Abstract
Technical advances in neuroimaging, notably in fMRI, have allowed distributed patterns of functional connectivity to be mapped in the human brain with increasing spatiotemporal resolution. Recent years have seen a growing interest in extending this approach to rodents and non-human primates to understand the mechanism of fMRI connectivity and complement human investigations of the functional connectome. Here, we discuss current challenges and opportunities of fMRI connectivity mapping across species. We underscore the critical importance of physiologically decoding neuroimaging measures of brain (dys)connectivity via multiscale mechanistic investigations in animals. We next highlight a set of general principles governing the organization of mammalian connectivity networks across species. These include the presence of evolutionarily conserved network systems, a dominant cortical axis of functional connectivity, and a common repertoire of topographically conserved fMRI spatiotemporal modes. We finally describe emerging approaches allowing comparisons and extrapolations of fMRI connectivity findings across species. As neuroscientists gain access to increasingly sophisticated perturbational, computational and recording tools, cross-species fMRI offers novel opportunities to investigate the large-scale organization of the mammalian brain in health and disease.
Similar content being viewed by others
Introduction
Mapping functional connectivity networks with fMRI
A large body of evidence suggests that, even at rest (i.e. without any overt cognitive or sensory stimulation) brain activity synchronously fluctuates in concert across distributed regions1,2,3. This important observation has promoted an emerging model whereby brain activity and complex cognitive functions can be conveniently operationalized as the result of a flexible and dynamic interaction between nodes that constitute distributed functional networks4,5.
A popular approach for studying how brain regions are intrinsically organized and communicate with each other is resting state fMRI (rsfMRI). Initial observations that rsfMRI signal is highly synchronous between functionally related regions6 have promoted a framework whereby statistical dependencies between spontaneous fluctuations in brain activity are interpreted as an index of interareal communication, or “functional connectivity” between regions7. Subsequent investigations have revealed reproducible patterns extending beyond motor sensory areas to encompass other sensory systems and distributed networks of regions that are highly synchronous at rest, and whose activity is modulated by cognitive tasks8. These spatiotemporal patterns of coordinated activity define a set of reproducible resting-state networks (RSNs) characterized by high intrinsic functional connectivity9. Importantly, RSNs topography has also been shown to spatially reconstitute several known functional systems of the human brain, thus suggesting that intrinsic and task-evoked brain activity may strongly influence each other1,8,10.
The accessibility and practicality of rsfMRI, along with recent progress in open data initiatives and the development of advanced analytical methods have greatly improved our understanding of the intrinsic functional organization of the human brain11,12. Prominent investigations of rsfMRI networks have characterized how patterns of intrinsic brain activity and functional connectivity are related to cognitive function and behavior10,13. Similarly, rsfMRI mapping in pathological states has revealed atypical network activity and altered functional connectivity in all major neurological and psychiatric diseases14,15.
While many ongoing brain mapping initiatives heavily rely on the use of rsfMRI, the correlative nature of this approach and our inability to interpret and physiologically decode the mechanisms underlying interareal fMRI coupling strongly limit the impact of this research, often relegating it to phenomenological evidence devoid of any mechanistic information. As a result, rsfMRI connectivity mapping is still considered by many as an imaging endophenotype, or proxy for interregional communication of uncertain physiological significance.
Bridging the explanatory gap: mechanistic studies of fMRI connectivity
Many key questions related to the fundamental principles of rsfMRI connectivity remain to date unaddressed, primarily as a consequence of our inability to mechanistically disambiguate a highly correlative phenomenon like rsfMRI in humans. While this issue has not prevented a proficient use of rsfMRI connectivity in cognitive neuroscience, a deeper understanding of the physiological basis of rsfMRI may equally benefit cognitive neuroscientists and researchers interested in the organization of functional networks in brain disorders. A better understanding of the mechanism of rsfMRI may for example prevent the erroneous (yet common) use of this metric as a monotonic index of connectivity16, or may help understand its relationship with internal states and physiological signals that bias network organization e.g. arousal and peripheral signals17. Furthermore, research into the physiological bases of rsfMRI coupling may help decode patterns of dysconnectivity in brain disorders and relate them to physiologically interpretable measures that could help diagnose or stratify patient populations for customized treatments.
More broadly, mechanistic investigations of fMRI connectivity can help bridge the big explanatory gap that exists between molecular and biophysical modeling of microscale neural activity, and network-level descriptions of brain function (Fig. 1a)18. Human studies have attempted to address this issue by relating patterns of fMRI activity to measures of anatomical connectivity or physiological measurements recorded concurrently with rsfMRI (reviewed by Suárez et al.4). Similarly, large-scale imaging initiatives have linked genetic variation to network features both in healthy populations19,20 as well as in carriers of genetic alterations associated with brain disorders21,22,23,24. However, the correlative nature of these investigations has not allowed a reliable mechanistic disambiguation of fMRI (dys)connectivity.
a A major gap exists between models of human brain function at different levels of inquiry. The use of neural systems measurements like fMRI in physiologically accessible animals can help bridge this gap by causally linking neural events at different investigational scales. Another major advantage of this approach is related to the possibility of probing the role of specific microscale processes in shaping whole brain patterns of fMRI connectivity. b Highly complementary experimental approaches can be implemented in humans, non-human primates (NHPs) and rodents to investigate the neural basis and organization of fMRI connectivity. The panel illustrates, for each species, experimental approaches or resources that are readily available and commonly used (green), currently unavailable or ethically inaccessible (red), and available under special circumstances (yellow), or for which only proof of concept studies have been described. For the latter group, references to recently published examples are reported here: [1]43,135,222, [2]223, [3]27, [4]224, [5]225, [6]205 [7]226. Non-invasive stimulation includes transcranial magnetic stimulation (TMS), transcranial alternating current stimulation (tACS) and related methods. EEG: electroencephalography; DBS: deep brain stimulation.
Attempts to establish causality using brain stimulation, or by relating patterns of altered rsfMRI connectivity to neurological lesions have revealed interesting associations between clinical scores and rsfMRI network dysfunction in humans, hence substantiating the role of distributed network activity in mediating complex behavioral traits and functions25,26,27,28,29. However, our poor comprehension of the complex physiological cascade underlying neurological damage and the similarly limited understanding of the physiological basis of brain stimulation limit the mechanistic significance of these investigations. Crucially, the recent extension of rsfMRI mapping to physiologically accessible species offers the unprecedented opportunity to bridge the “explanatory gap” (Fig. 1a). Specifically, the combination of rsfMRI with the ever-increasing set of advanced neuromanipulations and recording tools available in animals provide a means to causally deconstruct rsfMRI activity, relate central and peripheral physiological events to brain-wide patterns of fMRI activity, and investigate the anatomical correlates of brainwide network activity across investigational scales (Fig. 1b).
This experimental approach is coming of age30, and promises to add novel interpretative dimensions to rsfMRI and its use in cognitive and translational neuroscience. The impact and potential of mechanistic investigation in animals is epitomized by a series of investigations that we briefly summarize in the following sections, with the intent to make readers aware of the critical contribution and impact preclinical fMRI is having in the field. Crucially, these examples also highlight the need to identify reliable means to map and relate fMRI connectivity networks (and related findings) across species to ensure an efficacious transfer of information from and to human and animal studies.
Mechanistic studies in non-human primates
fMRI connectivity mapping was initially extended from human to non-human primates (NHP), where its combined use with invasive electrophysiological recordings and stimulations has produced a foundational understanding of the neural correlates of blood oxygenation level dependent (BOLD) fMRI activity31,32,33,34,35,36,37,38. Further electrophysiological studies have linked rsfMRI activity to resting local field potentials39, neural coupling40, and physiological signatures of arousal41.
fMRI studies in NHP have also benefitted from the availability of detailed anatomical information attainable via use of invasive axonal tracers42,43,44. By relating topographical patterns of fMRI connectivity to the organization of the NHP axonal connectome, NHP studies have supported the notion that fMRI network organization is strongly constrained by underlying patterns of anatomical connectivity37,45,46. The generalization of this notion has been further investigated in interventional studies encompassing the use of lesions46,47 or reversible inactivation of brain activity via infusion of drugs48 and chemogenetics ligands in virally-transfected animals49.
Further interventional studies in NHP have encompassed the use of electrical, ultrasound50,51, or infrared stimulation52 to understand how the activity of a single brain region causally affects global patterns of fMRI connectivity. The use of sedative and anesthetic agents has also been explored to map and compare patterns of fMRI activity as a function of brain state. For example, a study comparing wakeful versus unconscious states has shown that spontaneous functional connectivity patterns in awake monkeys show a rich repertoire of functional connections that is more dissimilar to structural connectivity compared to anesthesia53,54. Differences in cross-subjects variability of fMRI connectivity patterns has been described in the cortex of awake vs. anesthetized NHP55. Cortical BOLD responses to visual stimulation are instead consistent in awake vs. unconscious NHP56. Finally, it has also been shown that the intrinsic network structure of main primary and associative networks in the anesthetized macaque monkeys are topographically similar to those mapped in conscious humans37.
Finally, attempts to map fMRI connectivity in transgenic primate models of human disorders have also been recently described57.
The impact of NHP investigations of the neural correlates of BOLD signal and fMRI connectivity has been substantial. As novel recording and manipulation tools become available to NPH researchers, the added value of mechanistic investigation in these species is expected to exponentially grow, offering novel opportunities to bridge theoretical and experimental neuroscience, and reliably translate findings from and to human research. However, despite its tremendous potential, the implementation of fMRI in NHPs remains technically, logistically and procedurally very demanding, requiring highly specialized equipment and personnel, and high investment and running costs. Of equal importance, ethical constraints prevent a full exploitation of the ever-expanding repertoire of manipulations and recording methods available to experimental neuroscientists.
Mechanistic studies in rodents
To circumvent these limitations, recent years have seen a strong drive towards the implementation of fMRI connectivity mapping in more physiologically accessible species, such as rats and mice. The field of rodent fMRI has steadily grown, and now encompasses a large number of laboratories in the world58,59,60,61. Much of the interest in this approach lies in the possibility of combining fMRI connectivity mapping with recordings and experimental manipulations (including genetically modified lines) not directly applicable for technical or ethical reasons in higher mammalian species.
The possibility of combining fMRI connectivity mapping with cell-type specific neuromanipulations represents a very promising area of investigation in the field of rodent fMRI. Notable examples of this approach have investigated the contribution of ascending transmitter nuclei (e.g. noradrenergic, cholinergic, serotonergic or dopaminergic neurotransmission) in causally shaping the organization of the fMRI connectome via chemogenetic62,63,64 or optogenetic manipulations65,66. Further studies have employed chemogenetic manipulations to physiologically-decode fMRI connectivity signals67 and study how activity in one region causally affects whole-brain patterns of fMRI connectivity68. By linking neural signals to large-scale patterns of fMRI connectivity these studies are uncovering a set of fundamental principles governing communication between brain regions at the macroscale, and in some cases challenging prevailing interpretations of fMRI connectivity16.
