Abstract
Gtf2i encodes the general transcription factor II-I (TFII-I), with peak expression during pre-natal and early post-natal brain development stages. Because these stages are critical for proper brain development, we studied at the single-cell level the consequences of Gtf2i’s deletion from excitatory neurons, specifically on mitochondria. Here we show that Gtf2i’s deletion resulted in abnormal morphology, disrupted mRNA related to mitochondrial fission and fusion, and altered autophagy/mitophagy protein expression. These changes align with elevated reactive oxygen species levels, illuminating Gtf2i’s importance in neurons mitochondrial function. Similar mitochondrial issues were demonstrated by Gtf2i heterozygous model, mirroring the human condition in Williams syndrome (WS), and by hemizygous neuronal Gtf2i deletion model, indicating Gtf2i’s dosage-sensitive role in mitochondrial regulation. Clinically relevant, we observed altered transcript levels related to mitochondria, hypoxia, and autophagy in frontal cortex tissue from WS individuals. Our study reveals mitochondrial and autophagy-related deficits shedding light on WS and other Gtf2i-related disorders.
Similar content being viewed by others
Introduction
Proper brain development is accomplished through a precise sequence of developmental processes1, mediated mainly by the temporal and spatial regulation of gene expression. This is the result of transcription and transcription regulators such as the transcription factor called TFII-I, which is encoded by GTF2I. TFII-I is a vertebrate-specific multifunctional transcription factor, highly conserved and ubiquitously expressed. It is activated in response to a variety of extracellular signals and translocate to the nucleus2,3,4,5. As such, TFII-I can regulate the expression of genes4,6,7 through interactions with tissue-specific transcription factors and complexes related to chromatin remodeling2,5, and is capable of both promoting and inhibiting transcription7.
TFII-I was shown to be involved in various functions, among them regulating embryonic development8,9,10, cell cycle2,6,11,12, actin cytoskeleton and axon guidance11, endoplasmic reticulum stress response13, and epigenetic modulation14,15,16. TFII-I was also suggested to affect the excitatory/inhibitory balance of the brain17, a disruption known to be responsible for brain development impairments18. A study on human induced pluripotent stem cells showed that Gtf2i was responsible for 10–20% of the transcriptional dysregulation in pathways related to pathologies related to Williams syndrome (WS) and the 7Dup autism spectrum disorder19, both involved with altered expression of GTF2I. We recently studied the role of Gtf2i in post-natal excitatory neurons by deleting it using viral gene-editing techniques. This resulted in hyper-sociability, anxiety, impaired cognition, and hyper-mobility in young mice, suggesting Gtf2i’s importance in regulating behavior during the post-natal stage20. Together, these studies show that Gtf2i expression is critical for proper transcriptional regulation starting from early developmental stages and is essential for proper development of neural circuits involved in brain development and behavior9,18,21,22,23.
Altered transcriptional regulation, such as a result of malfunctioning transcription factor, may lead to cellular dysfunction, including defective energy production which is essential for cell viability and functionality. Due to the high energy demand of neurons, they are highly dependent on mitochondrial function to sustain their homeostasis24,25, support neural plasticity, and enable proper synaptic transmission26. Lack of sufficient energy support in neurons may therefore lead to defected neuronal plasticity and development, resulting in long-term changes in neural circuits activity.
In addition to providing essential energy for neurons, mitochondria maintain calcium balance, involved in metabolism of amino acids, lipids, and steroids, and generate free radicals, all are critical for neurons functionality26,27,28. To maximize functionality, mitochondria evolved as a cellular network that undergo fusion, or fission and biogenesis, in response to the ever-changing cellular environment, in order to meet the cells’ needs. This highly dynamic process is mediated by gene expression regulation at the transcription and translation levels, relying on tight coordination between the nuclear and mitochondrial genomes29. While the importance of mitochondrial properties in relation to proper neuronal development was observed by numerous studies, direct genetic evidence related to the role of TFII-I in mitochondrial health was never provided and is therefore the main aim of the current study.
Here, we chose to focus mainly on the consequences of Gtf2i deletion in forebrain excitatory neurons. Of the genes absent in WS, Gtf2i is of special interest as it has been linked in both humans30,31,32,33,34 and mice20,22,23,35,36,37,38 to the hypersociability and mental retardation associated with this condition. Because these impairments involve functions that are controlled by the cortex, a sub region of the forebrain, and because in the cortex excitatory neurons are major players in controlling the neuronal functions of the cortex, we studied in vitro primary excitatory neurons and the outcomes of Gtf2i deletion in those cells. Our in vitro studies revealed deficits in mitochondrial network morphology, alterations in molecular properties associated with mitochondrial fission and fusion, as well as autophagy and mitophagy, and increased levels of reactive oxygen species (ROS) at a single cell type resolution resulting from Gtf2i deletion in excitatory neurons. These alterations are further supported by our in vivo findings, as studied along the mouse cortex development.
To more accurately replicate the heterozygous deletion seen in individuals with WS, we developed a mouse model featuring heterozygous Gtf2i deletion and conducted subsequent in vitro investigations. Moreover, in order to gain deeper insights into the role of Gtf2i and its relevance to mitochondrial function, we explored the outcomes of a hemizygous neuronal Gtf2i deletion. Our findings unveiled meaningful mitochondrial alterations in both mouse lines, implying the presence of mitochondrial dysfunction in individuals with WS. These discoveries underscore the dosage-sensitive nature of Gtf2i and emphasize its crucial role as a key regulator of mitochondrial dynamics and overall mitochondrial health. Of clinical relevance, we confirmed mitochondrial abnormalities in terms of function, content, hypoxic levels, and autophagy in human frontal cortex brain samples from individuals with WS as compared to typically-developed (TD) controls. Together, our findings reveal new cellular pathways affected by Gtf2i deletion, and suggest new pathophysiological mechanisms that may explain neuronal dysfunction in WS and other GTF2I-related disorders.
Results
Basal levels of Gtf2i and TFII-I are highest at early developmental stages
To explore Gtf2i-encoded TFII-I functions at its natural peak of expression, we first characterized Gtf2i mRNA transcript and TFII-I expression levels in whole hemispheres of naïve mice over the course of development. We noted the highest degrees of Gtf2i transcription and TFII-I expression in developing naïve mice to be in the pre-natal and early post-natal stages (Fig. 1a, b, respectively. Full data are described in the Supplementary Data 1), indicating important roles in mediating embryonic development8,9,39.
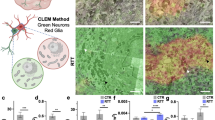
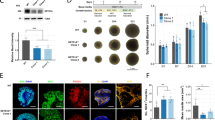
a Gtf2i mRNA levels at E17 and at 1, 2, and 5 months of age, in comparison to post-natal day 1 (P1) (ns, ns, *P = 0.047, *P = 0.011, respectively). b TFII-I levels at E17 and at 1, 2 and 5 months of age, in comparison to P1, in whole hemispheres of naïve mice show the highest basal levels at P1 (ns, *P = 0.031, *P = 0.031, ***P = 0.003, respectively). c Representative images of primary cortical cultures from P1 cKO and control mice (days in vitro; DIV14) showing no TFII-I expression in excitatory neurons (identified by labeling with antibodies to CaMKIIa) of cKO mice. Arrowheads show excitatory neurons. d Quantitation of TFII-I expressing cells from all CaMKIIa-expressing cells shows that only 9.7% of excitatory neurons express TFII-I in cKO mice. Co-localization of e microglia (labeled with anti-Iba1 antibodies, see arrowheads), f astrocytes (labeled with anti-S100B antibodies, see arrowheads), and g inhibitory neurons (labeled with anti-GAD67 antibodies, see arrowheads) and TFII-I showed that TFII-I levels in glia cells and inhibitory neurons of cKO mice remain as in controls. Statistical significance was determined by a, b Kruskal–Wallis tests with multiple comparisons, a n = 3 for E17, n = 3 for P1, n = 4 for 1 month-, n = 5 for 2 month and n = 3 for 5 month-old mice; b n = 3 for E17, n = 5 for P1, n = 5 for 1 month-, n = 5 for 2 month-, and n = 5 for 5 month-old mice. d n = 3 for cKO mice, with 80–120 cells being studied from each mouse. Values represent means ± SEM. ns non-significant.
Gtf2i deletion is specific to excitatory cortical neurons in cKO mice
These findings prompted us to study the role of Gtf2i at early post-natal stages, specifically focusing on neuronal functions. For this, we crossed Gtf2i loxP mice, which have Gtf2i floxed in both alleles40, with NEX-Cre mice41, a Cre line that selectively expresses Cre recombinase in excitatory neurons of the forebrain starting at embryonic day (E) 11.5. The resulting mice, subsequently referred to as cKO mice (Gtf2if/f;NEX-Cre+/−), presented homozygous deletion of Gtf2i selectively in excitatory forebrain neurons42. These mice and their littermate controls (Gtf2if/f;NEX-Cre-/-) were used to prepare primary cortical cultures, enabling us to study molecular and cellular properties at single cell resolution. As shown, TFII-I was absent in excitatory neurons of primary cKO cortical cultures (Fig. 1c), specifically from 90.3% of the excitatory neurons (Fig. 1d), yet was normally expressed in microglia (Fig. 1e), astrocytes (Fig. 1f), and inhibitory neurons (Fig. 1g).
Neuronal Gtf2i deletion alters mitochondrial dynamics and morphology
Mitochondria form interconnected networks to provide an efficient system for the delivery of energy, metabolites, and calcium channels to different areas of the cell43,44. These dynamic networks often change in shape and number, and are controlled by the fusion and fission of individual mitochondria. To study mitochondrial network dynamics as a result of Gtf2i deletion in excitatory neurons, we employed primary cortical cultures (Fig. 1c). Mitochondrial network morphology was visualized using MitoTracker Deep Red staining at DIV14 (Fig. 2a) and analyzed using the ImageJ MiNA macro tool45. Excitatory neurons were identified by staining with antibodies against CaMKIIa, and their soma and neurites were analyzed separately. MiNA was previously used to successfully detect and characterize morphological features of mitochondrial networks in cells in which mitochondria were labeled by MitoTracker staining45, including cells from primary cortical cultures46,47.
