Abstract
SARS-CoV-2 poses an unprecedented threat to the world as the causative agent of the COVID-19 pandemic. Among a handful of therapeutics developed for the prevention and treatment of SARS-CoV-2 infection, ensitrelvir is the first noncovalent and nonpeptide oral inhibitor targeting the main protease (Mpro) of SARS-CoV-2, which recently received emergency regulatory approval in Japan. Here we determined a 1.8-Å structure of Mpro in complex with ensitrelvir, which revealed that ensitrelvir targets the substrate-binding pocket of Mpro, specifically recognizing its S1, S2, and S1' subsites. Further, our comprehensive biochemical and structural data have demonstrated that even though ensitrelvir and nirmatrelvir (an FDA-approved drug) belong to different types of Mpro inhibitors, both of them remain to be effective against Mpros from all five SARS-CoV-2 variants of concern, suggesting Mpro is a bona fide broad-spectrum target. The molecular mechanisms uncovered in this study provide basis for future inhibitor design.
Similar content being viewed by others
Introduction
SARS-CoV-2 poses an unprecedented threat to the world as the causative agent of the COVID-19 pandemic. Currently, a handful of therapeutics have been approved for the prevention and treatment of SARS-CoV-2 infection, including vaccinations, monoclonal antibodies, and compounds that target key enzymes with crucial roles in the viral life cycle1,2. However, the long-lasting pandemic plus the error-prone nature of the RNA viral genome enables SARS-CoV-2 to accumulate a variety of mutations, giving rise to various mutant strains that could potentially impair the efficacy of existing therapies3,4. The World Health Organization (WHO) named COVID-19 variants with increased transmissibility or harmful changes in epidemiology variants of concern (VOCs). By March 2023, the WHO had identified five VOCs, including Alpha (B.1.1.7), Beta (B.1.351), Gamma (P.1), Delta (B.1.617.2), and Omicron (B.1.1.529). Since May 2021, these variants have become the dominant variants in more than 90 countries5,6.
Mpro, also named 3-chymotrypsin-like protease (3CLpro), is a viral-encoded cysteine protease that plays a fundamental role in viral replication. After invading the host cell, the positive-sense single-stranded viral RNA genome will be treated as mRNA by the host ribosome and generate two long polyproteins named pp1a and pp1ab, which will be further proteolytically processed and give rise to the various non-structural proteins (nsps) required for the subsequent viral life cycle7. During the proteolytic process of pp1a and pp1ab, Mpro is indispensable and cleaves polyproteins at no less than 11 conserved sites. Owing to the key role of the protein in viral replication, two oral antiviral drugs targeting Mpro received emergency use authorization (EUA): nirmatrelvir and ensitrelvir. Nirmatrelvir is the main ingredient of Paxlovid, developed by Pfizer, which has been officially approved by the FDA8. Paxlovid showed a greater reduction in the risk of hospitalization and death than a placebo. Ensitrelvir is the first oral noncovalent, nonpeptide inhibitor developed by Shionogi9. Unlike nirmatrelvir, ensitrelvir does not need pharmaceutical boosters such as ritonavir and can be directly used to treat patients with mild COVID-19. Ensitrelvir received emergency authorization in Japan after treatment showed rapid SARS-CoV-2 clearance in a phase 2/3 clinical trial and was well tolerated10. Last month, ensitrelvir just received Phase III clinical approval in the United States. It is critical to understand its appropriate molecular mechanism of inhibition. Besides, SARS-CoV-2 VOCs carry mutations at varying frequencies in the Mpro that are specific to Alpha, Beta, Gamma, Lambda, and Omicron. where Alpha, Beta, and Gamma have the substitution K90R, Lambda has the substitution G15S, and Omicron has the substitution P132H11. It remains unclear whether these changes in Mpro across different SARS-CoV-2 variants will affect the architecture of the reaction pocket and thus affect the inhibition of current compounds targeting Mpro.