The wide availability of transgenic models in rodents represents an important complement to mechanistic investigations in NHP, where the use of transgenes is still limited (Fig. 1b). A fertile area of application of fMRI in transgenic models is to investigate how individual disease-risk genetic mutations affect brain wiring and fMRI connectivity. For eample, this approach has been used by us and others to link autism-associated risk genes to specific macroscale signatures of fMRI dysconnectivity, thus corroborating a key role of genetics in shaping the functional connectome in developmental disorders69,70,71,72,73,74,75,76. By mechanistically relating dysfunctional fMRI activity to atypical axonal wiring70,72,74,77, synaptic abnormalities71 or electrophysiological alterations74, some of these studies have shed light on the neural mechanisms underlying connectional atypicality. Multisite extensions of this approach have revealed the effect of genetic variability across multiple genetic models of autism78. Notable applications of fMRI in rodents have also been extended to other neurological and psychiatric disorders (reviewed in79), to include Alzheimer’s disease80,81, Huntington disease82, schizophrenia83 and ADHD84.
Rodent fMRI studies have also leveraged the availability in the mouse of a high-resolution, directed, mesoscale axonal connectome85 to uncover the relationship between functional and anatomical connectivity. This resource has made it possible to extend investigations of the structural basis of network activity to finer spatial scales inaccessible to human investigators86,87, as well as to probe axonal correlates of sub-networks of the rodent brain, such as the mouse default mode network (DMN)88 or salience network (SN)89.
Recent years have also seen great advancements in linking macroscale fMRI mapping with mesoscale neural signals, particularly through the use of genetically encoded calcium indicators via fiber photometry. This innovative approach enabled the prediction of MRI network organization based on patterns of calcium neural activity in specific brain regions and the establishing of reciprocal causal interactions between networks systems of translational relevance via calcium-related signals90,91,92,93. More importantly, simultaneous mapping of whole brain fMRI and cortex-wide mesoscale calcium imaging has been recently demonstrated, offering novel opportunities to relate neural and hemodynamic signals across multiple investigational scales94.
We conclude this overview by mentioning the widespread use, both in NPH and rodent fMRI studies, of intravascular contrast agents aimed at increasing functional contrast to noise ratio95, reduce susceptibility artefacts, and in turn increase the sensitivity of both evoked and intrinsic functional imaging96,97,98. Superparamagnetic iron-oxide nanoparticles in particular have been widely used to map cerebral blood volume99, producing vascular contrast that reproduces the macroscale organization of conventional BOLD fMRI connectivity networks96,100. The high vascular specificity of this approach and its practicality makes it a robust approach for mapping fMRI networks in small mammals like rodents and NHP.
Collectively, complementary fMRI research in NHP and rodents offers the opportunity to mechanistically probe the neural basis of fMRI coupling in the mammalian brain. However, to best leverage and properly contextualize the wealth of information generated by animal fMRI studies, methods to extrapolate findings across species are required. At the core of this argument lies the critical need to understand whether the mechanistic inferences made in lower mammalian species, especially those pertaining to specific networks or circuits, are species-invariant (or not), and if so, to which extent they can be topographically related to network systems of the human brain. Addressing this question is of special importance for the investigation of mechanisms that influence fMRI network topography such as the contribution of central modulatory systems, or mechanism of dysconnectivity produced by genetic or developmental insult30.
Organization of fMRI networks across species
Species-invariant and species-specific fMRI connectivity networks
While the question of how to reliably match network organization across-species remains open, correspondences in the organization of fMRI network across the phylogenetic tree suggest that topographical organization of large-scale networks can be used to index patterns of regional activity and guide the extrapolation of fMRI finding to and from mammalian species. The ultimate goal of this line of investigation is to identify homologous functional networks enabling comparative brain mapping and the extrapolation of region- or network-specific findings across the phylogenetic tree. The cogency of this approach is supported by the results of recent studies revealing a set of evolutionary precursors of human RSNs in NHP and lower mammalian species.
Initial investigations in macaques37, followed by more recent studies in lower mammalian species (i.e. marmosets, lemur monkeys, rats and mice)100,101,102,103, have revealed that that the organization of spontaneous fMRI signal into distributed and bilateral spatiotemporal patterns is a foundational, evolutionarily-conserved property of the mammalian brain. Further investigations of the rodent and primate cortices have revealed the presence of phylogenetically-conserved RSNs, including interhemispheric synchronous visual, auditory104,105 and somatomotor networks55,88,100,105,106,107,108. The latter has been termed in rodents “latero-cortical network” (LCN) owing to its antero-lateral distribution107. Owing to their unambiguous location, and their anchoring on well characterized cortical territories (in each species), visual, auditory, and somatomotor networks can thus be considered “species-invariant” networks, i.e. networks that can be reliably identified and mapped across the phylogenetic tree. Clearly, species invariance does not imply here absolute spatial, cytoarchitectural or connectional homogeneity, but rather an unambiguous representation of these systems across multiple species, independent of their evolutionary complexity.
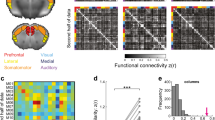
The observation of distributed fMRI networks in the human brain encompassing higher order areas such as the default mode or salience network8,109 led to the initial assumption that the organization of these systems, could reflect a distinctive organizational mode that is exclusive to humans. However, the discovery of similarly distributed connectivity networks also in anesthetized NHPs has refuted this hypothesis37,110. Further studies in NHP and rodents have identified RSNs that anatomically reconstitute the topography of the human DMN and salience networks (SN)60,88,89,100,110,111. Interestingly, similar to what is observed in humans2,60 somatomotor regions of the LCN exhibit anticorrelation with the DMN55,100,106,107,108 (Fig. 2a). It should be noted that all these network correspondences hinge on the presence of synchronous activity in phylogenetically conserved regions such as limbic and somatomotor cortical areas, or evolutionarily-ancient precursors of the medial prefrontal cortex like the anterior cingulate cortex112. Therefore, in lower species, the organization of these fMRI network precursors only partly recapitulate the richer topography of corresponding systems in humans88,103,113.
a Representative cortical fMRI networks in humans227 and their putative evolutionarily precursors in awake NHP55 and rodents106. Plausible homologs of the human default mode, salience, limbic, somatomotor, and visual networks have been described in NHP55 and mice106. A possible homolog of the executive control network has been identified in NHP162. To date, no evolutionarily plausible precursors of human dorsal attention and temporoparietal networks have been identified in animals. b Approximate location of hub constituents of default mode, salience, limbic, somato-motor and visual networks in the human, NHP and mouse brain as shown in later (top panel) and sagittal (bottom) panel. Please note that the Yeo parcellation227 of the human brain incorporates the auditory network within somato-motor areas. Evidence of dissociable auditory networks has been however reported in all species, including mice100 rats139 and NHP117. Default mode network is in red, salience network is in violet, limbic network is in yellow, somatomotor network is in blue, visual network is in purple, control network is in orange, temporoparietal network is in dark blue, and dorsal attention network is in green, according to Yeo’s parcellation227.
In keeping with this, plausible rodent and NHP precursors of human higher-order cognitive networks such as the dorsal-attention, temporoparietal or executive control networks remain to be firmly identified. Being these systems anchored on high-order cortical areas that have greatly expanded and that are highly specialized in humans with respect to primates and rodents, the possibility that these systems are human-specific seems highly plausible60,114. We note however that reports of a cortical system subserving executive control functions in NHP have been published115,116,117 (Fig. 2a). Further research into the phylogenetic evolution of these networks is required to address the question of whether and how these functions are represented at the network level in lower species.
Importantly, notable cross-species correspondences in fMRI network organization also encompass subcortical systems. Seed-based mapping or independent component analyses in rodents have allowed the break down of fMRI network activity into fine-grained subcortical functional systems, including hippocampal, basal ganglia, thalamic and basal-forebrain networks106,118,119,120. Analogous systems can be reliably identified both in NHP and humans121,122,123,124,125,126. While the evolutionarily ancient origin of these systems makes them more easily identifiable and directly related across species, subcortical systems may also exhibit increasingly complex topography along the phylogenetics trees similar to what is observed in cortical networks125, especially in terms of integration and cross-talk with other anatomical substrates. These factors should be accounted for when mapping and comparing activity in subcortical networks systems across species.
General organizational principles of fMRI connectivity across species
Irrespective of some expected differences in the organization of higher-order cortical networks along the phylogenetic tree, some generalizable, evolution-invariant principles underlying the functional organization of RSN are apparent, which can guide topographical mapping of networks systems across species.
The most conspicuous of these principles is the organization of cortical and subcortical RSN into exquisitely bilateral, homotopic systems exhibiting robust interhemispheric synchronization. This feature applies to virtually all fMRI networks so far identified in lower mammalian species and encompasses associative (i.e., DMN, salience, etc.) as well as sensory and somatomotor (i.e., visual, lateral cortical etc.) networks. Interhemispheric homotopicity is also a defining characteristic of subcortical rsfMRI network100,126,127.
The presence of a tight relationship between the topography of anatomical and functional connectivity is a second foundational organizational principle of fMRI connectivity across species. Tracer-based high-resolution connectome mapping in rodents and NHPs have expanded the results of initial investigations of the structural basis of fMRI connectivity in humans based on diffusion weighted MRI128,129,130, by allowing fMRI network architecture to be related to quantitative and directional measures of connectivity at the mesoscale85,131,132. These studies have revealed conserved rules of cortical connectivity across species133, including cell-class-specific projection patterns88,134. Importantly, this relationship appears to be independent of the spatial scale investigated135. These correspondences are of interest in the light of an emerging body of evidence supporting the notion that topographic organization of fMRI networks is shaped and constrained by the underlying anatomical organization of the brain, with contributions that go beyond and above axonal connectivity4,86,87, to also include laminar microstructure of myelination136,137 and the geometric organization of brain anatomy138.