Primary cortical cultures from P1 cKO and control mice were seeded on coverslips. After 14 days, the cells were incubated with MitoTracker Deep Red followed by fixation and staining with anti-CaMKIIa and anti-TFII-I antibodies. a Representative images of neurons at DIV14. Arrowheads show excitatory neurons. Neuronal soma (surrounded by white continuous line) and neurites (surrounded by white dashed lines) were analyzed. Morphological skeleton analysis performed using the MiNA tool revealed b individual mitochondria numbers, normalized per soma area, to be higher in cKO mice (*P = 0.044), c increased numbers of networks, normalized per soma area, in cKO mice (***P = 0.003), d network sizes in neurites in cKO mice were significantly bigger (***P = 0.004) and e the mean length of mitochondria in networks in neurites were significantly shorter, reflecting smaller mitochondria, in cKO mice (*P = 0.048), all relative to controls. f A scheme of mitochondrial network parameters, as analyzed using MiNA. Parameters analyzed in panels b-e are listed in the scheme in parentheses. g Mitochondrial fission properties were measured by examining Drp1 mRNA transcript levels in homogenates of cortex from P1 cKO and control mice (**P = 0.009). h Mitochondrial fusion was measured by examining Mfn2 mRNA levels in homogenates of cortex from P1 cKO and control mice (*P = 0.01). Statistical significance was determined by b–e Kolmogorov–Smirnov test or g, h Unpaired t-test. b–e n = 10 control mice, n = 10 cKO mice; 10–20 cells from each mouse. g n = 12 control mice, n = 15 cKO mice. h n = 13 control mice, n = 14 cKO mice. Values represent means ± SEM.
As shown in Fig. 2b, a lack of neuronal Gtf2i resulted in defective mitochondrial dynamics, as demonstrated by the significantly more individual mitochondria in the neuronal soma (normalized per area) in the deletion cultures than seen in the controls. Increased numbers of mitochondrial networks were also observed in cKO mice neuronal soma (Fig. 2c), while the mean number of branches per network was found to be significantly higher in cKO mouse neuronal neurites (Fig. 2d), as compared to controls. Finally, the mean branch length (i.e., the length of those lines that represent mitochondrial structures according to MiNA) was significantly shorter in cKO mouse neuronal neurites as compared to controls (Fig. 2e), suggesting altered mitochondrial structure in the former.
The observed alterations in mitochondrial size, number, and shape in excitatory neurons lacking Gtf2i (Fig. 2a–f) could be the result of altered fission and fusion dynamics, as these processes regulate mitochondrial network homeostasis48,49. Key proteins involved in mitochondrial fusion include the outer membrane GTPases mitofusin 1 and 2 (Mfn1 and Mfn2) and the inner membrane GTPase optic atrophy 1 (OPA1) that coordinate double membrane fusion with the activity of additional scaffolding proteins50,51. The key protein involved in mitochondrial fission is dynamin-related protein 1 (Drp1), which is crucial for the maintenance of mitochondrial morphology and dynamics50,51. To study how neuronal Gtf2i deletion affected mitochondrial fission and fusion dynamics in the mouse brain, we measured transcript levels of these components in whole cortex of P1 cKO mice, as compared to controls, using quantitative PCR (qPCR). We found that cKO mice had significantly higher Drp1 mRNA levels in the whole cortex of P1 mice as compared to controls (Fig. 2g), suggesting altered fission-related properties in the mutant mice. To study molecular properties associated with fusion, we quantified Mfn2 mRNA levels in the whole cortex of P1 mice and found significantly higher Mfn2 transcript levels in cKO mice as compared to controls (Fig. 2h). The changes in mitochondrial gene expression levels observed in cKO mice could potentially be attributed to aberrant transcriptional regulation resulting from the absence of TFII-I in neurons. TFII-I has been demonstrated to interact with various DNA binding motifs52, suggesting its potential direct or indirect role in regulating the expression of these mitochondrial genes (Supplementary Fig. 1).
To further examine mitochondrial function and content as a result of neuronal Gtf2i deletion, we measured transcript levels of components associated with mitochondrial function and content in the whole cortex of P1 cKO mice, as compared to controls, using qPCR. We found that cKO mice had significantly lower mRNA levels of nuclear-encoded succinate dehydrogenase subunit B 1 (Sdhb1), a component of the mitochondrial respiratory chain (mitochondrial complex II), as compared to controls (Fig. 3a). Sdhb1 is known for its function in the process of mitochondrial energy generation, thus, this downregulation may suggest altered mitochondrial function. Furthermore, we measured a marker of mitochondrial content, mitochondrially encoded cytochrome C oxidase I (Mtco1), in which no significant change was detected in the whole cortex of P1 cKO mice, as compared to controls (Fig. 3b).
a Mitochondrial functional properties were measured by examining Sdhb1 transcript levels in homogenates of cortex from P1 cKO and control mice (*P = 0.02). b Mitochondrial content was measured by examining Mtco1 levels (ns). Statistical significance was determined by unpaired t-test. a n = 8 control mice, n = 7 cKO mice. b n = 7 control mice, n = 8 cKO mice. Values represent means ± SEM. ns non-significant.
Neuronal Gtf2i deletion results in higher oxidative stress levels in primary cortical cultures compared to controls
A recent study suggested that aberrations in mitochondrial morphology, mitochondrial dysfunction, and high levels of ROS are interconnected53. Specifically, it was suggested that ROS plays a role in regulating mitochondrial morphology in the nervous system and specifically, in neurons54. To assess ROS levels in primary cultures prepared from the cortex of cKO mice, we performed a 2’,7’-dichlorodihydrofluorescein diacetate (DCF) fluorescent dye-based assay which relies on a ROS-sensitive probe to report on ROS levels in living cells. The DCF assay revealed ROS levels to be significantly higher as a result of neuronal Gtf2i deletion in the primary cKO cortical cultures, as compared to controls (Fig. 4a). To determine whether the higher ROS levels represent a by-product of abnormal mitochondrial respiratory function, we used the MitoSox red reagent, which is specifically targeted to mitochondria in living cells and emits fluorescence when oxidized (Fig. 4b). The finding confirmed that mitochondrial ROS levels were significantly higher in excitatory neurons derived from the primary cKO cortical cultures, as compared to controls (Fig. 4c).
Primary cortical cultures from P1 cKO and control mice were seeded and studied at DIV14. a ROS levels were examined by measuring DCF fluorescence by flow cytometry. H2O2 levels served as positive controls (*P = 0.01). b Mitochondrial ROS levels were measured using the MitoSox red reagent, specifically in the soma of excitatory neurons (arrowheads). c cKO mice showed higher ROS levels than did controls, as measured by assessing mean fluorescence intensity (*P = 0.014). d Primary cortical cultures from the cortex of cKO and control P1 mice were seeded on 96-well plates. Proton leak rates were determined at DIV14 using a Seahorse Mito-stress assay following pharmacological modifications and an XF96 analyzer. Results were normalized to cell number using a methylene blue assay according to the manufacturer’s instructions (***P < 0.0001). Statistical significance was determined by a Mann–Whitney test, c Kolmogorov–Smirnov test and d Unpaired t-test. a n = 8 control mice, n = 6 cKO mice, c n = 4 control mice, n = 3 cKO mice; 5–15 cells from each mouse. d n = 5 control mice, n = 6 cKO mice. Values represent means ± SEM.
To characterize molecular consequences associated with increased ROS levels in primary cortical cultures as a result of neuronal Gtf2i deletion, we next examined proton leak across the mitochondrial inner membrane. Proton leak can decrease ROS formation and, moreover, ROS can prompt proton leak, suggesting the existence of a feedback loop linking ROS and proton leak55. In addressing this putative relation, we found a significant reduction in proton leak in cKO mice, as compared to controls (Fig. 4d), suggesting that the negative regulation of ROS formation had been altered.
We next measured the mRNA levels in the whole cortex in P1 and P30 mice, of ROS-related genes; hypoxia-inducible factor 1 alpha (Hif1a), peroxisome proliferator-activated receptor gamma coactivator 1-alpha (Pgc1a) and apoptosis-related gene Jun proto-oncogene (Jun). Hif1a, which encodes a master regulator of cellular and systemic homeostatic response to hypoxia56, exhibited significantly higher mRNA levels at P1 in cKO mice (Fig. 5a), but no significant change in protein levels for Hif1a as compared to controls (Fig. 5b, c). There were no significant changes observed in mRNA and protein levels at P30 (Fig. 5a, b). Furthermore, Pgc1a, known for its role in inhibiting ROS production57 and acting as a master regulator of mitochondrial biogenesis58,59, exhibited significantly lower levels of mRNA expression in P30 cKO mice compared to controls, while no significant change was measured at P1 (Fig. 5d). Jun, known to be upregulated in response to oxidative stress60,61 and subsequently increase the release of the apoptotic Cyt-C from the mitochondria to the cytosol, exhibited significantly higher expression in cKO pups at P1 compared to controls. However, its expression normalized at P30 (Fig. 5e).
mRNA levels of ROS-related genes and protein levels in homogenates of cortex from cKO mice compared to control were examined. a Hypoxia-related properties were examined by measuring Hif1a mRNA transcript levels in homogenates of cortex from cKO and control mice. In P1 cKO mice showed significantly higher Hif1a mRNA levels (*P = 0.04), but not in P30 (ns), as compared to controls. b Whole cortex homogenates from P1 and P30 cKO compared to control samples were subjected to immunoblot, using the Hif1a antibody, and showed no significant difference in protein levels. c Representative results of the immunoblot of Hif1a. d ROS inhibition properties were examined by measuring Pgc1a mRNA levels in homogenates of cortex from cKO and control mice. In P1 cKO mice showed no significant difference (ns), while in P30 cKO mice showed significantly lower expression level of Pgc1a as compared to controls (*P = 0.01). e Apoptotic properties were examined by measuring Jun mRNA transcript levels in homogenates of cortex from cKO and control mice. In P1 cKO mice showed significantly higher Jun mRNA levels (*P = 0.01), but not in P30 (ns), as compared to controls. Statistical significance was determined by a Mann–Whitney test and b, d, e unpaired t-test. a n = 8 control mice, n = 7 cKO mice, n = 6 control mice, n = 8 cKO mice. b n = 8 control mice, n = 8 cKO mice. d n = 12 control mice, n = 12 cKO mice, n = 7 control mice, n = 8 cKO mice. e n = 8 control mice, n = 7 cKO mice, n = 11 control mice, n = 13 cKO mice. Values represent means ± SEM. ns non-significant.
Neuronal Gtf2i deletion results in decreased autophagy in primary cortical cultures under stress conditions
The altered mitochondrial properties, along with the increased ROS levels we observed due to neuronal Gtf2i deletion, prompted us to define a cellular mechanism that could explain these phenomena. One such mechanism is autophagy, namely, the process of defective organelles clearance62. In autophagy, LC3 (microtubule-associated protein 1 light chain 3) is converted into LC3-II (namely, the lipidated form of LC3), which then associates with the membrane of newly-generated autophagosomes, vesicles that engulf defective organelles, in this case defective mitochondria, destined for destruction63.
