera, Geometridae). (Collingwood, VIC CSIRO Stenstrup, Denmark Apollo Books, Collingwood, VIC Stenstrup, Denmark, 1999).
de Oliveira, H. N., Zanuncio, J. C., Pedruzzi, E. P. & Espindula, M. C. Rearing of Thyrinteina arnobia (Lepidoptera: Geometridae) on guava and eucalyptus in laboratory. Braz. Arch. Biol. Technol. 48, 801–806 (2005).
Holtz, A. M. et al. Adaptação de Thyrinteina arnobia em novo hospedeiro e defesa induzida por herbívoros em eucalipto. Pesqui. Agropecu. Bras. 38, 453–458 (2003).
Medauar, C. C. et al. Zoning the occurrence of Thyrinteina arnobia in eucalypts and its relationship with the spectral behaviour of canopies and soil attributes. South. For. J. For. Sci. 86, 53–64 (2024).
Oda, S. & Berti-FIlho, E. Incremento anual volumétrico de Eucalyptus saligna Sm. em áreas com diferentes níveis de infestação de lagartas de Thyrinteina arnobia (Stoll, 1782) (Lepidoptera, Geometridae). IPEF Piracicaba 17, 27–31 (1978).
dos Santos Bobadilha, G. et al. Effect of defoliator insect on growth and wood properties of eucalypt trees. Eur. J. Wood Wood Prod. 77, 861–868 (2019).
Fresia, P., Azeredo-Espin, A. M. L. & Lyra, M. L. The phylogeographic history of the New World screwworm fly, inferred by approximate Bayesian computation analysis. PLoS One 8, e76168 (2013).
Peretolchina, T. et al. Phylogeography and demographic history of the Chagas disease vector Rhodnius nasutus (Hemiptera: Reduviidae) in the Brazilian Caatinga biome. PLoS Negl. Trop. Dis. 12, e0006731 (2018).
Soares, P. L., Cordeiro, E. M. G., Santos, F. N. S., Omoto, C. & Correa, A. S. The reunion of two lineages of the Neotropical brown stink bug on soybean lands in the heart of Brazil. Sci. Rep. 8, 2496 (2018).
Ströher, P. R. et al. Phylogeography of ants from the Brazilian Atlantic Forest. Org. Divers. Evol. 19, 435–445 (2019).
Nagoshi, R. N. & Meagher, R. L. The Spodoptera frugiperda host strains: What they are and why they matter for understanding and controlling this global agricultural pest. J. Econ. Entomol. 115, 1729–1743 (2022).
Hickmann, F., Cordeiro, E. M. G., Aurélio, M. S. L., Saldanha, A. V. & Corrêa, A. S. Host preference and survivorship of Euschistus heros strains on cotton and soybean. Entomol. Exp. Appl. 171, 564–571 (2023).
Coates, B. S., Dopman, E. B., Wanner, K. W. & Sappington, T. W. Genomic mechanisms of sympatric ecological and sexual divergence in a model agricultural pest, the European corn borer. Curr. Opin. Insect Sci. 26, 50–56 (2018).
Avise, J. C. et al. Intraspecific phylogeography: The mitochondrial DNA bridge between population genetics and systematics. Annu. Rev. Ecol. Syst. 18, 489–522 (1987).
Beheregaray, L. B. Twenty years of phylogeography: The state of the field and the challenges for the Southern Hemisphere. Mol. Ecol. 17, 3754–3774 (2008).
da Silva Santana, G. et al. Climate suitability modeling for Anastrepha suspensa (Diptera: Tephritidae): Current and future invasion risk analysis. Int. J. Biometeorol. 67, 1185–1197 (2023).
Hugall, A., Moritz, C., Moussalli, A. & Stanisic, J. Reconciling paleodistribution models and comparative phylogeography in the Wet Tropics rainforest land snail Gnarosophia bellendenkerensis (Brazier 1875). Proc. Natl. Acad. Sci. 99, 6112–6117 (2002).