The presence of hub-like nodes in associative regions is an additional organizational feature that appears to underpin the global structure of fMRI connectivity in multiple mammalian species. fMRI connectivity hubs have been shown to be similarly anchored to midline components of the DMN in rodents107,139, NHPs140 and humans141,142, albeit with a shift from more anterior (medial prefrontal) to posterior (retrosplenial, posterior cingulate cortex) cortical areas occurring across the evolutionary timeline60,107 (Fig. 2b). Interestingly, a general overlap between the anatomical location of the functional and structural connectivity hubs has been observed in rodents86,143,144, NHP140,145 and humans129,146. These correspondences further corroborate the intimate relationship between anatomical and functional organization of the mammalian brain we have described above.
fMRI connectivity gradients
The spatial correspondence between structural and fMRI connectivity in cortical areas has been recently investigated in greater depth as part of a broader question of high evolutionary relevance: why is each brain region located where it is, and how does its anatomical organization impact interareal functional communication? This is an old question that has fascinated researchers for many years147. Classical investigations of cortical microarchitecture and cortico-cortical connectivity have proposed a hierarchical model whereby regions having similar features occupy the same position along a graded cortical axis148,149,150. According to the “tethering hypothesis”, high-order association cortices untethered from sensory hierarchies in the cortical expansion during evolution, resulting in their current topological location as in-between zones of the cortical mantle151,152,153. Interestingly, recent studies have expanded this framework by showing that multiple biological properties that are relevant to the organization of functional connectivity are also distributed along a primary axis in spatial variation ranging from sensory (i.e. unimodal) to transmodal cortical regions. These include cortical profiles of axonal connectivity86, gene expression154, receptor expression155, as well as cytoarchitectural properties such as local neuronal density156, dendritic spine density and size and dendritic tree complexity157, all of which are organized a common axis of spatial variation that is recognizable in humans158, NHP136 and rodents159.
Recently, Margulies and colleagues160 captured this unimodal to transmodal axis (i.e. gradient) with fMRI using a diffusion embedding approach, thus demonstrating that the topological organization of cortical fMRI connectivity is hierarchically organized along a spatial gradient spanning unimodal-transmodal cortical regions in both humans and NHP (Fig. 3a). This principal gradient is also found to be spatially aligned with T1-T2-weighted inferences of intracortical myelin content, thus relating macroscale RSN organization to classical sensory-transmodal hierarchy and its underlying microstructural foundations150,158,161. Similar fMRI-based connectivity gradients have also been recently identified in macaques, marmosets and rodents where they were found to recapitulate the unimodal-transmodal axis mapped in the human cortex (Fig. 3b), and to be exquisitely aligned with the organization of the axonal connectome86,162,163,164. Notably, it has been recently suggested that this gradient is representative of the course of cortical evolution, with phylogenetically newer areas emerging at particular points along the axis133,165.
a fMRI connectivity in the human cortex is arranged along two main axes. The first and dominant gradient spans unimodal-polymodal cortical regions. The second gradient reconstitutes the sensory specific arrangement of cortical regions. Adapted with permission from160. b The topography of the dominant fMRI gradient is evolutionarily conserved in the mouse86, macaque monkey162 and human162 cortex. Adapted with permission from86 © The Authors, some rights reserved; exclusive licensee AAAS. Distributed under a CC BY-NC 4.0 License (http://creativecommons.org/licenses/by-nc/4.0/”). Adapted with permission from162. c Reciprocal organization of unimodal-polymodal and modality-specific gradient in the three species. Gradients are color coded by fMRI networks. DMN, default mode network; Visual, visual network; Auditory, auditory network; Somatomotor, somatomotor network. Adapted with permission from86 © The Authors, some rights reserved; exclusive licensee AAAS. Distributed under a CC BY-NC 4.0 License (http://creativecommons.org/licenses/by-nc/4.0/”). Adapted with permission from162.
Collectively, these lines of evidence suggest that the spatial topology of cortical fMRI connectivity along preordered evolutionarily-relevant gradients is a fundamental organizational principle of mammalian fMRI connectivity across species86,162,164,166 Above and beyond the use of network-specific inferences, this organizational axis may also provide valuable evolutionarily-relevant landmark coordinates to enable functional alignment of fMRI networks across the phylogenetic tree (Fig. 3).
fMRI spatiotemporal modes
Our discussion of fMRI connectivity has so far been based on canonical descriptions of interareal fMRI coupling as a time-invariant measure of static functional connectivity. However, the brain is inherently a dynamical system, and a large body of evidence over the past ten years has demonstrated that the correlation structure of fMRI signal continually evolves over a time scale of seconds167,168,169,170,171. fMRI networks dynamics can be mapped using different computational approaches that are referred to under the umbrella term “dynamic functional connectivity” (dFC). A key goal of these approaches is the identification of recurring fMRI network configuration whose occurrence and temporal sequence determine the steady state structure of “static” fMRI connectivity networks172. Using dFC based on sliding correlation windows, the dynamic nature of fMRI connectivity networks have been initially identified in humans167,168,169, NHPs173 and later in rodents174.
While a thorough description of all the contemporary approaches used in dFC is beyond the scope of this review (see172), here we focus on a set of recently developed dFC methods, as they have proven effective in revealing evolutionarily relevant principles in the dynamic organization of fMRI networks across species175 The framework of these methods is designed to describe the intrinsic dynamics of BOLD fMRI signal with voxel-resolution (i.e. without the necessity of a brain parcellation), in non-correlational terms. Theoretical and experimental work suggest that resting fMRI activity emerges from instances of segregation between networks interspersed with transient states of temporally overlapping network interactions176,177,178. These dynamic states can be effectively captured by spatially clustering individual fMRI frames into recurring coactivation patterns (CAPs)177,179. or, alternatively, by mapping converging spatio-temporal patterns of fMRI activity termed Quasi-Periodic Patterns—QPPs180. Both these methods do not rely on second-order statistics (correlations), nor the arbitrary choice of a sliding-window, in addition to maintaining the highest temporal resolution allowed by the sampling of rsfMRI data172. These features are especially attractive for cross-species fMRI mapping as they allow for a direct comparison of dynamic fMRI state topography unconstrained by predetermined parcellation schemes.
Whole-brain CAP mapping, first described in humans179 and then further expanded in a mouse study181, represent a powerful framework to probe the organization of fMRI dynamics across species. Initial investigations of CAPs as the basis for fMRI dynamics have shown that time-varying fMRI connectivity is governed by a limited number of co-fluctuating network configurations, whose occurrence accounts for more than 60% of the variance in fMRI timeseries181. As CAPs capture peaks and throughs of recurring spatiotemporal patterns of BOLD activity, spatially opposed pairs of CAPs can be merged to describe coactivation modes (C-modes), i.e. recurring spatiotemporal patterns of infraslow BOLD activity59. Crucially, the topography and dynamic organization of C-modes encompass key evolutionarily relevant features that appear to be conserved across the phylogenetic tree59. The most prominent of these, is the topographic organization of a dominant coactivation mode of BOLD activity extending from somatomotor areas to midline components of the DMN, hence spatially reconstituting the principal unimodal-transmodal cortical gradient mapped with static fMRI connectivity (Fig. 4a). Similar patterns of fMRI BOLD activity have been described using QPPs and complex principal component analysis in humans182. These findings are of great importance, as they suggest that the microarchitectural organization of cortical areas critically shapes both the static and the dynamic fMRI connectome, a notion that has been formally explored more in detail in recent rodent86 and human studies182. This observation is also of interest in the light of the natural emergence of functional anticorrelation between somatomotor and DMN systems, a finding that has been observed in multiple species2,100,176. Consistent with the emergence of these topographies as a species-invariant phenomenon, the occurrence of C-modes in humans, NHP and mice is time-locked to the infraslow cycle of global fMRI signal59 (Fig. 4a, inset), and encompass network topographies that can be tracked across evolution in multiple species (Fig. 4b)59,106. Specifically, the presence of four phylogenetically related principal C-modes in NHP and humans, three of which exhibited plausible topographic correlates also in the mouse brain was recently demonstrated59. Figure 4b shows the prevalent C-mode in humans and its correspondent network topographies in macaques and mice. Beyond the C-mode framework, phylogenetically-relevant correspondences have also been described in anesthetized mice and macaques using region-of-interest approaches108, or via the detection of QPPs in anesthetized rats, mice, and awake humans180,183,184. Collectively, these features define an evolutionarily conserved set of dynamic rules governing fMRI network dynamics and offer opportunities to map and compare evolutionarily relevant dynamic patterns of fMRI activity across the phylogenetic tree.
a Resting state fMRI activity is dominated by recurring network configurations whose time-varying peaks and troughs can be effectively mapped using frame-wise clustering to produce CAP and anti-CAP pairs, respectively. Red and blue coloring represent peaks and troughs of coherent whole-brain BOLD signals, respectively. Collectively, CAPs and anti-CAPs define a set of evolutionary conserved coactivation modes (C-Modes) whose occurrence is linked to global fMRI signal phases (circular distributions in insets). b The spatial topography of C-Modes is largely conserved in awake humans, NHPs and rodents. Red and blue coloring on the mouse brains represent high and low values of BOLD signal, respectively (modified from59). Dashed lines denote the evolutionary trajectory of a network’s co-activation profile across species in the specified C-mode. aDMN anterior default mode network, pDMN posterior default mode network, SMN somatomotor network, VIS visual network, VAT ventral attention network.
Relating fMRI connectivity findings across species
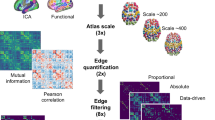
The identification of evolutionarily conserved principles underlying the organization of fMRI connectivity in the mammalian brain has prompted research aimed at comparing and relating fMRI network organization across the phylogenetic tree. This endeavor is part of the broader field of comparative neuroscience, an emergent area of investigation devoted to understanding how variation in the organization of different brains has unfolded across evolutionary history185,186. Attempts to relate fMRI connectivity across species are relatively recent and involve translation strategies characterized by different degrees of complexity. Univariate methods such as region-of-interest approaches (Fig. 5a) have been used to quantify and relate measurements of fMRI connectivity in pre-selected brain regions that are assumed to be homologous across species. This approach has been used to compare network organization and relate regional changes in fMRI connectivity across species both in healthy states37,117,118,187, as well as in models of brain disorders71,73,74. In these studies, regional homology has been typically established by selecting functionally related predefined regions of interest using parcellated anatomical atlases. Although useful in comparing network or connectivity attributes that can be quantified at the regional level, this simple approach fails to capture the multivariate nature of fMRI connectivity and as such it is not optimally suited to highlight the relationship between areas that underlie these interactions.
a Region-wise approaches are used to quantify rsfMRI connectivity across evolutionarily conserved brain regions. Green indicates the approximate location of visual cortex in mouse, NHP and human brain. b Network-level approaches can be used to relate whole fMRI network patterns of connectivity across species via fingerprints encompassing an extended set of evolutionarily relevant anchor points125. c Fusion-based approaches extend these methods to align fMRI connectivity between species (here referred to NPH, or rodents) using joint embedding of fMRI connectivity per se, or complementary measurements that covary with brain anatomy and connectivity. For example, functional connectivity can be measured in brain regions showing similar profiles of gene expression across species. A major advantage of this approach is the possibility of interpolating the whole cortical space via in between a set of evolutionarily conserved anchor regions.
