Abstract
Microglia are the resident immune cells of the central nervous system and are involved in brain development, homeostasis, and disease. New imaging and genomics technologies are revealing microglial complexity across developmental and functional states, brain regions, and diseases. We curated a set of publicly available gene expression datasets from human microglia spanning disease and health to identify sets of genes reflecting physiological and pathological microglial states. We also integrated multiple human microglial single-cell RNA-seq datasets in Alzheimer’s disease (AD), multiple sclerosis (MS), and Parkinson’s disease, and identified a distinct microglial transcriptional signature shared across diseases. Analysis of germ-line DNA identified genes with variants associated with AD and MS that are overrepresented in microglial gene sets, including the disease-associated transcriptional signature. This work points to genes that are dysregulated in disease states and provides a resource for the analysis of diseases in which microglia are implicated by genetic evidence.
Similar content being viewed by others
Introduction
Microglia are the resident immune cells of the central nervous system distinct from other tissue-resident macrophages1,2,3. Microglia originate in the yolk sac and colonize the brain at the early stages of development4, Microglia play an essential role in brain development5, via synaptic pruning and clearing of neural progenitor cells6. Microglia are also involved in brain homeostasis by sensing changes in the brain3 and can shift out of their homeostatic state to a variety of reactive states in case of tissue injury or inflammation.
Microglia have been implicated in multiple CNS diseases, including Alzheimer’s disease (AD)7, multiple sclerosis (MS)8, Parkinson’s disease (PD)9, amyotrophic lateral sclerosis10, stroke11, and glioblastoma12. Genes with established genetic associations with disease (for example, TREM2 and APOE with AD) are expressed in microglia and are dysregulated in patients and animal models13,14.
In both physiological and pathological conditions, transcriptional profiling—an established approach to elucidate the role of microglia in health and disease—has revealed considerable microglial complexity and heterogeneity15,16. This heterogeneity has multiple determinants, including developmental stage17, brain region18,19, sex20, age21,22, and disease state23,24,25. However, technical factors are likely to contribute to this heterogeneity, as inconsistencies have been reported across studies of the same condition or disease26. We set out to understand the main factors contributing to transcriptional heterogeneity in microglia, and how they relate to neurodegenerative disease. We collected, curated, and harmonized publicly available human microglia transcriptomics datasets and performed a systematic analysis to identify gene expression sets conserved across studies which we called microglial gene modules. We extended our work to human genetics by analyzing the overlap between human microglial modules and loci identified in genome-wide association studies (GWAS). We identified genes with genetic variation associated with neurodegenerative diseases that are also dysregulated in disease-associated microglia.
Results
Human microglia transcriptome profiles (bulk RNA-seq datasets)
We identified 21 published transcriptome studies focusing on human microglia (MGL, Fig. 1a and Supplementary Data 1) and selected the 12 containing purified microglial cells with relevant control samples and with available raw sequencing data for further analysis (Supplementary Data 1 and “Methods”), 10 datasets (6 bulk RNA-seq and 4 single-cell (sc) RNA-seq,) from 8 publications were included in the analysis (Figs. 1 and 2 and Supplementary Data 1).
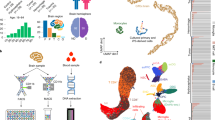
a Summary of datasets reviewed, processed, and used in this study. A total of 21 datasets were identified focusing on the human microglia transcriptome. After pre-processing and QC steps, 12 were selected and analyzed further. Subsequently, 8 of 12 were selected due to biological importance and relevancy. b Bar plot showing differentially upregulated (red) and downregulated (blue) genes in each dataset. Differential expression is determined by DESeq2 (false discovery rate (FDR) < 0.05 and fold change difference > 2). Error bars are standard errors of the mean. c Heatmap showing gene expression patterns of differentially expressed genes across datasets. Each row is a sample, each column is a gene. Black boxes mark sets of genes differentially expressed across multiple datasets.
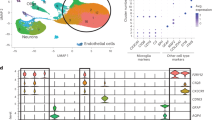
a Uniform manifold approximation and projection (UMAP) plot showing microglia clusters for integrated microglia analysis of 18,713 cells from four datasets. Non-microglial cells were excluded from this analysis. Each dot represents a single cell. Colors correspond to different clusters. b As in (a), but, colors correspond to different conditions. c Heatmap of top differentially expressed genes in each cluster compared to all other clusters. d Top gene ontology terms for differentially expressed genes in each cluster compared to all other clusters.
The six bulk RNA-seq datasets included: (1) purified microglia from human brain with no reported neurological disease, compared to other cell types in the cortex (Fig. S1a26); (2) purified microglia from human brain with no reported neurological disease compared to the whole cortex (Fig. S1b21); (3, 4) differentiating (stage 1, 10 days post-injection: maturing microglia) and differentiated (stage 2, 60 days post-injection: mature microglia) human induced pluripotent stem cell (iPSC)-derived microglia, compared to microglial progenitor cells (Fig. S1c27); (5) human iPSC-derived, lipopolysaccharide (LPS)-induced microglia compared to control microglia (Fig. S1d28); and (6) microglia purified from brains from patients with AD compared to control subjects (Fig. S1a26). After re-processing and differential expression analysis (“Methods”), differentially expressed genes (adjusted P-value < 0.01 and fold change > 2, Fig. 1b) were combined across datasets and the overlapping set (Fig. S1e) was evaluated to establish human microglial modules (Fig. 1c, Supplementary Data 2 and 3).