To characterize the extent of autophagy in primary cortical cultures, we induced Earle’s balanced salt solution (EBSS) starvation, a scenario that leads to autophagy64 and measured levels of the autophagic protein marker LC3. Under these stress conditions, we found significantly increased autophagy levels in control primary cultures (fold change (FC) = 1.55), as expected (Fig. 6a, b). However, in primary cKO cultures, there was no significant change in the response to stress (FC = 1.24), suggesting an inability to effectively induce autophagy (Fig. 6a) as a result of neuronal Gtf2i deletion.
Primary cortical cultures from P1 cKO and control mice were seeded. After 14 days, proteins were separated by SDS-PAGE and subjected to immunoblot, using the indicated antibodies. a Autophagy was measured by examining the levels of LC3 protein in the primary cortical cultures. A significant increase in the LC3-II/Tubulin ratio was seen under stress condition (3 h EBSS starvation) in the case of control cells, yet no significant such changes were seen with cKO mouse cultures (*P = 0.025). b Representative results of anti-LC3 antibody staining. c mRNA levels of LC3 are lower in homogenate cortex of cKO as compared to control mice at P1 (P = 0.0506). d Mitophagy was measured by examining Parkin protein levels. A significant reduction was detected in cKO cultures (**P = 0.006), while e assessing Pink1 protein levels revealed no significant changes (ns) between the two cultures. f Representative staining by the indicated antibodies. Statistical significance was determined by a, c Unpaired t-test and d, e One sample t-test. a n = 8 control mice, n = 7 control mice subjected to EBSS stress, n = 8 cKO mice, n = 8 cKO mice subjected to EBSS stress, c n = 8 control mice, n = 7 cKO mice. d n = 7 control mice, n = 8 cKO mice, e n = 7 control mice, n = 6 cKO mice. Values represent means ± SEM. ns non-significant, FC fold change, FL full length.
To further characterize autophagy under neuronal Gtf2i deletion, we measured transcript levels of LC3 in whole cortex of P1 cKO mice, as compared to controls, using qPCR. We found that cKO mice had lower LC3 mRNA levels (ns, P = 0.0506) in the whole cortex of P1 mice as compared to controls (Fig. 6c), suggesting altered autophagy.
Neuronal Gtf2i deletion alters mitophagy-associated protein levels in primary cortical cultures and transcript levels in whole cortex
The combination of mitochondrial dysfunction and decreased autophagy as a result of neuronal Gtf2i deletion prompted us to characterize mitophagy, a form of autophagy in which selective degradation of mitochondria occurs. One of the main pathways of mitophagy is the Pink1/Parkin pathway65. As such, we characterized Pink1 and Parkin protein levels in primary cultures and found a significant reduction in Parkin levels as a result of neuronal Gtf2i deletion, as compared to controls (Fig. 6d), which suggests a reduction in mitophagy in the cKO cultures. In contrast, Pink1 expression was unchanged (Fig.6e, f).
To further examine mitophagy and explore additional developmental stages, we conducted Western blot (WB) and qPCR experiments in whole cortex of P1 and P30 mice. We measured transcript levels of Pink1 which revealed significantly increased amounts of Pink1 mRNA levels in whole cortex of cKO mice at P1 and at P30, as compared to controls (Fig. 7a). We further measured Pink1 protein levels and did not find a significant difference between cKO mice and controls in P1 or P30 mice (Fig. 7b). We then measured transcript levels of Parkin and found significantly decreased levels of Parkin mRNA levels at P30, but no significant difference at P1 in cKO mice compared to controls (Fig. 7c). Parkin protein levels did not show a significant difference between cKO mice and controls (Fig. 7d, e). In addition, mRNA for Tbk1, which promotes mitophagy, was found significantly higher in cKO P1 mice, and significantly lower at P30 compared to controls (Fig. 7f).
mRNA levels of mitophagy-related genes and protein levels in homogenates of cortex from cKO mice compared to controls were examined. a Mitophagy-related properties were examined by measuring Pink1 mRNA transcript levels in homogenates of cortex from cKO and control mice. At P1 cKO mice showed significantly higher Pink1 mRNA levels (***P = 0.0008) as well as at P30 (*P = 0.04), as compared to controls. b Whole cortex homogenates from P1 and P30 cKO compared to control mice were subjected to immunoblot, using the Pink1 antibody, and showed no significant difference in protein levels (ns). c Parkin mRNA levels in homogenates of cortex from cKO and control mice were measured. At P1 cKO mice showed no significant difference (ns), while in P30 cKO mice showed significantly lower expression level of Parkin compared to controls (***P = 0.004). d Whole cortex homogenates from P1 and P30 cKO compared to control samples were subjected to immunoblot, using the Parkin antibody, and showed no significant difference in protein levels (ns). e Representative results of the immunoblot of Pink1 and Parkin. f Mitophagy-related properties were examined by measuring Tbk1 mRNA transcript levels in homogenates of cortex from cKO and control mice. P1 cKO mice showed significantly higher Tbk1 mRNA levels (***P = 0.0003) and P30 showed significantly lower Tbk1 mRNA levels (***P = 0.001), as compared to controls. Statistical significance was determined by Unpaired t-test/Mann–Whitney test. a n = 12 control mice, n =14 cKO mice, n = 9 control mice, n = 9 cKO mice, b n = 7 control mice, n = 6 cKO mice, n = 7 control mice, n = 6 cKO mice. c n = 13 control mice, n = 15 cKO mice, n = 7 control mice, n = 7cKO mice. d n = 6 control mice, n = 5 cKO mice, n =10 control mice, n = 6 cKO mice. f n = 8 control mice, n = 7 cKO mice, n = 12 control mice, n = 13 cKO mice. Values represent means ± SEM. ns non-significant. FL full length.
Heterozygous and hemizygous mouse models result in altered mitochondrial morphology and higher oxidative stress levels in primary cortical cultures compared to controls
We observed various mitochondrial dysfunctions in a homozygous Gtf2i neuronal deletion model. To further validate the relevance of Gtf2i dosage-sensitive function, we developed a mouse model to replicate the heterozygous deletion of Gtf2i observed in individuals with WS.
Mitochondrial network morphology was visualized using MitoTracker Deep Red staining at DIV14 (Fig. 8a) and analyzed using the ImageJ MiNA macro tool45. Excitatory neurons were identified by staining with antibodies against CaMKIIa, and their soma and neurites were analyzed separately. As shown in Fig. 8b, the heterozygous deletion of Gtf2i resulted in significantly higher number of branches per network in neuronal neurites as compared to controls. Additionally, the mean branch length was significantly shorter in heterozygous mouse neuronal neurites as compared to controls (Fig. 8c), suggesting altered mitochondrial structure.
Primary cortical cultures from P1 Gtf2i heterozygous (Gtf2i(+/-)) and control mice were seeded on coverslips. After 14 days, the cells were incubated with MitoTracker Deep Red followed by fixation and staining with anti-CaMKIIa and anti-TFII-I antibodies. a Representative images of neurons at DIV14. Arrowheads show excitatory neurons. Neuronal soma (surrounded by white continuous line) and neurites (surrounded by white dashed lines) were analyzed. Morphological skeleton analysis performed using the MiNA tool revealed b network sizes in neurites in heterozygous mice were significantly bigger (*P = 0.02) and c the mean length of mitochondria in networks in neurites was significantly shorter, reflecting smaller mitochondria, in heterozygous mice (*P = 0.03), all relative to controls. d Mitochondrial ROS levels were measured using the MitoSox red reagent, specifically in the soma of excitatory neurons (arrowheads). e Heterozygous mice showed higher ROS levels than did controls, as measured by assessing mean fluorescence intensity (***P = 0.0003). Statistical significance was determined by Kolmogorov-Smirnov test. b, c n = 4 control mice, n = 4 heterozygous mice, 2–15 cells from each mouse. e n = 7 control mice, n = 5 heterozygous mice, 2-30 cells from each mouse. Values represent means ± SEM.
To assess ROS levels in primary cultures prepared from cortex of heterozygous mice, we performed the MitoSox experiment (Fig. 8d). We found that mitochondrial ROS levels were significantly higher in excitatory neurons derived from the primary heterozygous cortical cultures, as compared to controls (Fig. 8e).
These findings provide compelling evidence for the importance of Gtf2i dosage in maintaining proper mitochondrial properties. Moreover, these results have important clinical implications for individuals with WS, as the heterozygous Gtf2i deletion closely resembles the genetic profile observed in individuals with WS66.
Next, we wanted to further explore whether the haploinsufficiency of Gtf2i specifically in forebrain excitatory neurons, is sufficient to induce mitochondrial dysfunction. To achieve this, we developed another mouse model of hemizygous Gtf2i neuronal deletion (Gtf2i(f/+); Nex-Cre(+/-)). Mitochondrial network morphology was once again visualized using MitoTracker Deep Red staining at DIV14 (Fig. 9a) In this model we observed that the mean network size in the soma, as was measured by MitoTracker, was significantly increased (Fig. 9b) and that the number of individual mitochondria in the neurites was significantly higher, as compared to controls (Fig. 9c). We further found that the mean branch length of the mitochondria in the neurites was significantly shorter in hemizygous mouse neuronal neurites as compared to controls (Fig. 9d). This indicates a consistent effect of Gtf2i dosage on mitochondrial morphology, particularly in neurites.
Primary cortical cultures from P1 hemizygous (Gtf2i(f/+);Nex-Cre(+/-)) and control mice were seeded on coverslips. After 14 days, the cells were incubated with MitoTracker Deep Red followed by fixation and staining with anti-CaMKIIa and anti-TFII-I antibodies. a Representative images of neurons at DIV14. Arrowheads show excitatory neurons. Neuronal soma (surrounded by white continuous line) and neurites (surrounded by white dashed line) were analyzed. Morphological skeleton analysis performed using the MiNA tool revealed b mean mitochondrial size in soma in hemizygous mice was significantly bigger (*P = 0.01), c individual mitochondria number in neurites was significantly higher (**P = 0.007) and d the mean length of mitochondria in neurites was significantly shorter, reflecting smaller mitochondria, in hemizygous mice (*P = 0.03), all relative to controls. e Mitochondrial ROS level was measured using the MitoSox red reagent, specifically in the soma of excitatory neurons (arrowheads). f Hemizygous mice showed significantly higher ROS level than did controls, as measured by assessing mean fluorescence intensity (*P = 0.02). Statistical significance was determined by Kolmogorov–Smirnov test. b–d n = 3 control mice, n = 3 hemizygous mice, 2–11 cells from each mouse. f n = 4 control mice, n = 4 hemizygous mice, 5–30 cells from each mouse. Values represent means ± SEM.
