Abstract
Our nervous system contains billions of neurons that form precise connections with each other through interactions between cell surface proteins. In Drosophila, the Dpr and DIP immunoglobulin protein subfamilies form homophilic or heterophilic interactions to instruct synaptic connectivity, synaptic growth, and cell survival. However, the upstream regulatory mechanisms of Dprs and DIPs are not clear. On the other hand, while transcription factors have been implicated in target recognition, their downstream cell surface proteins remain mostly unknown. We conduct an F1 dominant modifier genetic screen to identify regulators of Dprs and DIPs. We identify huckebein (hkb), a transcription factor previously implicated in target recognition of the dorsal Is motor neuron. We show that hkb genetically interacts with DIP-α and loss of hkb leads to complete removal of DIP-α expression specifically in dorsal Is motor neurons. We then confirm that this specificity is through the dorsal Is motor neuron specific transcription factor, even-skipped (eve), which acts downstream of hkb. Analysis of the genetic interaction between hkb and eve reveals that they act in the same pathway to regulate dorsal Is motor neuron connectivity. Our study provides insight into the transcriptional regulation of DIP-α and suggests that distinct regulatory mechanisms exist for the same CSP in different neurons.
Similar content being viewed by others
Introduction
The way animals perceive and respond to the environment relies on precise and robust neuronal connections. During development, each neuron must identify the correct synaptic partners among thousands of potential targets. A prevalent model for instructing synaptic recognition, repulsion, and self-avoidance is through the interaction between unique cell surface proteins (CSPs). A major subset of CSPs belong to the immunoglobulin superfamily (IgSF), which play important roles in synaptic development and maintenance in both vertebrates and invertebrates. In the well-studied vertebrate retina, retinal ganglion cells require Dscams and Sidekicks (Sdks) 1 and 2 to avoid self-synapses and form stereotyped connections, respectively1,2,3. In hard-wired invertebrate nervous systems, such as C. elegans, the heterophilic interaction between two IgSF proteins, Syg1 and Syg2, is required for HSNL motor neuron (MN) synapse formation4,5. In the Drosophila mushroom body, neurons utilize different isoforms of Dscam1 to discriminate self/non-self 6,7,8. Several IgSF CSPs have also been implicated in synaptic connectivity in the Drosophila larval neuromuscular system where two type-Is motor neurons (Is MNs) and ~29 type-Ib motor neurons (Ib MNs) form stereotyped connections with 30 muscles in each hemisegment9,10,11. For example, the immunoglobulin proteins Fasciclin 212,13 and Fasciclin 314,15 are required for specific larval MNs to recognize their muscle targets.
Recent biochemical studies revealed two Drosophila immunoglobulin protein subfamilies, the Defective proboscis response proteins (Dprs, 21 members) and Dpr-interacting proteins (DIPs, 11 members) families, that form homophilic or heterophilic interactions to instruct synaptic connectivity, synaptic growth, and cell survival16,17,18,19,20,21,22,23,24,25,26,27,28,29. For example, the well-studied Dpr10-DIP-α interaction is necessary for innervation of the dorsal Is MN on muscle 4 (m4) as loss of either dpr10 or DIP-α leads to complete loss of m4-Is innervation24. In addition, loss of dpr10 or DIP-α in the optic lobe causes significant mistargeting and cell death of Dm12 medulla neurons, suggesting multifaceted roles for Dpr10-DIP-α interactions19,25. Similarly, the recognition between yellow R7 photoreceptors (yR7) and yellow Dm8 neurons (yDm8) relies on the complementary expression of Dpr11 and DIP-γ, respectively, and the lack of either dpr11 or DIP-γ leads to the failure of yR7 and yDm8 to recognize each other and subsequent yDm8 cell death16,30,31. Although extensive studies have uncovered roles for Dpr-DIP interactions, the regulation and downstream mechanisms are severely understudied.
Transcription factors (TFs) are the fate determinants of all cell types, and in neurons, they are master regulators of synaptic wiring by determining the expression of many factors including CSPs. In the fly olfactory system, cell type-specific expression of the TF Acj6 controls expression of a cell-surface code that instructs neurons to identify correct synaptic partners32,33. In the visual system, the homeodomain TF, Brain-specific homeobox (Bsh), directly binds to the DIP-β locus and other L4 identity genes to specify L4 neuronal fate29. In addition, stochastic expression of Spineless (ss) determines the yR7 fate and controls the expression of dpr11, which is required for synaptic connectivity with yellow Dm8s30. Similarly, during embryonic development, several key TFs specify the neuroblast (NB) lineages, including huckebein (hkb)34,35,36,37, which is detected in 8 NB lineages and is required for the expression of the cell fate marker, even-skipped (eve), in NB4-238,39,40. In hkb mutant embryos, RP2 MNs (also known as the dorsal Is MNs) derived from NB4-2 show severe wiring defects as they do not reach the correct muscles, suggesting that hkb controls specific CSPs for synaptic recognition in dorsal Is MNs39. However, unlike Acj6, Bsh, and Ss, the CSP(s) downstream of Hkb that control synaptic recognition are not known.
In this study, we sought to identify genes involved in connectivity by developing a sensitized genetic background with known wiring CSPs that could be modified. Homozygous loss of dpr10 and DIP-α led to complete loss of innervation of m4 by the dorsal Is MNs but DIP-α/+;dpr10/+ trans-heterozygous larvae showed a 50% reduction of m4-Is innervation frequency. This trans-heterozygous background was used for an F1 deficiency screen to identify dominant enhancers or suppressors of Dpr10/DIP-α-mediated connectivity. We screened deficiency lines from the Bloomington Deficiency Kit covering the right arm of the third chromosome41,42, and within one interacting line, we identified hkb as a genetic regulator of DIP-α. DIP-α is expressed in both dorsal and ventral Is MNs, but interestingly, we found that hkb is only necessary for DIP-α expression and MN-muscle recognition in the dorsal Is MN, suggesting distinct regulatory mechanisms for DIP-α in dorsal and ventral Is MNs. Next, we showed that hkb functions through the dorsal Is MN specific TF, eve, as DIP-α expression and dorsal Is MN innervation are also disrupted in eve mutants. Genetic interaction tests between hkb and eve further confirmed that they act in the same pathway. In summary, our study reveals that Hkb acts through Eve to control DIP-α gene expression to regulate MN-muscle connectivity, bridging the gap between upstream TFs and downstream CSPs. Moreover, our study suggests that distinct regulatory mechanisms exist for the same CSP in different neurons.
