Abstract
The Qinghai-Tibet Plateau (QTP), renowned for its exceptional biological diversity, is home to numerous endemic species. However, research on the virology of vulnerable vertebrates like yaks remains limited. In this study, our objective was to use metagenomics to provide a comprehensive understanding of the diversity and evolution of the gut virome in yak populations across different regions of the QTP. Our findings revealed a remarkably diverse array of viruses in the gut of yaks, including those associated with vertebrates and bacteriophages. Notably, some vertebrate-associated viruses, such as astrovirus and picornavirus, showed significant sequence identity across diverse yak populations. Additionally, we observed differences in the functional profiles of genes carried by the yak gut virome across different regions. Moreover, the virus-bacterium symbiotic network that we discovered holds potential significance in maintaining the health of yaks. Overall, this research expands our understanding of the viral communities in the gut of yaks and highlights the importance of further investigating the interactions between viruses and their hosts. These data will be beneficial for revealing the crucial role that viruses play in the yak gut ecology in future studies.
Similar content being viewed by others
Introduction
As one of the most concentrated areas of global biodiversity, the Qinghai-Tibet Plateau (QTP) is home to numerous endemic species1. The yak (Bos grunniens), an ancient even-toed ungulate species in the family Bovidae, is native to the QTP and surrounding high-altitude regions, with approximately 90% of the global yak population distributed there2. The yak’s coat provides excellent thermal insulation, allowing it to thrive in extremely cold and oxygen-deficient conditions. In addition, studies of genetic adaptation suggest that, compared to their low-altitude relatives, the influence of adaptive evolution on energy metabolism genes further supports the survival of yaks in these harsh habitats3,4,5. However, in some regions, due to human activities, environmental changes, and disease issues, the yak population is continuously declining.
In recent decades, the application of metagenomics has greatly expanded our knowledge of the vast array of uncharacterized microbial nucleotide sequences present in the digestive tracts or tissues of various animals, including ruminants6,7,8, birds9, pigs10,11, cats12,13, rabbits14,15, and chickens16,17. Some of these studies have revealed associations with the health and diseases of their hosts. However, our understanding of the microbial ecology in the digestive tract or tissues of yaks living in the QTP is limited, with only a few studies conducted on their gut bacterial metagenomics and metabolomics of yaks so far18,19,20. Recent studies have also shown that the gastrointestinal tracts of ruminants harbor a rich diversity of prokaryotic and eukaryotic microorganisms21,22,23, which play essential physiological roles such as aiding in the digestion of feed, protecting their animal hosts from pathogens, producing volatile fatty acids (VFAs) that contribute to increased energy. Conversely, microbial imbalances can lead to metabolic disorders and negatively affect the health of their animal hosts24. Viruses constitute a substantial portion of the gut microbiome, with bacteriophages (phages) being the predominant constituents of the gut virome. They infect bacteria and play a crucial role in regulating the gut bacteriome by either lysing their host bacteria or modulating their physiological functions25. Therefore, identifying the viral hosts may reveal the effects of viruses on the gut microbiota, thereby advancing the development of related applications. For example, some phages (primarily lytic phages) are considered to have great potential for treating infections caused by Staphylococcus aureus26, Klebsiella oxytoca27, and Escherichia coli28 isolated from cases of bovine mastitis. Recently, a comprehensive study by Wu et al. on the gut virome of ruminants identified 109 phages that infect methanogenic archaea, 74 of which were lytic24. This finding provides new insights into reports that the rumen microbiomes of yaks living at high altitudes produce more VFAs and less methane4. Furthermore, a study indicates that the rumen virome can regulate microbial diversity and is associated with diet as well as several important animal production traits29. Therefore, exploring the gut virome of yaks on the QTP will provide valuable data for future research, particularly on the regulatory roles of viral communities in the guts of high-altitude yaks. In addition, some potential eukaryotic viruses that can infect QTP yaks and other vertebrates and cause diseases remain unexplored. Furthermore, the extensive use of antibiotics in human, veterinary, and agricultural practices has led to the continuous release of antibiotics and antibiotic resistance genes (ARGs) into the environment30. Recent studies have identified phage-associated ARGs from diverse sources, including cattle, pigs, and poultry8,31,32, as well as various wastewater environments33. However, ARGs from animals living in the QTP have yet to be explored.
In this study, we embarked on a comprehensive investigation of the gut virome of yaks living in the QTP to understand its potential uniqueness. A total of 122 fecal samples were collected from five sampling points in the QTP, revealing the presence of viruses within them. Furthermore, we compared the differences in viral community composition among yak populations from different regions and further explored the genetic relationships between known and novel viruses, as well as the functional profiles of phage-encoded genes. Additionally, virus-bacterium association analysis and virus-bacterium interaction analysis were also performed to reveal the virus-bacterium interaction mediated by viruses identified in this research.
Results
Analysing the yak gut virome
An extensive metagenomic investigation was carried out on fecal samples from 122 yaks (Supplementary Data 1), which were collected from five distinct altitude sampling points located in four provinces across the QTP and its neighboring regions in Chinese Mainland, with an average altitude of 4139.12 m (Fig. 1a–c). After quality control, a total of 225,929,680 paired-end reads were generated. Subsequently, 25,248,520 reads were assigned to viruses. After de novo assembly, a total of 3,343,456 contigs were generated, out of which 372,598 were assigned to viruses (Fig. 1d). The viruses accounted for approximately 11% in both the reads and contigs (Fig. 1e). The library labeled as “sichuanganzi119” has been removed due to its poor quality. The viral species richness of these 121 quality-controlled fecal samples was represented by the rarefaction curves (Fig. 1f). As the number of sampled reads increased, the curve gradually reached a plateau, indicating that the number of libraries collected in this study was sufficient. Additional data would only reveal a limited number of new species. According to the species accumulation curves estimation based on random sampling strategy, these 121 libraries contain approximately 800 different viral species (Fig. 1f). Surprisingly, the 30 samples from Deqin show lower overall species richness compared to other sampling points. While potential influences from the construction process of the libraries cannot be ruled out (all of these libraries were constructed in the same batch), it can cautiously be inferred that the composition of the gut virome in the Deqin region’s yak population may differ from that of other regions.
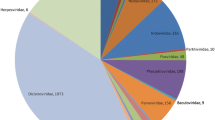
a An overview of the sampling points in four provinces of Chinese Mainland, with five distinct sampling points indicated by blue dots. b A detailed topographic map of the five sampling points, with the scale and corresponding elevation of different locations displayed on the right side of the figure. The source of the map is Geospatial Data Cloud (https://www.gscloud.cn), and the software used to create the map is ArcMap v10.5. All of these data are freely available to the public. c An elevation profile graph of the sampling points. d The scatter plot depicts the quantities of reads and contigs identified as viruses, produced by each individual library. e The light blue portion in both pie charts represents the reads (on the left) or contigs (on the right) annotated as viruses in all libraries. The light pink portion represents reads or contigs annotated as non-viral or unannotated. f The larger figure represents the species rarefaction curve plotted using Megan6 software, with a logarithmic scale transformation applied; The smaller figure depicts the species accumulation curve, where the horizontal axis represents the number of randomly sampled libraries, and the vertical axis represents the cumulative number of identified viral species. g An UpSet plot based on different sampling points as the classification criterion. This plot showcases the number of viruses shared or unique among different populations of yaks’ gut. The left-side bar chart represents the total number of viral species for each sampling point, while the top bar chart represents the number of viral phyla corresponding to shared or unique viruses.