Richards, C. L., Carstens, B. C. & Lacey Knowles, L. Distribution modelling and statistical phylogeography: An integrative framework for generating and testing alternative biogeographical hypotheses. J. Biogeogr. 34, 1833–1845 (2007).
Biber-Freudenberger, L., Ziemacki, J., Tonnang, H. E. Z. & Borgemeister, C. Future risks of pest species under changing climatic conditions. PLoS One 11, e0153237 (2016).
Overgaard, J., Kearney, M. R. & Hoffmann, A. A. Sensitivity to thermal extremes in Australian Drosophila implies similar impacts of climate change on the distribution of widespread and tropical species. Glob. Chang. Biol. 20, 1738–1750 (2014).
Shabani, F., Kumar, L. & Ahmadi, M. A comparison of absolute performance of different correlative and mechanistic species distribution models in an independent area. Ecol. Evol. 6, 5973–5986 (2016).
Turbelin, A. J. et al. Biological invasions as burdens to primary economic sectors. Glob. Environ. Chang. 87, 102858 (2024).
Simberloff, D. et al. Impacts of biological invasions: What’s what and the way forward. Trends Ecol. Evol. 28, 58–66 (2013).
Leathwick, J. R., Rowe, D., Richardson, J., Elith, J. & Hastie, T. Using multivariate adaptive regression splines to predict the distributions of New Zealand’s freshwater diadromous fish. Freshw. Biol. 50, 2034–2052 (2005).
Marske, K. A., Leschen, R. A. B., Barker, G. M. & Buckley, T. R. Phylogeography and ecological niche modelling implicate coastal refugia and trans-alpine dispersal of a New Zealand fungus beetle. Mol. Ecol. 18, 5126–5142 (2009).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Fu, Y. X. Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147, 915–925 (1997).
Peterson, A. T., Papeş, M. & Soberón, J. Rethinking receiver operating characteristic analysis applications in ecological niche modeling. Ecol. Model. 213, 63–72 (2008).
Jiang, W. et al. Taxonomic status and molecular phylogeography of two sibling species of Polytremis (Lepidoptera: Hesperiidae). Sci. Rep. 6, 20820 (2016).
Corrêa, A. S., Vinson, C. C., Braga, L. S., Guedes, R. N. C. & de Oliveira, L. O. Ancient origin and recent range expansion of the maize weevil Sitophilus zeamais, and its genealogical relationship to the rice weevil S. oryzae. Bull. Entomol. Res. 107, 9–20 (2017).
Liu, Y., Dietrich, C. H. & Wei, C. Genetic divergence, population differentiation and phylogeography of the cicada Subpsaltria yangi based on molecular and acoustic data: An example of the early stage of speciation?. BMC Evol. Biol. 19, 5 (2019).
Suzuki, H., Takenaka, M. & Tojo, K. Phylogeography of an insect inhabiting ‘Sky Islands’: The relationships among genetic structures and geographical characteristics, geohistorical characteristics, and cyclical climate changes. Biol. J. Linn. Soc. 141, 503–519 (2024).
Bonnet, T., Leblois, R., Rousset, F. & Crochet, P.-A. A reassessment of explanations for discordant introgressions of mitochondrial and nuclear genomes. Evolution (N Y) 71, 2140–2158 (2017).
Hinojosa, J. C. et al. A mirage of cryptic species: Genomics uncover striking mitonuclear discordance in the butterfly Thymelicus sylvestris. Mol. Ecol. 28, 3857–3868 (2019).
Stukel, M. et al. Phylogenomics improves the phylogenetic resolution and provides strong evidence of mito-nuclear discordance in two genera of a New Zealand cicada (Hemiptera: Cicadidae) species radiation. Syst. Entomol. 49, 237–257 (2024).
Rindge, F. H. A revision of the Nacophorini (Lepidoptera, Geometridae). Bull. Entomol. Res. 123, 91–153 (1961).