First, we determined the upregulated genes between MGL vs other brain cells (dataset 126) and MGL vs whole cortex (dataset 221) datasets to identify genes upregulated in microglia compared to other brain cells (‘human microglial gene set’). From that set, we selected transcripts that were not differentially expressed in any other conditions (differentiation, LPS, or AD) to define an ‘MGL Core module’ (n = 480, Fig. 1c, box 1). ‘MGL Core module’ contains genes such as TREM2 and SYK. Some of the known microglia-related genes such as P2RY12, BIN2, and TYROBP were not assigned to the ‘MGL Core module’ despite their upregulated expression in datasets 1 and 2 due to downregulated expression in LPS treatment. Similarly, from the human microglial gene set, we selected the microglial genes significantly regulated in the differentiation data set (dataset 3 and 427) to identify a ‘Differentiation- MGL’ module (box 2 in Fig. 1c). We identified human microglial genes upregulated (box 3 in Fig. 1c) and downregulated (box 4) by LPS treatment (dataset 528). These two groups represent microglia-enriched genes that are regulated upon LPS treatment. Likewise, we identified human microglial genes upregulated (box 5) and downregulated (box 7 in Fig. 1c) in AD (dataset 626). We also identified upregulated AD genes that were not defined as microglial markers in dataset 1 or 2 (box 6). Finally, we used Euclidean distance to cluster the features (Fig. S1f) and reveal a group of transcripts most highly expressed in microglia (MGL Top expressed module, n = 830), which contains known microglial genes such as TREM2 and SYK, as well as P2RY12, BIN2, TYROBP, and APOE. In summary, combining several different datasets allowed the identification of a core set of microglial markers, and condition-specific subsets of transcripts.
Sc transcriptomics identifies shared microglial states across conditions
We identified sc or single-nucleus (sn) publicly available RNA-seq datasets from microglial cells from subjects with neurodegenerative diseases including AD, MS, and PD. We first identified sc and sn RNA-seq datasets either profiling purified microglial samples (microglia-specific Fig. S2a, b) from AD29 and MS19 patients, or reporting sc brain expression data from PD30 and MS31 patients, from which microglial data could be computationally derived (Fig. S2c, d). Each dataset was individually processed for quality control, normalization, dimension reduction, clustering, annotation, and marker identification using the Seurat package (Fig. S2). The two MS datasets were combined using Seurat (“Methods”, Fig. S2e). In the AD dataset (Fig. S2b), 13 clusters were identified, including some non-microglial clusters (e.g., 5, 9, and 10 due to lack of microglia marker genes expression) and AD-associated clusters (3 and 6). 10 clusters were identified in PD (Fig. S2c), including clusters 1, 2, and 3 containing PD-associated cells. The combined analysis of MS datasets (Masuda et al. 19 and Schirmer et al. 31) identified ten clusters (Fig. S2e).
We then checked the expression of two microglia markers—CSF1R, which is broadly expressed in microglia, and P2RY12, which is expressed in homeostatic microglia32—to validate and further annotate microglial clusters in these datasets (Fig. S2f). CSF1R was expressed broadly in these cells as expected, regardless of their disease association. In contrast, P2RY12 was expressed less uniformly in disease-associated clusters. This helped us to further categorize diseased microglia in each dataset. For example, in the PD dataset clusters, 1, 2, and 3 contained diseased microglia, however, expression of P2RY12 was decreased only in clusters 2 and 3.
In order to identify a microglial gene expression subcluster associated with multiple neurodegenerative diseases, we integrated all four datasets and further excluded cells not expressing microglial markers (see “Methods” for details). This integrated analysis identified 13 clusters (Fig. 2a) containing healthy and disease-associated cells (Fig. 2b). In order to check for microglial specificity, we (1) reviewed the expression of known microglial markers including P2RY12, BIN1, CSF1R, and SLCO2B1 (Fig. S3a); (2) checked for the absence of common B and T-cell markers (including TRAC, TRBC2, CD52, IL32, CD3D, CD25, CD52, CD3, CD8, CD56, CD1, CD21, CD25, CD27, CD138, and CD4); and (3) checked for the expression of MRC1 (a macrophage-specific marker33). Although some cells expressed MRC1 at high levels, we didn’t observe a macrophage-specific cluster (Fig. S3b). A few clusters (7, 10, 12, and 13) clustered distantly from the others. Among those, clusters 10 and 13 expressed known neuronal markers (Fig. S3c), and cluster 13 expressed known oligodendrocyte markers (Fig. S3d), pointing to possible contaminating cells.
Cluster analysis (Fig. 2a, b) identified clusters 0, 1, 2, and 3, which included predominantly microglia from control samples from all 4 datasets (Fig. 2b and Fig. S3f), and a few clusters predominantly containing disease cells (cluster 4, 6, and 9). Cluster 4 contained cells from all 3 diseases and was defined as a cross-disease-associated microglia (CDAM) cluster (Fig. 2b). We annotated each cluster by identifying marker genes (Fig. 2c, see “Methods” for details) and gene ontology annotation (Fig. 2d). Immune response-related terms were significantly enriched in disease-associated clusters 4 (P = 1.3E-11) and 9 (P = 1.4E-7).