Additionally, elevated ROS level was detected under the hemizygous neuronal deletion, as compared to controls (Fig. 9e, f), further emphasizing the similarities among all three mouse models we studied in this article, reinforcing the notion of Gtf2i as a key regulator of mitochondrial properties.
Mitochondrial content, autophagy-related, and hypoxia-related transcripts expression levels in the brain are altered in individuals with WS
Our data suggest mitochondrial dysfunction as a result of neuronal Gtf2i deletion in mice, although mitochondrial properties in brain samples derived from individuals with WS as compared to TD controls are largely unknown.
To determine whether changes in mitochondrial properties occur in brain region relevant for the neurocognitive profile of WS, we extracted RNA from brain samples from Brodmann area 9 (BA9) sections, located in the frontal cortex, derived from individuals with typical deletion of the WS critical region (WSCR) and age-matched TD controls.
Our transcriptomic analysis revealed significant decrease in SDHB1 mRNA level in WS, as compared to TD controls, suggesting altered mitochondrial energy production (Fig. 10a). Additionally, individuals with WS showed a significant increase in COX1 (the human gene name for the mouse gene Mtco1) mRNA level compared to TD controls, which is a marker for mitochondrial content (Fig. 10b). This confirms our findings of an increased number of mitochondria in our mouse model.
Human brain samples from frontal cortex BA9 sections obtained from individuals with typical deletion of the WSCR and age-matched TD controls were used, and their mRNA levels were quantified. Mitochondrial transcripts showed altered expression, with individuals with WS exhibiting a significantly lower levels of SDHB1 mRNA (*P = 0.02), and b significantly higher levels of COX1 mRNA (*P = 0.01) as compared to TD controls. c Autophagy-related transcript was measured by comparing the mRNA levels of MAP1LC3A, which showed lower levels in individuals with WS compared to TD controls (ns). d Hypoxia-related transcript was measured by comparing the mRNA levels of HIF1a, which showed significantly higher levels in individuals with WS compared to TD controls (*P = 0.01). Statistical significance was determined by Unpaired t-test. a n = 4 TD controls, n = 3 WS. b n = 6 TD controls, n = 3 WS. c n = 4 TD controls, n = 3 WS. d n = 6 TD controls, n = 3 WS. Values represent means ± SEM. ns non-significant.
To study autophagy in the brain of individuals with WS, we measured the expression level of the autophagy-related MAP1LC3A transcript (known as LC3 in mice) and found that the mRNA level was lower in WS as compared to TD controls, although not significantly (Fig. 10c), similar to the reduced expression in cKO mice compared to controls.
Last, to study if hypoxia-related properties are altered in the brain of individuals with WS, we measured the expression level of the hypoxic-related HIF1a transcript and found that the mRNA level was significantly higher in WS as compared to TD controls (Fig. 10d), which was also detected in the mouse model.
These transcriptional alterations may be explained by abnormal transcriptional regulation due to the haploinsufficiency of GTF2I (Supplementary Fig. 2). To further investigate the relationship between these genes, see Supplementary Fig. 3. Additionally, for an unbiased overview of transcriptional changes in relevant pathways see Supplementary Fig. 4.
Discussion
Here, using primary cortical cultures and measuring mitochondrial properties, we sought to define Gtf2i roles in mitochondrial dynamics and function in excitatory neurons at the single-cell level. We provided several lines of evidence indicating that mitochondrial morphology and network are abnormal as a result of neuronal Gtf2i deletion, including an increased number of individual mitochondria and larger mitochondrial networks. These results suggest that TFII-I is critical in excitatory neurons for a balanced mitochondrial network and potentially in regulating energy distribution67. Moreover, we validated mitochondrial and autophagy-related deficits in human brain samples derived from individuals with WS, a neurodevelopmental disorder involved with GTF2I haploinsufficiency.
Defects in mitochondrial networks and morphology, such as we found, may be due to improper degradation of damaged mitochondria via mitophagy, which is crucial for energy homeostasis and mitochondrial function67. To investigate autophagy levels in our mouse model, we examined levels of LC3 protein in primary cortical cultures at basal levels and under stress conditions. We found that autophagy was not induced properly in neurons that lack Gtf2i expression under stress conditions as occurs in control cells. Stresses such as hypoxia, mitochondrial depolarization, or proteotoxicity can serve as a trigger for Pink1 accumulation on the mitochondrial surface and for recruitment of the ubiquitin E3 ligase Parkin, which in turn ubiquitinates outer mitochondrial membrane proteins to enable mitophagy50,68,69,70. Accordingly, the reduction in Parkin protein levels in cKO cultures, as compared to controls, suggests defective mitophagy, and may explain the increased number of individual mitochondria and larger mitochondrial networks we found in our model of neuronal Gtf2i deletion.
The defective mitochondrial morphology and decreased mitophagy that we measured in excitatory neurons unable to generate Gtf2i transcripts has the potential to affect oxidative phosphorylation in the defected neurons. ATP distribution over a wider cytoplasmic area could be facilitated by increasing mitochondrial length. Conversely, shorter mitochondrial lengths, as found for neurites in our model culture, could affect ATP distribution and lead to increased energy loss associated with respiration71. Moreover, shorter mitochondria could sequester ions less effectively over a given surface compared to longer mitochondria67.
The mitochondrial defects we described could also lead to generation of oxidative stress due to overproduction of ROS during the process of oxidative phosphorylation in the mitochondria72. Many different factors regulate ROS generation, including mild uncoupling of oxidative phosphorylation via proton leak through the inner mitochondrial membrane. Proton leak decreases ROS generation and ROS have been shown to induce proton leak, suggestive of a feedback loop between them55. In our mouse model, we showed reduced proton leak upon neuronal Gtf2i deletion, which may explain the up-regulation of ROS formation we detected, as a result of potential altered regulation.
To further characterize molecular aspects, we measured the transcript levels of Hif1a, a crucial regulator of cellular and systemic response to hypoxia, to assess whether hypoxia levels are higher due to neuronal Gtf2i deletion and elevated ROS levels. Interestingly, we observed elevated levels of Hif1a mRNA in the whole cortex of P1 cKO pups compared to controls, while protein levels remained unchanged. This finding suggests that following the complete deletion of neuronal Gtf2i, the increased levels of ROS may contribute to enhanced hypoxia.
Furthermore, we conducted an analysis of the transcript levels of Pgc1a, a recognized suppressor of ROS production, and identified a notable and statistically significant decrease. This prompted us to delve into the potential correlation between elevated oxidative stress and the upregulation of the apoptotic marker Jun. Subsequently, our findings substantiated this hypothesis. These discoveries enrich our comprehension of the intricate regulatory mechanisms governing cellular reactions to oxidative stress, encompassing phenomena like hypoxia and apoptosis. It is prudent to acknowledge that these genes harbor multifaceted roles beyond the scope of the observations outlined in this study.
The increased number of mitochondria we found as a result of neuronal Gtf2i deletion may lead to the increased ROS levels we measured, which, in turn, could lead to mitochondrial damage. It was previously shown that dysregulated ROS may contribute to processes that eventually lead to neurodegenerative diseases72 manifested by mtDNA mutations that cause neurological conditions73.
Our findings are supported by the recent suggestion that TFII-I’s targets are enriched in ROS-responsive genes74. Interestingly, previous studies suggested a mechanistic link between high ROS levels and the regulation of mitochondrial morphology and function, given that ROS regulates mitochondrial dynamics by affecting mitochondrial fusion and fission proteins53,75. More specifically, the balance between fusion and fission has been linked to ROS levels in neurons76.
To further validate the relevance of Gtf2i dosage, we sought to replicate the heterozygous microdeletion found in individuals with WS by inducing a heterozygous deletion of Gtf2i.
Our findings revealed substantial alterations in mitochondrial morphology and network, as well as an increase in ROS levels in these heterozygous mice, resembling the effects observed following the homozygous deletion of Gtf2i from excitatory neurons. These results provide compelling evidence for the involvement of Gtf2i in mitochondrial function and hold clinical relevance to individuals with WS, as the heterozygous Gtf2i deletion closely resembles the genetic profile observed in individuals with WS.
We proceeded with a mouse model in which Gtf2i was hemizygously deleted specifically from forebrain excitatory neurons to investigate whether the genetic haploinsufficiency of Gtf2i in these neurons alone is sufficient to induce mitochondrial dysfunction. Surprisingly, our findings once again demonstrated alterations in mitochondrial network and morphology, characterized by an increased number of individual mitochondria and mitochondrial networks. Furthermore, similar to the previous mouse lines studied in this research, we observed shorter mitochondrial branch lengths compared to the control group, and elevated levels of ROS in the excitatory neurons of hemizygous mice.
Notably, any type of Gtf2i deletion performed in this study led to significant changes, including increased mitochondrial number and shorter mitochondrial size. Moreover, the presence of elevated ROS levels, which are associated with altered mitochondrial function and can induce numerous neuronal alterations, was observed across all three mouse models. Mitochondrial dysfunction, and energy and metabolism deficits, are known features of numerous neurodevelopmental disorders77,78, including autism spectrum disorders79,80,81,82, Fragile X syndrome83,84, Angelman syndrome84,85, Rett syndrome84,86, and WS87,88. WS is caused by a heterozygous microdeletion of 25–27 genes89,90 on the long arm of chromosome 7 (7q11.23), which is part of the WSCR. WS individuals are known for their social abnormalities, particularly, they struggle with inhibiting their social behavior91,92, and exhibit non-social anxiety derived from fear and specific phobias93. WS individuals also have cognitive impairments94,95, with IQ scores typically ranging from 40 to 100, with a mean score of around 6096, which ranks them as mildly to moderately intellectually disabled.
To describe possible mitochondrial alterations in clinically relevant tissue samples, we examined mRNA levels in brain sections from the frontal cortex of individuals with WS and TD controls. Our data revealed reduced expression of SDHB1 in individuals with WS as compared to TD controls. The encoded mitochondrially-localized protein is a subunit of the succinate dehydrogenase enzyme complex, responsible for transferring electrons from succinate to ubiquinone, critical for mitochondrial function as mediated by the mitochondrial respiratory chain complex II. The reduced expression of SDHB1 in WS may decrease mitochondrial functionality in supporting cells energetically. Also, it may lead to increased ROS levels, similar to the increased ROS level we measured in our in vitro study, as a result of enhanced transfer of electrons to O2, due to the potential inhibited succinate-ubiquinone activity.