We also analyzed the species composition of the yak gut virome in each region. The results showed that the yaks in Ganzi had the highest number of viral species, with 499 species, followed by Naqu (393), Shannan (358), Haibei (291), and Deqin (221) (Supplementary Data 2). Surprisingly, despite the relatively lower number of viral species in the yak population of the Haibei, the presence of 100 unique viral species in this region’s yak population is second only to the 129 unique viral species detected in the Ganzi. This suggests the distinctiveness of viral communities harbored by yak populations in different regions (Fig. 1g). In addition, the Ganzi has the highest diversity of unique viral species in its yak population. The viruses belonging to the phylum Uroviricota have the highest proportion among the unique viral species in each region, except for Shannan. Overall, among the shared viral species in these five regions, the highest number belongs to the phylum Phixviricota, followed by the phylum Cressdnaviricota.
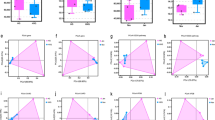
The richness and diversity of flora in specific regions or ecosystems are typically measured using ACE index, Chao1 index, Shannon index, and Simpson index. Chao1 and ACE indices are primarily used to estimate species richness. A higher Chao1 or ACE index indicates a more abundant flora in the sample. Shannon and Simpson indices, on the other hand, are mainly used to evaluate species diversity. A higher Shannon index or a lower Simpson index indicates a greater diversity of flora in the sample. In this study, we conducted α-diversity analysis on the gut viral communities of yaks in five regions. The results revealed that the Ganzi had the highest richness of gut viral communities in the yak population, followed by Naqu, Shannan, Haibei, and Deqin, which is consistent with the aforementioned findings. In terms of viral species diversity, the gut viral community of yaks in Ganzi exhibited the highest diversity, followed by Shannan, Naqu, Haibei, and Deqin (Fig. 2a and Table 1). It is noteworthy that there is some discrepancy between the Shannon index and the Simpson index, which could be attributed to the emphasis of the Simpson index on uniformity, while this study encounters variations in sample sizes among different regions. Further PCoA analysis revealed significant differences in the composition of viral communities in the gut of yaks among these five regions (PERMANOVA, P = 0.001) (Fig. 2b).
a Comparisons of richness and diversity of viral communities in the gut of yaks across different sampling points: Ganzi (n = 32), Naqu (n = 30), Shannan (n = 20), Haibei (n = 9), and Deqin (n = 30). The ACE, Shannon, Chao1, and Simpson indices were all analyzed using the Wilcoxon test. *P < 0.05, **P < 0.01, ***P < 0.001, between the two groups. b The PCoA analysis revealed significant differences among viral communities from different sampling points. The two principal component scores accounted for 20% (PC1) and 11% (PC2) of the total variations, respectively. Each symbol represents an individual sample. c A cluster heatmap of viral classes in the gut of yaks from five different sampling points. The different colored bands at the top of the heatmap represent the corresponding sampling points. The row names on the right side of the heatmap represent the names of viral families. The data is presented in logarithmic scale with a base of log10, and the legend is displayed in the upper right corner. d Filtered the viral taxonomy diagram to include the top 150 most abundant viral genera across all libraries. Different background colors represent distinct viral realms. The outermost green squares represent the relative abundance heatmap for each viral genus. Yellow triangles indicate viral genera with an abundance below 0.1% of all viruses. Blue squares represent viral genera with an abundance above 0.1% of all viruses.
It is evident and predictable that phages constitute a significant portion of the yak gut virome, particularly viruses belonging to the Caudoviricetes, Malgrandaviricetes and Faserviricetes (Fig. 2c). We then selected the top 150 most abundant viruses based on their genus-level abundance, retaining only those well-annotated across all seven levels from realm to genus, and constructed a viral taxonomy diagram (Fig. 2d and Supplementary Data 1). The unclassified Caudoviricetes family represents those taxa identified at the genus level within the class Caudoviricetes but not assigned to any specific family. It should be noted that some viruses, due to their novelty, may not be consistently classified across all seven levels, potentially leading to them being overlooked. The filtered viruses primarily belong to four viral realms: Monodnaviria, Duplodnaviria, Riboviria, and Varidnaviria. The viruses in the realm Duplodnaviria dominate in terms of quantity. We have also observed that viruses belonging to the families Astroviridae34,35, Caliciviridae36,37, Picornaviridae38,39 within the realm Riboviria, as well as viruses belonging to the families Circoviridae40,41, Parvoviridae42,43 within the realm Monodnaviria, have been reported to potentially be associated with numerous diseases in vertebrate animals. Therefore, these viruses are worth further analysis.
Surveying vertebrate-associated viruses in the QTP yaks
Animals may serve as natural hosts for certain viruses, and these viruses can potentially spread among different animal species. Investigating vertebrate-associated viruses in QTP yaks can help predict and control potential disease outbreaks, thereby contributing to the protection of the stability of the QTP animal population and the health of the ecosystem. Here, we have recovered and identified 6 parvoviruses, 24 astroviruses, and 12 picornaviruses from the yak’s metagenomic datasets, all of which contain complete or near-complete hallmark genes (Supplementary Data 1). Although viruses belonging to the family Caliciviridae are highly significant and associated with bovine diarrhea symptoms37, we have been unable to recover a sufficiently long fragment for further analysis. In general, most of these viruses show a considerable degree of identity to the currently known viruses. However, the host preferences of these viruses may not be the same for all. Based on phylogenetic analysis, some of the parvoviruses identified in yaks may be closely associated with lizards and mosquitoes (Fig. 3a), while the viruses belonging to Astroviridae and Picornaviridae were closely related only to vertebrates (Fig. 3b, c). Furthermore, we have identified nearly identical astroviruses in yaks from both Naqu and Haibei regions. Similarly, we found nearly identical picornaviruses in yaks from Naqu and Shannan, as well as from Naqu and Ganzi. Therefore, we can infer that these vertebrate-associated viral infections have already spread among different regions’ yak populations, although the pathogenicity of these viruses cannot be determined at present.
The maximum likelihood trees were constructed using the NS1 proteins of Parvoviridae (a) and the RdRp proteins of Astroviridae (b) and Picornaviridae (c), respectively. The red dots at the tips of the clades represent the viruses identified in this study. Lines of different colors represent the hosts of viruses, as detailed in the legend at the bottom right corner. All animal and other life form silhouettes are sourced from PhyloPic (https://www.phylopic.org) and are available for reuse under Creative Commons licenses.
Expansion the diversity of CRESS DNA viruses
Circular replication (Rep)-encoding single-stranded (CRESS)-DNA viruses are widespread and have been reported to infect nearly all eukaryotic organisms globally44. These viruses display an unforeseeable range of diversity and distribution, with their expansion showing no signs of abating45. In order to explore the diversity of CRESS DNA viruses in the gut of yaks, we attempted to recover the Rep protein sequences from the datasets. Sequence similarity network analysis revealed that the majority of sequences were well-clustered into several groups (Fig. 4a). Here, we present the detection of 176 circoviruses, 359 genomoviruses, 640 smacoviruses, and 91 unclassified CRESS DNA viruses from QTP yak fecal samples (Fig. 4b–e and Supplementary Data 1). The sequence analysis results indicated that there were 65 circoviruses showing less than 60% amino acid sequence identity in their Rep protein compared to known viruses. Similarly, there were 72 genomoviruses and 355 smacoviruses with less than 60% Rep amino acid sequence identity to known viruses. These sequences may represent potential novel viral species. Phylogenetic analysis revealed that viruses belonging to Genomoviridae exhibit a wide range of host diversity, including birds, reptiles, protozoa, plants, and arthropods; in contrast, circoviruses and smacoviruses are more closely associated with vertebrates. Furthermore, there are some CRESS DNA viral sequences that cannot currently be classified into established viral families. Similarly, these viruses demonstrate varying host preferences, indicating the uniqueness of the ecological environment on the QTP and the gut virome of yaks.
a Sequence similarity network of Reps associated with CRESS DNA viruses. The maximum likelihood trees were constructed using the Rep proteins of Circoviridae (b), Genomoviridae (c), Smacoviridae (d) and Unclassified CRESS DNA Viruses (e), respectively. Lines of different colors represent the hosts of viruses, as detailed in the legend at the bottom right corner. All animal and other life form silhouettes are sourced from PhyloPic (https://www.phylopic.org) and are available for reuse under Creative Commons licenses.