Characterization of the CDAM cluster
We examined the transcripts associated with cluster 4 (CDAM), containing cells from patients with AD, MS, and PD, by comparing its gene expression with that of ‘core’ microglial clusters (0, 1, 2, and 3, Fig. 3a). Transcripts upregulated in CDAM compared to core clusters 0,1,2,3 (Fig. 3a, b and Supplementary Data 4) included Apolipoprotein E (APOE), known to be associated with AD, Leucine-rich repeat kinase 2 (LRRK2), known to be associated with an increased risk in PD, and glycoprotein NMB (GPNMB), known to be upregulated in disease-associated microglia in mouse models of neurodegenerative disease, and recently implicated in PD pathogenesis34. By contrast, P2YR12, a homeostatic/healthy microglia marker35, was downregulated in CDAM.
a Differential expression analysis comparing genes from CDAM and healthy-microglia-associated clusters. The top significant genes are highlighted on the volcano plot. b Log-2 expression levels of representative genes up-regulated (APOE, LRRK2, and GPNMB) and down-regulated (P2RY12) in CDAM (cluster 4). c Boxplot showing gene expression in CDAM for microglia modules identified in bulk-RNA-seq analysis in Fig. 1. Red dots mark median values for each comparison. P-values between the groups were determined by the Wilcoxon test. UP stands for upregulation and DN stands for downregulation. Error bars on each box represent the highest and lowest values excluding outliers. d Overlap between CDAM and publicly annotated microglial gene sets. All gene sets are defined in the MGEnrichment database. Bar plot showing odds ratio (OR) and -log10 FDR for the most significant overlapping gene sets for CDAM upregulated and downregulated genes compared to core clusters.
We examined the overlap between the modules identified in the bulk RNA-seq and the clusters identified in the scRNA-seq analysis (Fig. 3c). We observed a significant overlap between disease-associated modules (AD-MGL upregulated (P = 1.5E-07) and AD-downregulated (P = 0.006)) and expression CDAM vs clusters 0–3. Overlapping genes include APOE, GPNMB, PARVG, TGFBI, and TMEM119. Genes that were upregulated in LPS microglia modules were also upregulated in CDAM.
We finally compared the overlap between CDAM genes and microglial gene sets reported in the literature and collected in the MGEnrichment database, which contains over 100 gene sets from 26 publications36 (Fig. 3d and Supplementary Data 5). As expected, both CDAM upregulated (CDAM > core clusters) and downregulated (CDAM < core clusters) genes showed a significant overlap with microglia core signals, though the downregulated set showed much higher overlap compared to the upregulated set (OR, 8.1 vs 3.7 and -log10 FDR 68 vs 25). CDAM upregulated genes showed a significant overlap with publicly annotated neurodegenerative disease signature microglial genes (MGnD signature increased, DAM > HOM MG), as well as LPS and age-related microglial genes (Fig. 3d, lower panel). On the other hand, CDAM downregulated genes overlapped with homeostatic/healthy state markers (Fig. 3d, upper panel).
We also annotated the CDAM module using the dataset reported by Geirsdottir et al. 37, a cross-species transcriptomic analysis of microglia. We observed a varied conservation pattern of the CDAM module across species (Fig. S4), supporting the notion that microglial genes relevant for neurodegeneration in humans might not be uniformly conserved across non-human species, and therefore conclusions from work in animal models might not readily transfer to human neurodegenerative conditions.
In summary, a combined analysis of human microglial gene expression identified modules associated with microglial states, and an integrated sc analysis identified a cross-disease-associated cluster that showed high expression of disease markers and downregulation of homeostatic/healthy microglia markers.
Microglial signature is enriched for common genetic variants associated with disease
GWAS have identified many genetic variants associated with traits and diseases over the past decades. Our transcription-level analysis revealed gene expression clusters associated with healthy and diseased-state microglia. We asked whether these clusters had any overlap with genetic signals associated with neurodegenerative disease. We used GWAS summary statistics for AD38, MS39, and PD40 and assigned aggregated variants to genes using the MAGMA tool41, which provides gene-level and gene set-level enrichment analysis for GWAS studies by aggregating all the variants while controlling for confounding effects (Fig. 4a). We identified multiple microglial modules significantly enriched in AD and MS GWAS (Fig. 4b and Supplementary Data 6). Interestingly, genes that were upregulated in AD microglia (AD-MGL upregulated module) were significantly associated with all three GWAS, while genes that were upregulated in AD, but not identified as microglia markers (AD-others upregulated) were not (Fig. 4b). CDAM was also significantly enriched in both AD and MS GWAS. Within this module, we identified many genes implicated in AD GWAS (Fig. 4c) including BIN1, APOE, and PLCG2. We then extended our variant search to exome sequencing data from the UK Biobank (UKB42) for the CDAM genes that were identified as significant in the MAGMA analysis, including PLCG2, which is upregulated in microglia compared to other cells in the cortex and upregulated in CDAM (Fig. 4d, left panel). We identified three variants associated with AD for PLCG2 (Fig. 4d, right). The most significant variant was a missense variant (rs72824905, P = 7.3E-05) associated with a decreased risk of an AD-related trait (Alzheimer’s in the mother, OR = 0.87). This variant was previously shown as an AD protective variant in another study43. Interestingly, an additional rare intronic variant (rs748729362) was nominally associated with an increased risk of AD (P-value: 8.46E-05). The evidence from our transcriptional analysis further supports PLCG2 as a microglial gene with relevance for AD.
a Schematic of common variant analysis. Microglial module enrichment for common variants (minor allele frequency (MAF) > 1%) was tested by MAGMA analysis using AD, MS, and PD GWAS summary statistics. b Summary of common variant enrichment analysis. Bubble plot showing enrichment P-values obtained from MAGMA analysis of each module for AD, MS, and PD GWAS studies. The size of the dot represents the number of genes in each module. P-values < 0.05 are shown in red. c Volcano plot showing genes in CDAM module that are significantly associated with the AD GWAS. d Bar plot showing PLCG2 gene expression changes in our datasets (left). Breakdown of PLCG2 variants associated with AD or related traits (right).