In this study, we combined the enzymatic activity assay and crystallography to study the structure-activity relationship (SAR) between SARS-CoV-2 Mpro and the inhibitors, respectively. We obtained a total of 10 crystal structures of Mpro, including the apo-forms of the variants and Mpro/variant-inhibitor complexes. Our results presented here will facilitate future antiviral design.
Results
Ensitrelvir is a potent inhibitor against WT SARS-CoV-2 Mpro
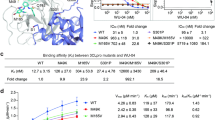
Based on the fluorescence-resonance-energy transfer (FRET)-based assay we previously established, the inhibition of ensitrelvir on SARS-CoV-2 Mpro was performed to examine the enzyme kinetics of WT. The half-maximal inhibitory concentrations (IC50) value of ensitrelvir against SARS-CoV-2 Mpro is 0.049 μM (Fig. 1a), while the IC50 of nirmatrelvir against WT is 0.044 μM, indicating that ensitrelvir exhibited as potent inhibition on viral protease as nirmatrelvir did in vitro.
a The IC50 of ensitrelvir and nirmatrelvir against WT SARS-CoV-2 Mpro. Data points are shown as the mean ± SD (n = 3). b The crystal structure of dimeric Mpro complex with ensitrelvir. c The interaction network of ensitrelvir in the substrate-binding pocket of Mpro. Ensitrelvir is shown in green, Mpro is shown in silver, and the residues of the neighboring protomer are labeled in yellow. Blue dotted lines represent hydrogen bonds, red dashed lines represent π-π stack, and red spheres represent water molecules. The substrate pocket is indicated separately. d Topology of the hydrogen-bond network between ensitrelvir and Mpro. Hydrogen bonds are indicated by green dashed lines, and dashed boxes indicate interaction with the stable S1 pocket. e Band diagram of the WT and ensitrelvir complex crystal structure superimposed on WT bond nirmatrelvir. f Analysis of the residues involved in the interactions between ensitrelvir and nirmatrelvir in the substrate-binding pocket. Ensitrelvir is highlighted in green, while nirmatrelvir is highlighted in yellow.
The crystal structure of SARS-CoV-2 Mpro in complex with ensitrelvir
In order to elucidate the molecular inhibition mechanism of ensitrelvir, we determined a 1.8-Å Mpro-ensitrelvir complex structure (PDB: 8HBK). In the crystals, there is merely one protomer in an asymmetric unit, and all 1–301 residues could be traced in the electron density map. Two protomers form a functional dimer by a crystallographic twofold axis of symmetry. The overall protomer of Mpro comprises three domains (Fig. 1b). Domain I (residues 8–101) and domain II (residues 102–184) possess an antiparallel β-barrel structure and take on a chymotrypsin-like fold, harboring the catalytic pocket between them. Domain III (residues 201–303) is a globular domain composed of five antiparallel α-helixes unique to Mpro and is crucial for the dimerization of Mpro, which is a prerequisite for its catalytic activity. The catalytic pocket containing the Cys-His dyad is located in the cleft between domain I and domain II (Fig. 1b). An ensitrelvir molecule could be identified in the substrate-binding pocket of each protomer, occupying S1, S2, and S1' subsites of Mpro. In the crystal structure of dimeric Mpro, the N-terminus of one promoter deeply inserts into the S1 subsite of the neighboring promoter and participates in the stabilization of the substrate pocket (Fig. 1c). In the S1 subsite, the Ser1 from the neighboring protomer stabilizes the pocket by forming four hydrogen bonds with E166 and F140. The 1-methyl-1H-1,2,4-triazole group forms a hydrogen bond with the sidechain imidazole group of H163. In the S2 subsite, the 2,4,5-trifluoromethyl forms a π-π stack with the sidechain of H41. In the S1' subsite, the 6-chloro-2-methyl-2H-indazole part interacts with the NH of the mainchain of T26 through a hydrogen bond. In addition, H163, C145, G143, and Q189 are also involved in the hydrogen-bond network to stabilize the binding of ensitrelvir (Fig. 1d). Nirmatrelvir forms a covalent bond with C145 of Mpro in the S1' subsite, whereas ensitrelvir forms no covalent bond in the substrate pocket. Instead, it interacts through hydrogen bonding with C145 and T26 in the S1' pocket. Furthermore, nirmatrelvir stabilizes in the S4 pocket through extensive hydrophobic interactions, whereas ensitrelvir has few interactions in this pocket. The interaction network in the S1 subsite is highly conserved for both inhibitors, as they both stabilize through hydrogen bonds with E166, F140, Ser1, and H172 in the S1 pocket (Fig. 1e, f).