Studying additional mitochondria-related transcript, we found that COX1 expression level in the frontal cortex of individuals with WS was higher, as compared to TD controls. COX1 encodes the protein cytochrome c oxidase I, which is the main subunit of the cytochrome c oxidase complex, part of the respiratory complex IV. This complex is the third and final enzyme, taking part in the electron transport chain of mitochondrial oxidative phosphorylation. COX is acting in creating the electrochemical gradient essential for ATP synthesis as part of the oxidative phosphorylation97,98. Therefore, it is possible that the increased expression of COX1 in WS represents the cells’ compensation machinery in order to increase their energy-generating capacity99. Interestingly, high level of COX1 expression was previously found in defective mitochondria, suggesting that mitochondrial genes can continue to function and be expressed in high levels, although their products are not functional properly100. COX1 is also a known marker for mitochondrial content, and the observed significant increase in its levels in individuals with WS aligns with the data obtained from the mouse model, in which mitochondrial numbers were higher. To characterize autophagy-related properties in the human brain, we studied MAP1LC3A and found that its expression level in the frontal cortex of individuals with WS was lower, as compared to TD controls, although not significantly due to the high variability in the control group. Microtubule-associated protein 1 light chain 3 alpha is involved in autophagy, by taking active part in the formation of autophagosomes101,102. The reduced expression of MAP1LC3A in WS may therefore result in insufficient degradation of unhealthy cells in the brain of individuals with WS, or that unnecessary or dysfunctional components in those cells will not be removed properly. Finally, to examine hypoxia-related properties in WS, we measured HIF1a transcript levels and found its expression level in the frontal cortex of individuals with WS was significantly higher, as compared to TD controls, as was also found in our mouse model. This implies that human individuals with WS may be suffering from higher ROS levels which eventually lead to hypoxia. Since GTF2I, a member of the WSCR, is known to be a key player in the neurocognitive profile of WS, our results may explain how a lack of GTF2I expression affects neuronal-mediated functions such as cognition, social behavior, and other WS-associated pathologies. Other important factors affecting cognition and behavior, traits which require intact levels of energy and metabolism, and which are altered in WS, are myelination and signal conduction in the brain42. Significant energy consumption by myelinated axons is devoted to axonal repolarization, axonal transport, and the maintenance of glial cells103. The mitochondrial alterations we defined as a result of Gtf2i neuronal deletion may therefore affect cognition through neuronal-specific mechanisms, but also through neuron-glia interactions, affecting not only gray matter, but also white matter properties essential for proper brain functionality104.
Gtf2i roles in mediating mitochondrial properties were unknown before our study, to the best of our knowledge. Nevertheless, previous WS-related studies suggested a connection between WS and mitochondrial abnormalities, involving other genes from the WSCR. For instance, Dnajc30 deletion in WS model mice resulted in dysfunctional mitochondria and functional differences in the oxidative phosphorylation pathway87. This study suggested that the protein encoded by Dnajc30 acts as a regulator of mitochondrial respiration through interaction with ATP synthase87. In a second study, the WSCR16 gene was demonstrated to be localized to mitochondria in HeLa cells88, while an earlier study suggested that WSCR16 could be co-precipitated with mitochondrial proteins and mitochondrial 16S rRNA105.
While widely studied in the context of the hypersociability and the neurocognitive aspects in WS23,30,31,32,35,42,106, Gtf2i mutations and altered expression are also related to autism spectrum disorders caused by duplication of 7q11.23107,108,109,110,111 and cancer112,113,114,115. TFII-I was also found to be involved in three autoimmune diseases: rheumatoid arthritis116, primary Sjogren’s syndrome117, and systemic lupus erythematosus118,119. Taken together, these studies demonstrate the wide diversity of functions TFII-I has in mediating proper cellular properties, and its wide clinical relevancy. In combination with our findings from the mouse model and the human brain samples, it is therefore conceivable to test in future studies whether targeting mitochondrial and autophagy-related deficits might be a beneficial therapeutic strategy for treating WS and other GTF2I-related pathologies.
Methods
Mice
Gtf2i loxP mice40 underwent more than 15 generations of back-crossed to pure C57Bl/6J mice (stock no. 000664; The Jackson Laboratory). All experimental protocols conformed to the guidelines of the Institutional Animal Care and Use Committee of Tel Aviv University, Tel Aviv, Israel. All efforts were made to minimize animal suffering and the number of animals used. Each cage contained 2–5 mice, regardless of genotype, within a controlled environment of 23 °C and a 12-h light–dark cycle (lights on at 7:00, lights off at 19:00) with free access to food and water. NEX-Cre mice are in a C57Bl/6 background41. NEX-Cre mice have exhibited normal behavior and development, with Cre activity commencing around E11.5, primarily limited to the excitatory neurons of the forebrain, particularly in the cortex and hippocampus41. Gtf2i-Het mice displayed germline deletion of a single Gtf2i allele. To induce hemizygous deletion of Gtf2i in excitatory neurons, Gtf2i homozygous loxP mice were bred with wild-type C57Bl/6J mice, resulting in the creation of mice with a single floxed Gtf2i allele and one wild-type allele [Gtf2i(f/+)mice]. Subsequently, these mice were crossed with NEX-Cre mice, leading to the generation of mice [Gtf2i(f/+); NEX-Cre(+/-)] with targeted hemizygous Gtf2i deletion exclusive to excitatory neurons.
All test mice were males. Therefore, the limitations of this article arise from the absence of considering the sex of the mice in the experimental design, potentially overlooking gender-specific differences in the results.
Primary cortical cultures
Pups were genotyped on P1 by PCR. Then, brains were removed and transferred to a culture dish containing ice-cold sterile PBS. Olfactory bulbs, basal ganglia, and cerebellum were removed, and the cortex was isolated, collected in digestion solution (HBSS-HEPES, CaCl2, EDTA, DNAse, l-cysteine, and papain (Sigma-Aldrich)), cut into smaller pieces with sterile scissors and transferred to a conical tube for 20 min, with shaking in room temperature (RT). After transfer to a conical tube containing neuronal plating medium (Neurobasal A (Rhenium), FBS, SM1 (Enco), glutamax (Rhenium), and Penicillin-Streptomycin), the tissue was mechanically digested by trituration. Cells were pelleted by centrifugation (1000 rpm for 4 min), resuspended in plating medium and seeded on matrigel (Lapidot)-coated coverslips in 6-, 12-, 96-well plates. Cultures were maintained at 37 °C in a humidified atmosphere of 95% air, 5% CO2. One half of the medium was replaced with fresh growth medium (without FBS) every 72 h. FUDR (Merck) was added at DIV4-8.
Immunofluorescent staining
Primary cortical cultures were washed with PBS, followed by 10 min fixation in RT with 4% paraformaldehyde (PFA) and then washed 3 times (5 min for each wash) with PBS. Blocking was performed with Triton X-100 in PBS and 2% normal goat serum (NGS) for 1 h at RT. Primary antibodies diluted in blocking buffer were applied for 1 h at RT. Wells were then washed 3 times with PBS for 10 min each time and stained with secondary antibodies conjugated to Alexa Fluor (1:1000) in blocking buffer for 1 h at RT. Wells were washed three times in PBS for 10 min each time, followed by 4′,6-Diamidine-2′-phenylindole dihydrochloride (DAPI; 1:1000, Sigma) staining in PBS for 10 min, and an additional 20 min wash in PBS. Vectashield hardset mounting medium (Zotal) was used to mount the coverslips on glass slides. Images were captured using a Leica TCS SP5 Olympus confocal microscope, and an Olympus IX83 inverted microscope. Commercial antibodies used included anti-TFII-I (1:1000; CST-4562S, Cell signaling), anti-CaM Kinase IIa (1:400; 50049, Cell signaling), anti-Iba1 (1:500; 234 006, SYSY) anti-S100B (1:600; S-2532, Sigma-Aldrich) and anti-Gad67 (1:1000; MAB5406, Millipore) antibodies.
Analysis of mitochondrial network morphology
Primary cortical cultures were incubated with 250 nM MitoTracker Deep Red FM (MTDR, Invitrogen) for 30 min at 37 °C. Wells were washed with PBS, followed by fixation with 100% ice-cold methanol at −20 °C for 20 min. Immunofluorescent staining was performed as described below. Mitochondrial network morphology of individual cells was visualized with a Leica TCS SP5 confocal microscope (×63 magnification) and analyzed using the ImageJ MiNA macro tool45. Analysis was performed in excitatory neurons alone and normalized to soma area. Neurites were all analyzed at the same distance from soma and at the same length.
Proton leak assessment
The Seahorse XF Cell Mito Stress Test Kit (Agilent) was utilized. Cells (20,000) were plated on matrigel-coated Seahorse assay plates, and analyzed after 14 days in culture. The assay medium was formulated by enriching Seahorse XF medium with precise quantities of sodium pyruvate (1 mM), glutamine (2 mM), and glucose (10 mM). The medium was then heated to 37 °C and adjusted to a pH of 7.4. The Agilent Seahorse XF Cell Culture Microplate was primed for the assay by replacing the growth medium with the prepared assay medium. This step is followed by an incubation period to ensure the cells are in optimal condition before commencing the assay. Normalization was performed to compare similar experiments across different 96-well plates. The results were normalized to cell number using a methylene blue assay according to the manufacturer’s instructions.
ROS measurement using DCF and MitoSox
DCF
For flow cytometry analysis of DCF staining, 250,000 cells were plated on matrigel-coated 6-well plates. After 14 days, the cells were washed and loaded with 10 μM of the dye (Sigma-Aldrich, D6883) for 30 min at 37 °C. After washing to remove excess dye, the cells were suspended in trypsin for 5 min at 37 °C and then in PBS and centrifuged. As a control, 1 µM of H2O2 was introduced to the cells for 2 h at 37 °C.
MitoSox
Primary cortical cultures at DIV14 in 12-well plates were washed and incubated with 2.5 µM MitoSOX Red (M36008, Rhenium) and Hoechst (33342) for 10 min at 37 °C. Coverslips with live cells were captured using an Olympus IX83 inverted microscope. Coverslips were fixed with 4% PFA and stained, as described.