Disease-associated rare genetic variation also affects the CDAM signature
While GWAS captures common variant associations efficiently, rare coding variation (especially if predicted damaging) could have stronger consequences on phenotypes. We mined exome sequencing data for rare (minor allele frequency (MAF) < 1%) coding variants and performed gene burden testing for genes with microglial expression (see “Methods”). We identified genes in each microglial module with nominally significant (P < 0.05) burden associations with AD, MS, and PD (Fig. 5a) and calculated the enrichment in each module (Fig. 5b). We only observed significant enrichment in AD ExWAS for disease-associated modules (AD- MGL upregulated, CDAM, CDAM UP, CDAM DN) (Fig. 5b and Supplementary Data 7). In CDAM, we identified 160 upregulated and 206 downregulated genes with nominally significant burden in AD ExWAS (Fig. 5c). For example, gamma Parvin (PARVG), encoding an actin-binding protein, is significantly upregulated in microglia compared to other cells in human cortex and upregulated in AD-MGL and CDAM (Fig. 5d, left panel), confirming reports of increased expression in AD mouse models and humans44. Although not reaching genome-wide statistical significance, a burden of rare LOF variants in PARVG is associated with protection from parental AD in the UK biobank (Fig. 5d, right panel). Taken together, these results point to a role for PARVG in AD and will need to be confirmed in a larger series.
a Schematic of rare variant analysis. Gene burden test was done for microglial genes for rare variants (MAF < 1%). Microglial module enrichment for rare variants was tested by Fisher’s Exact Test using AD, MS, and PD ExWAS summary statistics. b Summary of rare variant enrichment analysis. Bubble plot showing enrichment P-values obtained from MAGMA analysis of each module for AD, MS, and PD ExWAS studies. The size of the dot represents the number of genes in each module. P-values < 0.05 are shown in red. c Volcano plot showing genes in CDAM module that are significantly associated with AD ExWAS (rare variants). d Bar plot showing gene expression of PARVG in our datasets (left). Breakdown of gene burdens in PARVG associated with an AD-related trait (parental history of AD, right).
Similarly, Interleukin-1 receptor-associated kinase-like 2 (IRAK2), which is a microglia marker gene, is downregulated in CDAM (Fig. S5, left panel), confirming a literature report implicating IRAK2 in AD45. A missense variant in IRAK2 is associated with increased risk for AD in siblings (OR = 8.9, P = 1.28E-06) suggesting that a decrease in gene expression, possibly caused by variants in IRAK2, is a risk factor for AD.
Discussion
We have compiled multiple publicly available human microglia RNA-seq datasets profiling microglia in health and disease states. This analysis led to the identification of human microglial gene modules including core (high expression in human microglia (compared to other cells, but not in disease state), differentiation (high expression in differentiating human microglia), LPS (high expression in LPS-induced, IPSC-derived microglia), and AD (high expression in microglia from AD patients) modules. We examined the genes identified in our combined analysis to see if they were consistent with previous reports on similar studies. Gosselin et al. 46 identified a core microglial signature by comparing microglia to cortex, which is similar to our approach for datasets 126 and 221. Human microglial set (upregulated genes in both datasets 126 and 221) in our study overlaps with Gosselin et al. over 70%. We also observed a similar overlap with the microglia signature defined by Patir et al. 47. Genes that were previously identified as microglial markers, such as TREM2, TYROBP, P2RY12, CSF1R, were also upregulated in human microglial set (microglia vs other cells) in our analysis. We observed that disease-associated genes, such as APOE, were highly expressed in human microglia and upregulated in AD patients and LPS-induced microglia. In addition to confirming known microglial markers, this analysis provides potential novel targets for investigation. For example, we identified an upregulation of PARVG in human microglia compared to other cells and observed a slight, but significant PARVG increase in AD, LPS-induced microglia (Fig. 5d), and in our sc, CDAM cluster. PARVG expression has been recently reported as higher in microglia compared to other cells48, and upregulated in the brains of AD patients44. This and other genes identified in our combined analysis could be considered for further investigation.
Our study’s primary contribution lies in the integration of sc RNA sequencing (scRNA-seq) data to identify a CDAM cluster. While previous studies have defined disease-specific microglia clusters, our work does so by amalgamating multiple datasets across different diseases. This analysis revealed a specific cluster of microglia (the CDAM module) associated with neurodegenerative disease (AD, MS, PD), and showed upregulation of several markers, including APOE and LRRK2, and downregulation of the homeostatic microglia marker P2YR12. One of the top upregulated genes in CDAM was transforming growth factor beta-induced (TGFBI) (log2FC = 15.1 and P-value = 1.27E-217). We found that this gene is highly expressed in human microglia (compared to whole cortex), AD-MGL, and LPS-induced microglia in our bulk RNA-seq analysis. A mouse study showed that TGFBI promotes amyloid-beta clearance in microglia49, and a 2012 report showed that TGFBI is involved in amyloid aggregation in corneal dystrophies50. Considering AD is an amyloid aggregation disease, there could be a potential role of TGFBI in disease-associated microglia.
Dihydropyrimidine dehydrogenase (DPYD) has been recently reported as increased in human AD-associated microglia in bulk RNA-seq analysis26. Our sc analysis also identified DPYD as the top expressed and the most significant gene in CDAM (log2FC = 15.1 and P-value < 1E-304).