The G15S, K90R, and P132H substitutions have a limited impact on the structural and enzymatic properties of Mpro
To elaborate on the impact of the G15S, K90R, and P132H mutations on the structure of Mpro, the apo-form structures of G15S, K90R, and P132H were determined at 1.77, 1.66, and 1.82 Å resolution, respectively (Supplementary Table S1). There is only one protomer in an asymmetric unit in each crystal structure, and all 1–301 residues could be traced in the electron density map.
The mutant residues 15 and 90 are located in domain I and residue 132 is located in domain II. All these mutations are far away from the catalytic pocket (Fig. 2a and Supplementary Fig. S3); thus, it is reasonable to speculate that the G15S, K90R, and P132H mutations do not directly affect the architecture of the catalytic pocket. This was further confirmed when comparing the G15S, K90R, and P132H structures with the Mpro apo-form structure (PDB ID: 6Y2E). As shown in Fig. 2a, G15S, K90R, and P132H showed merely a slight difference from WT, with average root-mean-square deviation (RMSD) values of only 0.54 Å, 0.52 Å, and 0.58 Å, respectively, indicating that all these three mutations had a limited impact on the apo structure of Mpro. It is worth mentioning that the mutation of the amino acid at position 15 from glycine to serine did not cause any serious structural changes to the backbone, except that the substitute of serine formed an additional hydrogen bond with a surrounding water molecule (Fig. 2b). The mutation of the amino acid at position 90 from lysine to arginine results in the breaking of a hydrogen bond from lysine to the water molecule but does not significantly alter the conformations of surrounding residues (Fig. 2c). Similarly, the P132H substitution did not change the architecture of the mainchain at this site or the residues in the vicinity except that the sidechain of E240 was slightly “pushed” away from residue 132. It is presumably caused by the steric hindrance of the histidine, and it is noticed that the sidechain of H132 formed an extra hydrogen bond with a water molecule (Fig. 2d).
a Superposition of crystal structures of WT, G15S, K90R, and P132H. The color spectrum indicates the RMSD of the Cα atoms from the aligned structures. The position of residue 15 is shown as a yellow circle, 90 is shown as a red circle, and 132 is shown as a green circle. b–d Comparison of the region in the vicinity of residues 15, 90, and 132 between WT and G15S, K90R, and P132H, respectively. WT is shown in marine blue, and G15S, K90R, and P132H are shown in salmon. The 2Fo-Fc density map is shown as a black grid contour at 1.0σ. e Enumerated enzyme activity data of G15S, K90R, P132H, and WT calculated by the Michaelis equation, including Km, kcat, and kcat/Km. f Characterization of enzymatic activity shows that G15S, K90R, and P132H display very similar activity to WT. Data points are shown as the mean ± SD (n = 3).
To further validate whether these slight structural variations affect enzymatic efficiency, a previously established fluorescence-resonance-energy transfer (FRET)-based assay was used to compare the enzyme kinetic parameters of WT, G15S, K90R, and P132H. As shown in Fig. 2e, f, the catalytic efficiency (kcat/Km) values were determined to be 2.66 × 104 M−1s−1 for G15S, 3.05 × 104 M−1s−1 for K90R, 2.64 × 104 M−1s−1 for P132H, respectively, which are comparable to 2.71 × 104 M−1s−1 for WT. It is indicated that the G15S, K90R, P132H, and WT have similar enzymatic kinetic parameters and the impact of the three substitutions on Mpro could be neglected.