The QTP yak population harbors a highly diverse range of phages
Phages are abundant in diverse habitats and crucial for maintaining bacterial communities and ecosystem stability. Investigating the distribution and functionality of yak gut phages enables a profound understanding of their roles in the QTP ecosystem and reveals their evolutionary mechanisms and dynamics of diversity. In this study, we screened and obtained 109,461 phage-associated contigs, with the majority belonging to the classes Caudoviricetes and Malgrandaviricetes. However, approximately 8954 contigs were tentatively assigned to bacterial viruses. PhaGCN2, based on the GCN model, was used for further prediction and classification of these contigs. Consequently, 523 phage genomes were successfully classified into 19 different viral families (Fig. 5a and Supplementary Data 3). Additionally, by comparing with the RVD database, we matched 1715 phages and successfully annotated 176 of them at the family level, but no further annotations were obtained at the genus level (Supplementary Data 1). Using the DeePhage tool to predict the lifestyles of the viruses in this study’s dataset revealed that, overall, the proportions of temperate phages (50.6%) and lytic phages (49.4%) in QTP yaks were comparable. Regionally, the yaks in Deqin had the lowest proportion of lytic phages at 45.6%, while those in Haibei had the highest proportion at 62.7% (Supplementary Fig. 1). The region encoding the TerL in Caudoviricetes exhibits remarkably high evolutionary conservation, which may assist us in depicting the distinctive evolutionary patterns of Caudoviricetes in the gut of the QTP yaks. A total of 460 TerL sequences were detected and included in the phylogenetic analysis, along with closely related sequences from GenBank and other reference sources (Supplementary Data 1). The phylogenetic tree indicated that the TerL genes of the majority of identified viruses in this study showed substantial divergence from known sequences, making it impossible to include them within the established classification framework (Fig. 5b). Furthermore, it has been noted that certain viruses identified in this study form clusters with those already recognized for infecting specific bacteria, thus suggesting potential viral hosts. Nevertheless, additional research is imperative to authenticate these findings.
a PhaCGN2 was employed for the classification prediction of the 8954 contigs annotated as bacterial viruses identified in this study, with 523 contigs successfully assigned to 19 viral families. The network graph was generated using Gephi based on the node and edge files produced by PhaGCN2. b Phylogenetic analysis of Caudoviricetes based on the TerL amino acid sequences from yak fecal samples. The red lines represent newly identified viruses in this study. The lines of other colors indicate the hosts corresponding to the selected reference sequences, as shown in the legend on the left. The outermost circle represent the genera to which each virus belongs. c The scatter plot depicts the identity and coverage of 6351 MCPs generated in this study compared to their best match in the GenBank database. d Sequence similarity network of MCPs associated with Microviridae. Gray dots represent those sourced from the GenBank database, while purple dots represent those derived from the gut of yaks in the QTP.
Microviridae, a family of CRESS viruses that infect bacteria, is globally recognized as one of the most widespread and diverse viral families46. They inhabit diverse environments, including the guts of animals and humans47,48, insects49, freshwater50, seawater51, and sediments52. A total of 6347 distinct hallmark gene protein sequences, namely MCPs, were identified from the gut of yaks in the QTP. The average protein sequence length was 452 aa (Supplementary Data 1). Interestingly, the vast majority (over 5300) of MCPs exhibited a sequence identity lower than 60% with the best matches in the GenBank database (Fig. 5c and Supplementary Data 1). Furthermore, network clustering analysis revealed that a subset of MCPs identified in this study formed several major clusters with known sequences, most of which were not assigned to specific viral genera or species. On the other hand, another subset formed clusters comprising a few or several dozen MCPs, while some MCPs existed as individual clusters (Fig. 5d). These findings enhance our understanding of the hidden diversity of phages in the gut of yaks on the QTP, which may be shaped by the unique diet of yaks or the distinctive natural geographical environment of the plateau.
Gene functional analysis of yak gut phages
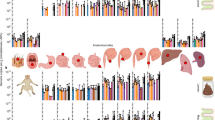
The KEGG search program in eggNOG-mapper v2 was employed to annotate detected genes in phage sequences, allowing for the exploration of their potential functions (Supplementary Data 4). The results revealed that the majority of annotated genes were associated with metabolic pathways, including those for nucleotide, amino acid, lipid, and carbohydrate metabolism. Among the QTP yak populations, the Naqu population had the highest abundance of genes involved in KEGG pathways, followed by the Ganzi and Deqin populations, while the Shannan and Haibei populations had fewer genes detected in these pathways. Notably, yak gut phages from the Ganzi population possessed the most diverse and abundant functional gene categories, including Cellular Processes, Organismal Systems, and Human Diseases (Fig. 6a). Among these genes, those involved in DNA replication, such as ssb and dnaB, as well as genes involved in nucleotide metabolism (thyX) and amino acid metabolism (ydiP), were prevalent in yak populations in the QTP. Furthermore, it was observed that genes involved in lipid metabolism (avrBs2) and DNA replication (pcrA) were more abundant in specific regions of yak populations (Fig. 6b). The widespread detection of genes involved in energy metabolism, DNA replication and repair subcategories in yaks may suggest the unique survival pressures faced by endemic species on the QTP, likely related to their adaptation to low-temperature and hypoxic environments. However, further research is needed to confirm this.
a Bubble plot of functional annotation results for phage proteins based on the eggNOG database. b Heatmap showing the relative abundance of genes annotated by eggNOG in yak gut phage communities across different regions. c Bar chart depicting the relative abundance of gut bacteria in yaks. d The upper panel represents the quantile-quantile plot of P values for the virus-bacterium association analysis (Pvirus-bacteria). The x-axis represents the expected −log10(Pvirus-bacterium) values from a uniform distribution. The y-axis represents the observed −log10(Pvirus-bacterium) values. The red diagonal line represents the line y = x, which corresponds to the null hypothesis. The horizontal red dashed line represents the Bonferroni-corrected threshold (α = 0.05), while the brown dashed line indicates the FDR threshold (FDR = 0.05) calculated using the Benjamini-Hochberg method. The x-axis in the lower panel represents the effect sizes in linear regression. The y-axis and the horizontal dashed lines are consistent with those shown in the panel above.