Results
Genetic screen identifies hkb, a genetic interactor of Dpr10-DIP-α pathway
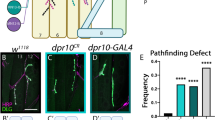
The Drosophila larval neuromuscular system provides an ideal model to study genetic programs that instruct synaptic recognition due to the ease of genetic manipulation and the stereotyped connectivity patterns. Each larval body wall hemisegment is innervated by one ventral and one dorsal Is MN that connect to the ventral or dorsal muscle groups, respectively, in a stereotyped manner (Fig. 1a). In a previous study, we showed that among all MNs, DIP-α is expressed exclusively in these two Is MNs, and its interacting partner, Dpr10, is expressed in a subset of muscles10,24. The interaction between Dpr10 and DIP-α is required for the recognition between dorsal Is MNs and several dorsal muscles. Specifically, loss of either dpr10 or DIP-α leads to complete loss of dorsal Is MN innervation on m4, suggesting that Dpr10-DIP-α interaction is absolutely required for m4-Is innervation. This easily scorable phenotype prompted us to ask what other genes are involved in this Dpr10-DIP-α-dependent synaptic recognition. Because loss of either CSP results in complete loss of m4-Is connectivity and single heterozygotes have either no or very mild phenotypes (see below), we created a sensitized genetic background in which one copy of dpr10 and DIP-α are removed. In this background, m4-Is innervation frequency is reduced to ~51% (Fig. 1b), compared to a 90% m4-Is innervation frequency in wild type animals24. We chose a dpr10 CRISPR (dpr10CR) allele and a GAL4 insertion allele of DIP-α (DIP-α-GAL4) derived from a MiMIC line, which disrupts endogenous DIP-α transcription and translation. Together with a UAS-2xEGFP construct, this DIP-α-GAL4 allele aids identification of Is MN axons and neuromuscular junctions (NMJs) on different muscles since DIP-α is exclusively expressed in Is MNs (Fig. 1b). The reduced m4-Is innervation frequency in the sensitized background allowed us to screen for genetic interactors of the Dpr10-DIP-α pathway by introducing other mutations – if a mutation exacerbates or suppresses the decreased Is MN innervation on m4, we hypothesize that the gene may be part of the Dpr10-DIP-α pathway.
a Cartoon depicting the innervation pattern of dorsal Is MN and ventral Is MN. b Representative images of muscle 4 with Is innervation or without Is innervation, in trans-heterozygotes of DIP-α (the DIP-α-GAL4 is also a null allele) and dpr10. 51% of m4s are innervated by the dorsal Is MN. GFP (green), DLG (magenta) and HRP (blue) are shown in the images. Arrow pointing to the Is NMJ. c Workflow of the deficiency screen. Male flies carrying the deficiency chromosome were crossed to females with DIP-α and dpr10 mutations. Female third instar larvae were selected against the second and third chromosome balancers (CyO,actin > GFP and TM6,Tb,Hu). Triple-heterozygous larvae were dissected and Is innervation frequency on m4, 3, 12 and 13 were scored. Cartoon is created with BioRender.com.
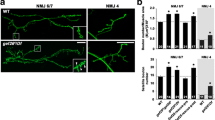
To improve the throughput, we utilized the Bloomington Deficiency (Df) kit and conducted an F1 dominant modifier screen (Fig. 1c). We screened 105 Df lines that cover the entire Drosophila chromosome 3R. Each Df line was combined into the sensitized background to create triple heterozygotes and the m4-Is innervation frequency was quantified (Fig. 2a). In addition, we also quantified the innervation frequency of Is MNs on other muscles, including dorsal muscle 3 (m3) and ventral muscle 12 (m12) and muscle 13 (m13) (Fig. 2b–d). Although the Is innervation frequencies on these muscles were not significantly decreased in the sensitized background, we hypothesized that genetic interactors or redundant molecules of the Dpr10-DIP-α pathway may be uncovered. Compared to the sensitized background (red columns), we identified several Df lines that significantly increased or decreased Is innervation frequency (yellow columns) (Fig. 2). The ED5100 Df line reduced m4-Is innervation frequency most significantly (p < 0.01, Chi-square test), but did not affect Is innervation frequency on m12, m13 and m3 (p > 0.05, Chi-square test), suggesting that it covers a gene(s) that may positively regulate the Dpr10-DIP-α pathway in the dorsal Is MNs for m4-Is recognition.
Is innervation frequency on (a) m4, (b) m3, (c) m12 and (d) m13. The red column indicates the control innervation frequency from the sensitized background (trans-heterozygotes of DIP-α and dpr10). Gray columns are non-significant from control whereas the yellow columns are the deficiency lines that show significantly different innervation frequencies compared to control. The cut-off p values are indicated by dashed lines. Asterisk indicates ED5100.
