Abstract
Oral mucosal wounds exhibit accelerated healing with reduced scarring compared to cutaneous wounds, representing an optimal wound healing paradigm. However, the specific cellular subtypes orchestrating the efficient healing of mucosal tissues remain elusive. Through a comprehensive analysis integrating bulk-mRNA and single-cell sequencing data during the wound healing process in oral mucosa and skin, we have delineated a distinct set of genes markedly upregulated during tissue repair. This collection of wound healing-associated genesets was highly enriched in a specific keratinocyte subpopulation identified as STAT3-activated SPRR1B+ keratinocytes. Notably, despite the inherent rapidity of oral mucosal healing, the induction of SPRR1B+ keratinocytes is evident in both skin and mucosal wound healing processes in murine model. Intriguingly, these wound healing-promoting SPRR1B+ keratinocytes, which are induced via STAT3 activation, inherently abundant in unwounded normal mucosa but absent in normal skin. SPRR1B knockdown significantly inhibits mucosal keratinocyte migration, a critical attribute for effective wound healing. In summary, through analysis of human oral and skin wound healing processes at single-cell resolution, coupled with validation in murine model, suggests STAT3-activated SPRR1B+ keratinocytes are associated with the rapid mucosal repair process. This discovery underscores the potential application of SPRR1B+ keratinocytes in the therapeutic management of chronic or non-healing wounds.
Similar content being viewed by others
Introduction
Wound healing is an intricate process that aims to restore the structure and function of damaged tissue following an injury1. In cases of deficient wound healing, patients may experience prolonged suffering, the development of ulcers, and, in some instances, life-threatening consequences2,3. Conversely, excessive wound healing can result in distorted scars and organ fibrosis4, inflicting significant psychological and physical harm on patients. Currently, the speed and effectiveness of wound healing remain unsatisfactory, necessitating urgent improvement in this area. Notably, oral mucosa exhibits distinct advantages over skin in the context of wound healing, including faster recovery5 and minimal residual scarring6,7. This characteristic is acknowledged as an excellent model of wound healing. Investigating the specific healing patterns of oral mucosa holds the potential to offer new insights into the clinical management of chronic or non-healing wounds.
Several studies have identified factors that contribute to oral better wound healing, including the presence of saliva8, more rapid immune responses9, and lower differentiation of keratinocytes10. However, wound healing represents a multifaceted process consisting of several distinct yet interconnected phases, encompassing hemostasis, inflammation, proliferation, and tissue remodeling1. Successful wound healing relies on the intricate coordination of different cell types in sequential stages. Currently, our understanding of the wound healing process primarily relies on conventional bulk transcriptome sequencing to identify genes associated with wound healing, which has limitations in dissecting the mechanism at the single-cell level9. The advent of single-cell RNA sequencing (scRNA-seq) technologies has enabled the examination of gene expression at a single-cell resolution, offering the potential to unravel specific cell types that govern the rapid wound healing observed in the oral mucosa11. Indeed, scRNA-seq has been profiled in normal oral12 and skin13,14,15. Some studies also performed scRNA-seq to track the skin wound healing process in mice16,17. However, these datasets are insufficient to elucidate which cell subtypes contribute to faster wound healing in human oral mucosa compared to skin.
To address this knowledge gap, we conducted an integrative analysis using publicly available transcriptome data from paired samples of uninjured and injured human skin and oral mucosa. Additionally, we utilized single-cell RNA sequencing (scRNA-seq) data from healthy human oral mucosa and skin. Our objective was to pinpoint key cell types and regulators contributing to the accelerated wound healing observed in oral mucosa. Through this approach, we identified a novel keratinocyte subpopulation, named SPRR1B+ keratinocytes, was rapidly induced during both skin and mucosa would healing, and act as the primary contributor to the wound healing process due to their heightened migration capacity. Furthermore, although not observed in normal skin, SPRR1B+ keratinocytes were specifically enriched in normal oral mucosa. This suggests that SPRR1B+ keratinocytes play a role in priming wound healing in unwounded oral mucosa, potentially contributing to an accelerated wound healing response compared to the skin.
Result
Constitutive expression of P2-Wound Healing-associated Gene Sets (P2-WHGs) in uninjured oral mucosa, but not in uninjured skin
In order to analyze the shared gene expression programs during the wound healing process in both human oral mucosa and skin, we conducted a comparative study using bulk mRNA sequencing data from injured and uninjured sites of human mucosa and skin, obtained from publicly available datasets (GSE97615). Our focus was on identifying genes that were upregulated during wound healing in both skin and mucosa, which were subsequently categorized as shared wound healing-associated gene sets (Fig. 1a, adjusted P < 0.05 and log2 fold change > 1). This analysis led to the identification of 485 differentially expressed genes (DEGs), defining them as shared wound healing-associated gene sets (Fig. 1a). Further analysis involved unsupervised clustering of these 485 genes, revealing two distinct gene modules labeled as Gene Panel 1 (P1) and Gene Panel 2 (P2). Both P1 and P2 gene panels exhibited increased expression during oral mucosa and skin wound healing. Notably, while P1 gene sets were expressed at similar levels in uninjured oral mucosa and skin, P2 gene sets were exclusively abundant in uninjured oral mucosa, but not in the skin (Fig. 1b). Given that 146 P2 wound healing genes (P2-WHGs) were constitutively expressed in uninjured oral mucosa, we hypothesize that P2 gene sets may serve as a precondition for the rapid healing of oral mucosa after injury. Indeed, gene ontology (GO) enrichment analysis of the P2 gene sets revealed that those genes were primarily involved in cornification and keratinization, immune response, and antimicrobial response (Fig. 1c, adjusted P < 0.05), which are indispensable programs for initiating early wound healing after injury18. These findings suggest that oral mucosa may possess a preexisting readiness for the initiation of wound healing-associated gene expression programs even before injury occurs, presenting an advantage not observed in the skin.
a Differentially expressed genes (DEGs) between injured skin (left) or injured oral (right) vs uninjured counterpart were selected for further analysis. (|logFC| > 1 and adjusted p < 0.05). b Heatmap showing expression of selected DEGs between injured and uninjured group in each GSE dataset. c Gene ontology (GO) enrichment analysis revealed the top 15 GO Terms of P2-WHGs.