Virus-bacterium coabundance in the yak dataset
Viruses, particularly phages, can alter the abundance and function of bacteria upon infection, disseminate virulence factors between bacterial hosts to modify the severity of bacterial infections, thereby indirectly impacting the stability of the gut microbiota53. We performed an association analysis of the abundances of viruses and bacteria in all collected yak libraries. The relative abundance of bacteria in all libraries is shown in Fig. 6c. After filtering, a total of 8 bacterial clades and 2 viral clades were included for further analysis. The abundances of 7 different bacterial clades were negatively correlated with the abundances of Microvirus and/or Siphovirus, respectively, satisfying FDR and Bonferroni thresholds Fig. 6d and Supplementary Data 5. Overall, the genus Prevotella, which belong to the family Prevotellaceae, had the most significant negative correlation with Siphovirus (effect size = –0.240, P = 4.29E–05). Prevotella is commonly considered a probiotic associated with a healthy plant-based diet. They are not only abundant in the human guts but also prevalent in the guts of animals54,55. Additionally, Zhang et al. found that Prevotella spp. were increased in the rumen of yaks compared to cattle4. However, some studies have indicated that Prevotella in the gut is also associated with inflammation56,57. Therefore, the virus-bacterium symbiosis network such as Prevotella-Siphovirus revealed by our results has important implications for maintaining the health of yaks.
Viral host detection based on the CRISPR spacer sequences
To further clarify the virus-bacterium interaction, MinCED was used to detect CRISPR sequences in bacterial sequences in all yak libraries and to determine the corresponding viral sequences in the same library. A total of 29 spacer sequences were detected and 9 unique virus-bacterium interactions were identified (Table 2). Among them, there are 7 clades of bacteria interacting with Myovirus, 1 clade interacting with Siphovirus, and 1 clade interacting with Arthrobacter phage Corgi (Supplementary Data 1). Consistent with the above results, the interaction between Siphovirus and Prevotella was observed.
Discussion
Recent metagenomic advancements have yielded vast genetic information on viruses, yet much remains unknown. According to some studies, mammals host at least 40,000 distinct viral species, significantly surpassing the viral species presently recognized by the ICTV58. Additionally, one study has shown that viruses are abundant in the rumen, with concentrations reaching 107 to 1010 virions per milliliter of rumen fluid59. Therefore, ongoing and extensive research into viral diversity is essential for addressing future epidemics. Sampling viruses from a broader range of vertebrate hosts should provide better evolutionary insights. Here, we primarily focus on the analysis of the gut virome of yaks living in the extreme environment of the QTP. To the best of our knowledge, this is the first comprehensive virological research conducted on the yak populations in this region.
In the past, the QTP was considered pristine due to its sparse population. However, recent industrial activities have introduced pollutants, posing a threat to the region’s ecological communities60. Previous studies have shown that the rumen microbiota differs between yaks and cattle raised at different altitudes4,61. It is evident that there are differences in the composition of the QTP yak gut viral communities across different regions, but these differences seem to be minimally affected by altitude. Interestingly, there is a significant correlation (P < 0.001) between the richness and diversity of the yak gut viral communities and the permanent resident population in the sampling regions (Supplementary Fig. 2). According to the data from the Seventh National Population Census of China, Ganzi has the highest permanent resident population, with approximately 1.1 million people, followed by Naqu with 500,000, Shannan with 350,000, Haibei with 300,000, and Deqin with 50,000 residents. This suggests that human activities may be one of the potential factors affecting the gut viral composition of endemic species in the QTP, while the gut of yaks in the Deqin region may have retained a relatively primitive viral composition. In addition, we have not observed any signs of vertebrate-related viral sharing between the yaks in Deqin region and those in other areas. Previous studies have shown that diet is the most influential factor affecting the bacterial composition in ruminants62; recent works have also revealed that diet can impact the rumen virome29,63. Unfortunately, because the yak samples in this study were collected from the wild, we cannot speculate on their dietary habits. A recent study characterized the lifestyles of phages in ruminants and noted that the proportion of lytic phages is higher in ruminants compared to other environments, where temperate phages constitute the majority24. Another study found that half of the rumen microbial genomes and metagenome-assembled genomes contain at least one prophage, highlighting the importance of lysogeny in the rumen ecosystem29. Lytic phages lyse host cells, releasing host cellular components and increasing nutrient cycling in the rumen, including carbohydrates, lipids, and proteins. Temperate (lysogenic) phages can grant their hosts novel metabolic capabilities, enhancing their ecological fitness and potentially aiding in their evolution. Consequently, rumen viruses can greatly influence the rumen microbiome, its functions, and overall animal productivity25. Although the influence of sample size differences cannot be entirely ruled out, our results support the aforementioned observation (Supplementary Fig. 1). However, the lifestyles of these viruses identified by DeePhage need further validation in the laboratory or confirmation of accuracy by training models on larger cohorts. In the field of virology research, over 60% of newly discovered viral sequences displayed substantial deviations from established reference sequences, defying categorization within a defined viral species. Such sequences even bordered on the creation of novel viral families, earning them the moniker ‘viral dark matter’64,65,66. Whether within the human gastrointestinal tract or the global oceans, the existence of these viral dark matters has been extensively confirmed, and there are abundant genetically diverse phage populations in the given environments67,68. Similarly, the yaks dwelling in the extreme environments of the QTP harbor an exceptionally rich and distinct collection of phages especially those associated with the family Microviridae. However, due to the limitations posed by the length of assembled sequences and the analytical methods used, the full extent of viral diversity cannot yet be completely resolved. Further in-depth research is required. While these modest advancements have broadened our comprehension of phage genomic diversity, they also suggest that our quest to discover new viruses has barely begun to scratch the surface of the iceberg.
A recent study on QTP wetland soil samples indicates that the composition of bacterial communities is the primary driving force affecting the diversity and geographical distribution of ARGs. Proteobacteria, Bacteroidetes, Actinobacteria, and Firmicutes comprised over 75% of the bacterial community structure in QTP wetlands. FCA and β-lactamase resistance genes also make up a significant proportion of ARG abundance in these regions60. Likewise, Bacteroidetes and Firmicutes have been confirmed as predominant bacteria in the yak’s gastrointestinal tract20, and our study also supports this conclusion. Additionally, research has shown that genera such as Prevotella, Ruminococcus, and Streptococcus, which can be infected by rumen viruses, dominate the core rumen microbiome8. Therefore, rumen viruses may play a role in influencing the diversity, metabolism, and functions of the rumen ecosystem. Previous research has stated that Firmicutes play a crucial role in energy absorption processes69. As the dietary energy levels and concentrate ratios increase, the relative abundance of Firmicutes may increase70,71. This may explain why this group of bacteria accounted for more than half of the bacterial composition in yak feces. Correspondingly, we have also detected a wide range of genes involved in energy metabolism and DNA replication in the yak’s gut. Here, we did not detect ARGs encoded by phages in the gut of QTP yaks. A recent study indicated that ARGs are rarely encoded in phages72. Yan et al. also identified only 24 viruses carrying ARGs out of 705,380 viral contigs in a large-scale rumen virome analysis8, which may explain this observation. However, including a larger sample size of QTP yaks may reveal new discoveries. Nevertheless, these annotated genes need further curation to confirm their accuracy. For example, (i) confirming that the candidate gene is actually encoded by a virus, and (ii) confirming that the candidate gene truly participates in cellular metabolic pathways or other cellular processes. Additionally, the specific genomic context surrounding each candidate gene should be carefully examined. Therefore, this necessitates the future improvement of in silico prediction tools, robust benchmarking, and high-throughput experimental methods73.
In conclusion, this study provides the first-ever depiction of the gut virome profile of yaks on the QTP, revealing a remarkably rich diversity, complexity, and novelty of the yak gut virome; and discusses their genetic similarities with known viruses. This study not only enhances our understanding of the health status of yaks but, more importantly, underscores the necessity of conducting such research within a broader ecological context.