Oliveira, M. R. C., Corrêa, A. S., de Souza, G. A., Guedes, R. N. C. & de Oliveira, L. O. Mesoamerican origin and pre- and post-Columbian expansions of the ranges of Acanthoscelides obtectus Say, a cosmopolitan insect pest of the common bean. PLoS One 8, e70039 (2013).
Leite, N. A. et al. Pan-American similarities in genetic structures of Helicoverpa armigera and Helicoverpa zea (Lepidoptera: Noctuidae) with implications for hybridization. Environ. Entomol. 46, 1024–1034 (2017).
Moraes, T., Santos, L. M. A., Schwertner, C. F. & Corrêa, A. S. Molecular insights on the historical dispersion of Piezodorus guildinii (Hemiptera: Pentatomidae) in Brazil. J. Econ. Entomol. 116, 2173–2183 (2023).
Fernandes, D. S. et al. Molecular characterization and demographic insights into soybean bud borer (Lepidoptera: Tortricidae) in Brazil. J. Insect Sci. 24, 5 (2024).
Domingues, F. A. et al. Genetic structure and gene flow among Brazilian populations of Heliothis virescens; (Lepidoptera: Noctuidae). J. Econ. Entomol. 105, 2136–2146 (2012).
Silva-Brandao, K. L., Santos, T. V., Consoli, F. L. & Omoto, C. Genetic Diversity and structure of Brazilian populations of Diatraea saccharalis (Lepidoptera: Crambidae): Implications for pest management. J. Econ. Entomol. 108, 307–316 (2015).
Albernaz, K. C., Silva-Brandão, K. L., Fresia, P., Cônsoli, F. L. & Omoto, C. Genetic variability and demographic history of Heliothis virescens (Lepidoptera: Noctuidae) populations from Brazil inferred by mtDNA sequences. Bull. Entomol. Res. 102, 333–343 (2012).
Silva-Brandão, K. L. et al. Loci under selection and markers associated with host plant and host-related strains shape the genetic structure of Brazilian populations of Spodoptera frugiperda (Lepidoptera, Noctuidae). PLoS One 13, e0197378 (2018).
Perini, C. R. et al. Genetic structure of two Plusiinae species suggests recent expansion of Chrysodeixis includens in the American continent. Agric. For. Entomol. 23, 250–260 (2021).
Nagoshi, R. N., Meagher, R. L. & Hay-Roe, M. Inferring the annual migration patterns of fall armyworm (Lepidoptera: Noctuidae) in the United States from mitochondrial haplotypes. Ecol. Evol. 2, 1458–1467 (2012).
Dourado, P. M. et al. Host plant use of Helicoverpa spp. (Lepidoptera: Noctuidae) in the Brazilian agricultural landscape. Pest Manag. Sci. 77, 780–794 (2021).
Paula-Moraes, S. V. et al. Continental-scale migration patterns and origin of Helicoverpa zea (Lepidoptera: Noctuidae) based on a biogeochemical marker. Environ. Entomol. 53, 487–497 (2024).
Peterson, A. T. et al. Ecological niches and geographic distributions (MPB-49) (Princeton University Press, 2011). https://doi.org/10.23943/princeton/9780691136868.001.0001.
Pantoja-Gomez, L. M., Corrêa, A. S., de Oliveira, L. O. & Guedes, R. N. C. Common origin of Brazilian and Colombian populations of the neotropical coffee leaf miner, Leucoptera coffeella (Lepidoptera: Lyonetiidae). J. Econ. Entomol. 112, 924–931 (2019).
Silva, C. S. et al. Population expansion and genomic adaptation to agricultural environments of the soybean looper, Chrysodeixis includens. Evol. Appl. https://doi.org/10.1111/eva.12966 (2020).