Keratinocytes are the major contributor of P2-WHGs expression in oral mucosa
To identify the specific cell types expressing the P2 wound healing gene sets in uninjured oral mucosa but not observed in skin, we conducted an integrative analysis of publicly available single-cell RNA-seq data of human mucosa and skin tissues. This dataset includes 5 skin samples (GSA accession: PRJCA006797, 5 normal skin) and 21 oral mucosal samples (GEO accession: GSE164241, 13 gingiva and 8 buccal sites). Rigorous quality control measures were applied, resulting in the retention of 103,758 cells for subsequent data normalization, scaling, sample integration, and cell clustering (See Methods, Supplementary Fig. 1). We identified nine distinct cell types with different gene expression patterns, including myeloid cells (Myeloid; CD68+ FCER1A+), melanocytes (MEL; MLANA+ DCT+), T cells (Tcell; CD3D+ CD3E+), keratinocytes (KC; KRT15+ KRT4+), B cells (Bcell; CD79A+ MS4A1+), fibroblasts (FB; PDGFRA+ SFRP2+), smooth muscle cells (Muscle; ACTA2+ TAGLN+), endothelial cells (Endo; CLDN5+ PECAM1+), and mast cells (MAST; TPSB2+ CPA3+) (Fig. 2a–c). A significant variation in cell proportions between oral mucosa and skin was observed, notably with a significant increase in endothelial cells in oral mucosa, explaining the enriched blood vessels in mucosal tissues (Fig. 2a).
a Unbiased clustering of 103,758 cells based on the single cell RNA-seq data from normal skin and oral mucosa reveals 9 cellular clusters. Clusters are distinguished by different colors. The identity of each cell cluster is shown at the top (Fig. 2a; left). The proportion of cell lineages in oral mucosa and skin (Fig. 2a; right). (O: oral mucosa; S: skin). b Violin plot analysis of cluster-specific genes for each cell type. c Heatmap of differentially expressed genes. For each cluster, the top 10 genes and their relative expression levels in all sequenced cells are shown (Fig. 2c; left). Bubble plot showing representative GO term enrichment of biological processes according to cell types (Fig. 2c: right). d UMAP plots based on the P2-WHGs score of all cells, and high P2-WHGs score cell clusters are highlighted. e Violin plots based on the P2-WHGs score of all cells. p values are from two-sided Wilcoxon rank-sum tests, ****p < 2.2e-16.
To identify cell types associated with P2 wound healing gene sets from single-cell RNA-seq data, we calculated P2 gene sets enrichment score for the cells of skin and oral mucosa based on the P2 gene sets (See Methods). The UMAP embedding of cell clustering was color-coded based on the P2 gene sets score for each cell, illustrating the activation of cell clusters in the P2 gene set (Fig. 2d). Notably, keratinocytes demonstrated significantly higher P2 gene sets scores, suggesting their potential involvement in the acute wound repair process (Fig. 2d). Interestingly, despite the lower abundance of keratinocytes in oral mucosa compared to skin, oral keratinocytes exhibited significantly higher P2 gene sets scores than their skin counterparts (Fig. 2e). This observation suggests that oral keratinocytes may possess a distinctive characteristic, potentially being equipped with a strengthened wound healing program compared to their counterparts in the skin, which is consistent with previous studies19,20.
Oral-specific SPRR1B+ KC4 is major contributor of P2 -WHGs expression
To decode which keratinocyte subtype is enriched with P2 gene set, we further classified keratinocytes into 5 cell clusters based on their gene expression patterns (KC1 to KC5) (Fig. 3a). Specifically, Ki67+ KC1 displayed high proliferation signature with high expression of TOP2A and MKI67; KRT14+ KC2 are featured as basal-like keratinocytes with the specific expression of basal cell marker KRT14 and DST; KRT5+ KC3 displayed a characteristic of early differentiation (KRT5, TXN); Both SPRR1B+ KC4 and KRT10+ KC5 are featured as terminally differentiated superbasal keratinocytes as KC5 expressed genes with spinous layer characteristics (KRT1, KRT10), while KC4 expressed genes involved in cornification (SPRR1B, CNFN, CLDN) and anti-inflammation (S100A8, S100A9, and IL36G), which are essential elements for skin wound healing and tissue remodeling (Fig. 3b, Supplementary Fig. 4).
a Identification of 5 distinct keratinocyte subtypes based on the single cell sequencing data (Fig. 3a; left). The proportion of each keratinocyte subtypes in mucosa and skin (Fig. 3a; right). (O: oral mucosa; S: skin). b Dot plot analysis of cluster-specific genes for each subtype of keratinocyte. c UMAP plots based on the P2-WHGs score of keratinocytes. High P2-WHGs score cell clusters are highlighted. d Violin plots based on the P2-WHGs score of each keratinocyte subclusters in oral mucosa and skin. Two-sided Wilcoxon rank-sum tests were performed. ****p < 2.2e-16. e GSEA enrichment plots for P2-WHGs signaling upregulated in oral KC4 compared to skin KC4.
As Ki67+ KC1 and SPRR1B+ KC4 were identified as subpopulations enriched in the oral mucosa (Fig. 3a), we proceeded to investigate whether these cells exhibit an enrichment of P2 wound healing gene sets. Remarkably, while P2 gene sets were elevated in all mucosal KC populations compared to their skin counterparts, the SPRR1B+ KC4 population displayed the highest scores (Fig. 3c, d). Constitutively, while a small subset of KC4 were also identified in the skin, those skin KC4 exhibited significant lower expression of the P2 gene set signature compared to mucosal KC4 (Fig. 3d, e). This suggests that SPRR1B+ KC4 cells are the primary contributors to the expression of P2 gene sets, potentially acting as more potent mediators for the rapid wound healing in the oral mucosa.
SPRR1B+ KC4 contribute to wound healing with characteristics of terminal differentiation and inflammation
Cellular trajectory analysis revealed distinct differentiation trajectories in keratinocyte subpopulations, which were composed of three trajectory paths (path-I, path-II, and path-III) (Fig. 4a). We found that the basal-like keratinocytes (KRT14+ KC2) were primarily enriched in the early phase of the trajectory, the proliferative keratinocytes (Ki67+ KC1) were located at the end of path-I, while the early differentiated keratinocytes (KRT5+ KC3) were distributed in the middle phases of path-II and path-III. The terminally differentiated superbasal keratinocyte subclusters, including SPRR1B+ KC4 and the KRT10+ KC5, were distributed in the late phases of path-II and path-III, respectively (Fig. 4b). Interestingly, the path-II was occupied by oral keratinocytes, while the path-III of the trajectory was mainly composed of skin keratinocytes (Fig. 4c). These results indicated that oral mucosal and cutaneous keratinocytes had similar origins named as KRT14+ basal-like keratinocytes, but differentiated into two different trajectories, dominated by cluster KRT10+ KC5 in skin and SPRR1B+ KC4 in oral mucosa, respectively. Indeed, the basal KC marker, KRT14, was primarily expressed at early phase of the differentiation trajectory, while the markers for the two highly differentiated KC subpopulations (KC4 and KC5), KRT10 and SPRR1B, were highly expressed at late phase (Fig. 4d).