At the network level, region of interest approaches can be extended to map fMRI connectivity across multiple regions belonging to the same network, or that are centered around a specific hub region, with the aim to identify and compare fMRI network fingerprints186 (Fig. 5b). A key advantage of such multivariate methods is that they allow for a comparison of whole-networks, and their corresponding connectivity profiles (i.e. interaction with regions within and outside the network) in a semi-quantitative way. This in turn may lead to a precise estimation of the evolutionary trajectory of specific network systems across the evolutionary landscape125,185,186. Importantly, the obtained functional interactions can then be related to their underlying anatomical connectivity using white matter fiber bundles188 or axonal wiring125, an approach that is particularly powerful in mice where the mesoscale connectome is available85 and can be directly compared to corresponding whole brain rsfMRI measurements86. An interesting extension of this approach is to propose novel functional homologies based on cross-species similarity of inter-regional functional connectivity fingerprints. For example, this method has been recently used to identify a cortical region homolog to macaque area F6 in the human brain, under the assumption that the best candidate for homology would be the human region exhibiting the functional connectivity fingerprint most similar to that observed in the macaque brain186. Interestingly, network level approaches can be used to carry out interspecies comparisons using sensory-driven fMRI responses. By using naturalistic visual stimulation, multivariate methods revealed topologically related convergences extending beyond the primary visual cortex between humans and macaque monkeys189, and humans and marmosets190.
Moreover, this approach can also encompass multivariate metrics of functional connectivity beyond Pearson’s correlation, thus extending this use to more complex measures of regional interaction and communication191. Overall, network-based multivariate methods appear to be optimally suited to compare the topographic organization of specific networks across the phylogenetic tree, with the possible pitfall of requiring the identification and preselection of anchor regions that need to be directly relatable in evolutionarily terms across species. This may not be easily achievable for regions (such as the neocortex) that have greatly expanded and have undergone major reorganization during evolution.
To overcome this issue and allow functional matching across species beyond evolutionarily conserved brain regions, recent years have seen the emergence of a set of complementary methods enabling the quantification of functional connectivity in homologous regions between humans and NHPs, even when their location is decoupled from brain anatomy. These approaches rely on the mono- or multimodal fusion of brain data to create a joint coordinate space, enabling a spatially continuous remapping of evolutionarily relevant anatomical or functional features across the phylogenetic tree. For example, by realigning cortical connectivity gradients of humans and macaque monkeys, joint embedding has been used to capture the common mesoscale brain functional architecture of the two species162. Evolutionarily conserved intrinsic coordinate systems can be also built with non-functional data and/or at other scales of investigation. In a recent study153, spectral embedding was used to align the cortices of pairs of closely related mammalian species, by creating an average feature space that retains shared geometrical and anatomical properties for each pair of species. The iteration of this approach over 90 brains of extant species enabled a reconstruction of the evolutionary distribution of primary sensory areas across the phylogenetic tree, ranging from ancestral mammals up to non-human primates. Fusion spaces like the one described by Schwartz and colleagues153 could in principle be used to map and compare fMRI connectivity networks within a unified coordinate system. Finally, leveraging the striking conservation of genetic architecture across mammals, recent attempts to relate brainwide transcriptomic data across species have been described. In one of such studies, regional transcriptomic landmarks in the mouse have been linked to corresponding measures in humans, thus providing a spatial index of anatomical homology between these two species192. This research revealed that, as expected, subcortical districts (e.g. caudoputamen) and sensorimotor subdivisions of the neocortex exhibit greater similarity between species compared with polymodal subdivisions of the cerebral cortex. Future extensions of fusion approaches to include multimodal datasets193 and fMRI connectivity-based metrics (such as gradients, or dynamic topographies) will offer further opportunities to relate anatomical and functional landmarks across species with increased precision (Fig. 5c).
Challenges of cross-species fMRI
In the preceding sections we have emphasized resemblances in the organization and dynamics of fMRI connectivity networks in rodent, NHPs and humans. Noteworthy evolutionary convergence exists in network organization across species, offering opportunities to extrapolate the results of mechanistic research along the phylogenetic tree, and narrow the “explanatory gap” that exists between microscale modeling and systems levels investigation of brain activity in humans (Fig. 1).
Moving along the phylogenetic tree
While increasingly sophisticated approaches are now available to map and compare fMRI connectivity networks and related mechanistic findings across species, a few methodological and conceptual caveats need to be carefully accounted for when extrapolating findings along the evolutionary axis. A first hurdle on the path of a reliable translation of network findings across species, is our fragmentary understanding of how cortical regions have evolved along the phylogenetic tree. This knowledge gap complicates the comparison of functional connectivity results that rely on the use of brain regions that are assumed to be functionally homologous across species. This problem is exemplified by the long-standing debate regarding the existence of a “prefrontal cortex” in rodents194,195. Supporting the presence of a prototypical medial prefrontal cortex in rats and mice, plausible cytoarchitectural convergence in the organization of the anterior cingulate have been described in rodents and humans112. This finding is consistent with the neuroanatomical organization of the rodent DMN, which includes the whole cingulate cortex as rostral hub60. However, the functional relevance of these regions and their role as bona fide substrates of the supramodal cognitive functions typically associated with the (medial) prefrontal cortex in humans remain debated196. These controversies underscore the need of exercising caution when specific neocortical locations, and in particular evolutionarily recent polymodal areas, are used as specific anatomical anchor-points for comparing fMRI connectivity networks across species.
Cross-species mapping based on qualitative approaches that are minimally dependent on discrete brain parcellations represents an attractive extension of region-based approaches. One example is to compare whole network systems, e.g. the entire DMN, as opposed to the sole medial prefrontal cortex, or the somatomotor network as opposed to specific motor-sensory sub-domains. The assumption here is that whole fMRI networks can embody more reliably cognate cognitive/behavioral functions than specific brain regions of unclear evolutionarily relevance. The efficacy of this simple approach has been recently demonstrated in neuropsychiatry research, where network-based inferences have facilitated mouse to human extrapolation of connectivity alterations of relevance to autism71,73,74.
Other methods have expanded this strategy to enable comparisons of RSN organization across species, while minimizing potential bias related to pre-imposed neuroanatomical or network constraints. A notable example of this approach is the recent work from Xu and colleagues162 in which cross-species matching of fMRI connectivity gradients has been used to project macaque network systems onto human cortical surface. Leveraging evolutionarily conserved hierarchical organization of cortical connectivity (Fig. 3), this work has documented the possibility of improving alignment and interpretation of imaging datasets across species through the use of common coordinate systems162. Extensions of this approach down the phylogenetic tree to include rodents are foreseeable, and could be facilitated by the incorporation of intermediate species characterized by lissencephalic neocortical architecture such as marmoset monkeys140.
Perhaps the most obvious, yet pervasive, caveat to consider when relating fMRI connectivity attributes across the evolutionarily timelines is the fact that higher order cortical functions in human and NHPs have undergone a conspicuous volumetric expansion during evolution. For these reasons, many of these functions (e.g. vocal communication, problem solving) are less specialized or underdeveloped in lower mammalian species, or are subserved by broad cortical substrates that are less specialized or just absent in NHPs and rodents103,197. This problem follows the general observation that size and complexity of brain regions are not merely proportional to the variation of total brain volume, area, and cortical folding across evolution198. Subcortical systems are somewhat less affected by this issue, as the organizational principles and neural wiring underlying these regions are phylogenetically older, and core to a number of lower-level behavioral domains and physiological responses that appear to be more strongly preserved across evolution. Nonetheless, examples of non-marginal differences in the organization of these systems across species have been reported. For one, basal ganglia connectivity networks exhibit only partly conserved connectivity profiles in rodents and NHPs125. Further comparative investigations of subcortical systems are required to provide a proper phylogenetic contextualization of fMRI connectivity topography of non-cortical systems across species.
fMRI in rodents and non-human primates: is anesthesia the lesser evil?
A methodological consideration of key importance when comparing or extrapolating fMRI connectivity changes across species is the widespread use of light anesthesia and sedation in rodent and NHP imaging. The use of anesthesia follows the need to minimize stress responses and head motion artifacts in fMRI scanning of non-compliant animal species199,200. The convenience and ease of use of anesthetized imaging along with the possibility of obtaining long-time series virtually devoid of motion artefacts have greatly contributed to the recent expansions of animal fMRI over the last decade60,201. By contrast, human imaging of fMRI connectivity mapping is carried out in awake resting conditions, without the use of sedatives or pharmacological agents.
These procedural discrepancies come with two potential problems: first, differences in fMRI network organization between animals and humans may be compounded by state-related changes related to the use of anesthesia in animals. A second possible issue pertains to the use of multiple anesthetic and sedative combinations in different species, each of which may bias the functional organization of fMRI network in different ways (reviewed in202). As a result, fMRI network comparisons between animal species may suffer from anesthetic-related bias.
While these issues complicate inferences that can be made at the level of networks across species, retrospective analyses of the fMRI literature suggests that their impact can be mitigated by choice of anesthetic regimes that preserve network organization100,199,203. Specifically, converging evidence suggests that several sedative regimens in rodents and NHP broadly preserve the functional organization of RSNs with respect to the structure of underlying brain anatomy37,86,106,135,204. Analogous observations have been reported in humans, where differences in awake versus sedated states produced by medetomidine appeared to be small, and very focal compared to the broad extension of brain networks in awake and sedated state205. Moreover, the spatial association between functional and axonal connectivity in anesthetized rodents and NHPs is robust86,131, recapitulating the general good agreement between these quantities observed in awake humans4.
These observations suggest that the use of anesthesia per se does not prevent meaningful comparisons of network features across species, provided the anesthetic protocol is judiciously chosen and validated with reference to the underlying anatomical structure of brain networks. fMRI imaging under light sedation is thus a practice that is commonly used in the preclinical fMRI community71,74,89,111, as it combines readiness of use with the potential of recording long time-series with negligible motion artifacts.
Importantly, the possibility of obtaining robust fMRI connectivity mapping with sedation in animals does not imply that pharmacological agents negligibly affect the organization of brain connectivity, or its underlying neurovascular coupling. Important interactions between anesthesia and hemodynamic signals have indeed been reported206,207,208,209 which may confound the interpretation and topological organization of fMRI networks. Moreover, different anesthetic mixtures can result in different patterns on connectivity100,210, and may exert different effects across species208. Owing to these differences, it is important that mechanistic studies, whenever possible, report the generalizability of findings across anesthetic regimens, or their extension to awake conditions16,106. Of equal importance, the administration of mixtures of anesthetic agents, a procedure frequently employed in both rodent and NHP studies203, can present latent interpretational and technical difficulties related to the stability of the ensuing neurophysiological parameters in multimodal investigations16. The main problem associated with the use of anesthetic cocktails is the presence of major physiological drifts in neural rhythms and brain states reflecting the pharmacokinetic properties of the employed drugs and ensuing receptor desensitization mechanisms211. This effect is often not patently detectable in terms of global network configuration but becomes apparent when fMRI is coupled with more sensitive measures of neural activity (such as electrophysiological recordings). In this respect, the application of single-anesthetic preparations200 may offer advantages over drug cocktails in terms of brain state stability.