One of the top markers of CDAM was a long noncoding RNA nuclear enriched abundant transcript 1 (NEAT1) (log2FC = 3.3 and P-value = 6E-304). The regulatory role of NEAT1 in inflammation in the central nervous system has been an interest of researchers in recent years51. A 2020 study showed NEAT1 expression is upregulated in oxygen–glucose deprivation/reoxygenation-induced microglial cell culture52. NEAT1 regulation has also been reported in stroke and PD studies51. Our study showed a NEAT1 upregulation in CDAM, warranting further study of this transcript.
In summary, our analysis provides valuable resources to be further explored to understand the transcription profile of microglia in health and disease.
One additional strength of this approach is the integration with human genetics data. We examined the overlap between the gene sets we identified based on transcriptional data with signals from GWAS and ExWAS in three neurological diseases. Multiple microglial modules were significantly enriched with GWAS variants for AD and MS, however, we didn’t observe significant enrichments in PD data. One reason could be that we did not observe PD-specific microglial modules, unlike AD. Another possibility is that this lack of overlap denotes a minor role of microglia in PD pathogenesis. One of the modules with significant enrichment for AD GWAS and ExWAS was CDAM. We focused on genes in this module and further discovered the relationship between transcription and genetic variants (Figs. 4 and 5). Other modules with significant enrichment in GWAS and ExWAS can be individually explored in further studies.
We acknowledge the limitations of this study. First, we exclusively used human microglia datasets, which limited the size of our meta-analysis. Another potential problem is the technical variability due to the heterogeneity of tissue/cell processing, library preparation, and RNA isolation techniques across studies. We addressed these issues by applying normalization methods and outlier elimination at each step of the analysis. On the other hand, since we used many different datasets from different labs, consistent findings across various datasets bring a level of confidence for the targets we identified. We also note that three out of four scRNA-seq studies were derived from the cortex, while the PD samples were extracted from the midbrain. This introduces an additional element of technical variability that we were unable to account for in the scope of this study. Finally, the datasets utilized in our study are derived from human postmortem brain tissue, which can introduce additional confounders compared to freshly isolated tissue53. In our sc analysis, we did not observe the artefactual signature described by Marsh et al. 53 within specific clusters or datasets. However, it is important to note that a direct comparison between postmortem tissue and freshly isolated tissue was not feasible in this study.
Another possible technical challenge in this study is the use of both sc and sn RNA-seq data. It has been recently suggested that sn RNA-seq may fail to detect microglial transcripts associated with disease states54, including APOE, APOC1, and CST3. We examined the expression levels of the transcripts characterized as “microglia activation genes depleted in the single nucleus” from Thrupp et al. 54. We detected expression of this set of transcripts in our disease-associated cluster (Fig. S6a), both in both sc (MS) and sn datasets (AD, PD, and MS, Fig. S6b).
In conclusion, we have performed a multi-level analysis of human microglia, identified microglial gene expression changes in health and disease, and discovered subclusters of microglia by combining several sc-level microglial disease data sets. Using expression-based microglial markers, we identified microglial genes with genetic associations for neurodegenerative diseases including AD, MS, and PD. This pipeline revealed known and novel genes that are regulated in microglia and associated with disease. Our findings provide a comprehensive resource to the field to investigate the complexity of microglia in health and disease.
Methods
Dataset selection, data processing, and analysis for bulk RNA-seq
Datasets were selected by applying two primary criteria. First, we required the dataset to pertain to health or disease states, meaning the experiment should represent a microglial state in a disease (AD, LPS, MS, and PD) or healthy condition, such as development or differentiation. The second criterion was technical suitability. We included RNA-seq datasets on RNA isolated from humans (for scRNA-seq), or ex-vivo (for bulk RNA-seq).
Raw fastq files were downloaded from the NCBI GEO database (see Supplementary Data 1 for accession numbers). Fastq files were preprocessed for adapter trimming and filtering short reads using the bbduk tool (http://jgi.doe.gov/data-and-tools/bb-tools/). STAR alignment tool was used to map fastq files to the human genome (GRCh38) with the following parameters: STAR—runThreadN 24—outFilterMultimapNmax 1—outSAMtype BAM SortedByCoordinate—quantMode TranscriptomeSAM55. RSeQC was used for QC56. Gene counts were calculated in R by applying the summarizeOverlaps function on reads mapping to exonic regions. Library size normalization was done by DESeq2 package by estimateSizeFactors57. Samples in each dataset were clustered using Euclidean distance and Pearson correlation using normalized read counts in order to identify outlier samples in R with the pheatmap function. We employed an outlier removal strategy to ensure the integrity of our RNA-seq data analysis. This strategy involved both statistical and qualitative assessments to identify and exclude outliers. Initially, we evaluated several quality control metrics for each sample, including library size, library preparation, the percentage of reads mapped to the genome and genes, and overall read quality scores. Samples exhibiting low performance in any of these metrics were flagged as potential outliers. We then performed clustering analysis on normalized gene counts by DESeq2 for each data set using both Euclidean distance and Pearson correlation metrics to visually observe samples that significantly deviated from their assigned biological group with pheatmap function in R. Samples exhibiting expression patterns that were inconsistent with those of their biological group, particularly those with a z-score exceeding ±3, were flagged as outliers. PCA plots were also generated in R using the prcomp function to further validate and visualize outlier samples in each dataset. ComBat batch effect adjustment was applied to each dataset, and clustering and PCA analysis were repeated.