Lysate preparation and immunoblot
Fourteen day-old primary cortical cultures were starved for 3 h with EBSS (Sigma-Aldrich) and then dissected. Protein contents were separated using 4% Laemmli loading buffer, boiled, and resolved by SDS–polyacrylamide gel electrophoresis through 10% and 15% gels. The separated proteins were electrophoretically transferred to a nitrocellulose membrane in transfer buffer (25 mM Tris-HCl, pH 7.5, 190 mM glycine, and 10% methanol, absolute). Membranes were blocked for 45 min in TBST buffer (0.05 M Tris-HCl, pH 7.5, 0.15 M NaCl, and 0.1% Tween 20) with 6% skimmed milk, and blotted overnight with the indicated antibodies in TBST buffer, followed by incubation with secondary antibodies linked to horseradish peroxidase. Immunoreactive bands were detected with enhanced chemiluminescence reagent. Commercial antibodies used included rabbit anti-Hif1a (1:1000, PA1-16601, Rhenium), rabbit anti-LC3 (1:1000; 12741, Cell signaling), rabbit anti-Pink1 (1:1000; ab23707, Abcam) mouse anti-Parkin (1:100; 4211, Cell signaling) and mouse anti-beta-tubulin (1:5000; AB44928, Abcam) antibodies.
RNA extraction
Upon ice thawing, extracted and frozen cortices underwent homogenization in 1 mL of cold TRIzol reagent (Thermo Fisher Scientific) utilizing a handheld electric homogenizer (Pro Scientific). Following a 5 min incubation at RT, 200 μL of chloroform was introduced to each sample, with subsequent manual tube shaking for 15 seconds. After another 3 min incubation at RT, the tubes were centrifuged for 20 min at 4 °C (13,800 rpm; Eppendorf Centrifuge 5430 R). Upon the separation of the mixture into three layers, the uppermost clear layer containing RNA was transferred to a fresh tube, to which 1:1 (v/v) isopropanol was added to induce RNA precipitation. After brief shaking and a 5 min RT incubation, the tubes were subjected to a 15 min centrifugation at 4 °C (13,800 rpm) to precipitate the RNA. The ensuing steps included removal of isopropanol, two washes with 1 mL of 80% ethanol (Sigma-Aldrich) in DEPC-treated water (Biological Industries), followed by a 5 min 4 °C centrifugation. After ethanol removal, the tubes were air-dried for 15–25 min, and subsequently, 20–35 μL of DEPC-treated water was added to each tube. Final RNA concentrations were determined using a Thermo Scientific NanoDrop One device (Thermo Fisher Scientific).
mRNA expression
Total RNA extraction was used as the starting material for mRNA complementary deoxyribonucleic acid (cDNA) synthesis. The reverse transcription of mRNA was performed employing random primers and a High-Capacity cDNA Reverse Transcription Kit (Thermo Fisher Scientific). The C1000 Touch thermal cycler (Bio-Rad) was employed with the following parameters: 10 min at 25 °C, 120 min at 37 °C, 5 min at 85 °C, followed by a final step at 4 °C until completion.
Real-time mRNA quantification
mRNA levels were determined using a Fast SYBR Green PCR Master Mix (Thermo Fisher Scientific), according to the manufacturer’s instructions, and the Bio-Rad CFX Connect Real-Time PCR Detection System (Bio-Rad). Thermal cycler conditions were as follows: 20 seconds at 95 °C, 40 amplification cycles (3 seconds at 95 °C to denature, and 30 seconds at 60 °C to anneal and extend), and a melt curve: 60 °C for 5 seconds, and an increase of 0.5 °C every 5 seconds (including a plate read) until reaching 95 °C. Expression values were calculated based on the comparative cycle threshold method120. mRNA levels were normalized to those of glyceraldehyde 3-phosphate dehydrogenase (Gapdh) mRNA. mRNA levels are shown as fold change, relative to levels in the control group. Specific primers for the detection of mRNA levels were ordered from Hy Laboratories (see supplementary table 1) and diluted to 10 mM in DEPC-treated water according to the manufacturer’s instructions.
Human brain samples
All human tissue samples were kindly provided by the NIH NeuroBioBank at the University of Maryland, School of Medicine (NBB request #1068), in accordance with all legal provisions and relevant ethical considerations. Blocks of fixed control and WS human cortical brain samples from BA9 were examined.
Human RNA isolation, cDNA preparation, and real-time polymerase chain reaction
RNA from BA9 cortex was isolated according to Kumar et al.121 with minor modifications, briefly: 650 mg of fixed tissue was lysed in 400 µl TES buffer (500 mM Tris pH 8, 100 mM EDTA, 100 mM NaCl, 1%SDS, 500 µg/mL proteinase K) and decrossed in 56 °C for 3 h. RNA was isolated from the lysates using TRI Reagent (Molecular Research Center) according to the manufacturer’s instructions. RNA was reverse-transcribed to single-stranded cDNA by Super Script II Reverse Transcriptase (Thermo Fisher Scientific) and random primers were mixed with 1 µM gene-specific primers using a mix of reverse primers from the quantified genes; qPCR was performed in a CFX Connect Real-Time PCR Detection System (Bio-Rad) with PerfeCTa SYBR Green FastMix (Quanta BioSciences). Dissociation curves were analyzed following each real-time qPCR to confirm the presence of only one product and the absence of primer dimer formation. The threshold cycle number (Ct) for each tested gene (X) was used to quantify the relative abundance of that gene using the formula 2-(Ct geneX – Ct standard). Tubulin (TUBA1B) was used as the standard for mRNA expression. Primers used for the real-time qPCR and gene-specific amplification are listed in supplementary table 2.
Multi correlation analysis
Correlation analysis between a set of genes driven from major biological pathways of mitochondrial functions, differential DNA methylation, and differentially expressed genes (DEGs) was performed by GeneOverlap R package. First, we used ToppGene Suite database in order to compile lists of genes specifically associated with the following biological pathways: Mitochondrial fusion, Pink1-PRKN mediated mitophagy, Mitochondrial biogenesis, Response to oxidative stress, Mitophagy, Mitochondrial fission, Negative regulation of oxidative stress-induced cell death, Response to mitochondrial depolarization and Apoptotic mitochondrial changes. As a negative control, we employed a set of genes driven from the biological pathway Structural constituent of cytoskeleton, which is not directly associated with mitochondrial function.
Next correlation analysis was performed between these gene lists and DEGs or genes that were annotated to differential DNA methylation. Transcriptional data were obtained from human brain samples of individuals with WS42, and were presented as either DEGs calculated based on a p-value < 0.01 or false discovery rate (FDR) < 0.05. Further transcriptional data (DEGs, FDR < 0.05) were collected from brain samples of mice with Gtf2i homozygous deletion specifically from excitatory neurons42. Last, DNA methylation data were taken from human brain samples from individuals with WS122. P adjusted value (presented in numbers) and odds ratio (color scale) from Fisher’s exact are presented in Supplementary Fig. 3.
Statistics and reproducibility
Data are presented as means ± standard error of the mean (SEM), as calculated by GraphPad Prism 8.4.3. p-values were calculated using Student’s t-test (two-sided unpaired t-test or one sample t-test), with p < 0.05 considered as significant (*<0.05, **<0.01, ***<0.005). Normality of distributions and equality of variances were checked and addressed accordingly using the appropriate statistical analysis. Outliers were determined via the extreme studentized deviate (ESD) method.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data and source data are included in the manuscript and the supplementary information and will be accessible to anyone interested upon reasonable request. The numerical source data for all graphs in the manuscript can be found in Supplementary Data 1. All uncropped blots images generated and analyzed during the current study are available in the supplementary information. All materials are commercialized and will be available upon reasonable request.
References
Jiang, X. & Nardelli, J. Cellular and molecular introduction to brain development. Neurobiol. Dis. 92, 3–17 (2016).
Roy, A. L. Biochemistry and biology of the inducible multifunctional transcription factor TFII-I: 10 years later. Gene 492, 32–41 (2012).
Roy, A. L. Signal-induced functions of the transcription factor TFII-I. Biochimica et Biophysica Acta (BBA)-. Gene Struct. Expr. 1769, 613–621 (2007).
Cheriyath, V., Desgranges Zp Fau Roy, A. L. & Roy, A. L. c-Src-dependent transcriptional activation of TFII-I. J. Biol. Chem. 277, 22798–22805 (2002).
Roy, A. L. Biochemistry and biology of the inducible multifunctional transcription factor TFII-I. Gene 274, 1–2 (2001).
Desgranges, Z. P. et al. Inhibition of TFII-I-dependent cell cycle regulation by p53. Mol. Cell. Biol. 25, 10940–10952 (2005).
Hakre, S. et al. Opposing functions of TFII-I spliced isoforms in growth factor-induced gene expression. Mol. Cell 24, 301–308 (2006).
Makeyev, A. V. & Bayarsaihan, D. New TFII-I family target genes involved in embryonic development. Biochem. Biophys. Res. Commun. 386, 554–558 (2009).
Enkhmandakh, B. et al. Essential functions of the Williams-Beuren syndrome-associated TFII-I genes in embryonic development. Proc. Natl Acad. Sci. USA 106, 181–186 (2009).
Enkhmandakh, B., Bitchevaia, N., Fau, -, Ruddle, F., Ruddle, F., Fau, -, Bayarsaihan, D. & Bayarsaihan, D. The early embryonic expression of TFII-I during mouse preimplantation development. Gene Expr. Patterns 4, 25–28 (2004).
Chimge, N. O., Makeyev, Fh. R. & Bayarsaihan, D. Identification of the TFII-I family target genes in the vertebrate genome. Proc. Natl Acad. Sci. USA 105, 9006–9010 (2008).
Ashworth, T. & Roy, A. L. Phase specific functions of the transcription factor TFII-I during cell cycle. Cell Cycle 8, 596–605 (2009).
Hong, M. et al. Transcriptional regulation of the Grp78 promoter by endoplasmic reticulum stress: role of TFII-I and its tyrosine phosphorylation. J. Biol. Chem. 280, 16821–16828 (2005).
Bayarsaihan, D. What role does TFII-I have to play in epigenetic modulation during embryogenesis? Epigenomics 5, 9–11 (2013).
Tussie-Luna, M. I., D, B., Fau, S. E., Fh, R. & Roy, A. L. Physical and functional interactions of histone deacetylase 3 with TFII-I family proteins and PIASxbeta. Proc. Natl Acad. Sci. USA 99, 12807–12812 (2002).
Strong, E. et al. DNA methylation profiles in individuals with rare, atypical 7q11. 23 CNVs correlate with GTF2I and GTF2IRD1 copy number. NPJ Genom. Med. 8, 25 (2023).