Ensitrelvir remains strong inhibition on SARS-CoV-2 Mpro variants
Next, we tested the half-maximal inhibitory concentrations (IC50) of ensitrelvir against SARS-CoV-2 Mpro and variants. The results showed that ensitrelvir exhibited similar inhibition on WT and its variants with an IC50 of approximately 0.04 μM (Fig. 3a). To obtain the structural basis for the inhibition of SARS-CoV-2 Mpro and variants of the ensitrelvir, we eventually determined the structures of three variants individually with ensitrelvir for a total of three complex structures (Supplementary Table S1). Superimposition of the structures of all three complexes has shown that all three variant mutant sites (G15, K90, P132) are more than 20 Å away from the binding sites of ensitrelvir (Fig. 2a and Supplementary Fig. S3), indicating that these changes in Mpro between different SARS-CoV-2 variants may not affect the architecture of the substrate-binding pocket and thus would not impair the efficacy of the current compounds targeting Mpro. Among the structures of mutant complexes of ensitrelvir, H163, E166, C145, G143, and T26 are involved in hydrogen bonding (Fig. 3b), and certain residues in the substrate-binding pocket participate in stabilizing ensitrelvir, which has demonstrated a conservative binding mode among Mpro from the WT and other variants. To summarize, all the data obtained above strongly support that Mpro from SARS-CoV-2 WT and its variants have similar structural features and kinetic characters, and the G15S, K90R, and P132H substitutions do not impair the inhibition of ensitrelvir in vitro.
a, c The IC50 of ensitrelvir and nirmatrelvir against Mpro and variants, respectively. Data points are shown as the mean ± SD (n = 3). b, d Superimposed crystal structures of WT Mpro and variants complexed with ensitrelvir and nirmatrelvir, respectively. The key residues in the pocket are shown in sticks.
Nirmatrelvir is also a potent inhibitor against SARS-CoV-2 Mpro variants
Nirmatrelvir is also an inhibitor targeting SARS-CoV-2 Mpro, which has just been fully approved to treat mild to moderate COVID-19 in adults at risk of severe infections. Next, we tested IC50 values of nirmatrelvir against SARS-CoV-2 Mpro and variants to see whether these mutations affect nirmatrelvir binding. The results showed that IC50 of nirmatrelvir against WT was 0.044 μM while IC50 of ensitrelvir against WT was 0.049 μM (Fig. 3c), suggesting that nirmatrelvir strongly inhibits WT Mpro like ensitrelvir. In addition, the IC50 values of nirmatrelvir against all variants are approximately 0.04 μM, indicating that these mutations did not obviously affect nirmatrelvir binding.
To obtain the structural basis for the inhibition of SARS-CoV-2 Mpro and variants of nirmatrelvir, we eventually determined the structures of three variants individually with nirmatrelvir for a total of three complex structures (Supplementary Table S1). Superimposition of the structures of all three complexes has shown all three variant mutant sites (G15, K90, P132) were more than 20 Å away from the binding site of nirmatrelvir (Fig. 2a and Supplementary Fig. S3), indicating that these changes in Mpro would not impair the in vitro efficacy of nirmatrelvir. All three mutant Mpro structures show little difference from that of WT, with an average RMSD of only 0.25–0.35 Å, and neither the binding pose of the compound nor the conformation of residues participating in drug binding exhibits significant differences. As shown in our previous analysis of the complex structure of WT with nirmatrelvir (PDB ID: 7VH8)12, the nitrile group of nirmatrelvir is attached to the Sγ atom of C145 through a standard 1.8 Å C-S covalent bond, the classical (S)-γ-lactam ring at the P1 position fits into the S1' subsite, and a hydrogen bond is formed between the oxygen atom of the lactam ring and the Nε2 atom of H163. In addition, the Oε1 atom of E166 interacts with the NH group to stabilize nirmatrelvir. The rigid dimethylcyclopropyl proline (DMCP) located at the S2 subsite is surrounded by extensive hydrophobic interactions. Most of the amino acids used to stabilize nirmatrelvir near the substrate pocket described above show great similarity in the complex structures of Mpro (Fig. 3d), which may imply that nirmatrelvir may exhibit similar inhibition for WT Mpro and other variants, which is consistent with the in vitro enzyme activity inhibitory of nirmatrelvir (Fig. 3a, c).