To circumvent these problems, recent years have seen a major drive towards the implementation of fMRI connectivity mapping in awake NHP53,212 and rodents102,106,213. Most awake fMRI protocols in animals deal with the problem of head-related motion and restraint-induced stress via the use of progressive habituation to scanner environment and the use of customized helmet214, head or body-restraint devices106,215,216. In NHPs, these strategies have been recently combined with the use of naturalistic viewing paradigms, with very encouraging results in terms of reduced head motion, and the possibility of harmonizing states between human and NHP fMRI data189,190,217. These approaches have shown the possibility of reliably mapping fMRI networks with minimal contamination by motion and stress, paving the way for a broader use of awake rsfMRI mapping in the preclinical field. Not surprisingly, head-to-head comparisons of awake and anesthetized animals have revealed subtle but important differences in the organization of fMRI networks. Specifically, while the overall structure of RSN networks is broadly comparable between anesthesia and awake states, focal changes in fMRI network organizations have been consistently reported in rodents and primates53,106,218. These include a stronger involvement of arousal-related forebrain nuclei, an increased anti-correlation between network sub-systems, and a weakening of structure-function coupling in awake states such to maximize between-network communication53,106,218. Moreover, time-varying fMRI connectivity presents stereotypical trajectories that are predictive of the consciousness in rodents, NHP, and humans53,106,218,219,220. These are important factors that can aid the interpretation of results from investigations regarding rsfMRI network dynamics in humans and animal models.
As promising as awake animal imaging is, it also comes with some notable limitations. For example, head-fixation usually involves invasive surgeries that may result in susceptibility artefacts that may degrade image quality. Similarly, habituation to the scanner is typically achieved through a lengthy and progressive exposure to restraint, hence making the whole procedure long and labor intensive106,221. Awake imaging is also predictably associated with higher levels of head motion, hence potentially polluting the quality of resulting images. More importantly, even though corticosterone levels in head-fixed, scanner acclimated rodents suggest extensive habituation results in marginal stress levels106, questions remain as to whether the “resting-state” that is attainable upon scanner acclimation does actually recapitulate the quiet wakefulness that characterizes human fMRI experiments. In this respect, the differences observed between static fMRI network organization in awake and anesthetized humans205 seem to be qualitatively more focal and smaller than those found between awake and anesthetized rodents or NHP53,106, suggesting that this might not be the case. Thereby, the possibility that awake animal imaging entails considerably higher arousal conditions than human fMRI mapping should be given serious consideration and needs to be accounted for when comparing network topography across species. fMRI mapping in high arousal-related states could be especially noxious in brain stimulation studies, where stimulus-locked motion can occur in response to stimulus-unrelated arousing cues (such as light detection in optogenetics, or interoceptive signals), hence contaminating fMRI timeseries. Because of these issues, the use of anesthesia is a practice that still has (and will retain) value in the animal fMRI community, where it is regarded by some as lesser (methodological) evil, owing to its readiness of use and the high quality of the data obtainable. However, despite its growing pains, awake fMRI in animals will soon reach methodological maturity, and will thus soon steadily complement the use of anesthesia in animal fMRI.
Concluding remarks
Advancements in the implementation of fMRI connectivity mapping across species offer a privileged angle of investigation to probe the evolutionary trajectory or the mammalian brain at the network level, and to mechanistically decode the elusive physiological mechanisms governing large scale network organization. The recent inception of methods enabling reliable fMRI network mapping in rodents is a welcome addition to the methodological repertoire available to neuroscientists interested in these research themes.
Form a comparative neuroscience standpoint, major correspondences have been identified in the large-scale organization of fMRI networks across the mammalian realm, revealing phylogenetically conserved network systems (e.g. salience, default, motor-sensory etc.), a dominant axis of cortical fMRI connectivity, and topographically-consistent dominant fMRI substates. These general principles are overlaid onto species-specific systems that are evolutionarily more recent, and that as such cannot be straightforwardly translated along the phylogenetic axis. Notwithstanding these evolutionary constraints, and the inevitable growing pains of this relatively new field of comparative neuroscience, the future of cross-species fMRI is bright, and poised to greatly advance our understanding of the physiological basis and organizational principles of fMRI connectivity in health and disease.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
References
Power, J., Schlaggar, B. & Petersen, S. Studying brain organization via spontaneous fMRI signal. Neuron 84, 681–696 (2014).
Fox, M. D. et al. The human brain is intrinsically organized into dynamic, anticorrelated functional networks. Proc. Natl. Acad. Sci. 102, 9673–9678 (2005).
Raichle, M. E. The restless brain. Brain Connect. 1, 3–12 (2011).
Suárez, L. E., Markello, R. D., Betzel, R. F. & Misic, B. Linking structure and function in macroscale brain networks. Trends Cognit. Sci. 24, 302–315 (2020).
Deco G., Tononi G., Boly M., Kringelbach M. L. & Rolls, E. T. Rethinking segregation and integration: Contributions of whole-brain modelling. Nat. Rev. Neurosci. 16, 430–439 (2015).
Biswal, B., Zerrin Yetkin, F., Haughton, V. M. & Hyde, J. S. Functional connectivity in the motor cortex of resting human brain using echo‐planar MRI. Magn. Reson. Med. 34, 537–541 (1995).
Friston, K. J. Functional and effective connectivity in neuroimaging: a synthesis. Hum. Brain Mapp. 2, 56–78 (1994).
Smith S. M. et al. Correspondence of the brain’s functional architecture during activation and rest. Proc. Natl. Acad. Sci. 2, 56–78 (2009).
Smith, S. M. et al. Functional connectomics from resting-state fMRI. Trends Cogn. Sci. 17, 666–682 (2013).
Pezzulo, G., Zorzi, M. & Corbetta, M. The secret life of predictive brains: what’s spontaneous activity for? Trends Cogn. Sci. 25, 730–743 (2021).
Sudlow, C. et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Casey, B. J. et al. The adolescent brain cognitive development (ABCD) study: Imaging acquisition across 21 sites. Dev. Cogn. Neurosci. 32, 43–54 (2018).
Finn, E. S. et al. Functional connectome fingerprinting: identifying individuals using patterns of brain connectivity. Nat. Neurosci. 18, 1664–1671 (2015).
Whitfield-Gabrieli, S. & Ford, J. M. Default mode network activity and connectivity in psychopathology. Annu. Rev. Clin. Psychol. 8, 49–76 (2012).
Fornito, A. & Bullmore, E. T. Connectomics: A new paradigm for understanding brain disease. Eur. Neuropsychopharmacol. 2014/04/15, (2014).
Rocchi, F. et al. Increased fMRI connectivity upon chemogenetic inhibition of the mouse prefrontal cortex. Nat. Commun. 13, 1–15 (2022).
Raut R. V. et al. Global waves synchronize the brain’s functional systems with fluctuating arousal. Sci. Adv. (2021).
Liska, A. & Gozzi, A. Can mouse imaging studies bring order to Autism connectivity chaos? Front Neurosci. 10, 484 (2016).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Volkow, N. D. et al. The conception of the ABCD study: From substance use to a broad NIH collaboration. Dev. Cogn. Neurosci. 32, 4–7 (2018).
Moreau, C. A. et al. Mutations associated with neuropsychiatric conditions delineate functional brain connectivity dimensions contributing to autism and schizophrenia. Nat. Commun. 11, 5272 (2020).
Zhao B. et al. Common variants contribute to intrinsic human brain functional networks. Nat. Genet. 1-10 (2022).
Trachtenberg, A. J. et al. The effects of APOE on the functional architecture of the resting brain. Neuroimage 59, 565–572 (2012).
Schreiner, M. J. et al. Default mode network connectivity and reciprocal social behavior in 22q11. 2 deletion syndrome. Soc. Cogn. Affect. Neurosci. 9, 1261–1267 (2014).
Corbetta, M. Functional connectivity and neurological recovery. Dev. Psychobiol. 54, 239–253 (2012).
Grefkes, C. & Fink, G. R. Connectivity-based approaches in stroke and recovery of function. Lancet Neurol. 13, 206–216 (2014).
Siddiqi, S. H., Kording, K. P., Parvizi, J. & Fox, M. D. Causal mapping of human brain function. Nat. Rev. Neurosci. 23, 361–375 (2022).
Horn, A., Al-Fatly, B., Neumann, W-J & Neudorfer, C. Connectomic DBS: An introduction. In: Connectomic Deep Brain Stimulation (ed Horn A). Academic Press (2022).
Ramsey, L. E. et al. Normalization of network connectivity in hemispatial neglect recovery. Ann. Neurol. 80, 127–141 (2016).
Gozzi A. & Zerbi, V. Modelling brain dysconnectivity in rodents. Biol. Psychiatry, (2022).
Logothetis, N. K. Neural-Event-Triggered fMRI of large-scale neural networks. Curr. Opin. Neurobiol. 31, 214–222 (2015).
Logothetis, N. K. et al. The effects of electrical microstimulation on cortical signal propagation. Nat. Neurosci. 13, 1283–1291 (2010).
Logothetis, N. K. et al. Hippocampal-cortical interaction during periods of subcortical silence. Nature 491, 547–553 (2012).
Logothetis, N. K., Pauls, J., Augath, M., Trinath, T. & Oeltermann, A. Neurophysiological investigation of the basis of the fMRI signal. Nature 412, 150–157 (2001).
Magri, C., Schridde, U., Murayama, Y., Panzeri, S. & Logothetis, N. K. The amplitude and timing of the BOLD signal reflects the relationship between local field potential power at different frequencies. J. Neurosci. 32, 1395–1407 (2012).
Shmuel, A. & Leopold, D. A. Neuronal correlates of spontaneous fluctuations in fMRI signals in monkey visual cortex: Implications for functional connectivity at rest. Hum. Brain Mapp. 29, 751–761 (2008).
Vincent, J. L. et al. Intrinsic functional architecture in the anaesthetized monkey brain. Nature 447, 83–86 (2007).
Hutchison, R. M. & Everling, S. Monkey in the middle: why non-human primates are needed to bridge the gap in resting-state investigations. Front. Neuroanat. 6, 29 (2012).
Schölvinck, M. L., Maier, A., Frank, Q. Y., Duyn, J. H. & Leopold, D. A. Neural basis of global resting-state fMRI activity. Proc. Natl. Acad. Sci. 107, 10238–10243 (2010).
Wang, L., Saalmann, YuriB., Pinsk, MarkA., Arcaro, MichaelJ. & Kastner, S. Electrophysiological low-frequency coherence and cross-frequency coupling contribute to BOLD connectivity. Neuron 76, 1010–1020 (2012).
Chang, C. et al. Tracking brain arousal fluctuations with fMRI. Proc. Natl. Acad. Sci. 113, 4518 (2016).
Stephan, K. E. et al. Advanced database methodology for the Collation of Connectivity data on the Macaque brain (CoCoMac). Philos. Trans. R. Soc. Lond. Ser. B: Biol. Sci. 356, 1159–1186 (2001).