Wang, Y. et al. Dlx5 and Dlx6 regulate the development of parvalbumin-expressing cortical interneurons. J. Neurosci. 30, 5334–5345 (2010).
Barak, B. & Feng, G. Neurobiology of social behavior abnormalities in autism and Williams syndrome. Nat. Neurosci. 19, 647–655 (2016).
Sanders, S. J. et al. Multiple recurrent de novo CNVs, including duplications of the 7q11.23 Williams syndrome region, are strongly associated with autism. Neuron 70, 863–885 (2011).
Ophir, O. et al. Deletion of Gtf2i via systemic administration of AAV-PHP. eB virus increases social behavior in a mouse model of a neurodevelopmental disorder. Biomedicines 11, 2273 (2023).
Adamo, A. et al. 7q11.23 dosage-dependent dysregulation in human pluripotent stem cells affects transcriptional programs in disease-relevant lineages. Nat. Genet. 47, 132–141 (2015).
Lucena, J. et al. Essential role of the N-terminal region of TFII-I in viability and behavior. BMC Med. Genet. 11, 61 (2010).
Sakurai, T. et al. Haploinsufficiency of Gtf2i, a gene deleted in Williams Syndrome, leads to increases in social interactions. Autism Res. 4, 28–39 (2011).
Katsenelson, M. et al. IGF-1 receptor regulates upward firing rate homeostasis via the mitochondrial calcium uniporter. Proc. Natl Acad. Sci. USA 119, e2121040119 (2022).
Ruggiero, A., Katsenelson, M. & Slutsky, I. Mitochondria: new players in homeostatic regulation of firing rate set points. Trends Neurosci. 44, 605–618 (2021).
Mattson, M. P., Gleichmann, M., Fau, -, Cheng, A. & Cheng, A. Mitochondria in neuroplasticity and neurological disorders. Neuron 60, 748–766 (2008).
Gleichmann, M. & Mattson, M. P. Neuronal calcium homeostasis and dysregulation. Antioxid. Redox Signal. 14, 1261–1273 (2011).
MacAskill, A. F., Atkin Ta Fau Kittler, J. T. & Kittler, J. T. Mitochondrial trafficking and the provision of energy and calcium buffering at excitatory synapses. Eur. J. Neurosci. 32, 231–240 (2010).
Nisoli, E. et al. Mitochondrial biogenesis as a cellular signaling framework. Biochem. Pharmacol. 67, 1–15 (2004).
Antonell, A. et al. Partial 7q11.23 deletions further implicate GTF2I and GTF2IRD1 as the main genes responsible for the Williams-Beuren syndrome neurocognitive profile. J. Med. Genet. 47, 312–320 (2010).
Dai, L. et al. Is it Williams syndrome? GTF2IRD1 implicated in visual-spatial construction and GTF2I in sociability revealed by high resolution arrays. Am. J. Med. Genet. 149a, 302–314 (2009).
Morris, C. A. et al. GTF2I hemizygosity implicated in mental retardation in Williams syndrome: genotype-phenotype analysis of five families with deletions in the Williams syndrome region. Am. J. Med. Genet. 123a, 45–59 (2003).
Tassabehji, M. et al. GTF2IRD1 in craniofacial development of humans and mice. Science 310, 1184–1187 (2005).
Danoff, S. K., Taylor, H. E., Blackshaw, S. & Desiderio, S. TFII-I, a candidate gene for Williams syndrome cognitive profile: parallels between regional expression in mouse brain and human phenotype. Neuroscience 123, 931–938 (2004).
Borralleras, C., Sahun, I., Perez-Jurado, L. A. & Campuzano, V. Intracisternal Gtf2i gene therapy ameliorates deficits in cognition and synaptic plasticity of a mouse model of Williams-Beuren syndrome. Mol. Ther. 23, 1691–1699 (2015).
Li, H. H. et al. Induced chromosome deletions cause hypersociability and other features of Williams-Beuren syndrome in mice. EMBO Mol. Med. 1, 50–65 (2009).
Osborne, L. R. Animal models of Williams syndrome. Am. J. Med. Genet. C, Semin. Med. Genet. 154c, 209–219 (2010).
Segura-Puimedon, M. et al. Heterozygous deletion of the Williams-Beuren syndrome critical interval in mice recapitulates most features of the human disorder. Hum. Mol. Genet. 23, 6481–6494 (2014).
Fijalkowska, I., Sharma, D., Bult, C. J. & Danoff, S. K. Expression of the transcription factor, TFII-I, during post-implantation mouse embryonic development. BMC Res. Notes 3, 203 (2010).
Enkhmandakh, B. et al. Generation of a mouse model for a conditional inactivation of Gtf2i allele. Genesis 54, 407–412 (2016).
Goebbels, S et al. Genetic targeting of principal neurons in neocortex and hippocampus of NEX-Cre mice. Genesis 44, 611–621 (2006).
Barak, B. et al. Neuronal deletion of Gtf2i, associated with Williams syndrome, causes behavioral and myelin alterations rescuable by a remyelinating drug. Nat. Neurosci. 22, 700–708 (2019).
De Giorgi, F., Lartigue, L. & Ichas, F. Electrical coupling and plasticity of the mitochondrial network. Cell Calcium 28, 365–370 (2000).
Karbowski, M. & Youle, R. J. Dynamics of mitochondrial morphology in healthy cells and during apoptosis. Cell Death Differ. 10, 870–880 (2003).
Valente, A. J., Maddalena, L. A., Robb, E. L., Moradi, F. & Stuart, J. A. A simple ImageJ macro tool for analyzing mitochondrial network morphology in mammalian cell culture. Acta Histochem. 119, 315–326 (2017).
Wang, L. et al. Enhancing S-nitrosoglutathione reductase decreases S-nitrosylation of Drp1 and reduces neuronal apoptosis in experimental subarachnoid hemorrhage both in vivo and in vitro. Brain Res. Bull. 183, 184–200 (2022).
Cymerys, J., Chodkowski, M., Slonska, A., Krzyzowska, M. & Banbura, M. W. Disturbances of mitochondrial dynamics in cultured neurons infected with human herpesvirus type 1 and type 2. J. Neurovirol. 25, 765–782 (2019).
Zamponi, N. et al. Mitochondrial network complexity emerges from fission/fusion dynamics. Sci. Rep. 8, 363 (2018).
van der Bliek, A. M., Shen, Q. & Kawajiri, S. Mechanisms of mitochondrial fission and fusion. Cold Spring Harb. Perspect. Biol. 5, a011072 (2013).
Liu, Y. J., McIntyre, R. L., Janssens, G. E. & Houtkooper, R. H. Mitochondrial fission and fusion: a dynamic role in aging and potential target for age-related disease. Mech. Ageing Dev. 186, 111212 (2020).
Palmer, C. S., Osellame, L. D., Stojanovski, D. & Ryan, M. T. The regulation of mitochondrial morphology: intricate mechanisms and dynamic machinery. Cell. Signal. 23, 1534–1545 (2011).
Makeyev, A. V. et al. Diversity and complexity in chromatin recognition by TFII-I transcription factors in pluripotent embryonic stem cells and embryonic tissues. PLoS ONE 7, e44443 (2012).
Willems, P. H., Rossignol, R., Dieteren, C. E., Murphy, M. P. & Koopman, W. J. Redox homeostasis and mitochondrial dynamics. Cell Metab. 22, 207–218 (2015).
Cid-Castro, C., Hernandez-Espinosa, D. R. & Moran, J. ROS as regulators of mitochondrial dynamics in neurons. Cell. Mol. Neurobiol. 38, 995–1007 (2018).
Brookes, P. S. Mitochondrial H(+) leak and ROS generation: an odd couple. Free Radic. Biol. Med. 38, 12–23 (2005).
Sharp, F. R. & Bernaudin, M. HIF1 and oxygen sensing in the brain. Nat. Rev. Neurosci. 5, 437–448 (2004).
Luo, C., Widlund, H. R. & Puigserver, P. PGC-1 coactivators: shepherding the mitochondrial biogenesis of tumors. Trends Cancer 2, 619–631 (2016).
Cheng, A. et al. Involvement of PGC-1α in the formation and maintenance of neuronal dendritic spines. Nat. Commun. 3, 1250 (2012).
Lehman, J. J. et al. Peroxisome proliferator–activated receptor γ coactivator-1 promotes cardiac mitochondrial biogenesis. J. Clin. Investig. 106, 847–856 (2000).
Martindale, J. L. & Holbrook, N. J. Cellular response to oxidative stress: signaling for suicide and survival. J. Cell. Physiol. 192, 1–15 (2002).
Su, B. et al. Oxidative stress signaling in Alzheimer’s disease. Curr. Alzheimer Res. 5, 525–532 (2008).
Kelekar, A. Autophagy. Ann. New York Acad. Sci. 1066, 259–271 (2005).
Liu, H. et al. From autophagy to mitophagy: the roles of P62 in neurodegenerative diseases. J. Bioenerg. And Biomembr. 49, 413–422 (2017).
Munafo, D. B. & Colombo, M. I. A novel assay to study autophagy: regulation of autophagosome vacuole size by amino acid deprivation. J. Cell Sci. 114, 3619–3629 (2001).
Pickles, S., Vigie, P. & Youle, R. J. Mitophagy and quality control mechanisms in mitochondrial maintenance. Curr. Biol. 28, R170–R185 (2018).
Kozel, B. A. et al. Williams syndrome. Nat. Rev. Dis. Primers. 7, 42 (2021).
Chang, D. T. & Reynolds, I. J. Mitochondrial trafficking and morphology in healthy and injured neurons. Prog. Neurobiol. 80, 241–268 (2006).
Harper, J. W., Ordureau, A. & Heo, J. M. Building and decoding ubiquitin chains for mitophagy. Nat. Rev. Mol. Cell Biol. 19, 93–108 (2018).
Matsuda, N. et al. PINK1 stabilized by mitochondrial depolarization recruits Parkin to damaged mitochondria and activates latent Parkin for mitophagy. J. Cell Biol. 189, 211–221 (2010).
Ordureau, A. et al. Quantitative proteomics reveal a feedforward mechanism for mitochondrial PARKIN translocation and ubiquitin chain synthesis. Mol. Cell 56, 360–375 (2014).
Skulachev, V. P. Mitochondrial filaments and clusters as intracellular power-transmitting cables. Trends Biochem. Sci. 26, 23–29 (2001).