Discussion
In the 21st century, three previously unknown coronaviruses have spread globally, including severe acute respiratory syndrome (SARS) caused by SARS-CoV in 2003, Middle East respiratory syndrome (MERS) caused by MERS-CoV in 2012, and the current COVID-19 caused by SARS-CoV-2. During the first two rounds of the coronavirus outbreak, no approved targeted therapies, vaccines, or compounds were available for treatment13. However, for the COVID-19 pandemic, it is the first time newly developed numerous approved vaccines, targeted compounds, etc., that we can use to battle against the coronavirus14,15. Unfortunately, due to the long-term prevalence of the virus in the population worldwide and the error-prone nature of RNA viruses, an increasing number of SARS-CoV-2 variants have been reported worldwide, raising concerns about the effect of current vaccines and compounds.
Mpro has received much attention in the last three years as an ideal target for drugs against SARS-CoV-2. As the first noncovalent and non-peptidomimetic drug candidate16, ensitrelvir has just received Phase 3 clinical approval in the United States. It is critical to understand its molecular mechanism of inhibition. In this work, a 1.8-Å Mpro-ensitrelvir complex structure (PDB: 8HBK) was determined to elucidate the precise molecular inhibition mechanism of ensitrelvir. We found that ensitrelvir mainly recognizes the S1, S2, and S1' subsites of Mpro and relies on the stability of the hydrogen bonding network in the substrate-binding pocket, unlike the covalent inhibition mechanism of nirmatrelvir. In addition, the functional Mpro exists in a homo-dimer form in the physiological state, and the N-terminus residues 1–7 penetrate deeply into the substrate-binding pocket of the neighboring promoter and the N-terminus serine residue stabilizes the S1 subsite by hydrogen bonding, contributing to the stability of the substrate pocket and ensitrelvir binding (Fig. 2c, d). This differs from the previously reported model (PDB ID:7VU6)9, in which the N-terminal of Mpro lacks two residues (Ser1 and Gly2), while these two N-terminal residues can be traced in our complex structure (PDB ID:8HBK) based on the clear electron density (Supplementary Fig. S1). Moreover, the electron density of the Serl and Gly2 at the N-terminus is also clearly visible in all the complex structures of nirmatrelvir we solved (Supplementary Fig. S2).
The disruption of the N-terminus of Mpro may result from their construction of in vitro preparations of the protein. In previous research, we identified that the N-terminus of Mpro of SARS had excess amino acids, which will significantly impact on in vitro enzyme activity. The enzyme activity of Mpro with two extra residues at the N-terminus (GS-Mpro) decreased by about 24 times compared to the clean N-terminus Mpro 17. The additional amino acids (Gly-2 and Ser-1) may have led to a difference in Mpro activity compared to the physiological state, with the protein being less active in vitro. This may explain why ensitrelvir exhibits stronger inhibition of enzyme activity than nirmatrelvir in vitro in that work, while our results show that the enzyme activity inhibition of ensitrelvir is similar to that of nirmatrelvir. Furthermore, there is a big gap in binding affinity between the two types of proteins, and the Kd of GS-Mpro was 418 and 8.43 nM compared to the clean N-terminus Mpro (Supplementary Fig. S4). Under physiological conditions, residues in the N-terminus of Mpro penetrate deep into the substrate pocket of the neighboring protomer and are involved in stabilizing the S1 subsite18. The extra residues may spatially block ensitrelvir from entering the substrate pocket.