a–c Reconstruction of a trajectory with keratinous subpopulation; Results of pseudotime analysis of keratinocytes, colored by subclusters (Fig. 4a), pseudotime (Fig. 4b) and samples (Fig. 4c). d Expression levels for each cell are colored and overlaid onto pseudotime. Genes were selected as specifically expressed in celltypes. Bright yellow indicates maximum relative expression, and dark blue indicates low or no expression of this gene, each were accompanied by color-coded pseudotime feature plots. e, f Representative immunofluorescent images of normal skin and oral mucosa stained with the basal cell marker keratin 14 (K14, red), differentiation markers keratin 10 (K10, green), and Small Proline-Rich Protein IB (SPRR1B, green). Scale bar 50 µm. g Gene Ontology analysis of marker genes of the KC4 keratinocyte cluster. h The violin plot represents the gene expression levels in different group. i Gene scoring analysis of KC4 between oral and skin using the indicated molecular signatures. Two-sided Wilcoxon rank-sum tests were performed (h, i). ****p < 2.2e-16.
To validate the distinct differentiation trajectories observed between skin and oral mucosa as identified by scRNA-seq, we performed immunofluorescent staining of KRT14, KRT10, and SPRR1B in human normal skin and oral mucosa biopsies (Fig. 4e, f). Consistent with the scRNA-seq findings, KRT14+ basal-like keratinocytes were located at the basal layer of both skin and oral mucosa. Notably, oral mucosa exhibited multiple layers of KRT14+ keratinocytes compared to skin tissue, which typically displayed a single layer of basal keratinocytes. As these basal keratinocytes serve as progenitor cells for differentiated keratinocytes during tissue regeneration in both skin and mucosa, this observation suggests a heightened regenerative capacity in oral mucosa due to its abundance of basal keratinocytes (Fig. 4e, f). In accordance with the findings from scRNA-seq data, the terminal differentiation marker KRT10 was more abundant in the skin, but rarely detected in oral mucosa (Fig. 4e). In contrast, SPRR1B was expressed higher in mucosal keratinocytes as compared with skin (Fig. 4f). These results suggested that the two terminally differentiated keratinocyte subpopulations were tissue specific, with SPRR1B+ KC4 mainly located in oral mucosa.
Gene set overrepresentation enrichment analysis of marker genes in oral mucosa KC4 revealed associations predominantly with keratinocyte differentiation, cornification, neutrophil activation, and responses to molecules of bacterial origin (Fig. 4g). This observation is highly consistent with the P2 gene signature as shown in Fig. 1c. Notably, compared with skin KC4, oral KC4 has shown increased expression of genes involved in inflammation (S100A8, S100A9, and IL36G) and epithelial-to-mesenchymal transition (EMT) (VIM, LUM, and COL1A1) (Fig. 4h). Furthermore, gene signature score analysis revealed elevated scores for EMT, wound healing, and inflammatory response in oral mucosa KC4 compared to their skin counterparts (Fig. 4i). Collectively, these findings demonstrate the pre-existence of wound healing-stimulated genes in oral keratinocytes within uninjured mucosa, with SPRR1B+ KC4 potentially playing a pivotal role in the primary repair of oral mucosa through its anti-inflammatory and promigratory effects.
STAT3 regulates SPRR1B expression of KC4 keratinocytes in oral mucosa
To unravel the regulatory mechanisms governing SPRR1B and other P2 gene sets in oral mucosa-specific KC4 keratinocytes, we first analyzed the transcription factors (TFs) accountable for regulating the expression of the P2 gene set within KC4 subpopulations. By utilizing SCENIC21 to estimate TF regulon activities based on the oral KC4 gene expression pattern, we observed heightened activation of TF regulons, including STAT3, KLF5, PRDM1, and GRHL1 in oral KC4 (Fig. 5a and Supplementary Table 1). To elucidate potential interactions among the transcription factors within KC4, we employed STRING, an online tool for retrieving interacting genes, to map TFs to a Protein–protein interactions (PPI) network22. Notably, STAT3 emerged as a hub TF in the PPI network, connecting with 6 nodes (Fig. 5b). STAT3 is well-recognized for its role in the control of inflammation and immunity23, aligning with the molecular characteristics of KC4. Moreover, STAT3 has been extensively established as a key transcription factor in promoting wound healing21,22.
a Heatmaps showing the transcriptional factors (TFs) which were differently expressed in each keratinocyte clusters. b The 10 TF genes with the highest scores in the Protein–Protein Interaction (PPI) network obtained by using the MCC algorithm of the CytoHubba plugin. c The predicted binding sites of STAT3 at the promoter region of SPRR1B gene. d Schematic of SPRR1B transcriptional start site (TSS) (red) and 0 to 500 bp upstream sequence. Design of PCR products for chromatin immunoprecipitation (ChIP) experiments (orange and red vertical lines) and predicted STAT3 binding sites (gray band) are indicated. e ChIP-PCR detection of STAT3 binding to the SPRR1B promoter region. f Western blot of STAT3, p-STAT3, JAK1, p-JAK1 and SPRR1B in HOK cells with JAK1/2 inhibitor ruxolitinib (Ruxo).
To validate whether the expression of SPRR1B is transcriptionally regulated by STAT3 regulate, we re-analyzed public CHIP-seq data of epithelial cells and revealed that STAT3 could bind to the SPRR1B promoter region (Fig. 5c). The detailed sequence of binding region was provided in Supplementary Table 2. CHIP-PCR confirming the direct interaction of STAT3 with the SPRR1B promoter within a region spanning 0 to 500 base pairs upstream of the transcriptional start site (TSS) (Fig. 5d, e). Furthermore, pharmacological inhibition of STAT3 utilizing a JAK inhibitor (Ruxolitinib) led to a decrease in STAT3 phosphorylation levels, accompanied by a corresponding reduction in SPRR1B protein levels (Fig. 5f). In summary, these findings support the notion that activation of the STAT3 pathway is pivotal for the induction of oral mucosa-specific SPRR1B+ keratinocytes, which facilitate as the preparatory steps for the rapid wound healing process within the oral mucosa.