Angelova, P. R. & Abramov, A. Y. Functional role of mitochondrial reactive oxygen species in physiology. Free Radic. Biol. Med. 100, 81–85 (2016).
Angelova, P. R. & Abramov, A. Y. Role of mitochondrial ROS in the brain: from physiology to neurodegeneration. FEBS Lett. 592, 692–702 (2018).
Kopp, N. D. et al. Functions of Gtf2i and Gtf2ird1 in the developing brain: transcription, DNA binding and long-term behavioral consequences. Hum. Mol. Genet. 29, 1498–1519 (2020).
Jendrach, M., Mai, S., Pohl, S., Voth, M. & Bereiter-Hahn, J. Short- and long-term alterations of mitochondrial morphology, dynamics and mtDNA after transient oxidative stress. Mitochondrion 8, 293–304 (2008).
Knott, A. B., Perkins, G., Schwarzenbacher, R. & Bossy-Wetzel, E. Mitochondrial fragmentation in neurodegeneration. Nat. Rev. Neurosci. 9, 505–518 (2008).
Uittenbogaard, M. & Chiaramello, A. Mitochondrial biogenesis: a therapeutic target for neurodevelopmental disorders and neurodegenerative diseases. Curr. Pharm. Des. 20, 5574–5593 (2014).
A, O., U, M., Lf, B. & A, G. C. Energy metabolism in childhood neurodevelopmental disorders. EBioMedicine 69, 103474 (2021).
Wen, Y. & Yao, Y. Autism Spectrum Disorders (ed A. M. Grabrucker) (2021).
Anitha, A. et al. Brain region-specific altered expression and association of mitochondria-related genes in autism. Mol. Autism 3, 12 (2012).
Haas, R. H. Autism and mitochondrial disease. Dev. Disabil. Res. Rev. 16, 144–153 (2010).
Rossignol, D. A. & Frye, R. E. Mitochondrial dysfunction in autism spectrum disorders: a systematic review and meta-analysis. Mol. Psychiatry 17, 290–314 (2012).
Griffiths, K. K. et al. Inefficient thermogenic mitochondrial respiration due to futile proton leak in a mouse model of fragile X syndrome. FASEB J. 34, 7404–7426 (2020).
Ortiz-Gonzalez, X. R. Mitochondrial dysfunction: a common denominator in neurodevelopmental disorders. Dev. Neurosci. 43, 222–229 (2021).
Santini, E. et al. Mitochondrial superoxide contributes to hippocampal synaptic dysfunction and memory deficits in angelman syndrome model mice. J. Neurosci. 35, 16213–16220 (2015).
Valenti, D. et al. Stimulation of the brain serotonin receptor 7 rescues mitochondrial dysfunction in female mice from two models of Rett syndrome. Neuropharmacology 121, 79–88 (2017).
Tebbenkamp, A. T. N. et al. The 7q11.23 Protein DNAJC30 Interacts with ATP Synthase and Links Mitochondria to Brain Development. Cell 175, 1088–1104 e1023 (2018).
Koyama, M., Sasaki, T., Sasaki, N. & Matsuura, Y. Crystal structure of human WBSCR16, an RCC1-like protein in mitochondria. Protein Sci. 26, 1870–1877 (2017).
Korenberg, J. R. et al. VI. Genome structure and cognitive map of Williams syndrome. J. Cogn. Neurosci. 12, 89–107 (2000).
Osborne, L. R. Williams-Beuren syndrome: unraveling the mysteries of a microdeletion disorder. Mol. Genet. Metab. 67, 1–10 (1999).
Pober, B. R. Williams-Beuren syndrome. New Engl. J. Med. 362, 239–252 (2010).
Laws, G. & Bishop, D. Pragmatic language impairment and social deficits in Williams syndrome: a comparison with Down’s syndrome and specific language impairment. Int. J. Lang. Commun. Disord. 39, 45–64 (2004).
Dykens, E. M. Anxiety, fears, and phobias in persons with Williams syndrome. Dev. Neuropsychol. 23, 291–316 (2003).
Mervis, C. B. et al. The Williams syndrome cognitive profile. Brain Cogn. 44, 604–628 (2000).
Mervis, C. B. & Klein-Tasman, B. P. Williams syndrome: cognition, personality, and adaptive behavior. Ment. Retard. Dev. Disabil. Res. Rev. 6, 148–158 (2000).
Bellugi, U., Lichtenberger, L., Mills, D., Galaburda, A. & Korenberg, J. R. Bridging cognition, the brain and molecular genetics: evidence from Williams syndrome. Trends Neurosci. 22, 197–207 (1999).
Schmidt, T. R. et al. Rapid electrostatic evolution at the binding site for cytochrome c on cytochrome c oxidase in anthropoid primates. Proc. Natl Acad. Sci. USA 102, 6379–6384 (2005).
Wikstrom, M. K. Proton pump coupled to cytochrome c oxidase in mitochondria. Nature 266, 271–273 (1977).
Barrett, T. et al. A murine dopamine neuron-specific cDNA library and microarray: increased COX1 expression during methamphetamine neurotoxicity. Neurobiol. Dis. 8, 822–833 (2001).
Terman, A. et al. Mitochondrial recycling and aging of cardiac myocytes: the role of autophagocytosis. Exp. Gerontol. 38, 863–876 (2003).
Suzuki, H. et al. Structural basis of the autophagy-related LC3/Atg13 LIR complex: recognition and interaction mechanism. Structure 22, 47–58 (2014).
Cherra, S. J. 3rd et al. Regulation of the autophagy protein LC3 by phosphorylation. J. Cell Biol. 190, 533–539 (2010).
Harris, J. J. & Attwell, D. The energetics of CNS white matter. J. Neurosci. 32, 356–371 (2012).
Nir, A. & Barak, B. White matter alterations in Williams syndrome related to behavioral and motor impairments. Glia 69, 5–19 (2021).
Arroyo, J. D. et al. A genome-wide CRISPR death screen identifies genes essential for oxidative phosphorylation. Cell Metab. 24, 875–885 (2016).
Grad, M. et al. Altered White Matter and microRNA Expression in a Murine Model Related to Williams Syndrome Suggests That miR-34b/c Affects Brain Development via Ptpru and Dcx Modulation. Cells. 11, 158 (2022).
López-Tobón, A. et al. GTF2I dosage regulates neuronal differentiation and social behavior in 7q11.23 neurodevelopmental disorders. Sci Adv. 9, eadh2726 (2023).
Malenfant, P. et al. Association of GTF2i in the Williams-Beuren syndrome critical region with autism spectrum disorders. J. Autism And Dev. Disord. 42, 1459–1469 (2012).
Morris, C. A. et al. 7q11.23 Duplication syndrome: physical characteristics and natural history. Am. J. Med. Genet. Part A 167A, 2916–2935 (2015).
Osborne, L. R. & Mervis, C. B. 7q11.23 deletion and duplication. Curr. Opin. Genet. Dev. 68, 41–48 (2021).
Mervis, C. B. et al. Children with 7q11.23 duplication syndrome: psychological characteristics. Am. J. Med. Genet. Part A 167, 1436–1450 (2015).
Higuchi, R et al. Primary driver mutations in GTF2I specific to the development of thymomas. Cancers. 12, 2032 (2020).
Feng, Y. et al. GTF2I mutation frequently occurs in more indolent thymic epithelial tumors and predicts better prognosis. Lung Cancer 110, 48–52 (2017).
Nathany, S., Tripathi, R. & Mehta, A. Gene of the month: GTF2I. J. Clin. Pathol. 74, 1–4 (2021).
Kim, D. W., Cheriyath, V., Roy, A. L. & Cochran, B. H. TFII-I enhances activation of the c-fos promoter through interactions with upstream elements. Mol. Cell. Biol. 18, 3310–3320 (1998).
Kim, K. et al. Association-heterogeneity mapping identifies an Asian-specific association of the GTF2I locus with rheumatoid arthritis. Sci. Rep. 6, 27563 (2016).
Li, Y. et al. A genome-wide association study in Han Chinese identifies a susceptibility locus for primary Sjögren’s syndrome at 7q11.23. Nat. Genet. 45, 1361–1365 (2013).
Sun, C. et al. High-density genotyping of immune-related loci identifies new SLE risk variants in individuals with Asian ancestry. Nat. Genet. 48, 323–330 (2016).
Meng, Y. et al. Association of GTF2I gene polymorphisms with renal involvement of systemic lupus erythematosus in a Chinese population. Medicine 98, e16716 (2019).
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108 (2008).
Kumar, K., Oli, A., Hallikeri, K., Shilpasree, A. S. & Goni, M. An optimized protocol for total RNA isolation from archived formalin-fixed paraffin-embedded tissues to identify the long non-coding RNA in oral squamous cell carcinomas. MethodsX 9, 101602 (2022).
Trangle, S. S. et al. In individuals with Williams syndrome, dysregulation of methylation in non-coding regions of neuronal and oligodendrocyte DNA is associated with pathology and cortical development. Mol. Psychiatry 28, 1112–1127 (2023).
Acknowledgements
The authors would like to thank the members of the Barak laboratory for their support along the project. Also, we would like to thank Prof. Inna Slutsky, Prof. Eitan Okun, Prof. Ronit Pinkas-Kramarski, and Dr. Avi Ashkenazi for their insightful comments on this study. This research was supported by the ISRAEL SCIENCE FOUNDATION (grant No. 2305/20, awarded to B.B.) Williams France Federation and Autour des Williams (awarded to B.B.).
Author information
Authors and Affiliations
Contributions
Conceptualization: A.N., S.S.T., G.E.S., A.A., O.E.S., and B.B.; methodology: A.N., G.L., S.S.T., G.E.S., E.B., A.M., A.A., O.E.S., and B.B.; analysis: A.N., G.L., H.P., T.R., A.M., S.S.T., and B.B.; research: A.N., G.L., S.S.T., G.E.S., O.O., I.F., M.R., A.A., A.M., H.P., T.R., O.E.S., and B.B.; writing: A.N., G.L., S.S.T., M.R., A.M., A.A., and B.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All experimental protocols conformed to the guidelines of the Institutional Animal Care and Use Committee of Tel Aviv University, Tel Aviv, Israel. We have complied with all relevant ethical regulations for animal use.
Informed consent
All authors consent the publication of this article by the publisher.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Joao Valente.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nir Sade, A., Levy, G., Schokoroy Trangle, S. et al. Neuronal Gtf2i deletion alters mitochondrial and autophagic properties. Commun Biol 6, 1269 (2023). https://doi.org/10.1038/s42003-023-05612-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-05612-5
- Springer Nature Limited