Methods
Sample collection, processing, and quality control
From May to June 2021, three teams departed respectively from Nyingchi in Tibet, Xining in Qinghai, and Ganzi in Sichuan to collect a total of 122 fresh fecal samples from the gut of yaks in five different habitats on the QTP. Specifically, there were 30 samples collected from Naqu, Tibet (altitude: 4724.41 m), 20 samples from Shannan, Tibet (5013.41 m), 30 samples from Deqin, Yunnan (3760.57 m), 33 samples from Ganzi, Sichuan (4197.20 m), and 9 samples from Haibei, Qinghai (3000.00 m) (Fig. 1 a–c). Most of the yaks involved in this study were inhabiting areas near the snowy mountains of the QTP, with a few scattered at the foothills. These areas have no access restrictions. In areas without roads, we observed the yaks from a distance and collected samples immediately after they defecated. None of the yaks exhibited any evident signs of illness or disease. All samples were preserved in sterile containers and transported using dry ice. Prior to viral metagenomic analysis, each 10-gram sample was submerged in 0.5 mL of Dulbecco’s phosphate-buffered saline (DPBS) and vigorously vortexed for 5 min. Subsequently, they were incubated at 4 °C for 30 min. After centrifugation at 15,000 × g for 10 min, the resulting supernatants were collected in 1.5 mL centrifuge tubes and stored at –80 °C for future use46. The collection of samples was carried out in compliance with the Wildlife Protection Law of the People’s Republic of China. All experiments were conducted following the guidelines of a Biosafety Level 2 laboratory. For each library, 100 µL of the supernatant was pipetted from a single sample and subsequently collected in a new 1.5 mL tube. These samples were centrifuged at 12,000 × g for 5 min at 4 °C and filtered through a 0.45 µm filter to enrich viral particles. The filtrates were treated with RNase and DNase, and the unprotected nucleic acids were subsequently digested at 37 °C for 60 min74. Total nucleic acids were then extracted using the manufacturer’s protocol provided with the QIAamp MinElute Virus Spin Kit (Qiagen). These nucleic acid samples containing DNA and RNA viral sequences were used for reverse transcription reactions with the SuperScript III reverse transcriptase (Invitrogen) and 100 pmol of a random hexamer primer, followed by a single round of DNA synthesis using Klenow fragment polymerase (New England BioLabs). Libraries were constructed using the Nextera XT DNA Sample Preparation Kit (Illumina) and sequenced on the Illumina NovaSeq 6000 platform with 250 base-paired ends with dual barcoding.
During the experiment, all procedures were conducted with necessary precautions to avoid sample cross-contamination and degradation of nucleic acids. We used aerosol filter tips to reduce the likelihood of sample cross-contamination. Additionally, all other experimental materials, such as microcentrifuge tubes and tips, that came into direct contact with nucleic acid samples were free of DNase and RNase. The samples were dissolved in DEPC-treated water containing RNase inhibitors. For blank controls, sterile ddH2O was prepared simultaneously and further processed under the same experimental conditions. Quality testing was performed using agarose gel electrophoresis and Agilent bioanalyzer 2100. While sequencing on the Illumina NovaSeq platform, the control pool generated a very small number of reads.
Metagenome assembly
In order to minimize host contamination, we downloaded the reference genome sequences (GCA_005887515.3) of yak (Bos grunniens) from NCBI. Subsequently, we employed Bowtie2 v2.4.575,76 to align and remove potential host sequences (https://www.metagenomics.wiki/tools/short-read/remove-host-sequences) from the 122 libraries. Primers and low-quality sequences were trimmed using Trim Galore v0.6.5 (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore), and the files were quality controlled with specific options as follows ‘--phred33 --length 100 --stringency 3 --paired’. Duplicated reads were marked using PRINSEQ-lite v0.20.4 (-derep 1)77. The paired-end reads were assembled using MEGAHIT v1.2.978 with default parameters. The results were then imported into Geneious Prime v2022.0.1 (https://www.geneious.com) for sorting and renaming. To reduce false negatives during sequence assembly, additional semi-automated assembly was conducted on the unmapped contigs and singlets shorter than 500 bp, and contigs that were >1500 bp long after reassembly were retained. Moreover, mixed assembly was performed using MEGAHIT combined with BWA v0.7.1779 to search for unused reads and low-abundance contigs.
Identification of viral genomes in yak libraries
We conducted the identification of viral sequences in the yak libraries through a series of steps. Firstly, a specialized local viral database was created for screening the assembled contigs, which included the non-redundant protein (nr) database (downloaded in May 2022) and IMG/VR v380. The contigs initially annotated as eukaryotic viruses, including those shorter than 1500 bp, were imported into Geneious Prime for manual assembly and examination, and used as the reference for mapping to the raw data using the Low Sensitivity/Fastest parameter. The resulting sequences were screened for potential vector contamination using VecScreen (https://www.ncbi.nlm.nih.gov/tools/vecscreen) and subjected to genome clustering using MMseqs2 (-k 0 -e 0.001 --min-seq-id 0.95 -c 0.9 --cluster-mode 0)81. Subsequently, these sequences were incorporated into the yak virus dataset along with those further identified as phage contigs.
Phage contigs were recognized in accordance with the viral sequence identification SOP (https://doi.org/10.17504/protocols.io.bwm5pc86). Contigs were validated using VirSorter282, and were then subjected to CheckV83 to remove host sequences flanking prophages. The potential phage contigs were screened based on data from VirSorter2 and CheckV outcomes, which took into account the counts of viral and host genes, VirSorter2 viral scores, and the presence of hallmark genes. Furthermore, we identified conserved motifs within candidate phage contigs, such as the large terminase subunit (TerL) and major capsid protein (MCP), and confirmed them through manual validation. These phage contigs was subsequently clustered at 95% average nucleotide identity (ANI) across 85% of the shortest contig per MIUViG standards84, utilizing a custom script from the CheckV repository, resulting in phage populations.
The non-redundant yak virus dataset was then compared against the local database using the BLASTx program built in DIAMOND v2.0.1585, and significant sequences with a cut-off E-value of <10–5 were filtered. The coverage of each sequence was computed using pileup, a tool within BBMap, and the relative abundance of each sequence was determined via a custom Bash shell script. Taxonomic identification of the yak virus dataset was performed using TaxonKit86 software and the rma2info program within MEGAN687. PhaGCN288 was employed for potential further classification of phages that cannot be classified through alignment with known sequences, and the generated node and edge files were integrated into Gephi v0.10 (https://gephi.org), resulting in the creation of a network graph. Furthermore, we used the BLASTn tool (v2.15.0)89 to compare these viruses with the rumen virome database (RVD)8 to obtain additional taxonomic information. We retained sequences with both alignment identity and coverage greater than 90% with the subject sequences. Coverage was calculated by merging the alignment fraction length of BLASTn high-scoring pair sequences. Additionally, due to the absence of viral sequences longer than 500 bp, sichuanganzi119 library was excluded from the analysis.
Virus genome annotation
Geneious Prime was used with parameters (minimum size: 100; start codon: ATG) to predict potential open reading frames (ORFs). These ORFs were subsequently validated by comparing them to similar viruses in the GenBank database. The annotations of these ORFs were assigned based on comparisons with the built-in CDD v3.21 database within the Conserved Domain Database (CDD)90. This database includes domains curated by NCBI, as well as data imported from Pfam, SMART, COG, PRK, and TIGRFAM. GraPhlAn was used to visualize the viral taxonomy diagram from the realm to the genus level, following the methodology provided in the GraPhlAn tutorial available at https://huttenhower.sph.harvard.edu/GraPhlAn.