Overall, the apo-form structures and in vitro enzyme activity assays of several variants suggest that minor changes in Mpro have little effect on the overall structure, particularly the substrate pocket and the active center, which is consistent with our earlier findings. In addition, mutations in variants do not alter the properties of Mpro, so they may not impair the in vitro enzymatic inhibition of nirmatrelvir and ensitrelvir, which further suggests that Mpro is an ideal drug target because it is one of the least variable viral components. Finally, we determined the in vitro inhibitory effects of ensitrelvir and nirmatrelvir on Mpro enzymatic activities, and the results showed that ensitrelvir and nirmatrelvir exhibit consistent in vitro enzymatic inhibition against SARS-CoV-2 Mpro and its variants, which is consistent with their excellent clinical efficacy. Although these two inhibitors belong to different types, both of them remain to be effective against Mpros from all 5 SARS-CoV-2 variants of concern, suggesting Mpro is a bona fide broad-spectrum target.
Recently, a 2.2 Å resolution complex structure of Mpro-ensitrelvir was reported19. In their functional dimer, one protomer has clear electron density for its N-terminus, but its neighboring protomer does not. In the 1.8 Å resolution structure in this work, clear electron density can be observed for Mpro N-terminus, and we provide an accurate model for elucidating the ensitrelvir binding mode. The molecular mechanisms uncovered in this study provide the basis for future inhibitor design.
Methods
Cloning, protein expression, and purification of SARS-CoV-2 variant Mpro
The expression plasmid used to produce full-length Mpro was obtained by using site-directed mutagenesis to introduce G15S, K90R, and P132H substitutions into the expression plasmid of full-length WT Mpro. The expression plasmid was transformed into E. coli BL21 (DE3) cells and then cultured in a 2-L shaking flask with 1 L Luria broth medium containing 0.1 g/L ampicillin at 37 °C. When the optical density at 600 nm of the bacteria reaches 0.6–0.8, a final concentration of IPTG was added to the culture to induce protein expression at 16 °C. After 10 h, the bacteria were pelleted by centrifugation at 3000×g for 15 min. Then the bacteria were resuspended in lysis buffer (50 mM HEPES, pH 8.0, 300 mM NaCl), and the supernatant was incubated with Ni-NTA agarose gravity column (GE) after centrifuging at 18,000 rpm for 30 min. The fusion proteins were washed with wash buffer (50 mM HEPES, pH 8.0, 300 mM NaCl, 20 mM imidazole). After washing 10–20 column volume, the PreScission protease was added to remove the His-tag. Then the samples of the protein were loaded onto a Hitrap Q HP column and then purified by size-exclusion chromatography with a Superdex 75 increase column with storage buffer (50 mM HEPES, pH 7.5, 150 mM NaCl, and 4 mM DTT). The fractions were concentrated to 10 mg/mL for the next test20.
Crystallization, data collection, and structure determination
The crystals of Mpro apo forms were directly screened by crystallization kits, and the crystals of the complexes with inhibitors were screened through co-crystallization of the variants Mpro at a concentration of 5 mg/ml with nirmatrelvir or ensitrelvir at 0.5 mM. The crystals were obtained at 18 °C by the sitting method. Crystals were cryoprotected using the reservoir solution with 20% ethylene glycol and flash-frozen in liquid nitrogen. Diffraction data were collected at 100 K at a wavelength of 0.9785 Å. The structures were solved by molecular replacement with the program CCP4 using the complex structure of WT Mpro and N3 (PDB ID: 7VH8) as a search model12. The model was refined using PHENIX21 and manually constructed using Coot22.
IC50 measurement
The method of IC50 measurement has been previously demonstrated20. In brief, the fluorescent substrate was applied to measure the hydrolytic activity of Mpro. Mpro (0.1 μM) was incubated with different concentrations of nirmatrelvir or ensitrelvir for 90 s before 10 μM substrate was added. Fluorescence intensity was monitored by an EnVision multimode plate reader (Perkin Elmer) using wavelengths of 320 nm for excitation and 405 nm for emission. The changes in initial rates when adding different concentrations of nirmatrelvir or ensitrelvir were calculated to evaluate the inhibitory effect. The dose-response curve for IC50 values was determined by nonlinear regression using GraphPad Prism.