Markov, N. T. et al. A weighted and directed interareal connectivity matrix for macaque cerebral cortex. Cereb. Cortex 24, 17–36 (2014).
Majka, P. et al. Open access resource for cellular-resolution analyses of corticocortical connectivity in the marmoset monkey. Nat. Commun. 11, 1133 (2020).
Adachi, Y. et al. Functional connectivity between anatomically unconnected areas is shaped by collective network-level effects in the macaque cortex. Cereb. Cortex 22, 1586–1592 (2012).
Adam, R., Johnston, K., Menon, R. S. & Everling, S. Functional reorganization during the recovery of contralesional target selection deficits after prefrontal cortex lesions in macaque monkeys. Neuroimage 207, 116339 (2020).
O’Reilly, J. X. et al. Causal effect of disconnection lesions on interhemispheric functional connectivity in rhesus monkeys. Proc. Natl. Acad. Sci. 110, 13982–13987 (2013).
Turchi, J. et al. The basal forebrain regulates global resting-state fMRI fluctuations. Neuron 97, 940–952.e944 (2018).
Grayson, D. S. et al. The rhesus monkey connectome predicts disrupted functional networks resulting from pharmacogenetic inactivation of the Amygdala. Neuron 91, 453–466 (2016).
Folloni, D. et al. Manipulation of subcortical and deep cortical activity in the primate brain using transcranial focused ultrasound stimulation. Neuron 101, 1109–1116.e1105 (2019).
Verhagen, L. et al. Offline impact of transcranial focused ultrasound on cortical activation in primates. Elife 8, e40541 (2019).
Xu, A. G. et al. Focal infrared neural stimulation with high-field functional MRI: A rapid way to map mesoscale brain connectomes. Sci. Adv. 5, eaau7046 (2019).
Barttfeld, P. et al. Signature of consciousness in the dynamics of resting-state brain activity. Proc. Natl. Acad. Sci. 112, 887–892 (2015).
Hahn, G. et al. Signature of consciousness in brain-wide synchronization patterns of monkey and human fMRI signals. NeuroImage 226, 117470 (2021).
Xu, T. et al. Interindividual variability of functional connectivity in awake and anesthetized rhesus macaque monkeys. Biol. Psychiatry. Cogn. Neurosci. Neuroimaging 4, 543–553 (2019).
Logothetis, N. K., Guggenberger, H., Peled, S. & Pauls, J. Functional imaging of the monkey brain. Nat. Neurosci. 2, 555–562 (1999).
Cai, D.-C. et al.MECP2 duplication causes aberrant GABA pathways, circuits and behaviors in transgenic monkeys: Neural mappings to patients with autism.J. Neurosci.40, 3799–3814 (2020).
Grandjean, J. et al. Common functional networks in the mouse brain revealed by multi-centre resting-state fMRI analysis. Neuroimage 205, 116278 (2020).
Gutierrez-Barragan, D., Panzeri, S., Xu, T. & Gozzi, A. Evolutionarily conserved fMRI network dynamics in the human, macaque and mouse brain. Submitted, (2023).
Gozzi, A. & Schwarz, A. J. Large-scale functional connectivity networks in the rodent brain. Neuroimage 127, 496–509 (2016).
Grandjean J. et al. A consensus protocol for functional connectivity analysis in the rat brain. Nat. Neurosci. 1-9 (2023).
Zerbi V. et al. Rapid reconfiguration of the functional connectome after chemogenetic locus coeruleus activation. Neuron, (2019).
Oyarzabal, E. A. et al. Chemogenetic stimulation of tonic locus coeruleus activity strengthens the default mode network. Sci. Adv. 8, eabm9898 (2022).
Giorgi, A. et al. Brain-wide mapping of endogenous serotonergic transmission via chemogenetic fMRI. Cell Rep. 21, 910–918 (2017).
Hiro Taiyo H. et al Optogenetic activation of dorsal raphe serotonin neurons induces a brain-wide response in reward network. bioRxiv, 2022.2008.2007.503074 (2022).
Grandjean, J. et al. A brain-wide functional map of the serotonergic responses toacute stress and fluoxetine. Nat. Commun. 10, 350 (2019).
Trakoshis, S. et al. Intrinsic excitation-inhibition imbalance affects medial prefrontal cortex differently in autistic men versus women. Elife 9, e55684 (2020).
Tu W., Ma Z., Ma Y., Dopfel D. & Zhang, N. Suppressing anterior cingulate cortex modulates default mode network and behavior in awake rats. Cerebral Cortex, (2020).
Zerbi, V. et al. Dysfunctional autism risk genes cause circuit-specific connectivity deficits with distinct developmental trajectories. Cereb. Cortex 28, 2495–2506 (2018).
Pagani M. et al. Deletion of autism risk gene Shank3 disrupts prefrontal connectivity. J. Neurosci. 39, 5299-5310 (2019).
Pagani, M. et al. mTOR-related synaptic pathology causes autism spectrum disorder-associated functional hyperconnectivity. Nat. Commun. 12, 1–15 (2021).
Haberl, M. G. et al. Structural-functional connectivity deficits of neocortical circuits in the Fmr1GêÆ/y mouse model of autism. Sci. Adv. 1, e1500775 (2015).
Shofty, B. et al. Autism-associated Nf1 deficiency disrupts corticocortical and corticostriatal functional connectivity in human and mouse. Neurobiol. Dis. 130, 104479 (2019).
Bertero, A. et al. Autism-associated 16p11.2 microdeletion impairs prefrontal functional connectivity in mouse and human. BRAIN 141, 2055–2065 (2018).
Balasco L. et al. Abnormal whisker-dependent behaviors and altered cortico-hippocampal connectivity in Shank3b−/− mice. Cerebral Cortex, (2021).
Awad P. N. et al. CDKL5 sculpts functional callosal connectivity to promote cognitive flexibility. Mol. Psychiatry, 1–12 (2023).
Liska, A. et al. Homozygous loss of autism-risk gene CNTNAP2 results in reduced local and long-range prefrontal functional connectivity. Cereb. Cortex 10, 1–13 (2018).
Zerbi V. et al. Brain mapping across 16 autism mouse models reveals a spectrum of functional connectivity subtypes. Mol. Psychiatry, 26, 7610–7620 (2021).
Xu, N. et al. Functional connectivity of the brain across rodents and humans. Front. Neurosci. 16, 816331 (2022).
Tudela, R., Muñoz-Moreno, E., Sala-Llonch, R., López-Gil, X. & Soria, G. Resting state networks in the TgF344-AD rat model of Alzheimer’s disease are altered from early stages. Front. Aging Neurosci. 11, 213 (2019).
Green, C. et al. Functional networks are impaired by elevated tau-protein but reversible in a regulatable Alzheimer’s disease mouse model. Mol. Neurodegeneration 14, 1–13 (2019).
Vasilkovska, T. et al. Resting-state fMRI reveals longitudinal alterations in brain network connectivity in the zQ175DN mouse model of Huntington’s disease. Neurobiol. Dis. 181, 106095 (2023).
Domínguez-Iturza, N. et al. The autism-and schizophrenia-associated protein CYFIP1 regulates bilateral brain connectivity and behaviour. Nat. Commun. 10, 3454 (2019).
Huang, S.-M. et al. Inter-strain differences in default mode network: a resting state fmri study on spontaneously hypertensive rat and Wistar Kyoto rat. Sci. Rep. 6, 21697 (2016).
Oh, S. W. et al. A mesoscale connectome of the mouse brain. Nature 508, 207–214 (2014).
Coletta, L. et al. Network structure of the mouse brain connectome with voxel resolution. Sci. Adv. 6, eabb7187 (2020).
Grandjean, J., Zerbi, V., Balsters, J. H., Wenderoth, N. & Rudin, M. Structural basis of large-scale functional connectivity in the mouse. J. Neurosci. 37, 8092–8101 (2017).
Whitesell, J. D. et al. Regional, layer, and cell-type-specific connectivity of the mouse default mode network. Neuron 109, 545–559.e548 (2021).
Tsai, P. J. et al. Converging structural and functional evidence for a rat salience network. Biol. Psychiatry 88, 867–878 (2020).
Ma, Z., Zhang, Q., Tu, W. & Zhang, N. Gaining insight into the neural basis of resting-state fMRI signal. Neuroimage 250, 118960 (2022).
Pais-Roldán, P. et al. Indexing brain state-dependent pupil dynamics with simultaneous fMRI and optical fiber calcium recording. Proc. Natl. Acad. Sci. 117, 6875–6882 (2020).
Menon, V. et al. Optogenetic stimulation of anterior insular cortex neurons in male rats reveals causal mechanisms underlying suppression of the default mode network by the salience network. Nat. Commun. 14, 866 (2023).
Chao, T.-H. H. et al. Neuronal dynamics of the default mode network and anterior insular cortex: Intrinsic properties and modulation by salient stimuli. Sci. Adv. 9, eade5732 (2023).
Lake, E. M. et al. Simultaneous cortex-wide fluorescence Ca2+ imaging and whole-brain fMRI. Nat. methods 17, 1262–1271 (2020).
Mandeville, J. B. et al. Evidence of a cerebrovascular postarteriole windkessel with delayed compliance. JCerebBlood Flow. Metab. 19, 679–689 (1999).
Leite, F. P. et al. Repeated fMRI using iron oxide contrast agent in awake, behaving macaques at 3 Tesla. Neuroimage 16, 283–294 (2002).
Xu, T. et al. Delineating the macroscale areal organization of the macaque cortex in vivo. Cell Rep. 23, 429–441 (2018).
Goense, J. B., Zappe, A.-C. & Logothetis, N. K. High-resolution fMRI of macaque V1. Magn. Reson. imaging 25, 740–747 (2007).
Wu, E. X., Wong, K. K., Andrassy, M. & Tang, H. High‐resolution in vivo CBV mapping with MRI in wild‐type mice. Magn. Reson. Med. 49, 765–770 (2003).
Sforazzini, F., Schwarz, A. J., Galbusera, A., Bifone, A. & Gozzi, A. Distributed BOLD and CBV-weighted resting-state networks in the mouse brain. Neuroimage 87, 403–415 (2014).
Ghahremani, M., Hutchison, R. M., Menon, R. S. & Everling, S. Frontoparietal functional connectivity in the common marmoset. Cereb. Cortex 27, 3890–3905 (2017).
Liang, Z., King, J. & Zhang, N. Uncovering intrinsic connectional architecture of functional networks in awake rat brain. J. Neurosci. 31, 3776–3783 (2011).
Garin, C. M. et al. An evolutionary gap in primate default mode network organization. Cell Rep. 39, 110669 (2022).
Jonckers, E., Van Audekerke, J., De Visscher, G., Van der Linden, A. & Verhoye, M. Functional connectivity fMRI of the rodent brain: comparison of functional connectivity networks in rat and mouse. PloS one 6, e18876 (2011).