ED5100 is a 900 kb deletion that spans several genes and long non-coding RNAs. To narrow down the genomic region that covers our gene(s) of interest, we conducted a sub-screen using additional Df lines, ED5142 and ED5046, which partially overlap with the deletion in ED5100 (Fig. 3a). We observed a similar decrease of m4-Is innervation frequency when the sensitized line was crossed to ED5046, but not to ED5142 (Fig. 3b), suggesting that our gene(s) of interest is located in the region of ED5046 that overlaps with ED5100 but not ED5142, from 4197 kb to 4453 kb (Fig. 3a). This candidate region covers 58 genes including protein coding genes and non-coding RNAs. We further assayed 8 candidate genes with known or putative neuronal functions and an available mutant stock, including auxilin, abstrakt, complexin, vps24, hkb, contactin, tube and lost. We screened each candidate by combining a heterozygous mutant allele into our sensitized background and examined m4-Is innervation frequency. Of these candidates, we found that heterozygous loss of hkb (hkb2/+) exacerbated the m4-Is innervation defect when combined with the sensitized background (Fig. 3c), suggesting that hkb genetically interacts with the Dpr10-DIP-α pathway.
a Cartoon depicting the deleted regions within deficiency lines ED5100, ED5142, and ED5046. b Quantification shows a significant reduction of m4-Is innervation when combining ED5046, but not ED5142, with the sensitized background, suggesting the shared region between the original deficiency line (ED5100) and ED5046 covers the candidate gene(s). N (NMJs) = 177, 196, 155 and 155. p values are indicated. c Quantification of sub-screen of individual genes from candidate region shown in (a). Alleles used to create triple-heterozygotes are, auxD128, abs00620, cpxMI00784, vps24EY04708, hkb2, contG5080, tub2, lostEY11645. Note that huckebein (hkb2) further reduced m4-Is innervation frequency. N (NMJs) = 177, 138, 62, 58, 29, 135, 39, 77, 78 and 57. p values are indicated.
hkb genetically interacts with DIP-α, but not dpr10
Our sensitized background is heterozygous for both dpr10 and DIP-α. Therefore, hkb may genetically interact with either or both CSPs. Here, we examined genetic interaction between hkb and dpr10 or DIP-α in trans-heterozygous animals. We combined two different hkb mutant alleles (hkb2 or hkbA321R1) with a heterozygous dpr10 mutant or DIP-α mutant and examined the m4-Is innervation frequency. In this and the following experiments, we used a DIP-α CRISPR (DIP-αCR) allele since we will primarily focus on m4-Is innervation and no longer need to identify Is NMJs on different muscles using DIP-α-GAL4.
In wild type animals, m4s are innervated about 90% of the time by the dorsal Is MN, and single heterozygotes of dpr10 or hkb did not significantly decrease this innervation frequency (Fig. 4a). We then examined trans-heterozygotes of dpr10 and hkb and found that the m4-Is innervation frequency was not significantly changed compared to single heterozygotes (Fig. 4a), suggesting that hkb is not a genetic interactor for dpr10. In contrast, heterozygous loss of DIP-α reduced m4-Is innervation frequency to 71% (Fig. 4b), but the trans-heterozygotes of DIP-α and hkb further reduced the m4-Is innervation frequency to about 50%. Comparing the trans-heterozygous data with single heterozygotes (Fig. 4b) suggests that hkb and DIP-α are in the same genetic pathway. Overall, these data suggest that hkb genetically interacts with DIP-α but not dpr10.
a Genetic interaction assay between hkb and dpr10. Single heterozygotes of dpr10 or hkb did not have altered m4-Is innervation, and neither did the trans-heterozygotes. N (NMJs) = 93, 154, 83, 125, 102 and 149. p values are indicated. b Genetic interaction assay between hkb and DIP-α. Single heterozygotes of DIP-α or hkb had slightly decreased m4-Is innervation frequency, while the trans-heterozygotes showed a further reduction, suggesting that hkb genetically interacts with DIP-α. N (NMJs) = 114, 105, 106, 103, 99 and 103. p values are indicated.
hkb controls DIP-α expression in the dorsal Is MNs
Next, we asked how hkb genetically interacts with DIP-α. DIP-α is expressed in both dorsal and ventral Is MNs but not in muscles, and interestingly, prior studies found that hkb is expressed in NB4-2, which produces the dorsal Is MN38,39. Therefore, we wondered if the TF hkb is required for DIP-α expression in dorsal Is MNs. To visualize DIP-α expression, we used an endogenously tagged DIP-α-EGFP allele18. In wild type animals, DIP-α is highly expressed in both dorsal and ventral Is MNs by stage 16 (Fig. 5a, b, b’). However, in hkb2 mutant embryos, expression of DIP-α in dorsal Is MNs was completely lost (Fig. 5c). We confirmed loss of DIP-α in heteroallelic hkb mutant embryos (hkb2/hkbA321R1) (Fig. 5d). Notably, the dorsal Is MN marker, eve, was also lost in hkb mutants as hkb is required for eve expression (Fig. 5c, d)39. These findings could be explained by the loss of the dorsal Is MNs; however, prior studies confirmed that dorsal Is MNs remain in hkb mutants even though they are eve negative39,43. Therefore, the lack of DIP-α-EGFP is not due to missing Is MNs, but to the loss of hkb. In addition, DIP-α-EGFP was not affected in the ventral Is MNs (Fig. 5c’, d’) or in other DIP-α-EGFP positive neurons (arrowheads in Fig. 5), suggesting that hkb only controls DIP-α expression in the dorsal Is MNs. This result is consistent with the unchanged ventral Is MN innervation frequency for ED5100 in our genetic screen. Taken together, these data indicate that different mechanisms regulate the same CSP in different neurons, and that hkb is required for DIP-α expression specifically in the dorsal Is MNs.
a Cartoon depicting the focal planes in (b–d). Representative images of dorsal Is MN cell bodies (arrows in b) and ventral Is MN cell bodies (arrows in b’) labeled with GFP (green), Eve (red), and FasII (magenta) in control embryos. DIP-α-EGFP is expressed in both dorsal and ventral Is MNs. Note that there are also two interneurons on the dorsal side of each hemisegment that express DIP-α-EGFP (arrowheads in b). c’ Representative images of dorsal Is MN cell bodies (dashed circles in c) and ventral Is MN cell bodies (arrows in c’) in hkb2 mutant embryos. Eve and DIP-α-EGFP are missing in dorsal Is MNs, whereas DIP-α-EGFP expression in ventral Is MNs is not affected. In addition, DIP-α-EGFP expressing interneurons are not affected (arrowheads in c), suggesting that Hkb function is specific to dorsal Is MNs. Representative images of dorsal Is MN cell bodies (dashed circles in d) and ventral Is MN cell bodies (arrows in d’) in heteroallelic hkb mutant embryos (hkb2 /hkbA321R1). Eve and DIP-α-EGFP are missing in dorsal Is MNs, whereas DIP-α-EGFP expression in ventral Is MNs is not affected. In addition, DIP-α-EGFP expressing interneurons are not affected (arrowheads in d), confirming that Hkb function is specific to dorsal Is MNs.