Phylogenetic analysis and sequence similarity network analysis
To elucidate phylogenetic relationships, sequences belonging to different groups of corresponding viruses were downloaded from the GenBank database, along with sequences of proposed species pending ratification. Nucleotide or protein sequences were aligned using MUSCLE in MEGA-X91. Sites containing more than 50% gaps were temporarily removed from the alignments. Maximum likelihood trees were then constructed using IQ-TREE v1.6.1292. All phylogenetic trees were created using IQ-TREE with 1,000 bootstrap replicates (-bb 1000) and the ModelFinder function (-m MFP). Interactive Tree Of Life (iTOL) was used for visualizing and editing phylogenetic trees93.
We have also assembled a dataset comprising the protein sequences of the MCP obtained in this study, which serves as a hallmark gene for the family Microviridae, along with all available MCPs from the GenBank database. We employed MMseqs2 to cluster the dataset and conducted sequence similarity network analysis on the non-redundant dataset using EFI-EST94, with an alignment score threshold of 100, corresponding to 35% sequence identity. The obtained network was visualized in Cytoscape V3.10 for subsequent analysis95. Similarly, a dataset comprising replication-associated proteins (Reps) of circoviruses, genomoviruses, smacoviruses, and other unclassified CRESS DNA viruses was also generated, with an alignment score threshold of 27.
Functional annotation of phages
The ORFs of the viral contigs were functionally annotated by comparing them to the eggNOG v5.096 database using eggNOG-mapper v297 with default parameters, which is a tool for functional annotation based on precomputed orthology assignments. The functional annotations from KEGG, COG, and Pfam were derived from the results of the eggNOG-mapper analysis. The abundance of each filtered gene was calculated by mapping the clean reads to the datasets using BWA, the sum of the abundances of those genes with the same KO annotation was used to represent the relative abundance of each gene category. Additionally, we aligned phage-associated protein sequences against the Comprehensive Antibiotic Resistance Database (CARD) using default parameters to predict the profiles of ARGs98. However, we did not detect any phage-related ARGs.
Prediction of viral lifestyles
DeePhage99, which uses a deep neural network to learn features from both DNA and protein sequences and thus has better generalization ability for phages, was used to analyze the lifestyles of phages identified in this study. The virtual machine file for DeePhage was obtained from https://cqb.pku.edu.cn/zhulab/info/1006/1174.htm and opened using VirtualBox v7.0 (https://www.virtualbox.org). DeePhage classifies phages into four categories based on a scoring system: temperate (≤0.3), uncertain temperate (0.3–0.5), uncertain virulent (0.5–0.7), and virulent (>0.7), with higher scores indicating greater virulence.
Virus-bacterium association analysis
We extracted bacterial sequences from MEGAN6 to obtain bacterial abundance and normalised the relative abundance using log transformation. All sequences are aligned and annotated with the nr database. We only retained the clades that were detected in all 121 libraries for further analysis. After selection, we assessed 8 bacterial clades (2 phyla, 2 classes, 2 orders, 1 family and 1 genus) and 2 viral clades (Siphovirus and Microvirus). Virus-bacterium association analysis was performed separately for each virus-bacterium pair using the lm function in RStudio and the effect size of the viral abundance was evaluated100.
Virus-bacterium interaction analysis based on CRISPR spacers
CRISPR sequences in bacterial sequences were predicted using MinCED v0.4.2 (-minNR 2) (https://github.com/ctSkennerton/minced). Spacers sequences within the predicted CRISPR sequences were searched against the viral sequences from the same library using blastn with a cut-off E-value of <10–5, nucleotide identity of >95%, and coverage of spacers of >90%100. Then, we summarised the virus-bacterium pair in each library.
Statistics and reproducibility
Statistical analyses and normalization were performed using MEGAN6 and R. Alpha-diversity and beta-diversity analysis were performed using the vegan package, with statistical significance set at P < 0.05. The ACE, Shannon, Chao1, and Simpson indices were all analyzed using the Wilcoxon test. Visual presentation utilized the ggplot2 and ggpubr packages. Principal coordinate analysis (PCoA) based on Bray-Curtis dissimilarity was carried out using the Permute, lattice, vegan, and ape packages. The PERMANOVA analysis was performed using the adonis() function from the vegan package.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The library data involved in this study has been deposited into the National Genomics Data Center (NGDC) of China (https://ngdc.cncb.ac.cn) and the Short Read Archive (SRA) of the GenBank database under the BioProject accession no. PRJCA018020/PRJNA994540. The involved sequences have been submitted to the GenBank database without any access restrictions, with accession numbers detailed in Supplementary Data 1.
References
Wu, S., Wang, Y., Wang, Z., Shrestha, N. & Liu, J. Species divergence with gene flow and hybrid speciation on the Qinghai-Tibet Plateau. N. Phytol. 234, 392–404 (2022).
Liu, Y. et al. The sequence and de novo assembly of the wild yak genome. Sci. Data 7, 66 (2020).
Wang, Z. et al. Domestication relaxed selective constraints on the yak mitochondrial genome. Mol. Biol. Evol. 28, 1553–1556 (2011).
Zhang, Z. et al. Convergent evolution of Rumen microbiomes in high-altitude mammals. Curr. Biol. 26, 1873–1879 (2016).
Qiu, Q. et al. The yak genome and adaptation to life at high altitude. Nat. Genet 44, 946–949 (2012).
Stewart, R. D. et al. Compendium of 4,941 rumen metagenome-assembled genomes for rumen microbiome biology and enzyme discovery. Nat. Biotechnol. 37, 953–961 (2019).
Glendinning, L., Genc, B., Wallace, R. J. & Watson, M. Metagenomic analysis of the cow, sheep, reindeer and red deer rumen. Sci. Rep. 11, 1990 (2021).
Yan, M. et al. Interrogating the viral dark matter of the rumen ecosystem with a global virome database. Nat. Commun. 14, 5254 (2023).
Shan, T. et al. Virome in the cloaca of wild and breeding birds revealed a diversity of significant viruses. Microbiome 10, 60 (2022).
Xiao, L. et al. A reference gene catalogue of the pig gut microbiome. Nat. Microbiol. 1, 16161 (2016).
Chen, C. et al. Expanded catalog of microbial genes and metagenome-assembled genomes from the pig gut microbiome. Nat. Commun. 12, 1106 (2021).
Zhang, W. et al. Identification and genomic characterization of a novel species of feline anellovirus. Virol. J. 13, 146 (2016).
Van Brussel, K. et al. The enteric virome of cats with feline panleukopenia differs in abundance and diversity from healthy cats. Transbound. Emerg. Dis. 69, e2952–e2966 (2022).
Ning, S. Y. et al. Viromic analysis of feces from laboratory rabbits reveals a new Circovirus. Virus Res. 319, 198861 (2022).
Tsoleridis, T. et al. Discovery and prevalence of divergent RNA viruses in European field voles and rabbits. Viruses 12, 47 (2019).
Maki, J. J., Bobeck, E. A., Sylte, M. J. & Looft, T. Eggshell and environmental bacteria contribute to the intestinal microbiota of growing chickens. J. Anim. Sci. Biotechnol. 11, 60 (2020).
Glendinning, L., Stewart, R. D., Pallen, M. J., Watson, K. A. & Watson, M. Assembly of hundreds of novel bacterial genomes from the chicken caecum. Genome Biol. 21, 34 (2020).