Hutchison, R. M. et al. Resting-state networks in the macaque at 7 Tesla. NeuroImage 56, 1546–1555 (2011).
Gutierrez-Barragan, D. et al. Unique spatiotemporal fMRI dynamics in the awake mouse brain. Curr. Biol. 32, 631–644.e636 (2022).
Liska, A., Galbusera, A., Schwarz, A. J. & Gozzi, A. Functional connectivity hubs of the mouse brain. Neuroimage 115, 281–291 (2015).
Mandino, F. et al. A triple-network organization for the mouse brain. Mol. Psychiatry 27, 865–872 (2022).
Buckner, R. L. & Vincent, J. L. Unrest at rest: Default activity and spontaneous network correlations. NeuroImage 37, 1091–1096 (2007).
Mantini, D. et al. Default mode of brain function in monkeys. J. Neurosci. 31, 12954–12962 (2011).
Lu, H. et al. Rat brains also have a default mode network. Proc. Natl. Acad. Sci. USA 109, 3979–3984 (2012).
Vogt, B. A. & Paxinos, G. Cytoarchitecture of mouse and rat cingulate cortex with human homologies. Brain Struct. Funct. 219, 185–192 (2012).
Buckner, R. L., Andrews-Hanna, J. R. & Schacter, D. L. The brain’s default network. Ann. NY. Acad. Sci. 1124, 1–38 (2008).
Lyamzin, D. & Benucci, A. The mouse posterior parietal cortex: Anatomy and functions. Neurosci. Res. 140, 14–22 (2019).
Corbetta, M. & Shulman, G. L. Control of goal-directed and stimulus-driven attention in the brain. Nat. Rev. Neurosci. 3, 201–215 (2002).
Stoet, G. & Snyder, L. H. Neural correlates of executive control functions in the monkey. Trends Cogn. Sci. 13, 228–234 (2009).
Mantini, D., Corbetta, M., Romani, G. L., Orban, G. A. & Vanduffel, W. Evolutionarily novel functional networks in the human brain? J. Neurosci. 33, 3259–3275 (2013).
Schwarz, A. J. et al. The low-frequency blood oxygenation level-dependent functional connectivity signature of the hippocampal-prefrontal network in the rat brain. Neuroscience 228, 243–258 (2013).
Liang, Z., Li, T., King, J. & Zhang, N. Mapping thalamocortical networks in rat brain using resting-state functional connectivity. NeuroImage 83, 237–244 (2013).
Zerbi, V., Grandjean, J., Rudin, M. & Wenderoth, N. Mapping the mouse brain with rs-fMRI: An optimized pipeline for functional network identification. NeuroImage 123, 11–21 (2015).
Alves, P. N. et al. An improved neuroanatomical model of the default-mode network reconciles previous neuroimaging and neuropathological findings. Commun. Biol. 2, 370 (2019).
Zhang, S. & Li, C. R. Functional connectivity parcellation of the human thalamus by independent component analysis. Brain Connect 7, 602–616 (2017).
Kim, D. J., Park, B. & Park, H. J. Functional connectivity-based identification of subdivisions of the basal ganglia and thalamus using multilevel independent component analysis of resting state fMRI. Hum. Brain Mapp. 34, 1371–1385 (2013).
Hwang, K., Bertolero, M. A., Liu, W. B. & D’Esposito, M. The human thalamus is an integrative hub for functional brain networks. J. Neurosci. 37, 5594–5607 (2017).
Balsters, J. H., Zerbi, V., Sallet, J., Wenderoth, N. & Mars, R. B. Primate homologs of mouse cortico-striatal circuits. Elife 9, e53680 (2020).
Yacoub, E. et al. Ultra-high field (10.5 T) resting state fMRI in the macaque. Neuroimage 223, 117349 (2020).
Liu, X. et al. Functional parcellation of human and macaque striatum reveals human-specific connectivity in the dorsal caudate. Neuroimage 235, 118006 (2021).
Honey, C. J. et al. Predicting human resting-state functional connectivity from structural connectivity. Proc. Natl. Acad. Sci. 106, 2035–2040 (2009).
Hagmann, P. et al. Mapping the structural core of human cerebral cortex. PLoS Biol. 6, 1479–1493 (2008).
Honey, C. J., Kötter, R., Breakspear, M. & Sporns, O. Network structure of cerebral cortex shapes functional connectivity on multiple time scales. Proc. Natl Acad. Sci. USA 104, 10240–10245 (2007).
Hori, Y. et al. Comparison of resting-state functional connectivity in marmosets with tracer-based cellular connectivity. NeuroImage 204, 116241 (2020).
Shen, K. et al. A macaque connectome for large-scale network simulations in TheVirtualBrain. Sci. Data 6, 123 (2019).
Goulas, A., Majka, P., Rosa, M. G. P. & Hilgetag, C. C. A blueprint of mammalian cortical connectomes. PLOS Biol. 17, e2005346 (2019).
Harris, K. D. & Shepherd, G. M. The neocortical circuit: themes and variations. Nat. Neurosci. 18, 170–181 (2015).
Hori, Y. et al. Altered resting-state functional connectivity between awake and isoflurane anesthetized marmosets. Cereb. Cortex 30, 5943–5959 (2020).
Valk, S. L. et al. Genetic and phylogenetic uncoupling of structure and function in human transmodal cortex. Nat. Commun. 13, 2341 (2022).
Paquola, C. & Hong, S.-J. The potential of myelin-sensitive imaging: Redefining spatiotemporal patterns of myeloarchitecture. Biol. psychiatry 93, 442–454 (2023).
Pang, J. C. et al. Geometric constraints on human brain function. Nature 618, 566–574 (2023).
Ma, Z. et al. Functional atlas of the awake rat brain: A neuroimaging study of rat brain specialization and integration. Neuroimage 170, 95–112 (2018).
Belcher A. M. et al. Functional connectivity hubs and networks in the awake marmoset brain. Front. Integrative Neurosci. 10, 10–19 (2016).
Buckner, R. L. et al. Cortical hubs revealed by intrinsic functional connectivity: Mapping, assessment of stability, and relation to Alzheimer’s Disease. J. Neurosci. 29, 1860–1873 (2009).
Cole, M. W., Pathak, S. & Schneider, W. Identifying the brain’s most globally connected regions. NeuroImage 49, 3132–3148 (2010).
van den Heuvel, M. P., Scholtens, L. H. & de Reus, M. A. Topological organization of connectivity strength in the rat connectome. Brain Struct. Funct. 221, 1719–1736 (2016).
Rubinov, M., Ypma, R. J. F., Watson, C. & Bullmore, E. T. Wiring cost and topological participation of the mouse brain connectome. Proc. Natl. Acad. Sci. 112, 10032–10037 (2015).
Harriger L., Van Den Heuvel M. P. & Sporns, O. Rich club organization of macaque cerebral cortex and its role in network communication. PLoS One 7, e46497 (2012).
Van Den Heuvel, M. P. & Sporns, O. Rich-club organization of the human connectome. J. Neurosci. 31, 15775–15786 (2011).
Jones, E. G. & Powell, T. P. S. An anatomical study of converging sensory pathways within the cerebral cortex of the monkey. Brain 93, 793–820 (1970).
Mesulam, M. The evolving landscape of human cortical connectivity: Facts and inferences. NeuroImage 62, 2182–2189 (2012).
Hilgetag, C. C. & Goulas, A. Hierarchy’ in the organization of brain networks. Philos. Trans. R. Soc. Lond. B Biol. Sci. 375, 20190319 (2020).
Mesulam, M.-M. From sensation to cognition. Brain: A J. Neurol. 121, 1013–1052 (1998).
Buckner, R. L. & Krienen, F. M. The evolution of distributed association networks in the human brain. Trends Cogn. Sci. 17, 648–665 (2013).
Krubitzer, L. In search of a unifying theory of complex brain evolution. Ann. NY Acad. Sci. 1156, 44–67 (2009).
Schwartz, E. et al. Evolution of cortical geometry and its link to function, behaviour and ecology. Nat. Commun. 14, 2252 (2023).
Burt, J. B. et al. Hierarchy of transcriptomic specialization across human cortex captured by structural neuroimaging topography. Nat. Neurosci. 21, 1251–1259 (2018).
Froudist-Walsh S. et al. Gradients of receptor expression in the macaque cortex. BioRxiv, 2021.2002. 2022.432173 (2021).
Fulcher, B. D., Murray, J. D., Zerbi, V. & Wang, X.-J. Multimodal gradients across mouse cortex. Proc. Natl. Acad. Sci. 116, 4689–4695 (2019).
Hilgetag, C. C. & Goulas, A. Hierarchy’in the organization of brain networks. Philos. Trans. R. Soc. B 375, 20190319 (2020).
Huntenburg, J. M., Bazin, P. L. & Margulies, D. S. Large-scale gradients in human cortical organization. Trends Cogn. Sci. 22, 21–31 (2018).
Wang, X. J. Macroscopic gradients of synaptic excitation and inhibition in the neocortex. Nat. Rev. Neurosci. 21, 169–178 (2020).
Margulies, D. S. et al. Situating the default-mode network along a principal gradient of macroscale cortical organization. Proc. Natl. Acad. Sci. 113, 12574–12579 (2016).
Demirtaş, M. et al. Hierarchical heterogeneity across human cortex shapes large-scale neural dynamics. Neuron 101, 1181–1194.e1113 (2019).
Xu, T. et al. Cross-species functional alignment reveals evolutionary hierarchy within the connectome. Neuroimage 223, 117346 (2020).
Tong, C. et al. Multimodal analysis demonstrating the shaping of functional gradients in the marmoset brain. Nat. Commun. 13, 6584 (2022).
Ngo, G. N., Hori, Y., Everling, S. & Menon, R. S. Joint-embeddings reveal functional differences in default-mode network architecture between marmosets and humans. NeuroImage 272, 120035 (2023).
Hilgetag, C. C., Goulas, A. & Changeux, J.-P. A natural cortical axis connecting the outside and inside of the human brain. Netw. Neurosci. 6, 950–959 (2022).
Huntenburg, J. M., Yeow, L. Y., Mandino, F. & Grandjean, J. Gradients of functional connectivity in the mouse cortex reflect neocortical evolution. NeuroImage 225, 117528 (2021).
Hutchison, R. M. et al. Dynamic functional connectivity: Promise, issues, and interpretations. NeuroImage 80, 360–378 (2013).
Allen, E. A. et al. Tracking whole-brain connectivity dynamics in the resting state. Cereb. Cortex 24, 663–676 (2012).
Chang, C. & Glover, G. H. Time–frequency dynamics of resting-state brain connectivity measured with fMRI. Neuroimage 50, 81–98 (2010).
Keilholz, S. D. The neural basis of time-varying resting-state functional connectivity. Brain Connect. 4, 769–779 (2014).
Di, X. & Biswal, B. B. Dynamic brain functional connectivity modulated by resting-state networks. Brain Struct. Funct. 220, 37–46 (2015).