hkb functions through eve to regulate DIP-α and dorsal Is MN innervation
As a TF, hkb may directly instruct DIP-α expression, or alternatively, function through other intermediate TFs. Prior studies reported that hkb is expressed early in the NB4-2 lineage, which gives rise to dorsal Is MNs, and turned off by stage 12, before synaptic recognition occurs in the neuromuscular system39. However, a recent study found that hkb continues to be expressed in dorsal Is MNs during larval development44. Therefore, we decided to differentiate between the direct or indirect models of regulation. In NB4-2, a well-studied role of hkb is to trigger expression of the fate determinant TF, eve. Loss of hkb completely abolished eve expression in dorsal Is MNs (Fig. 5) (ref. 39). Thus, we wondered if hkb functions through eve to regulate DIP-α expression. Utilizing the DIP-α-EGFP allele and a conditional eve knock-out (eveΔRN2) which only lacks eve in dorsal Is MNs and siblings43, we observed that DIP-α expression was lost in the dorsal Is MN, and the expression in ventral Is MNs was not affected, in both embryos and 1st instar larvae (Fig. 6a–e). Consequently, eve mutants lacked m4-Is innervation by the dorsal Is NMJs, whereas innervation by the ventral Is MNs was not affected (Fig. 6f–k). To further investigate the role of eve in synaptic recognition, we created eve and DIP-α trans-heterozygotes and examined m4-Is innervation frequency. Compared to single heterozygotes of eve or DIP-α, trans-heterozygotes significantly reduced m4-Is innervation frequency to about 60%, confirming that eve regulates m4-Is innervation by driving DIP-α expression in the dorsal Is MNs (Fig. 7a). Finally, we examined the trans-heterozygotes of hkb and eve and found that partial loss of both hkb and eve reduced m4-Is innervation frequency to 75%, suggesting that hkb and eve indeed act in the same pathway to control synaptic recognition of the dorsal Is MNs (Fig. 7b). Taken together, our results reveal a transcriptional cascade that regulates expression of wiring CSPs to guide MN-muscle recognition.
Representative images of dorsal Is MN cell bodies (arrows in a) and ventral Is MN cell bodies (arrows in a’) labeled with GFP (green), Eve (red), and FasII (magenta) in control embryos. DIP-α-EGFP is expressed in both dorsal and ventral Is MNs. Representative images of dorsal Is MN cell bodies (dashed circles in b) and ventral Is MN cell bodies (arrows in b’) in eveΔRN2 mutant embryos. Eve and DIP-α-EGFP are missing in dorsal Is MNs, whereas DIP-α-EGFP expression in ventral Is MNs is not affected. Representative images of dorsal Is MN cell bodies (arrows in c) and ventral Is MN cell bodies (arrows in c’) labeled with GFP (green), Eve (red), and HRP (magenta) in control 1st instar larvae. DIP-α-EGFP is expressed in both dorsal and ventral Is MNs. Representative images of dorsal Is MN cell bodies (dashed circles in d) and ventral Is MN cell bodies (arrows in d’) in eveΔRN2 mutant 1st instar larvae. Eve and DIP-α-EGFP are missing in dorsal Is MNs, whereas DIP-α-EGFP expression in ventral Is MNs is not affected. e Quantification of DIP-α-EGFP expression in dorsal Is MN cell bodies in control and eveΔRN2 mutant larvae. N (ROI with two cell bodies) = 25 and 25. p value is indicated. Error bar indicates standard error of the mean (SEM). Representative images of NMJs formed by Ib MN, dorsal Is MN, and ventral Is MN (arrows) labeled with GFP (green), DLG (red), HRP (magenta), and Phalloidin (Blue), in 1st instar DIP-α-EGFP expressing larvae in (f) control and (g) eveΔRN2 mutant background. h Quantification of m4-Is innervation frequency in 1st instar control and eveΔRN2 mutant larvae. N (NMJs) = 27 and 25. p value is indicated. Representative images of NMJs formed by the Ib MN, dorsal Is MN, and ventral Is MN (arrows) labeled with GFP (green), DLG (red), HRP (magenta), and Phalloidin (Blue), in 2nd instar DIP-α-EGFP expressing larvae in (i) control and (j) eveΔRN2 mutant background. k Quantification of m4-Is innervation frequency in 2nd instar control and eveΔRN2 mutant larvae. N (NMJs) = 53 and 44. p value is indicated.
a Genetic interaction assay between eve and DIP-α. Single heterozygotes of eve or DIP-α did not have altered m4-Is innervation, while the trans-heterozygotes showed a further reduction, suggesting that eve genetically interacts with DIP-α. N (NMJs) = 102, 104, 84 and 104. p values are indicated. b Genetic interaction assay between hkb and eve. Single heterozygotes of hkb or eve did not have altered m4-Is innervation, while the trans-heterozygotes showed a further reduction, suggesting that hkb genetically interacts with eve to regulate Is innervation. N (NMJs) = 103, 87, 105, 81, 112 and 86. p values are indicated.