ITC assay
The ITC experiment was performed on a MicroCal PEAQ-ITC. After cleaning the sample cell and needle, the OMpro (20 μM) protein was carefully injected into the sample cell using a micro-syringe without any bubbles, and ensitrelvir (200 μM) was filled into a 40 μL titration syringe. OMpro and ensitrelvir were diluted with TB (1% DMSO, pH = 8.0), and deionized water was injected into the reference cell as a heat balance control. After 35 times titrations of ensitrelvir into the sample cell at a constant rate of 150 s, the One Set of Sites is selected as the fitting model.
Statistics and reproducibility
Statistical analysis was carried out using Prism software. The amount of enzyme was determined by the initial experiment, and the enzyme kinetic and half-inhibition tests consisted of three replicate trials.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All experimental data are provided in the manuscript. The structures determined in this study have been deposited to the Protein Data Bank (PDB) under accession codes: 8HBK (WT-Ensitrelvir), 8HOM (P132H-Ensitrelvir), 8INX (G15S-Ensitrelvir), 8INY (K90R-Ensitrelvir), 8HOZ (P132H-Nirmatrelvir), 8INU (G15S-Nirmatrelvir), 8INW (K90R-Nirmatrelvir), 8HOL (P132H), 8INQ (G15S), 8INT (K90R). The structures of the Mpro-nirmatrelvir complex and Mpro-Ensitrelvir complex were downloaded from PDB under accession codes 7VH8 and 7VU6, respectively. Source data for figures can be found in Supplementary Data 1.
Materials availability
Materials used in this study will be made available under an appropriate Materials Transfer Agreement.
Code availability
No code was used for this study.
References
He, X., He, C., Hong, W., Yang, J. & Wei, X. Research progress in spike mutations of SARS-CoV-2 variants and vaccine development. Med. Res. Rev. 43, 932–971 (2023).
Huang, C. et al. A new generation M(pro) inhibitor with potent activity against SARS-CoV-2 Omicron variants. Signal Transduct. Target. Ther. 8, 128 (2023).
Davies, N. G. et al. Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England. Science 372, eabg3055 (2021).
Faria, N. R. et al. Genomics and epidemiology of the P.1 SARS-CoV-2 lineage in Manaus, Brazil. Science 372, 815–821 (2021).
Al Kaabi, N. et al. Effect of 2 inactivated SARS-CoV-2 vaccines on symptomatic COVID-19 infection in adults: a randomized clinical trial. JAMA 326, 35–45 (2021).
Walensky, R. P., Walke, H. T. & Fauci, A. S. SARS-CoV-2 variants of concern in the United States: challenges and opportunities. JAMA 325, 1037–1038 (2021).
Mondal, S. et al. Dual inhibitors of main protease (M(Pro)) and cathepsin L as potent antivirals against SARS-CoV2. J. Am. Chem. Soc. 144, 21035–21045 (2022).
Owen, D. R. et al. An oral SARS-CoV-2 M(pro) inhibitor clinical candidate for the treatment of COVID-19. Science 374, 1586–1593 (2021).
Unoh, Y. et al. Discovery of S-217622, a noncovalent oral SARS-CoV-2 3CL protease inhibitor clinical candidate for treating COVID-19. J. Med. Chem. 65, 6499–6512 (2022).
Mukae, H. et al. A randomized phase 2/3 study of ensitrelvir, a novel oral SARS-CoV-2 3C-like protease inhibitor, in Japanese patients with mild-to-moderate COVID-19 or asymptomatic SARS-CoV-2 infection: results of the phase 2a part. Antimicrob. Agents Chemother. 66, e0069722 (2022).