In vivo validation of SPRR1B+ keratinocytes in uninjured oral mucosa and during skin wound healing
To validate if the identified SPRR1B+ keratinocytes pre-exsits both in uninjured oral mucosa and during skin wound healing, we implemented wound healing models in both skin and the oral cavity of mice. The experimental design for both the oral mucosa tongue and skin of mice is elucidated in the schematic diagram (Fig. 6a, b). The tongue was selected for the wound model due to the feasibility of performing wound surgery within the limited space of the mice’s oral cavity. Although the tongue differs from the buccal and gingival mucosa used in single-cell sequencing analysis, literature supports the similarity in cell composition and the oral bacterial microenvironment between the tongue and buccal/gingival mucosa24,25. To establish wounds in both oral and skin tissues, we adjusted the sectioning angle to ensure a complete presentation of the wound edges on slides. This adjustment allows for meticulous observation of growth and healing progression over time at the histological level (Fig. 6c). In accordance with Fig. 4f, we detected a higher proportion of SPRR1B+ KC in mouse oral mucosa compared to the skin in the uninjured state by Immunofluorescence staining (Fig. 6c). Furthermore, elevated expression of SPRR1B was seen during wound healing process in both wounded skin and wounded oral mucosa (Fig. 6d, e). The documented elevation in SPRR1B expression following injury in both skin and oral keratinocytes implies it may play indispensable role in the wound healing process.
a, b Schematic diagram of the experimental design in mice wound healing model. (Fig. 6a: skin; Fig. 6b: oral. Created with BioRender.com). The skin and tongue tissue of mice during the wound healing process are shown on the right (skin: n = 3, oral: n = 3). c Immunofluorescence images of oral and skin samples on different days during wound healing. Scale bar: 100 µm. d, e Quantification of the mean fluorescence intensity of SPRR1B over a 500-pixel area on either side of the skin and oral wound (n = 3). Error bars are shown as mean ± SEM from n = 3. One-way analysis of variance (ANOVA) were performed (d, e). *p < 0.05, **p < 0.01, ***p < 0.001. ns not significant.
SPRR1B enhances the migration ability of oral mucosal keratinocytes
SPRR1B (small proline-rich protein 1B), identified as a cornified envelope protein, stands out as the marker of KC4 in oral mucosa. Previous study has underscored the role of SPRR1B in promoting cell invasion and migration in lung adenocarcinoma26. Since motility of keratinocytes is vital for wound healing process, we hypothesize that SPRR1B+ KC4 accelerate mucosal wound healing while its increased cell motility and invasion driven by SPRR1B expression. To elucidate the contribution of SPRR1B to accelerated wound healing in the oral cavity, we initially assessed its expression levels in oral and skin keratinocyte cell line. Indeed, HOK cells, an oral keratinocyte line, has shown significantly higher SPRR1B expression compared to HaCaT cells, a skin keratinocytes line (Fig. 7a). To further investigate the functional role of SPRR1B in these cells, we employed lentiviral vectors to either knock down SPRR1B in HOK cells using three independent shRNAs (sh#1, sh#2, sh#3), or overexpress SPRR1B cDNA in HaCaT cells. Western blotting validated the successful knockdown of SPRR1B in HOK cells and overexpression of SPRR1B in HaCaT cells (Fig. 7b, e). To assess the impact of SPRR1B on keratinocyte migration, a wound healing assay was conducted using HaCaT cells overexpressing SPRR1B and HOK cells with SPRR1B knocked down, each compared with their respective control groups. The assay involved recording the migration of cells into the wounded area and those with extended protrusions from the wound borders at intervals immediately and every 3 h post-incubation. Photo recording and subsequent data analysis were performed automatically using Livecyte® Cell Analysis System (Fig. 7c, f). The results demonstrated that SPRR1B knockdown in HOK cells led to a reduction in their migration capacity, while SPRR1B overexpression in HaCaT cells enhanced their migration capacity (Fig. 7d, g). These findings underscore the role of SPRR1B in augmentation of keratinocyte migration capacity solely in oral mucosa, which contribute to the rapid wound healing in oral mucosa.
a Western blot of SPRR1B and internal control (GAPDH) between HOK and HaCat cells. b Western blot of SPRR1B in HOK cells expressing control shRNA or either of three independent shRNAs targeting SPRR1B. c Migrating HOK cells transduced with shRNAs of SPRR1B and shNC (control) by lentiviral delivery. Images were taken every 3 h from 0 to 12 h after scratching. d Line plot representing relative change in wound area size of HOK cells as a function of time (n = 3). e Western blot to show overexpression of SPRR1B after transfection with lentiviruses in HaCat cells. f Migrating HaCat cells transduced with SPRR1B and GFP (control) by lentiviral delivery. Images were taken every 3 h from 0 to 21 h after scratching. g Line plot representing relative change in wound area size of HaCat cells as a function of time (n = 3). Error bars are shown as mean ± SEM from n = 3. Two-way analysis of variance (ANOVA) were performed (d, g). *p < 0.05, **p < 0.01, ***p < 0.001. ns not significant.
Discussion
Wound healing has been classically described in skin wounds as occurring in sequential and interdependent phases such as hemostasis, inflammation, proliferation, and remodeling27, with the same phases having been reported for oral mucosa wound healing5. Despite the strong similarities between skin and oral mucosa regarding the temporal sequence of histologic and functional changes during wound healing, evidence has documented that oral mucosal wounds heal at a rate faster than skin wounds19, and typically without scar formation5. Understanding the biological mechanisms underlying oral mucosa wound healing will provide useful insights in efforts to enhance wound healing in other sites and organs.
To identify the mechanisms that define accelerated oral wound healing, we screened 485 DEGs that were significantly upregulated during wound healing process, which hold the key to the activation of the molecular events responsible for wound. Interestingly, 485 DEGs were unsupervised clustered into two modules. Module 2 has been significantly higher expressed in the oral mucosa than skin before the wound occurred and was defined as P2-Wound Healing-associated Gene Sets (P2-WHGs). These findings suggested that specific wound healing program has already been constitutively expressed in normal oral mucosa before wound initiation, but such healing program can only be gradually induced after wound production.
However, since wound healing is a complex multicellular cooperative process, the lack of high-resolution exploration of oral mucosal wound healing in vivo compared to skin wound healing limit our understanding of the mechanisms that accelerate wound healing. In this regard, we thoroughly explore the differences in the composition of Buccal and gingival mucosa as well as skin cell types with the advantage of single cell sequencing technology. We identified 9 cell types and the cell proportion varied greatly in the two tissue types. Endothelial cells, T cells, fibroblasts, and muscle cells, were significantly increased in oral mucosa, while subsets of keratinocytes and melanocytes were significantly decreased in oral mucosa. Indeed, Szpadersk et al. found that there are more blood vessels in the oral mucosa than in the skin based on the expression of α-smooth muscle actin (α-SMA) and CD31, which is consistent with our conclusion20. Integrated analysis of the bulk and scRNA-sequencing revealed that the oral-specific keratinocytes significantly upregulated wound healing transcriptional profiles compared to normal skin (Fig. 2e). Thus, our next work focused on the keratinocyte subset.