Discussion
Synaptic recognition requires the interaction of CSPs, highlighting the critical role of regulatory programs to instruct the expression of CSPs in specific cells. Most studies have focused on the roles of TFs and CSPs independently, but less is known about the CSPs downstream of specific TFs in synaptic target recognition. For example, hkb and eve were implicated in pathfinding and target recognition of the dorsal Is MNs, but the molecules acting downstream of hkb and eve were unknown. On the other hand, the well-studied Dpr10-DIP-α interaction was found to guide the recognition between the dorsal Is MN and m4, but the regulatory mechanisms controlling the expression of these CSPs were not known24. To identify additional components in the Dpr10-DIP-α pathway, including transcriptional regulators, we conducted a dominant modifier genetic screen and found that hkb exacerbates the decrease of m4-Is innervation frequency when introduced into a DIP-α and dpr10 trans-heterozygous background. Notably, our genetic screen identified many Df lines and candidate genes that rescue the m4-Is innervation frequency. This unexpected finding suggests that genes in these Df lines may be repulsive cues, repressors for synaptic growth, or are involved in the pruning process. However, all of these candidates may not act in the Dpr10-DIP-α pathway. For example, the triple heterozygote of complexin (cpx), DIP-α, and dpr10 shows a wild type m4-Is innervation frequency (Fig. 3c), yet cpx mutants do not show connectivity phenotypes. Instead, loss of cpx revealed an increase of NMJ size and elevated activity45,46. The rescue in innervation observed in the triple heterozygote may be due to the perturbed synaptic activity which has been implicated in synaptic pruning47,48. A smaller subset of Df lines and candidate genes exacerbated the loss of m4-Is connectivity, and we chose to focus on these dominant modifiers. Specifically, we showed that hkb is required for DIP-α expression in dorsal Is MNs. Further examination revealed that hkb functions through eve to regulate DIP-α expression and dorsal Is MN innervation of m4, revealing a pathway linking TFs to specific CSPs and circuit assembly.
Interestingly, hkb is a gap gene originally implicated in embryonic development. In the early embryo, three pattern organizing centers, the anterior, the posterior, and the terminal, establish the anterior-posterior body plan by spatiotemporally regulating the expression of gap genes49. In the terminal control center, torso controls the expression of terminal gap genes including hkb34. hkb is expressed in the terminal cap during embryonic stages 5-6 and functions as a negative regulator to suppress gene expression in the terminal band, such as odd-paired (opa), Dichaete (D), and caudal (cad)50. In later embryonic stages, Hkb is expressed in a subset of NBs where it is required for glial development51, serotoninergic neuron differentiation52,53, and for Eve expression to control motor axon pathfinding39. However, the molecules downstream of Hkb and Eve for axon pathfinding are not known, and additionally, the role of Hkb after the motor axon pathfinding stage has not been examined likely due to the lethality of null mutant embryos. In this study, we identified hkb as a DIP-α genetic interactor, and utilizing trans-heterozygous hkb hypomorph animals, we found that hkb is required for DIP-α expression to instruct innervation by the dorsal Is MN. Notably, our data suggests that hkb indirectly regulates DIP-α expression through the cell fate determinant TF, Eve. However, in a previous ChIP-Seq study profiling Eve target genes, many CSPs required for synaptic development were found, but DIP-α was not identified54. This could be due to DIP-α only being expressed in a small subset of Eve-expressing cells, or more likely, because Eve also indirectly regulates DIP-α expression since it mostly functions as a transcriptional repressor43. Nevertheless, our genetic analyses focused on a single cell type and revealed regulatory relationships that would be obscured in sequencing-based profiling.
Excitatory MNs in Drosophila larvae are classified into type-Ib and type-Is MNs due to their terminal bouton size and innervation patterns. Notably, DIP-α is selectively expressed in the dorsal and ventral Is MNs but is absent in Ib MNs. We therefore initially hypothesized that a common regulatory program may be responsible for the expression of DIP-α in both Is MNs but absent in Ib MNs. However, we found that hkb and eve regulate DIP-α expression specifically in dorsal Is MNs. hkb and eve are not expressed in ventral Is MNs (derived from NB3-1)38, indicating that distinct mechanisms regulate DIP-α expression in different MNs. Single-cell transcriptomics or candidate approaches in ventral Is MNs will aid in identifying other TFs that instruct DIP-α expression.
DIP-α is a member of the Dpr/DIP subfamilies of immunoglobulin CSPs. In a previous study, we mapped the expression of dpr and DIP genes in larval MNs and found that dprs were shared among many MNs and DIPs were more selectively expressed10. Interestingly, each of the 33 MNs expresses a unique subset of dprs and DIPs to reveal a cell-specific cell surface code. These data suggest that a highly complex regulatory transcriptional network is required to instruct the expression of these CSPs. An alternative but not mutually exclusive model is that one TF may regulate different CSPs in distinct neurons. A recent study in the fly olfactory circuit described a divergent “transcription factor to CSPs” relationship where the same TF, acj6, regulates many different CSPs in different cell types32. Further identification of the TF network in the larval nervous system will help to understand how the TF code is transmitted into a CSP code to guide synaptic recognition.
In summary, the regulatory programs controlling expression of circuit wiring molecules are more complex than we hypothesized. The plethora of recent single-cell RNA-seq data will undoubtedly shed light on candidate TFs, but follow-up genetic analyses will be required to confirm causal relationships between TFs and CSPs that underlie circuit wiring.
Methods
Genetics
The following Drosophila lines were used in this study: w111816; DIP-α-GAL424; UAS-2×EGFP; dpr10CR19; DIP-αCR19; DIP-α-EGFP18; hkbA321R1 (BL#2059)35; hkb2 (BL#5457)55; eveΔRN243. Df lines discussed in this paper are: ED5100 (BL#9226), ED5046 (BL#9197), ED5142 (BL#9198). All lines used for screen and sub-screen are listed in Supplementary Table 141,42. For the genetic screen, males from the Df lines or mutant lines were crossed to sensitized females (DIP-α-GAL4; UAS-2×EGFP/CyO,actin > GFP; dpr10CR/TM6,Tb,Hu) and triple-heterozygous female larvae were selected to examine innervation frequency. For controls, w1118 males were crossed to sensitized females to create trans-heterozygotes of dpr10 and DIP-α. Screening workflow is illustrated in Fig. 1c (created with BioRender.com).