Aidoo, O. F. et al. Model-based prediction of the potential geographical distribution of the invasive coconut mite, Aceria guerreronis Keifer (Acari: Eriophyidae) based on MaxEnt. Agric. For. Entomol. 24, 390–404 (2022).
Gardner, A. S., Maclean, I. M. D. & Gaston, K. J. Climatic predictors of species distributions neglect biophysiologically meaningful variables. Divers. Distrib. 25, 1318–1333 (2019).
Corrêa, A. S., Cordeiro, E. M. G. G. & Omoto, C. Agricultural insect hybridization and implications for pest management. Pest Manag. Sci. 75, 2857–2864 (2019).
Arce-Valdés, L. R. & Sánchez-Guillén, R. A. The evolutionary outcomes of climate-change-induced hybridization in insect populations. Curr. Opin. Insect Sci. 54, 100966 (2022).
Doyle, J. J. & Doyle, J. L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 19, 11–15 (1987).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 3, 294–299 (1994).
Caterino, M. S. & Sperling, F. A. H. Papilio phylogeny based on mitochondrial cytochrome oxidase I and II genes. Mol. Phylogenet. Evol. 11, 122–137 (1999).
Hall, T. BioEdit: An important software for molecular biology. GERF Bull. Biosci. 2, 60–61 (2011).
Lopez, J. V., Yuhki, N., Masuda, R., Modi, W. & O’Brien, S. J. Numt, a recent transfer and tandem amplification of mitochondrial DNA to the nuclear genome of the domestic cat. J. Mol. Evol. 39, 174–190 (1994).
Tamura, K. et al. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Rozas, J. et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 34, 3299–3302 (2017).
Huelsenbeck, J. P. & Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, 754–755 (2001).
Leigh, J. W. & Bryant, D. POPART: Full-feature software for haplotype network construction. Methods Ecol. Evol. 6, 1110–1116 (2015).
Nylander, J. A. A. MrModeltest v2 Program distributed by the author. Preprint at (2004).
Akaike, H. A new look at the statistical model identification. IEEE Trans. Autom. Control 19, 716–723 (1974).
Excoffier, L., Laval, G. & Schneider, S. Arlequin ver. 3.0: An integrated software package for population genetics data analysis. Evol. Bioinform. 1, 47–50 (2005).
Slatkin, M. A measure of population subdivision based on microsatellite allele frequencies. Genetics 139, 457–462 (1995).
Mantel, N. The detection of disease clustering and a generalized regression approach. Cancer Res. 27, 209–220 (1967).
Rogers, A. R. & Harpending, H. Population growth makes waves in the distribution of pairwise genetic differences. Mol. Biol. Evol. 9, 552–569 (1992).
Papadopoulou, A., Anastasiou, I. & Vogler, A. P. Revisiting the insect mitochondrial molecular clock: The mid-aegean trench calibration. Mol. Biol. Evol. 27, 1659–1672 (2010).
Borer, M. et al. The phylogeography of an alpine leaf beetle: Divergence within Oreina elongata spans several ice ages. Mol. Phylogenet. Evol. 57, 703–709 (2010).
Pereira, F. et al. Thermal requirements and estimate number of generations of Palmistichus elaeisis (Hymenoptera: Eulophidae) in different Eucalyptus plantations regions. Braz. J. Biol. 71, 431–436 (2011).
GBIF.org. GBIF Occurrence. Download https://doi.org/10.15468/dl.ca6pz5 (2024).
Aiello-Lammens, M. E., Boria, R. A., Radosavljevic, A., Vilela, B. & Anderson, R. P. spThin: An R package for spatial thinning of species occurrence records for use in ecological niche models. Ecography 38, 541–545 (2015).
Ramos, R. S. et al. Climate model for seasonal variation in Bemisia tabaci using CLIMEX in tomato crops. Int. J. Biometeorol. 63, 281–291 (2019).