We further investigated different subpopulations of keratinocytes, which play an important role in the early stages of wound healing28. Interestingly, significant differences exist in keratinocytes when comparing skin and oral mucosa. First, keratinocytes were classified into five categories according to their functional and anatomical characteristics, and we found that the keratinocyte population KC1, represented by its proliferative characteristics, was significantly enriched in the oral mucosa. Research has shown that the proliferation of KC is beneficial to skin regeneration and wound re-epithelialization29. Moreover, Cellular trajectory analysis revealed a clear cell state transition from basal KC to the terminal differentiation keratinocytes that express keratinized and cornification proteins (KC4) and spinosum gene markers (KC5). Interestingly, KC4 and KC5 were at completely different branches in the late stage of the trajectory. The characteristics of KC4 only appeared in the oral mucosa, while the KC5 existed in the skin only. Our findings suggest that the oral keratinocytes diverge in very different trajectories after a brief parallel origin compared to skin (Fig. 4a–c), which may contribute to the mechanisms leading to distinct healing phenotypes in response to injury. Surprisingly, when integrated with the scRNA-seq data of homeostatic skin and oral mucosa, the expression of these 146 P2-WHGs was much higher in oral keratinocytes than in skin. Those genes were significantly enriched in KC4, indicating the KC4 population as a key cell subpopulation for accelerating oral mucosal healing.
Inflammatory memory, or “trained immunity”, endows cells with a newfound ability to respond more rapidly and powerfully to a wide variety of secondary attacks30. Initially thought to be unique to the immune system, inflammatory memory was recently identified in epithelial tissues31. It is known that the integrity of epithelial barrier provides the first line of defense against chemical, physical, or microbial damage32. Terminally differentiated keratinocytes tend to be closer to pathogens and external stimuli33. As expected, our data showed that the expression features of KC4 were mainly related to defense response to bacterial, neutrophil activation, and inflammatory response (Fig. 4g, h). Among the signature genes of KC4, S100A8 and S100A9 are actively released during inflammation and plays a key role in early stage of wound healing by stimulating leukocyte recruitment and inducing cytokine secretion34,35. We speculate that oral inflammatory microenvironment may induce the conversion of keratinocytes into inflammatory KC4 subsets, thus enhancing the ability of rapid response to oral injury or other secondary attacks in oral mucosa.
By single-cell transcriptional factors prediction and PPI network analysis, we identified STAT3 as a key transcription factor regulating wound healing-associated keratinocytes (KC4) differentiation. Signal transducer and activator of transcription 3 (STAT3) is a transcription factor that is activated by a variety of cytokines, such as the IL-22 and epidermal growth factor36,37. STAT3 has been shown to be important in wound healing. The current study proved that an antimicrobial peptide can accelerate wound healing by promoting angiogenesis and cell migration through FGFR/JAK2/STAT3 signaling38. Increased STAT3 phosphorylation in the corneal epithelium significantly accelerated wound healing39. To further explore the regulation of STAT3 on the KC4 cluster in specific differentiation as well as wound healing pathways, we verified the targeted regulation of the transcription factor STAT3 to KC4 signature genes in the chip-seq database. We speculate that the KC4 signature gene SPRR1B, its transcription was activated by STAT3 through promoter binding. And subsequent CHIP experiments strongly supported the speculation. SPRR1B (small proline-rich protein 1B) is a cornified envelope protein. Studies have shown that SPRR family genes can accelerate wound healing by acting as antimicrobial proteins to protect intestinal epithelial cells from bacterial invasion40. Proinflammatory cytokines IL-1β and IFN-gamma can activate SPRR1B expression via p38 MAPK-mediated signaling pathways41. We also verified the increased expression of SPRR1B during oral and skin wound healing in mouse wound healing model. The result demonstrated the specific expression of SPRR1B on differentiated oral keratinocytes relative to skin histologically, as well as significant transcriptomic upregulation in skin after injury. Further functional experiments showed that, the overexpression of SPRR1B increased the migratory ability in Keratinocytes, while the genetic knockdown of SPRR1B resulted in the opposite effects. These results indicated that SPRR1B may facilitate the migration of keratinocyte, suggesting that STAT3-induced overexpression of SPRR1B may be a potential strategy to promote wound healing. Nevertheless, the current functional studies were conducted exclusively in vitro, lacking in vivo validation. Establishing a mouse model with keratinocyte-specific SPRR1B knockout would be essential to determine the role of SPRR1B-positive keratinocytes in mucosal fast healing, which requires further investigation.
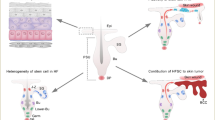
In summary, by integrative analysis of bulk mRNA seq and scRNA seq data during the oral mucosa and skin wound healing process, we found a new keratinocytes subtype, named SPRR1B+ keratinocytes, that was induced during both oral mucosa and skin wound healing. The SPRR1B+ keratinocytes expressed the major wound healing genes regulated by Transcription factor STAT3. Moreover, these wound healing related SPRR1B+ keratinocytes were constitutively enriched in human oral mucosa but not in skin, proving that SPRR1B+ keratinocytes prime oral mucosa for faster wound healing when compared with skin (Fig. 8), may have a critical application in the treatment of chronic or non-healing wounds in the future.
Methods
Human and sample information
Skin and oral mucosa tissue discards of human used for immunofluorescence staining were obtained from subjects of Shanghai Ninth People’s Hospital respectively. All human tissues were collected after informed consent for use of tissues in research, and further detailed information is available in Supplementary Table 3. Informed consent was obtained from all participants. The study was conducted in accordance with the Declaration of Helsinki, and approved by the Ethics Committee of the ninth People’s Hospital of Shanghai (grant numbers: SH9H-2022-TK379-1). All ethical regulations relevant to human research participants were followed.
Mice
Experiments involving C57BL/6J mice purchased from the biological company ShengChang (Shanghai, China). 6–8 weeks old male mice were used for this study. The mice were fed ad libitum, and handled and raised under standard conditions in the animal facility of Shanghai Institute of Precision Medicine. The methods were performed in accordance with relevant guidelines and regulations and approved by the Ethics Committee of the ninth People’s Hospital of Shanghai (grant numbers: SH9H-2022-A709-SB). We have complied with all relevant ethical regulations for animal use.