After applying these initial criteria, we analyzed the data for differential expression between groups, within each dataset, and only included those with significant gene expression changes that overlapped with microglial-enriched genes defined by Galatro et al. 21. and Srinivasan et al. 26 (Supplementary Data 1 and Notes). Differential expression analysis was carried out by DESeq2 when there were 2 conditions (Fig. S1a, d), using default parameters. Genes with an adjusted P-value less than 0.05 and a fold change greater than 1.5-fold were included in the downstream analysis (Fig. 1b, c)57. ANOVA analysis was carried out for multiple groups (Fig. S1a, b heatmaps), comparing one group (e.g., microglia) vs other groups (e.g., neuron, endothelial, and astrocytes). Significant genes were selected for downstream analysis using the same criteria as DESeq2.
To combine datasets, genes with adjusted P-values less than 0.05 and fold change greater than 1.5 in each dataset were included. UpSet plots were generated to identify common genes across datasets (Fig. S1e).
Sc RNA-seq analysis
For each dataset, we used preprocessed 10X Genomics Cell Ranger output aligned to GRCh38 human reference genome if available. For Masuda et al., the processed gene count table was used. Gene matrices were loaded into R using the Seurat package58. Each dataset was individually processed for quality control, normalization, dimension reduction, clustering, annotation, and marker identification using the Seurat package (Fig. S2). The ‘LogNormalize’ function in the Seurat package was used for initial normalization. Filters for Feature_RNA > 500 and nFeature_RNA < 2500 and percent.mt < 20 were applied to exclude low-quality cells. PCA, Uniform Manifold Approximation and Projection (UMAP), and tSNE were used for dimension reduction. If the dataset was not microglia-specific, we only used identified microglia from each dataset. We further filtered cells by requiring the expression of either CSF1R or P2RY12 before moving to the next steps. We identified subclusters in each microglia dataset.
Two MS datasets were combined using SCTransform in the Seurat package. MS-associated clusters in individual-level analysis (cluster 5 in Fig. S1a and cluster 1 in Fig. S1d) clustered closely in the integrated analysis (Fig. S2e), indicating that the two MS datasets were consistent with each other. We then integrated all datasets using the SCTransform method in Seurat. Clusters were identified after integration using the “FindClusters” function with a resolution of 0.4. Cluster markers were identified using the FindMarkers algorithm algorithm with “min.pct = 0.25” and “logfc.threshold = 0.25” parameters.
To assess cluster robustness, we used chooseR59 and clustAssess60 (Fig. S7). We calculated Element-Centric similarity for different algorithms in Seurat (Fig. S7a). In addition, we calculated co-clustering frequency and silhouette score using chooseR (Fig. S7b).
Gene Ontology analysis was performed for each cluster marker using GOStats and redundant terms were removed if there was more than 75% overlap between terms61. Top Biological Process GO terms are reported in Fig. 3c.
Common variant analysis
To aggregate common variants in each GWAS and apply gene sets analysis we used the MAGMA (Multi-marker Analysis of GenoMic Annotation) tool41,62. AD GWAS summary statistics were based on Jansen et al. 38. PD summary statistics were based on Nalls et al. 40 and MS summary statistics were based on the International MS Genetics Consortium 2019 publication39. For MAGMA analysis, first, we annotated gene windows and SNPs by following arguments: window = 35, 10. Then we performed the gene-based analysis with the following parameters: --gene-model snp-wise = top,1,000,000,000 use = Name, Pval ncol = N. Finally, we performed pathway analysis by MAGMA using gene-based results which provided a P-value for each gene set and each gene within the gene set.
Rare variant analysis
Exome sequencing data from the UK Biobank were processed as previously described42,63.
To perform gene burden analysis, rare coding variants were aggregated per gene, based on different categories; (i) predicted loss of function (pLoF), referred to as an “M1” burden mask; (ii) pLoF or missense (“M2”); (iii) pLoF or missense variants predicted to be deleterious by 5/5 prediction algorithms (“M3”); (iv) pLoF or missense variants predicted to be deleterious by 1/5 prediction algorithms (“M4”)64.
Each of these variant-burden categories was tested at four thresholds based on minor allele frequencies: less than 1%, less than 0.5%, less than 0.1%, and less than 0.01%. We then selected the most significant P-value for each gene across these thresholds. We recognize that selecting the lowest P-value result for each gene skews the distribution. However, our enrichment analysis does not directly consider the gene burden test P-values. Instead, it compares them in the gene set of interest versus the rest of the microglial genes. By using microglial genes as a background, we reduce bias in the enrichment analysis. We then performed the Fisher’s Exact test by comparing the significant burden in the microglial module vs the significant burden in the other gene. The OR (greater than 1) and P-value (less than 0.01) were used to determine whether there was an overrepresentation of rare burden variants in each gene set. This analysis was performed for each microglial module and all three ExWAS (AD, MS, PD).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The integrated sc RNA-seq data is available at https://aysegulguvenek.shinyapps.io/shinyapp/, a web application created with R ShinyCell65. The bulk RNA-seq datasets that were used in this study can be accessed through NCBI GEO database with accession numbers; GSE125583 (Srinivasan et al.), GSE99074 (Galatro et al.), GSE139194 (Svoboda et al.), GSE133434 (Hasselman et al.). For the sc RNA-seq data, accession numbers for GEO are GSE120747 (Masuda et al.), GSE160936 (Smith et al.), GSE157783 (Smajic et al.), and for the Sequence Read Archive (SRA) is PRJNA544731 (Schirmer et al.). Source data to generate all the figures and plots are available as Supplementary Data 8.
Code availability
Scripts used in this manuscript are available at: github.com/aguvenek/human-microglia66.
References
Saijo, K. & Glass, C. K. Microglial cell origin and phenotypes in health and disease. Nat. Rev. Immunol. 11, 775–787 (2011).