Greasley, S. E. et al. Structural basis for the in vitro efficacy of nirmatrelvir against SARS-CoV-2 variants. J. Biol. Chem. 298, 101972 (2022).
Zhao, Y. et al. Crystal structure of SARS-CoV-2 main protease in complex with protease inhibitor PF-07321332. Protein Cell 13, 689–693 (2022).
de Wit, E., van Doremalen, N., Falzarano, D. & Munster, V. J. SARS and MERS: recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 14, 523–534 (2016).
Wang, M. Y. et al. SARS-CoV-2: structure, biology, and structure-based therapeutics development. Front. Cell. Infect. Microbiol. 10, 587269 (2020).
Yang, H. & Rao, Z. Structural biology of SARS-CoV-2 and implications for therapeutic development. Nat. Rev. Microbiol. 19, 685–700 (2021).
Lamb, Y. N. Nirmatrelvir plus ritonavir: first approval. Drugs 82, 585–591 (2022).
Xue, X. et al. Production of authentic SARS-CoV M(pro) with enhanced activity: application as a novel tag-cleavage endopeptidase for protein overproduction. J. Mol. Biol. 366, 965–975 (2007).
Noske, G. D. et al. An in-solution snapshot of SARS-COV-2 main protease maturation process and inhibition. Nat. Commun. 14, 1545 (2023).
Noske, G. D. et al. Structural basis of nirmatrelvir and ensitrelvir activity against naturally occurring polymorphisms of the SARS-CoV-2 main protease. J. Biol. Chem. 299, 103004 (2023).
Jin, Z. et al. Structure of M(pro) from SARS-CoV-2 and discovery of its inhibitors. Nature 582, 289–293 (2020).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Acknowledgements
This work was supported by grants from the R&D Program of Guangzhou Laboratory (grant No. SRPG22-003 and SRPG22-011); Science and Technology Commission of Shanghai Municipality (grant No. YDZX20213100001556 and 20XD1422900 to H.Y.); Shanghai Municipal Science and Technology Major Project (grant No. ZD2021CY001); National Natural Science Foundation of China (grant No. 92169109 to H.Y. and grant No. 32171259 to X.L.) This work was supported by Shanghai Frontiers Science Center for Biomacromolecules and Precision Medicine, ShanghaiTech University. We would like to acknowledge the staff members from beamlines BL02U1, BL10U2, BL18U1, and BL19U1 at the Shanghai Synchrotron Radiation Facility and I04 beamline of Diamond Light Source for the beam time and operation at the facility.
Author information
Authors and Affiliations
Contributions
X.L. conceived this project. M.L., X.Z., and Z.Y. cloned, expressed, purified, and crystallized proteins. M.L. and X.Z. collected the diffraction data. X.L. and M.L. solved the crystal structures. M.L., X.Z., and Z.Y. performed enzymatic activity and inhibition assays. X.L., X.Y., H.Y., Y.M., and M.L. contributed to discussions of the data and analysis. M.L., Y.D., H.Y., and X.L. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
This manuscript has been previously reviewed in another Nature Portfolio journal. The manuscript was considered suitable for publication without further review at Communications Biology. Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary handling editor: Gene Chong.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lin, M., Zeng, X., Duan, Y. et al. Molecular mechanism of ensitrelvir inhibiting SARS-CoV-2 main protease and its variants. Commun Biol 6, 694 (2023). https://doi.org/10.1038/s42003-023-05071-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-05071-y
- Springer Nature Limited
This article is cited by
-
Computational evaluation and benchmark study of 342 crystallographic holo-structures of SARS-CoV-2 Mpro enzyme
Scientific Reports (2024)
-
Study of key residues in MERS-CoV and SARS-CoV-2 main proteases for resistance against clinically applied inhibitors nirmatrelvir and ensitrelvir
npj Viruses (2024)
-
SARS-CoV-2 Resistance to Small Molecule Inhibitors
Current Clinical Microbiology Reports (2024)
-
Ensitrelvir Fumaric Acid: First Approval
Drugs (2024)