Zhang, S. et al. AHP-GIS and MaxEnt for delineation of potential distribution of Arabica coffee plantation under future climate in Yunnan China. Ecol. Indic. 132, 108339 (2021).
Riahi, K. et al. The Shared Socioeconomic Pathways and their energy, land use, and greenhouse gas emissions implications: An overview. Glob. Environ. Chang. 42, 153–168 (2017).
Tatebe, H. et al. Description and basic evaluation of simulated mean state, internal variability, and climate sensitivity in MIROC6. Geosci. Model. Dev. 12, 2727–2765 (2019).
Steven, J. P., Miroslav, D. & Robert, E. S. [Internet] Maxent software for modeling species niches and distributions (Version 3.4.4). Available from http://biodiversityinformatics.amnh.org/open_source/maxent/ (2021).
Brown, J. L., Bennett, J. R. & French, C. M. SDMtoolbox 2.0: The next generation Python-based GIS toolkit for landscape genetic, biogeographic and species distribution model analyses. PeerJ 5, e4095 (2017).
Jarnevich, C. S., Holcombe, T. R., Thomas, C. C., Frid, L. & Olsson, A. Simulating long-term effectiveness and efficiency of management scenarios for an invasive grass. AIMS Environ. Sci. 2, 427–447 (2015).
Merow, C., Smith, M. J. & Silander, J. A. A practical guide to MaxEnt for modeling species’ distributions: What it does, and why inputs and settings matter. Ecography 36, 1058–1069 (2013).
Elith, J. et al. A statistical explanation of MaxEnt for ecologists. Divers. Distrib. 17, 43–57 (2011).
Zhang, K., Yao, L., Meng, J. & Tao, J. Maxent modeling for predicting the potential geographical distribution of two peony species under climate change. Sci. Total Environ. 634, 1326–1334 (2018).
Ansari, M. & Ghoddousi, A. Water availability limits brown bear distribution at the southern edge of its global range. Ursus 29, 13–24 (2018).
Mahatara, D. et al. Maxent modelling for habitat suitability of vulnerable tree Dalbergia latifolia; in Nepal. Silva Fenn. 55, 10441 (2021).
Liu, C., Berry, P. M., Dawson, T. P. & Pearson, R. G. Selecting thresholds of occurrence in the prediction of species distributions. Ecography 28, 385–393 (2005).
Ramos, R. S., Kumar, L., Shabani, F. & Picanço, M. C. Risk of spread of tomato yellow leaf curl virus (TYLCV) in tomato crops under various climate change scenarios. Agric. Syst. 173, 524–535 (2019).
Acknowledgements
We thank the colleagues Talita A. Menezes, Deivide Pereira, and Sidinei Dallacor for helping collect the insects.
Funding
Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (Finance Code 001), and Conselho Nacional de Desenvolvimento Científico e Tecnológico (310376/2021-1). This research was partially financed by Suzano S.A.
Author information
Authors and Affiliations
Contributions
F.N., E.P.S., E.A.V.Z., and A.S.C. conceived and designed the study. F.N., P.G.C.S., E.P.S., M.M.D., F.A.S., C.F.W., and R.S.S. collected the data. C.F.W., E.P.S., E.A.V.Z., and A.S.C. provided reagents and analytical tools. F.N., P.G.C.S., R.S.S., and A.S.C. analyzed the data. F.N., P.G.C.S., R.S.S., and A.S.C. wrote the manuscript. All authors read, corrected, made suggestions, and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
FN, PGCS, FAS, CFW, RSS, and ASC declare no competing interests. EPS, EAVZ, and MMD are employed by Suzano S.A and they declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nanini, F., Souza, P.G.C., Soliman, E.P. et al. Genetic diversity, population structure and ecological niche modeling of Thyrinteina arnobia (Lepidoptera: Geometridae), a native Eucalyptus pest in Brazil. Sci Rep 14, 20963 (2024). https://doi.org/10.1038/s41598-024-71816-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-71816-2
- Springer Nature Limited