Utz, S. G. et al. Early fate defines microglia and non-parenchymal brain macrophage development. Cell 181, 557–573.e518 (2020).
De Schepper, S., Crowley, G. & Hong, S. Understanding microglial diversity and implications for neuronal function in health and disease. Dev. Neurobiol. 81, 507–523 (2021).
Frost, J. L. & Schafer, D. P. Microglia: architects of the developing nervous system. Trends Cell Biol. 26, 587–597 (2016).
Paolicelli, R. C. et al. Synaptic pruning by microglia is necessary for normal brain development. Science 333, 1456–1458 (2011).
Prinz, M., Jung, S. & Priller, J. Microglia biology: one century of evolving concepts. Cell 179, 292–311 (2019).
Hemonnot, A. L., Hua, J., Ulmann, L. & Hirbec, H. Microglia in Alzheimer disease: well-known targets and new opportunities. Front Aging Neurosci. 11, 233 (2019).
Kamma, E., Lasisi, W., Libner, C., Ng, H. S. & Plemel, J. R. Central nervous system macrophages in progressive multiple sclerosis: relationship to neurodegeneration and therapeutics. J. Neuroinflammation 19, 45 (2022).
Sanchez-Guajardo, V., Tentillier, N. & Romero-Ramos, M. The relation between alpha-synuclein and microglia in Parkinson’s disease: recent developments. Neuroscience 302, 47–58 (2015).
Spiller, K. J. et al. Microglia-mediated recovery from ALS-relevant motor neuron degeneration in a mouse model of TDP-43 proteinopathy. Nat. Neurosci. 21, 329–340 (2018).
Pannell, M. et al. The subpopulation of microglia expressing functional muscarinic acetylcholine receptors expands in stroke and Alzheimer’s disease. Brain Struct. Funct. 221, 1157–1172 (2016).
Sankowski, R. et al. Mapping microglia states in the human brain through the integration of high-dimensional techniques. Nat. Neurosci. 22, 2098–2110 (2019).
McQuade, A. et al. Gene expression and functional deficits underlie TREM2-knockout microglia responses in human models of Alzheimer’s disease. Nat. Commun. 11, 5370 (2020).
Krasemann, S. et al. The TREM2-APOE pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581.e569 (2017).
Butovsky, O. & Weiner, H. L. Microglial signatures and their role in health and disease. Nat. Rev. Neurosci. 19, 622–635 (2018).
Vecchiarelli, H. A. & Tremblay, M. E. Microglial transcriptional signatures in the central nervous system: toward a future of unraveling their function in health and disease. Annu Rev. Genet. 57, 65–86 (2023).
Matcovitch-Natan, O. et al. Microglia development follows a stepwise program to regulate brain homeostasis. Science 353, aad8670 (2016).
Lopes, K. P. et al. Genetic analysis of the human microglial transcriptome across brain regions, aging and disease pathologies. Nat. Genet. 54, 4–17 (2022).
Masuda, T. et al. Spatial and temporal heterogeneity of mouse and human microglia at single-cell resolution. Nature 566, 388–392 (2019).
Young, A. M. H. et al. A map of transcriptional heterogeneity and regulatory variation in human microglia. Nat. Genet. 53, 861–868 (2021).
Galatro, T. F. et al. Transcriptomic analysis of purified human cortical microglia reveals age-associated changes. Nat. Neurosci. 20, 1162–1171 (2017).
Olah, M. et al. A transcriptomic atlas of aged human microglia. Nat. Commun. 9, 539 (2018).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169,1276–1290.e1217 (2017).
Mathys, H. et al. Temporal tracking of microglia activation in neurodegeneration at single-cell resolution. Cell Rep. 21, 366–380 (2017).
Mrdjen, D. et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, aging, and disease. Immunity 48, 380–395.e386 (2018).
Srinivasan, K. et al. Alzheimer’s patient microglia exhibit enhanced aging and unique transcriptional activation. Cell Rep. 31, 107843 (2020).
Svoboda, D. S. et al. Human iPSC-derived microglia assume a primary microglia-like state after transplantation into the neonatal mouse brain. Proc. Natl. Acad. Sci. USA 116, 25293–25303 (2019).
Hasselmann, J. et al. Development of a chimeric model to study and manipulate human microglia in vivo. Neuron 103, 1016–1033 e1010 (2019).
Smith, A. M. et al. Diverse human astrocyte and microglial transcriptional responses to Alzheimer’s pathology. Acta Neuropathol. 143, 75–91 (2022).
Smajic, S. et al. Single-cell sequencing of human midbrain reveals glial activation and a Parkinson-specific neuronal state. Brain 145, 964–978 (2022).
Schirmer, L. et al. Neuronal vulnerability and multilineage diversity in multiple sclerosis. Nature 573, 75–82 (2019).
Gomez Morillas, A., Besson, V. C. & Lerouet, D. Microglia and neuroinflammation: What place for P2RY12? Int. J. Mol. Sci. https://doi.org/10.3390/ijms22041636 (2021).
Van Hove, H. et al. A single-cell atlas of mouse brain macrophages reveals unique transcriptional identities shaped by ontogeny and tissue environment. Nat. Neurosci. 22, 1021–1035 (2019).
Diaz-Ortiz, M. E. et al. GPNMB confers risk for Parkinson’s disease through interaction with alpha-synuclein. Science 377, eabk0637 (2022).
Zarb, Y. et al. Microglia control small vessel calcification via TREM2. Sci. Adv. https://doi.org/10.1126/sciadv.abc4898 (2021).