Dissection, immunofluorescence, and imaging
To examine Is MN innervation frequency, wandering first, second, and third instar larvae were dissected as previously described56. Briefly, larvae were collected and dissected on sylgard plates in PBS. Dissected fillets were fixed by 4% paraformaldehyde or Zamboni’s solution for 20 min at room temperature and then washed three times in PBT (PBS with 0.05% Triton X-100). Samples were then blocked for 1 h in 5% goat serum (5% goat serum diluted in PBT) and incubated with primary antibodies at 4 °C overnight. Primary antibodies were then removed, and samples were washed for three times before 2 h incubation at room temperature (or overnight at 4 °C) with secondary antibodies. Finally, secondary antibodies were washed out and samples were washed and mounted in Vectashield (Vector Laboratories).
To examine DIP-α expression in embryos, stage 16 embryos were dissected as previous described57. Egg-laying chambers were set up with 40 females and 30 males and capped with grape juice plates. Eggs were collected for 2 h and grape juice plates covered in embryos were placed at 25 °C for 16 h for development. Embryos were staged under a Zeiss V20 stereoscope using the autofluorescence and morphology of the gut. Embryos at the correct stage were dechorionated and transferred onto a Superfrost Plus slides (Thermo Fisher Scientific, #22-037-246) covered by PBS. Embryos were then dissected with an electrolytically sharpened tungsten wire and stained similar to third instar samples. Antibodies used in this study were: rabbit anti-GFP (1:40k, gift from Michael Glozter, University of Chicago); rabbit anti-Eve (1:1000, gift from Ellie Heckscher, University of Chicago); mouse anti-Dlg (1:100, Developmental Studies Hybridoma Bank [DSHB] #4F3); mouse anti-FasII (1:100, DSHB #1D4); goat anti-rabbit Alexa 488 (1:500, Invitrogen #A11008); goat anti-rabbit Alexa 568 (1:500, Invitrogen #A11036); goat anti-mouse Alexa 568 (1:500, Invitrogen #A11031); goat anti-mouse Alexa 647 (1:500, Invitrogen #A32728); goat anti-HRP Alexa 405 (1:100, Jackson Immunological Research #123-475-021); goat anti-HRP Alexa 647 (1:100, Jackson Immunological Research #123-605-021).
Images were acquired on a Zeiss LSM800 confocal microscope using a 40X plan-neofluar 1.3 NA objective, or a 63X plan-apo 1.4 NA objective. The same imaging parameters were applied to samples from the same set of experiments. Images were then analyzed and processed in ImageJ.
Quantification of Is MN innervation frequency
To examine Is MN innervation frequency, at least 6 third instar larvae were dissected and stained with anti-GFP (marker for Is MNs), anti-DLG and anti-HRP. Samples were visualized under a Zeiss AxioImager M2 scope with a Lumen light engine with a 20× Plan Apo 0.8 NA objective or an Olympus BX43 with an X-Cite 120LEDmini LED fluorescent illuminator. Is and Ib NMJs can be distinguished by bouton size, DLG intensity and whether it was GFP positive. If there was at least one Is bouton present on the muscle, it is scored as “innervated”, otherwise it is scored as “not innervated”. For each animal, m12, 13, 4 and 3 from abdominal hemisegments A2-A6 were assayed, and we collected a sample size of 50–60 hemisegments for each muscle. Innervation frequency was calculated as the percentage of “innervated” muscles in all muscles examined.
Quantification of DIP-α-EGFP expression
First instar larval brain pulls were conducted in PBS and mounted on poly-lysine coverslips. Intact brains were fixed and stained as described above. A Z-stack of the VNC was taken under 40X objective and a 2-slice projection covering the dorsal Is MN cell bodies was created in ImageJ (Sum slices). An area ROI was applied to the neuromere region spanning two MN cell bodies and EGFP and HRP intensity were measured. The same ROI size was used in this entire experiment. EGFP intensity was then normalized to HRP intensity to better reflect EGFP signal density.
Statistics and reproducibility
As we were mostly comparing the innervation frequency between two groups, we performed the Chi-square test followed by Yates’ correction using Prism 8 software. Innervation frequencies and p values were reported in the figure legends. For the comparison of DIP-α-EGFP expression level in Fig. 6, unpaired t test with Welch’s correction was performed (two-sided). Data were assumed to follow a Gaussian distribution.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The source data behind the graphs in the paper can be found in Supplementary Data 1.
References
Krishnaswamy, A., Yamagata, M., Duan, X., Hong, Y. K. & Sanes, J. R. Sidekick 2 directs formation of a retinal circuit that detects differential motion. Nature 524, 466–470 (2015).
Yamagata, M. & Sanes, J. R. Expression and roles of the immunoglobulin superfamily recognition molecule Sidekick1 in Mouse Retina. Front. Mol. Neurosci. 11, 485 (2019).
Garrett, A. M., Khalil, A., Walton, D. O. & Burgess, R. W. DSCAM promotes self-avoidance in the developing mouse retina by masking the functions of cadherin superfamily members. Proc. Natl Acad. Sci. 115, 201809430 (2018).
Shen, K., Fetter, R. D. & Bargmann, C. I. Synaptic specificity is generated by the synaptic guidepost protein SYG-2 and its receptor, SYG-1. Cell 116, 869–881 (2004).
Shen, K. & Bargmann, C. I. The immunoglobulin superfamily protein SYG-1 determines the location of specific synapses in C. elegans. Cell 112, 619–630 (2003).
Hattori, D. et al. Robust discrimination between self and non-self neurites requires thousands of Dscam1 isoforms. Nature 461, 644–648 (2009).
Zhan, X.-L. et al. Analysis of Dscam diversity in regulating axon guidance in drosophila mushroom bodies. Neuron 43, 673–686 (2004).