Liu, L. et al. Multi-omics analyses reveal that the gut microbiome and its metabolites promote milk fat synthesis in Zhongdian yak cows. PeerJ 10, e14444 (2022).
Zhao, C. et al. Yak rumen microbiome elevates fiber degradation ability and alters rumen fermentation pattern to increase feed efficiency. Anim. Nutr. 11, 201–214 (2022).
Ma, J. et al. Comparing the bacterial community in the gastrointestinal tracts between growth-retarded and normal Yaks on the Qinghai-Tibetan Plateau. Front Microbiol. 11, 600516 (2020).
Tong, F. et al. The microbiome of the buffalo digestive tract. Nat. Commun. 13, 823 (2022).
Xie, F. et al. An integrated gene catalog and over 10,000 metagenome-assembled genomes from the gastrointestinal microbiome of ruminants. Microbiome 9, 137 (2021).
Sun, Y. et al. Elevated testicular apoptosis is associated with elevated sphingosine driven by gut microbiota in prediabetic sheep. BMC Biol. 20, 121 (2022).
Wu, Y. et al. A compendium of ruminant gastrointestinal phage genomes revealed a higher proportion of lytic phages than in any other environments. Microbiome 12, 69 (2024).
Yu, Z., Yan, M. & Somasundaram, S. Rumen protozoa and viruses: The predators within and their functions-A mini-review. JDS Commun. 5, 236–240 (2024).
Varela-Ortiz, D. F. et al. Antibiotic susceptibility of Staphylococcus aureus isolated from subclinical bovine mastitis cases and in vitro efficacy of bacteriophage. Vet. Res Commun. 42, 243–250 (2018).
Amiri Fahliyani, S., Beheshti-Maal, K. & Ghandehari, F. Novel lytic bacteriophages of Klebsiella oxytoca ABG-IAUF-1 as the potential agents for mastitis phage therapy. FEMS Microbiol. Lett. 365, https://doi.org/10.1093/femsle/fny223 (2018).
Porter, J., Anderson, J., Carter, L., Donjacour, E. & Paros, M. In vitro evaluation of a novel bacteriophage cocktail as a preventative for bovine coliform mastitis. J. Dairy Sci. 99, 2053–2062 (2016).
Yan, M. & Yu, Z. Viruses contribute to microbial diversification in the rumen ecosystem and are associated with certain animal production traits. Microbiome 12, 82 (2024).
Pazda, M., Kumirska, J., Stepnowski, P. & Mulkiewicz, E. Antibiotic resistance genes identified in wastewater treatment plant systems - A review. Sci. Total Environ. 697, 134023 (2019).
Xu, L. et al. Risk of horizontal transfer of intracellular, extracellular, and bacteriophage antibiotic resistance genes during anaerobic digestion of cow manure. Bioresour. Technol. 351, 127007 (2022).
Ji, Y. et al. Metagenomics analysis reveals potential pathways and drivers of piglet gut phage-mediated transfer of ARGs. Sci. Total Environ. 859, 160304 (2023).
Lekunberri, I., Villagrasa, M., Balcazar, J. L. & Borrego, C. M. Contribution of bacteriophage and plasmid DNA to the mobilization of antibiotic resistance genes in a river receiving treated wastewater discharges. Sci. Total Environ. 601-602, 206–209 (2017).
Selimovic-Hamza, S., Boujon, C. L., Hilbe, M., Oevermann, A. & Seuberlich, T. Frequency and pathological phenotype of bovine astrovirus CH13/NeuroS1 infection in neurologically-diseased cattle: Towards assessment of causality. Viruses 9, 12 (2017).
Castells, M. et al. Bovine astrovirus surveillance in uruguay reveals high detection rate of a novel mamastrovirus species. Viruses 12, 32 (2019).
Deng, Y. et al. Studies of epidemiology and seroprevalence of bovine noroviruses in Germany. J. Clin. Microbiol. 41, 2300–2305 (2003).
Lu, X. et al. Comparison of gut viral communities in diarrhoea and healthy dairy calves. J. Gen. Virol. 102, https://doi.org/10.1099/jgv.0.001663 (2021).
Wang, L., Lim, A. & Fredrickson, R. Genomic characterization of a new bovine picornavirus (boosepivirus) in diarrheal cattle and detection in different states of the United States, 2019. Transbound. Emerg. Dis. 69, 3109–3114 (2022).
Hao, L., Chen, C., Bailey, K. & Wang, L. Bovine kobuvirus-A comprehensive review. Transbound. Emerg. Dis. 68, 1886–1894 (2021).
Li, Y. et al. Porcine circovirus 3 in cattle in Shandong province of China: A retrospective study from 2011 to 2018. Vet. Microbiol. 248, 108824 (2020).
Zhu, J. et al. First detection and complete genome analysis of porcine circovirus-like virus P1 and porcine circovirus-2 in yak in China. Vet. Med Sci. 8, 2553–2561 (2022).
Kailasan, S. et al. Structure of an enteric pathogen, bovine parvovirus. J. Virol. 89, 2603–2614 (2015).
Wang, M. et al. Simultaneous detection of bovine rotavirus, bovine parvovirus, and bovine viral diarrhea virus using a gold nanoparticle-assisted PCR assay with a dual-priming oligonucleotide system. Front Microbiol. 10, 2884 (2019).
Desingu, P. A. & Nagarajan, K. Genetic diversity and characterization of circular replication (Rep)-encoding single-stranded (CRESS) DNA viruses. Microbiol Spectr. 10, e0105722 (2022).
Zhao, L., Rosario, K., Breitbart, M. & Duffy, S. Eukaryotic circular rep-encoding single-stranded DNA (CRESS DNA) viruses: Ubiquitous viruses with small genomes and a diverse host range. Adv. Virus Res. 103, 71–133 (2019).
Wang, H. et al. Gut virome of mammals and birds reveals high genetic diversity of the family Microviridae. Virus Evol. 5, vez013 (2019).
Walters, M. et al. Novel single-stranded DNA virus genomes recovered from chimpanzee feces sampled from the Mambilla Plateau in Nigeria. Genome Announc 5, e01715 (2017).
Shkoporov, A. N. et al. The human gut virome is highly diverse, stable, and individual specific. Cell Host Microbe 26, 527–541 e525 (2019).
Kraberger, S., Schmidlin, K., Fontenele, R. S., Walters, M. & Varsani, A. Unravelling the single-stranded DNA virome of the New Zealand Blackfly. Viruses 11, 532 (2019).
Tseng, C. H. et al. Microbial and viral metagenomes of a subtropical freshwater reservoir subject to climatic disturbances. ISME J. 7, 2374–2386 (2013).
Cheng, R. et al. Virus diversity and interactions with hosts in deep-sea hydrothermal vents. Microbiome 10, 235 (2022).
Deng, Z. et al. Phage-prokaryote coexistence strategy mediates microbial community diversity in the intestine and sediment microhabitats of shrimp culture pond ecosystem. Front Microbiol. 13, 1011342 (2022).
Bodner, K., Melkonian, A. L. & Covert, M. W. The enemy of my enemy: New insights regarding bacteriophage-mammalian cell interactions. Trends Microbiol. 29, 528–541 (2021).
Li, X. et al. Construction and characterization of Juglans regia L. polyphenols nanoparticles based on bovine serum albumin and Hohenbuehelia serotina polysaccharides, and their gastrointestinal digestion and colonic fermentation in vitro. Food Funct. 12, 10397–10410 (2021).