Lurie, D. J. et al. Questions and controversies in the study of time-varying functional connectivity in resting fMRI. Netw. Neurosci. 4, 30–69 (2020).
Hutchison, R. M., Womelsdorf, T., Gati, J. S., Everling, S. & Menon, R. S. Resting-state networks show dynamic functional connectivity in awake humans and anesthetized macaques. Hum. Brain Mapp. 34, 2154–2177 (2013).
Grandjean, J. et al. Dynamic reorganization of intrinsic functional networks in the mouse brain. NeuroImage 152, 497–508 (2017).
Gutierrez-Barragan, D., Ramirez, J. S. B., Panzeri, S., Xu, T. & Gozzi, A. Evolutionarily conserved fMRI network dynamics in the mouse, macaque, and human brain. bioRxiv, 2023.2007. 2019.549681 (2023).
Fasoli, D., Coletta, L., Gutierrez-Barragan, D., Gozzi, A. & Panzeri, S. A model of the mouse cortex with attractor dynamics explains the structure and emergence of rsfMRI co-activation patterns. bioRxiv, (2022).
Karahanoğlu, F. I. & Van De Ville, D. Transient brain activity disentangles fMRI resting-state dynamics in terms of spatially and temporally overlapping networks. Nat. Commun. 6, 775 (2015).
Tagliazucchi, E., Balenzuela, P., Fraiman, D. & Chialvo, D. R. Criticality in large-scale brain FMRI dynamics unveiled by a novel point process analysis. Front Physiol. 3, 15 (2012).
Liu, X., Chang, C. & Duyn, J. H. Decomposition of spontaneous brain activity into distinct fMRI co-activation patterns. Front. Syst. Neurosci. 7, (2013).
Yousefi, B., Shin, J., Schumacher, E. H. & Keilholz, S. D. Quasi-periodic patterns of intrinsic brain activity in individuals and their relationship to global signal. Neuroimage 167, 297–308 (2018).
Gutierrez-Barragan, D., Basson, M. A., Panzeri, S. & Gozzi, A. Infraslow state fluctuations govern spontaneous fMRI network dynamics. Curr. Biol. 29, 2295–2306.e2295 (2019).
Bolt, T. et al. A parsimonious description of global functional brain organization in three spatiotemporal patterns. Nat. Neurosci. 25, 1093–1103 (2022).
Belloy, M. E. et al. Dynamic resting state fMRI analysis in mice reveals a set of Quasi-Periodic Patterns and illustrates their relationship with the global signal. Neuroimage 180, 463–484 (2018).
Majeed, W. et al. Spatiotemporal dynamics of low frequency BOLD fluctuations in rats and humans. Neuroimage 54, 1140–1150 (2011).
Mars, R. B., Jbabdi, S. & Rushworth, M. F. S. A common space approach to comparative neuroscience. Annu Rev Neurosci. 44, 69–86 (2021).
Mars, R. B. et al. Comparing brains by matching connectivity profiles. Neurosci. Biobehav. Rev. 60, 90–97 (2016).
Bergmann, E., Zur, G., Bershadsky, G. & Kahn, I. The organization of mouse and human cortico-hippocampal networks estimated by intrinsic functional connectivity. Cereb. Cortex 26, 4497–4512 (2016).
Mars, R. B. et al. Whole brain comparative anatomy using connectivity blueprints. Elife 7, e35237 (2018).
Mantini, D. et al. Interspecies activity correlations reveal functional correspondence between monkey and human brain areas. Nat. Methods 9, 277–282 (2012).
Hori, Y. et al. Interspecies activation correlations reveal functional correspondences between marmoset and human brain areas. Proc. Natl. Acad. Sci. 118, e2110980118 (2021).
Griffa A. et al. The evolution of information transmission in mammalian brain networks. bioRxiv, 2022.2005.2009.491115 (2022).
Beauchamp, A. et al. Whole-brain comparison of rodent and human brains using spatial transcriptomics. eLife 11, e79418 (2022).
Markello, R. D. et al. neuromaps: structural and functional interpretation of brain maps. Nat. Methods 19, 1472–1479 (2022).
Preuss, T. M. Do rats have prefrontal cortex? The rose-woolsey-akert program reconsidered. J. Cogn. Neurosci. 7, 1–24 (1995).
Carlén, M. What constitutes the prefrontal cortex? Science 358, 478–482 (2017).
Laubach M., Amarante L. M., Swanson K. & White, S. R. What, If Anything, Is Rodent Prefrontal Cortex? eNeuro 5, ENEURO.0315-0318.2018 (2018).
Buckner R. L. & Krienen, F. M. The evolution of distributed association networks in the human brain. Trends Cognit. Sci. 17, 648-665.
Mota, B. & Herculano-Houzel, S. Cortical folding scales universally with surface area and thickness, not number of neurons. Science 349, 74–77 (2015).
Mandino, F. et al. Animal functional magnetic resonance imaging: Trends and path toward standardization. Front Neuroinform 13, 78 (2019).
Ferrari, L. et al. A robust experimental protocol for pharmacological fMRI in rats and mice. J. Neurosci. Methods 204, 9–18 (2012).
Grandjean J. et al. StandardRat: A multi-center consensus protocol to enhance functional connectivity specificity in the rat brain. bioRxiv, (2022).
Mandino, F. et al. Animal functional magnetic resonance imaging: trends and path toward standardization. Front. Neuroinf. 13, 78 (2020).
Grandjean J. et al. StandardRat: A multi-center consensus protocol to enhance functional connectivity specificity in the rat brain. bioRxiv, 2022.2004.2027.489658 (2022).
Liang, Z., King, J. & Zhang, N. Intrinsic organization of the anesthetized brain. J. Neurosci. 32, 10183–10191 (2012).
Akeju, O. et al. Disruption of thalamic functional connectivity is a neural correlate of dexmedetomidine-induced unconsciousness. eLife 3, e04499 (2014).
Rungta, R. L., Osmanski, B.-F., Boido, D., Tanter, M. & Charpak, S. Light controls cerebral blood flow in naive animals. Nat. Commun. 8, 14191 (2017).
Gozzi, A., Schwarz, A. J., Reese, T., Crestan, V. & Bifone, A. Drug-anaesthetic interaction in phMRI: The case of the pyschotomimetic agent phencyclidine. Magn. Reson Imag. 26, 999–1006 (2008).
Sirmpilatze, N. et al. Spatial signatures of anesthesia-induced burst-suppression differ between primates and rodents. Elife 11, e74813 (2022).
Zhang, Z. et al. Isoflurane-induced burst suppression increases intrinsic functional connectivity of the monkey brain. Front. Neurosci. 13, 296 (2019).
Grandjean, J., Schroeter, A., Batata, I. & Rudin, M. Optimization of anesthesia protocol for resting-state fMRI in mice based on differential effects of anesthetics on functional connectivity patterns. NeuroImage 102, 838–847 (2014).
Nasrallah, F. A., Tay, H. C. & Chuang, K. H. Detection of functional connectivity in the resting mouse brain. NeuroImage 1, 417–424 (2014).
Klink, P. C. et al. Combining brain perturbation and neuroimaging in non-human primates. Neuroimage 235, 118017 (2021).
Stenroos P. et al. Awake rat brain functional magnetic resonance imaging using standard radio frequency coils and a 3D printed restraint kit. Front. Neurosci. 12, 548 (2018).
Hung, C.-C. et al. Functional MRI of visual responses in the awake, behaving marmoset. Neuroimage 120, 1–11 (2015).
Han, Z. et al. Awake and behaving mouse fMRI during Go/No-Go task. NeuroImage 188, 733–742 (2019).
Yoshida, K. et al. Physiological effects of a habituation procedure for functional MRI in awake mice using a cryogenic radiofrequency probe. J. Neurosci. Methods 274, 38–48 (2016).
Russ, B. E. & Leopold, D. A. Functional MRI mapping of dynamic visual features during natural viewing in the macaque. Neuroimage 109, 84–94 (2015).
Demertzi, A. et al. Human consciousness is supported by dynamic complex patterns of brain signal coordination. Sci. Adv. 5, eaat7603 (2019).
Ma, Y., Hamilton, C. & Zhang, N. Dynamic connectivity patterns in conscious and unconscious brain. Brain Connectivity 7, 1–12 (2017).
Huang, Z., Zhang, J., Wu, J., Mashour, G. A. & Hudetz, A. G. Temporal circuit of macroscale dynamic brain activity supports human consciousness. Sci. Adv. 6, eaaz0087 (2020).
Chen, G. et al. Functional magnetic resonance imaging of awake monkeys: Some approaches for improving imaging quality. Magn. Reson. imaging 30, 36–47 (2012).
Majka, P. et al. Towards a comprehensive atlas of cortical connections in a primate brain: Mapping tracer injection studies of the common marmoset into a reference digital template. J. Comp. Neurol. 524, 2161–2181 (2016).
Wildenberg, G. A. et al. Primate neuronal connections are sparse in cortex as compared to mouse. Cell Rep. 36, 109709 (2021).
Feng, G. et al. Opportunities and limitations of genetically modified nonhuman primate models for neuroscience research. Proc. Natl. Acad. Sci. 117, 24022–24031 (2020).
Stephan M., Volkmann P. & Rossner, M. J. Assessing behavior and cognition in rodents, nonhuman primates, and humans: where are the limits of translation? Dialog. Clin. Neurosci. (2022).
Yu, Y. et al. Sleep fMRI with simultaneous electrophysiology at 9.4 T in male mice. Nat. Commun. 14, 1651 (2023).
Thomas Yeo, B. et al. The organization of the human cerebral cortex estimated by intrinsic functional connectivity. J. Neurophysiol. 106, 1125–1165 (2011).
Acknowledgements
This work has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation program (#DISCONN; no 802371 to AG). AG also acknowledges funding from the Telethon foundation (GGP19177), Simons Foundation (SFARI 982347). MP is supported by European Union’s Horizon 2020 research and innovation programme (Marie Sklodowska-Curie Global Fellowship - CANSAS, GA845065). EDG is supported by the Canadian Institute of Health Research (Postdoctoral fellowship—MFE187902). TX is supported by Brain Initiative National Institutes of Health (RF1MH128696).
Author information
Authors and Affiliations
Contributions
We confirm contribution to the paper as follows: study conception: A.G.; literature review: M.P., D.G.-B., E.D.G., T.X., A.G. draft manuscript preparation: M.P., D.G.B., E.D.G., A.G. All authors reviewed the results and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Julie Royo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Michel Thiebaut de Schotten and Joao Valente. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pagani, M., Gutierrez‐Barragan, D., de Guzman, A.E. et al. Mapping and comparing fMRI connectivity networks across species. Commun Biol 6, 1238 (2023). https://doi.org/10.1038/s42003-023-05629-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-05629-w
- Springer Nature Limited