Wang, J. et al. Transmembrane/Juxtamembrane domain-dependent Dscam distribution and function during mushroom body neuronal morphogenesis. Neuron 43, 663–672 (2004).
Hoang, B. & Chiba, A. Single-cell analysis of drosophila larval neuromuscular synapses. Dev. Biol. 229, 55–70 (2001).
Wang, Y. et al. Systematic expression profiling of dprs and DIPs reveals cell surface codes in Drosophila larval motor and sensory neurons. Development 149, dev200355 (2022).
Lnenicka, G. A. & Keshishian, H. Identified motor terminals in Drosophila larvae show distinct differences in morphology and physiology. J. Neurobiol. 43, 186–197 (2000).
Davis, G. W., Schuster, C. M. & Goodman, C. S. Genetic analysis of the mechanisms controlling target selection: target-derived Fasciclin II regulates the pattern of synapse formation. Neuron 19, 561–573 (1997).
Winberg, M. L., Mitchell, K. J. & Goodman, C. S. Genetic analysis of the mechanisms controlling target selection: complementary and combinatorial functions of Netrins, Semaphorins, and IgCAMs. Cell 93, 581–591 (1998).
Chiba, A., Snow, P., Keshishian, H. & Hotta, Y. Fasciclin III as a synaptic target recognition molecule in Drosophila. Nature 374, 166–168 (1995).
Kose, H., Rose, D., Zhu, X. & Chiba, A. Homophilic synaptic target recognition mediated by immunoglobulin-like cell adhesion molecule Fasciclin III. Dev. Camb. Engl. 124, 4143–4152 (1997).
Carrillo, R. A. et al. Control of synaptic connectivity by a network of drosophila IgSF cell surface proteins. Cell 163, 1770–1782 (2015).
Sergeeva, A. P. et al. DIP/Dpr interactions and the evolutionary design of specificity in protein families. Nat. Commun. 11, 2125 (2020).
Tan, L. et al. Ig superfamily ligand and receptor pairs expressed in synaptic partners in drosophila. Cell 163, 1756–1769 (2015).
Xu, S. et al. Interactions between the Ig-Superfamily Proteins DIP-α and Dpr6/10 regulate assembly of neural circuits. Neuron 100, 1369–1384.e6 (2018).
Brovero, S. G. et al. Investigation of Drosophila fruitless neurons that express Dpr/DIP cell adhesion molecules. Elife 10, e63101 (2021).
Cheng, S. et al. Molecular basis of synaptic specificity by immunoglobulin superfamily receptors in Drosophila. Elife 8, e41028 (2019).
Cosmanescu, F. et al. Neuron-subtype-specific expression, interaction affinities, and specificity determinants of DIP/Dpr cell recognition proteins. Neuron 100, 1385–1400.e6 (2018).
Bornstein, B. et al. Transneuronal Dpr12/DIP‐δ interactions facilitate compartmentalized dopaminergic innervation of Drosophila mushroom body axons. Embo J. e105763 https://doi.org/10.15252/embj.2020105763 (2021).
Ashley, J. et al. Transsynaptic interactions between IgSF proteins DIP-α and Dpr10 are required for motor neuron targeting specificity. eLife 8, e42690 (2019).
Xu, S. et al. Affinity requirements for control of synaptic targeting and neuronal cell survival by heterophilic IgSF cell adhesion molecules. Cell Rep. 39, 110618 (2022).
Barish, S. et al. Combinations of DIPs and Dprs control organization of olfactory receptor neuron terminals in Drosophila. Plos Genet. 14, e1007560 (2018).
Özkan, E. et al. An extracellular interactome of immunoglobulin and LRR proteins reveals receptor-ligand networks. Cell 154, 228–239 (2013).
Venkatasubramanian, L. et al. Stereotyped terminal axon branching of leg motor neurons mediated by IgSF proteins DIP-α and Dpr10. Elife 8, e42692 (2019).
Xu, C., Ramos, T. B., Rogers, E. M., Reiser, M. B. & Doe, C. Q. Homeodomain proteins hierarchically specify neuronal diversity and synaptic connectivity. eLife 12, e90133 (2024).
Courgeon, M. & Desplan, C. Coordination between stochastic and deterministic specification in the Drosophila visual system. Science 366, eaay6727 (2019).
Menon, K. P., Kulkarni, V., Takemura, S., Anaya, M. & Zinn, K. Interactions between Dpr11 and DIP-γ control selection of amacrine neurons in Drosophila color vision circuits. Elife 8, e48935 (2019).
Xie, Q. et al. Transcription factor Acj6 controls dendrite targeting via a combinatorial cell-surface code. Neuron, https://doi.org/10.1016/j.neuron.2022.04.026 (2022).
Li, H. et al. Single-cell transcriptomes reveal diverse regulatory strategies for olfactory receptor expression and axon targeting. Curr. Biol. 30, 1189–1198.e5 (2020).
Weigel, D., Jürgens, G., Klingler, M. & Jäckle, H. Two gap genes mediate maternal terminal pattern information in drosophila. Science 248, 495–498 (1990).
Gaul, U. & Weigel, D. Regulation of Krüppel expression in the anlage of the Malpighian tubules in the Drosophila embryo. Mech. Dev. 33, 57–67 (1990).
Brönner, G. & Jäckle, H. Control and function of terminal gap gene activity in the posterior pole region of the Drosophila embryo. Mech. Dev. 35, 205–211 (1991).
Brönner, G. & Jäckle, H. Regulation and function of the terminal gap gene huckebein in the Drosophila blastoderm. Int. J. Dev. Biol. 40, 157–165 (1996).
McDonald, J. A. & Doe, C. Q. Establishing neuroblast-specific gene expression in the Drosophila CNS: huckebein is activated by Wingless and Hedgehog and repressed by Engrailed and Gooseberry. Dev. Camb. Engl. 124, 1079–1087 (1997).