Cell culture
Human immortalized keratinocyte cell line (HaCaT cells) was a gift from Zheng Yang, Shanghai Jiao Tong University. Primary human oral keratinocytes (HOK cells) were purchased from ScienCell (Carlsbad, USA) and cells passages from 3 to 5 were used for the experiments. All the cell lines used in this study were tested for mycoplasma contamination and authenticated by genetic profiling using polymorphic short tandem repeats. HaCaT cells were cultured in Dulbecco’s modified eagle medium (DMEM, high glucose, Gibco), supplemented with 1% penicillin/streptomycin (P/S, PB180120, Pricella) and 10% fetal bovine serum (FBS, A5669701, Gibco). HOK cells were cultured in Oral keratinocyte medium (OKM, Cat.2611, ScienCell) consisted of 500 mL basal medium, 5 ml of oral keratinocyte growth supplement (OKGS, Cat.2652, ScienCell) and 5 ml of Antibiotic solution (P/S, Cat.0503, ScienCell). For passaging, cells at 70%–80% confluence were detached through a 2–5 min incubation with 0.05% Trypsin/EDTA and further resuspended in complete medium. All cells were cultured at 37 °C under 5% CO2 atmosphere.
Short hairpin RNA (shRNA) infection assay
HOK cells were transfected with SPRR1B-shRNA plasmid (h) (sh-SPRR1B-#1, sh-SPRR1B-#2, sh-SPRR1B-#3) or control-shRNA plasmid (shControl) which were purchased from HeYuan BioCompany. The sequences of the shRNAs were provided in the supplementary Table 4. HOK cells were transfected with SPRR1B-shRNA plasmid (h). 293 T cells were transfected using Lipofectamine™3000 (Invitrogen, L3000015, Thermo Fisher Scientific) according to the manufacturer’s protocol. The following ratio was used for vector transfection: Lentiviral vectors: packaging plasmid: envelope plasmid; 10 µg: 7.5 µg: 3 µg for 72 h at 37 °C in an incubator before the 293 T lentiviral supernatant was harvested by centrifugation at 200 × g for 5 min at room temperature. HOK and HaCat cells (2 × 105 cells/ml) in six-well plates were infected with the 293 T lentiviral supernatant in the presence of 5 µg/ml polybrene (40804ES86, Yeasen Biotechnology, Shanghai). Cells were collected 48 h after infection, and after removal of the particles containing medium, cells were incubated in culture medium with 10 µg/ml puromycin to positively select transduced cells. The efficiency of the shRNA knockdown was determined by Western blot analysis. The SPRR1B-KD HOK cells and SPRR1B-OE HaCaT cells were then used for additional experiments.
Chromatin immunoprecipitation
Experiments were performed using SimpleChIP Enzymatic Chromatin IP Kit according to the procedures provided by the manufacturer starting with 5 × 106 HOK cells. DNA and associated proteins on chromatin in cultured cells were crosslinked by 1% formaldehyde for 15 min at 37 °C. Cells were then scraped and collected in cellular lysis buffer (5 mM Pipes, 85 mM KCl, 0.5% NP-40, and protease inhibitors). Cytoplasmic lysates were discarded, and nuclear components were resuspended in nuclear lysis buffer (50 mM Tris pH 8, 10 mM EDTA pH 8, 0.2% SDS, and protease inhibitors), and sonicated for 10 min (Covaris). Approximately 4 mg of p-STAT3 antibody (#9145, diluted 1:1000, CST) or control IgG (#2729, diluted 1:1000, CST) were incubated with 25 ml of protein G magnetic beads for 6 h at 4°C, and then incubated with 100 mg of cleared chromatin overnight at 4 °C. After three washes, immunoprecipitated material was eluted at 55 °C for 1 h with 10 µg/ml proteinase K, and then decrosslinked at 65 °C for 4 h. Figure 5d shows the schematic of SPRR1B transcriptional start site (TSS) (red) and 0 to 500 bp upstream sequence. Design of predicted STAT3 binding sites (gray band) and PCR primers for ChIP experiments (orange and red vertical lines) and are also indicated. And the primers for detecting STAT3 binding to SPRR1B promoter region are listed in the Supplementary Table 5.
Migration assay
Confluent cell monolayers in 6-well plate were scratched with a 200 µl pipette tip to create a cell-free area. Debris were removed by washing once with PBS, then cells were incubated in DMEM supplemented with 10% FBS and 1% P/S. Images were taken at 3-hour intervals for three wells per condition by the Livecyte® Cell Analysis System (Phasefocus, Britain), until the cell fusion reached 100%. Subsequently, Single cell migration was analyzed using Livecyte’s label-free, quantitative phase imaging (QPI) mode and Scratch Wound Dashboard automatically. Three independent experiments were performed.
Mouse model of wound healing
The mice were allowed to acclimate for three days before the experiments. Mice were anesthetized with Tribromoethanol (Sigma Aldrich, T48402-25G, 200U/25 g), and 18 healthy mice were randomly divided into six groups of 3 mice each. For skin, a surgical knife blade (LianHui, 20010243, Shanghai) was used to generate a full-thickness incisional wound on the back of the mouse. The unwounded group was sacrificed after the first biopsy, while the two wounded groups were sacrificed on day 2 and day 4, respectively. For oral cavity, the wounds were made in the dorsum of the mouse tongues with the same surgical knife blade. The unwounded group was sacrificed without treatment to obtain tongue tissue, and the two wounded groups were sacrificed on day 2 and day 4 after the trauma. Depth is controlled by marking a mark on the knife, 1 mm from the knifepoint. And length is controlled to be approximated at 5 mm. Skin and tongue samples were harvested in 4% paraformaldehyde for fixation and immunefluorescence.
Integration and analysis of scRNA-Seq data
Publicly available scRNA-seq data of human oral mucosa samples (GSE164241) were obtained from Gene Expression Omnibus (GEO) database and can be accessed via the following link: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE164241. Publicly available human skin scRNA-seq datasets were retrieved from the Genome Sequence Archive (GSA) under accession number PRJCA006797.
Both sequencing datasets were from 10X Genomics and analyzed by the Seurat R package42. In the first dataset (GSE164241), 8 buccal mucosa and 13 gingival mucosa were used to represent healthy oral samples12 and integrated with 5 health skin samples43 from the control group in the second dataset (PRJCA006797). The Merge function was used to merge multiple datasets. The percentage of mitochondrial genes was calculated by the PercentageFeatureSet function, and the quality control parameters were set as follows: 500 <= nCount_RNA <= 25,000; 300 <= nFeature_RNA <=;3000; and a mitochondrial gene percentage <= 15%. (Supplementary Fig. 1).