Jao, J. & Ciernia, A. V. MGEnrichment: a web application for microglia gene list enrichment analysis. PLoS Comput. Biol. 17, e1009160 (2021).
Geirsdottir, L. et al. Cross-species single-cell analysis reveals divergence of the primate microglia program. Cell 179, 1609–1622.e1616 (2019).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet 51, 404–413 (2019).
International Multiple Sclerosis Genetics, C. Multiple sclerosis genomic map implicates peripheral immune cells and microglia in susceptibility. Science. https://doi.org/10.1126/science.aav7188 (2019).
Nalls, M. A. et al. Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet Neurol. 18, 1091–1102 (2019).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Backman, J. D. et al. Exome sequencing and analysis of 454,787 UK Biobank participants. Nature 599, 628–634 (2021).
Sims, R. et al. Rare coding variants in PLCG2, ABI3, and TREM2 implicate microglial-mediated innate immunity in Alzheimer’s disease. Nat. Genet. 49, 1373–1384 (2017).
Guennewig, B. et al. Defining early changes in Alzheimer’s disease from RNA sequencing of brain regions differentially affected by pathology. Sci. Rep. 11, 4865 (2021).
Cui, J. G., Li, Y. Y., Zhao, Y., Bhattacharjee, S. & Lukiw, W. J. Differential regulation of interleukin-1 receptor-associated kinase-1 (IRAK-1) and IRAK-2 by microRNA-146a and NF-kappaB in stressed human astroglial cells and in Alzheimer disease. J. Biol. Chem. 285, 38951–38960 (2010).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science. https://doi.org/10.1126/science.aal3222 (2017).
Patir, A., Shih, B., McColl, B. W. & Freeman, T. C. A core transcriptional signature of human microglia: derivation and utility in describing region-dependent alterations associated with Alzheimer’s disease. Glia 67, 1240–1253 (2019).
McKenzie, A. T. et al. Brain cell type specific gene expression and co-expression network Architectures. Sci. Rep. 8, 8868 (2018).
Wyss-Coray, T. et al. TGF-beta1 promotes microglial amyloid-beta clearance and reduces plaque burden in transgenic mice. Nat. Med. 7, 612–618 (2001).
Yam, G. H. et al. In vitro amyloid aggregate forming ability of TGFBI mutants that cause corneal dystrophies. Invest Ophthalmol. Vis. Sci. 53, 5890–5898, (2012).
Pan, Y. et al. Novel insights into the emerging role of Neat1 and its effects downstream in the regulation of inflammation. J. Inflamm. Res 15, 557–571 (2022).
Ni, X. et al. Knockdown lncRNA NEAT1 regulates the activation of microglia and reduces AKT signaling and neuronal apoptosis after cerebral ischemic reperfusion. Sci. Rep. 10, 19658 (2020).
Marsh, S. E. et al. Dissection of artifactual and confounding glial signatures by single-cell sequencing of mouse and human brain. Nat. Neurosci. 25, 306–316 (2022).
Thrupp, N. et al. Single-nucleus RNA-seq is not suitable for detection of microglial activation genes in humans. Cell Rep. 32, 108189 (2020).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Wang, L., Wang, S. & Li, W. RSeQC: quality control of RNA-seq experiments. Bioinformatics 28, 2184–2185 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 e1821 (2019).
Patterson-Cross, R. B., Levine, A. J. & Menon, V. Selecting single cell clustering parameter values using subsampling-based robustness metrics. BMC Bioinform. 22, 39 (2021).
Shahsavari, A., Munteanu, A. & Mohorianu, I. ClustAssess: tools for assessing the robustness of single-cell clustering. Preprint at https://doi.org/10.1101/2022.01.31.478592 (2022).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007).
de Leeuw, C. A., Neale, B. M., Heskes, T. & Posthuma, D. The statistical properties of gene-set analysis. Nat. Rev. Genet. 17, 353–364 (2016).
Van Hout, C. V. et al. Exome sequencing and characterization of 49,960 individuals in the UK Biobank. Nature 586, 749–756 (2020).
Kosmicki, J. A. et al. A catalog of associations between rare coding variants and COVID-19 outcomes. Preprint at https://doi.org/10.1101/2020.10.28.20221804 (2021).
Ouyang, J. F., Kamaraj, U. S., Cao, E. Y. & Rackham, O. J. L. ShinyCell: simple and sharable visualisation of single-cell gene expression data. Bioinformatics. https://doi.org/10.1093/bioinformatics/btab209 (2021).
Guvenek, A. aguvenek/human-microglia: v1.0 (v1.0). https://doi.org/10.5281/zenodo.12699948 (2024).
Author information
Authors and Affiliations
Contributions
All authors reviewed the manuscript for important intellectual content and approved the manuscript submitted for publication. A.G. and G.C. conceived the study. A.S. contributed to the study supervision. A.G. performed the analysis. E.S., A.M. and L.D. contributed to common and rare variant analysis. N.P., D.Z. and S.G. contributed to manuscript supervision. A.G. and G.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.G., A.M., L.D., D.Z., N.P., S.H., E.S., A.S., and G.C. are current employees and/or stockholders of Regeneron Genetics Center or Regeneron Pharmaceuticals. All other authors declare no conflict of interest.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Gracjan Michlewski and Joao Valente.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Guvenek, A., Parikshak, N., Zamolodchikov, D. et al. Transcriptional profiling in microglia across physiological and pathological states identifies a transcriptional module associated with neurodegeneration. Commun Biol 7, 1168 (2024). https://doi.org/10.1038/s42003-024-06684-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06684-7
- Springer Nature Limited