Chu-LaGraff, Q. et al. huckebein specifies aspects of CNS precursor identity required for motoneuron axon pathfinding. Neuron 15, 1041–1051 (1995).
Broadus, J. et al. New neuroblast markers and the origin of the aCC/pCC neurons in the Drosophila central nervous system. Mech. Dev. 53, 393–402 (1995).
Cook, R. K. et al. The generation of chromosomal deletions to provide extensive coverage and subdivision of the Drosophila melanogaster genome. Genome Biol. 13, R21 (2012).
Roote, J. & Russell, S. Toward a complete Drosophiladeficiency kit. Genome Biol. 13, 149 (2012).
Fujioka, M. et al. Even-skipped, acting as a repressor, regulates axonal projections in Drosophila. Development 130, 5385–5400 (2003).
Jetti, S. K. et al. Molecular logic of synaptic diversity between Drosophila tonic and phasic motoneurons. Neuron 111, 3554–3569.e7 (2023).
Choi, B. J. et al. Miniature neurotransmission regulates drosophila synaptic structural maturation. Neuron 82, 618–634 (2014).
Banerjee, S. et al. Miniature neurotransmission is required to maintain Drosophila synaptic structures during ageing. Nat. Commun. 12, 4399 (2021).
Carrillo, R. A., Olsen, D. P., Yoon, K. S. & Keshishian, H. Presynaptic activity and CaMKII modulate retrograde semaphorin signaling and synaptic refinement. Neuron 68, 32–44 (2010).
Jarecki, J. & Keshishian, H. Role of neural activity during synaptogenesis in Drosophila. J. Neurosci. 15, 8177–8190 (1995).
Nasiadka, A., Dietrich, B. H. & Krause, H. M. Anterior-posterior patterning in the Drosophila embryo. Adv. Dev. Biol. Biochem. 12, 155–204 (2002).
Clark, E., Battistara, M. & Benton, M. A. A timer gene network is spatially regulated by the terminal system in the Drosophila embryo. Elife 11, e78902 (2022).
Iaco, R. D. et al. Huckebein‐mediated autoregulation of Glide/Gcm triggers glia specification. Embo J. 25, 244–254 (2006).
Dittrich, R., Bossing, T., Gould, A. P., Technau, G. M. & Urban, J. The differentiation of the serotonergic neurons in the Drosophila ventral nerve cord depends on the combined function of the zinc finger proteins Eagle and Huckebein. Dev. Camb. Engl. 124, 2515–2525 (1997).
Lundell, M. J., Chu-LaGraff, Q., Doe, C. Q. & Hirsh, J. TheengrailedandhuckebeinGenes are essential for development of serotonin neurons in theDrosophilaCNS. Mol. Cell Neurosci. 7, 46–61 (1996).
Kudron, M. M. et al. The modERN resource: genome-wide binding profiles for hundreds of drosophila and caenorhabditis elegans transcription factors. Genetics 208, genetics.300657.2017 (2017).
Bossing, T., Technau, G. M. & Doe, C. Q. huckebein is required for glial development and axon pathfinding in the neuroblast 1-1 and neuroblast 2-2 lineages in the Drosophila central nervous system. Mech. Dev. 55, 53–64 (1996).
Wang, Y., Lobb-Rabe, M., Ashley, J., Anand, V. & Carrillo, R. A. Structural and functional synaptic plasticity induced by convergent synapse loss in the Drosophila neuromuscular circuit. J. Neurosci. JN-RM-1492-20, https://doi.org/10.1523/jneurosci.1492-20.2020 (2021).
Lobb-Rabe, M. et al. Dpr10 and Nocte are required for Drosophila motor axon pathfinding. Neural Dev. 17, 10 (2022).
Acknowledgements
This work is supported by NSF IOS-2048080, NINDS R01 NS123439 01, and a UChicago Faculty Diversity Grant to R.A.C., NINDS R15 NS101692 01 & 02 to C.G.V., and HHMI Gilliam GT16464 to R.J.S. This work is also supported by funds from the UChicago Biological Science Division, Committee of Developmental Biology and Department of Molecular Genetics & Cellular Biology, and by the Skidmore College Summer Collaborative Research Program. We thank the Bloomington Drosophila Stock Center (NIH P40OD018537) for fly lines. The monoclonal antibodies 4F3 and 1D4 were developed by Goodman, C. and they were obtained from the Developmental Studies Hybridoma Bank, created by the NICHD of the NIH and maintained at the University of Iowa, Department of Biology. We would like to thank Ellie Hecksher and Michael Glotzer for sharing resources. We thank the Skidmore Microscopy Imaging Center (SMIC) for the use of their microscopy resources. For their contributions to the screen, we also would like to thank the Spring 2018 Skidmore Neurophysiology students (Emily Blunt, Haoyang Huang, Jessy Idemoto, Dilhan Sirtalan, Julie Wang, Rob Warden, and Mary Beth Zahnleuter) and the Spring 2022 Skidmore Neurophysiology students (Jessica Auerbach, Margaret Besthoff, Gene Choi, Joa Comellas, DJ Flam, Melaina Gilbert, Kaylee Hua, Alexander Nardone, Bryan Taylor, and Victoria Thorpe). We would also like to thank Richard Fehon, David Pincus, Sunny Quinn, and members from the Carrillo laboratory for valuable discussions and comments.
Author information
Authors and Affiliations
Contributions
Y.W. and R.A.C designed research; Y.W., R.J.S., L.T.S., V.S., T.J.G., and B.W. performed experiments; Y.W. and R.J.S. analyzed data; Y.W. wrote the manuscript; R.J.S., C.G.V., and R.A.C. edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Chris Doe and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Ivo Lieberam and Benjamin Bessieres. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Y., Salazar, R.J., Simonetta, L.T. et al. hkb is required for DIP-α expression and target recognition in the Drosophila neuromuscular circuit. Commun Biol 7, 507 (2024). https://doi.org/10.1038/s42003-024-06184-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06184-8
- Springer Nature Limited