The sequencing data after quality control were standardized by LogNormalize. Then the normalized expression was scaled and the top 2000 highly variant genes were screened according to the FindVariableFeatures function. Subsequently, principal component analysis was performed based on the 2000 variable genes as the input and identified significant principal components (PCs) based on the ElbowPlot function. The top 30 PCs were selected as the input to further reduce the dimension using uniform manifold approximation and projection (UMAP).
To minimize batch effects between datasets, R package “Harmony” was used for batch correction44 (Supplementary Fig. 2a, b). Cluster analysis was performed using the FindNeighbors function (dims = 1:30) and the FindClusters function (resolution = 0.5). Next, FindAllMarkers function was used to identify marker genes in each cluster. Cluster identification was performed based on the top DEGs of each cluster. For sub clustering of keratinocytes, we first identified keratinocyte clusters based on marker genes and then subset out the appropriate keratinocyte clusters. These keratinocyte clusters were then run with Seurat pipline using 20 PCs with resolution = 1 and corrected with RunHarmony function again.
Bulk RNA-Seq data analysis
The publicly available bulk RNA-seq datasets were downloaded from the GEO database (GSE97615).The DEGs between the health and wounded groups were analyzed by the limma package45 and were screened under |log2 (Fold Change)| > 1 and adjusted p < 0.05. The Volcano plot shows these DEGs during skin wound healing process (Supplementary Fig. 3). The list of these DEGs can be found in Supplementary Data 1.
Identification of the transcription binding sites of STAT3
The transcription binding sites of STAT3 was identified from a publicly available Chromatin immunoprecipitation (ChIP-seq) data (National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA) accession: SRP070705). The paired-end reads (sample accessions: SRR3184130, SRR3184127, SRR3184126) were mapped to human reference genome (hg38 version) by bowtie v2.3.4.346 with default options. The peaks were called by MACS v2.1.247 with default options, and were annotated by R annotatr package48. The genes with promoter binding sites of STAT3 were predicted as transcriptionally regulatory target genes of STAT3.
Gene set score analysis
The “AddModuleScore” function of the Seurat R package was used to calculate module scores for gene expression programs at the single-cell level. First, all the analyzed genes were binned based on the average expression, and the control genes were randomly selected from each bin. Then, calculate the average expression value of the gene set at the single-cell level minus the aggregated expression of the control gene set. The wound healing-associated genes were defined by analysis of the bulk RNA-seq data described above, and other gene sets were obtained from the MSigDB database (https://www.gsea-msigdb.org/gsea/msigdb/).
Functional enrichment analysis
Normalized gene expression data were used for pathway analysis. Gene Ontology functional enrichment49 of DEGs was performed by R package “clusterProfiler”50 (https://github.com/YuLab-SMU/clusterProfiler) and visualized with the R package “ggplot2” (https://github.com/tidyverse/ggplot2). GO terms were identified with a cutoff of P < 0.05 and a false discovery rate (FDR) < 0.05. Representative terms selected from the top 15 ranked GO terms or pathways were displayed.
Western blot analysis
Total proteins were extracted from the keratinocytes and quantified by using a BCA Protein Assay kit (Beyotime, P0009, China). The extracted proteins were separated on 10 or 15% sodium dodecyl sulfate-polyacrylamide gels and then transferred onto polyvinylidene fluoride (PVDF) membranes. The membranes were incubated overnight at 4 °C with the following primary antibodies: JAK1 (CST, # 3344, diluted 1:1000); phospho-JAK1 (CST, # 3331, diluted 1:1000); STAT3 (HUABIO, # ET1607-38, diluted 1:1000); phospho-STAT3 (CST, # 9145, diluted 1:1000); SPRR1B (invitrogen, PA5-107277, Thermo Fisher Scientific, diluted 1:1000) and GAPDH (Abcam, ab263962, diluted 1:2000). Finally, the membranes were incubated with corresponding secondary antibodies for 60 min at room temperature, and densitometry measurements of the western blot bands were taken by using Image-Pro Plus 6.0 software (Media Cybernetics, USA).
Immunofluorescence staining
Immunofluorescence staining was performed on 4% paraformaldehyde-fixed, OCT-embedded human normal skin and oral mucosa tissues. Fresh tissue was soaked in 4% paraformaldehyde and incubated overnight at 4 °C, then transferred into 30% sucrose for gradient dehydration until settling. Next, the tissue was washed in PBS, embedded in Optimal cutting temperature compound (SAKURA Tissue-Tek® O.C.T. Compound 4583), and frozen. Subsequently, section the tissue at a range of 6–10 µm and place on slides in preparation for tissue staining. In brief, sections were fixed in 4% PFA (Biosharp; BL539A) for 5 min, permeabilized with 22 mg/ml Glycine solution (Adamas-beta; 56-40-6) for 15 min, and blocked with PBS–BSA 2% for 60 min with Blocking solution (Meilunbio; 9048-46-8), all at room temperature. Primary antibody, diluted in Blocking solution, was added to skin sections overnight at 4 °C (KRT14: Abcam, abcam7800, dilution 1:1000; KRT10: Abcam, abcam76318, dilution 1:5000; SPRR1B: Thermo Fisher, PA5-107277, dilution 1:400). The next day, sections were washed 3 times with PBS and then labeled with Alexa Fluor 488 (Thermo Fisher, A-11001, dilution 1:2000) and Alexa Fluor 568 (Thermo Fisher, A11011, dilution 1:2000) labeled secondary antibodies. Cover slips using aqueous mounting medium containing DAPI. Images were captured using a Zeiss 880 confocal microscope (ZEISS, LSM880 with Airyscan) and analyzed with Zeiss Blue (ZEISS).
Construction of PPI and screening of hub genes
The network of PPI was constructed by STRING22, an online search tool for retrieving interacting genes. Protein–protein interactions of genes were analyzed using Cytoscape software (https:// cytoscape.org/)51. The top 10 hub genes were identified by the Monte Carlo Collisions (MCC) algorithm of the CytoHubba plugin52.
Pseudotime trajectory analysis
Pseudotime trajectory was constructed using matrix of cells and gene expressions with the R package Monocle253. First, ordering genes were identified by the“VariableFeatures” function of R package Seurat. Next, the discriminative dimensionality reduction with trees (DDRTree) method was performed for data interpretation. To model how keratinocytes cycle and assume different cell fates, the cells with basal characteristics were chosen as the root population. Finally, the trajectory was visualized through the “plot_cell_trajectory” function.