Edwards, R. A. Prodigious Prevotella phages. Nat. Microbiol. 4, 550–551 (2019).
Dillon, S. M. et al. Gut dendritic cell activation links an altered colonic microbiome to mucosal and systemic T-cell activation in untreated HIV-1 infection. Mucosal Immunol. 9, 24–37 (2016).
Scher, J. U. et al. Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. Elife 2, e01202 (2013).
Kawasaki, J., Kojima, S., Tomonaga, K. & Horie, M. Hidden viral sequences in public sequencing data and warning for future emerging diseases. mBio 12, e0163821 (2021).
Lobo, R. R. & Faciola, A. P. Ruminal phages - A review. Front Microbiol. 12, 763416 (2021).
Yang, Y., Liu, G., Ye, C. & Liu, W. Bacterial community and climate change implication affected the diversity and abundance of antibiotic resistance genes in wetlands on the Qinghai-Tibetan Plateau. J. Hazard Mater. 361, 283–293 (2019).
Xin, J. et al. Comparing the Microbial Community in Four Stomach of Dairy Cattle, Yellow Cattle and Three Yak Herds in Qinghai-Tibetan Plateau. Front Microbiol. 10, 1547 (2019).
Henderson, G. et al. Rumen microbial community composition varies with diet and host, but a core microbiome is found across a wide geographical range. Sci. Rep. 5, 14567 (2015).
Anderson, C. L., Sullivan, M. B. & Fernando, S. C. Dietary energy drives the dynamic response of bovine rumen viral communities. Microbiome 5, 155 (2017).
Brum, J. R. & Sullivan, M. B. Rising to the challenge: accelerated pace of discovery transforms marine virology. Nat. Rev. Microbiol. 13, 147–159 (2015).
Reyes, A., Semenkovich, N. P., Whiteson, K., Rohwer, F. & Gordon, J. I. Going viral: next-generation sequencing applied to phage populations in the human gut. Nat. Rev. Microbiol. 10, 607–617 (2012).
Roux, S., Hallam, S. J., Woyke, T. & Sullivan, M. B. Viral dark matter and virus-host interactions resolved from publicly available microbial genomes. Elife 4, e08490 (2015).
Dion, M. B., Oechslin, F. & Moineau, S. Phage diversity, genomics and phylogeny. Nat. Rev. Microbiol. 18, 125–138 (2020).
Chevallereau, A., Pons, B. J., van Houte, S. & Westra, E. R. Interactions between bacterial and phage communities in natural environments. Nat. Rev. Microbiol. 20, 49–62 (2022).
Ley, R. E., Turnbaugh, P. J., Klein, S. & Gordon, J. I. Microbial ecology: Human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Ahmad, A. A. et al. Effects of dietary energy levels on rumen fermentation, microbial diversity, and feed efficiency of Yaks (Bos grunniens). Front Microbiol. 11, 625 (2020).
Liu, C. et al. Dynamic Alterations in Yak rumen bacteria community and metabolome characteristics in response to feed type. Front Microbiol. 10, 1116 (2019).
Enault, F. et al. Phages rarely encode antibiotic resistance genes: a cautionary tale for virome analyses. ISME J. 11, 237–247 (2017).
Pratama, A. A. et al. Expanding standards in viromics: in silico evaluation of dsDNA viral genome identification, classification, and auxiliary metabolic gene curation. PeerJ 9, e11447 (2021).
Zhang, W. et al. Faecal virome of cats in an animal shelter. J. Gen. Virol. 95, 2553–2564 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Langmead, B., Wilks, C., Antonescu, V. & Charles, R. Scaling read aligners to hundreds of threads on general-purpose processors. Bioinformatics 35, 421–432 (2019).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011).
Li, D. et al. MEGAHIT v1.0: A fast and scalable metagenome assembler driven by advanced methodologies and community practices. Methods 102, 3–11 (2016).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Roux, S. et al. IMG/VR v3: an integrated ecological and evolutionary framework for interrogating genomes of uncultivated viruses. Nucleic Acids Res. 49, D764–D775 (2021).
Mirdita, M., Steinegger, M. & Soding, J. MMseqs2 desktop and local web server app for fast, interactive sequence searches. Bioinformatics 35, 2856–2858 (2019).
Guo, J. et al. VirSorter2: a multi-classifier, expert-guided approach to detect diverse DNA and RNA viruses. Microbiome 9, 37 (2021).
Nayfach, S. et al. CheckV assesses the quality and completeness of metagenome-assembled viral genomes. Nat. Biotechnol. 39, 578–585 (2021).
Roux, S. et al. Minimum information about an uncultivated virus genome (MIUViG). Nat. Biotechnol. 37, 29–37 (2019).
Buchfink, B., Reuter, K. & Drost, H. G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods 18, 366–368 (2021).
Shen, W. & Ren, H. TaxonKit: A practical and efficient NCBI taxonomy toolkit. J. Genet Genomics 48, 844–850 (2021).
Gautam, A., Felderhoff, H., Bagci, C. & Huson, D. H. Using AnnoTree to get more assignments, faster, in DIAMOND+MEGAN microbiome analysis. mSystems 7, e0140821 (2022).
Shang, J., Jiang, J. & Sun, Y. Bacteriophage classification for assembled contigs using graph convolutional network. Bioinformatics 37, i25–i33 (2021).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinforma. 10, 421 (2009).
Wang, J. et al. The conserved domain database in 2023. Nucleic Acids Res. 51, D384–D388 (2023).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Minh, B. Q. et al. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
Oberg, N., Zallot, R. & Gerlt, J. A. EFI-EST, EFI-GNT, and EFI-CGFP: Enzyme Function Initiative (EFI) Web Resource for Genomic Enzymology Tools. J. Mol. Biol. 435, 168018 (2023).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Huerta-Cepas, J. et al. eggNOG 5.0: A hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Cantalapiedra, C. P., Hernandez-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: Functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 38, 5825–5829 (2021).
Alcock, B. P. et al. CARD 2020: Antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res 48, D517–D525 (2020).
Wu, S. et al. DeePhage: Distinguishing virulent and temperate phage-derived sequences in metavirome data with a deep learning approach. Gigascience 10, giab056 (2021).
Tomofuji, Y. et al. Whole gut virome analysis of 476 Japanese revealed a link between phage and autoimmune disease. Ann. Rheum. Dis. 81, 278–288 (2022).
Acknowledgements
This research was supported by National Key Research and Development Programs of China No.2023YFD1801301 to WZ; Funding for Kunlun Talented People of Qinghai Province, High-end Innovation and Entrepreneurship talents—Leading Talents No. 202208170046 to WZ; National Natural Science Foundation of China No.32060792 to GG and National Modern Agricultural Industry Technology System (CARS-37) to SS.
Author information
Authors and Affiliations
Contributions
All authors participated in the design, interpretation of the studies and analysis of the data and review of the manuscript; S.S., X.M., T.S. and W.Z. contributed to the conception and design; H.W., M.Z., X.W., Q.S., L.J., Y.L., Y.W. and J.L. contributed to the collection and assembly of data; X.L., G.G., Q.Z. and S.Y. contributed to the data analysis and interpretation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Antonio Charlys da Costa, Jiming Chen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Tobias Goris
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lu, X., Gong, G., Zhang, Q. et al. Metagenomic analysis reveals high diversity of gut viromes in yaks (Bos grunniens) from the Qinghai-Tibet Plateau. Commun Biol 7, 1097 (2024). https://doi.org/10.1038/s42003-024-06798-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06798-y
- Springer Nature Limited