SCENIC analysis
The SCENIC analysis was performed using default and recommended parameters as directed on the SCENIC vignette (https://github.com/aertslab/SCENIC)21. Transcription factors (TFs) of hg19 were used as reference TFs and downloaded using RcisTarget. The expression matrix firstly extracted from Seurat to construct expression matrix of desired states. Based on these matrices, co-expression modules were identified between transcription factors (TFs) and potential target genes filtered by importance through GENIE3. Genes with significant motif enrichment were considered as direct target genes while each transcription factor and its direct target genes were defined as a regulon. Ranked genes were then used to score each regulatory gene based on activity values. Networks of the TF modules were visualized by Cytoscape (version 3.9.1)51,54.
Statistical analysis
All statistical analysis were performed using R (4.0.1). The statistical significance between two groups was analyzed using Student’s t test (parametric) and Wilcoxon rank-sum test (non-parametric). For all figures, asterisks denote statistical significance, *, **, *** and **** indicate p < 0.05, p < 0.01, p < 0.001 and p < 0.0001, respectively. Statistical significance was defined as p < 0.05.
Data availability
The bulk gene expression data are from the NCBI Gene Expression Omnibus (GEO) with accession code GSE97615. Publicly human oral mucosa scRNA-seq datasets are from the NCBI Gene Expression Omnibus (GEO) with accession code GSE164241. Publicly available human skin scRNA-seq datasets were retrieved from the Genome Sequence Archive (GSA) under accession number PRJCA006797. GO gene-sets are from MSigDB (http://www.gsea-msigdb.org/gsea/ downloads.jsp). All source data behind the graphs are available within the article or supplementary Information. The uncropped and unedited blots can be found in Supplementary Fig. 5. Key plasmids generated from this work are available from Addgene: pSLenti-U6-SPRR1B-shRNA1 (Addgene ID: 225919); pSLenti-U6-SPRR1B-shRNA2 (Addgene ID: 225920), pSLenti-U6-SPRR1B-shRNA3 (Addgene ID: 225921), PLVX-Zsgreen-IRES-SPRR1B (Addgene ID: 225922).
References
Martin, R. F. Wound healing. Surg. Clin. North Am. 100, ix–xi (2020).
Choi, M. et al. The case : a nonhealing skin ulcer in a patient 5 years after successful transplantation. Kidney Int. 100, 1357–1358 (2021).
Kuntz, T., Koushk-Jalali, B. & Kreuter, A. Sunitinib-induced pyoderma gangrenosum-like skin ulcer. Cmaj 192, E552 (2020).
Pugliese, E., Coentro, J. Q., Raghunath, M. & Zeugolis, D. I. Wound healing and scar wars. Adv. Drug Deliv. Rev. 129, 1–3 (2018).
Waasdorp, M. et al. The bigger picture: why oral mucosa heals better than skin. Biomolecules 11, 1165 (2021).
Roh, J. L., Lee, J., Kim, E. H. & Shin, D. Plasticity of oral mucosal cell sheets for accelerated and scarless skin wound healing. Oral. Oncol. 75, 81–88 (2017).
Mak, K. et al. Scarless healing of oral mucosa is characterized by faster resolution of inflammation and control of myofibroblast action compared to skin wounds in the red Duroc pig model. J. Dermatol. Sci. 56, 168–180 (2009).
Waasdorp, M. [How does saliva contribute to wound healing?]. Ned. Tijdschr. Tandheelkd. 129, 275–278 (2022).
Iglesias-Bartolome, R. et al. Transcriptional signature primes human oral mucosa for rapid wound healing. Sci. Transl. Med. 10, eaap8798 (2018).
Turabelidze, A. et al. Intrinsic differences between oral and skin keratinocytes. PLoS ONE 9, e101480 (2014).
Wagner, A., Regev, A. & Yosef, N. Revealing the vectors of cellular identity with single-cell genomics. Nat. Biotechnol. 34, 1145–1160 (2016).
Williams, D. W. et al. Human oral mucosa cell atlas reveals a stromal-neutrophil axis regulating tissue immunity. Cell 184, 4090–4104.e4015 (2021).
Wang, S. et al. Single cell transcriptomics of human epidermis identifies basal stem cell transition states. Nat. Commun. 11, 4239 (2020).
Zou, Z. et al. A single-cell transcriptomic Atlas of human skin aging. Dev. Cell 56, 383–397.e388 (2021).
Li, Q. et al. Single-cell transcriptome profiling reveals vascular endothelial cell heterogeneity in human skin. Theranostics 11, 6461–6476 (2021).
Haensel, D. et al. Defining epidermal basal cell states during skin homeostasis and wound healing using single-cell transcriptomics. Cell Rep. 30, 3932–3947.e3936 (2020).
Guerrero-Juarez, C. F. et al. Single-cell analysis reveals fibroblast heterogeneity and myeloid-derived adipocyte progenitors in murine skin wounds. Nat. Commun. 10, 650 (2019).
Eming, S. A., Martin, P. & Tomic-Canic, M. Wound repair and regeneration: mechanisms, signaling, and translation. Sci. Transl. Med. 6, 265sr266 (2014).
Szpaderska, A. M., Zuckerman, J. D. & DiPietro, L. A. Differential injury responses in oral mucosal and cutaneous wounds. J. Dent. Res. 82, 621–626 (2003).
Szpaderska, A. M., Walsh, C. G., Steinberg, M. J. & DiPietro, L. A. Distinct patterns of angiogenesis in oral and skin wounds. J. Dent. Res. 84, 309–314 (2005).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–d613 (2019).
Hillmer, E. J., Zhang, H., Li, H. S. & Watowich, S. S. STAT3 signaling in immunity. Cytokine Growth Factor Rev. 31, 1–15 (2016).
Akintoye, S. O. & Mupparapu, M. Clinical evaluation and anatomic variation of the oral cavity. Dermatol Clin. 38, 399–411 (2020).
Yu, R. et al. IL-22 mediates the oral mucosal wound healing via STAT3 in keratinocytes. Arch. Oral. Biol. 72, 14–20 (2016).
Zhang, Z. et al. Identification of small proline-rich protein 1B (SPRR1B) as a prognostically predictive biomarker for lung adenocarcinoma by integrative bioinformatic analysis. Thorac. Cancer 12, 796–806 (2021).
Glim, J. E., van Egmond, M., Niessen, F. B., Everts, V. & Beelen, R. H. Detrimental dermal wound healing: what can we learn from the oral mucosa? Wound Repair Regen. 21, 648–660 (2013).
Moll, R., Divo, M. & Langbein, L. The human keratins: biology and pathology. Histochem Cell Biol. 129, 705–733 (2008).
Gandarillas, A. & Freije, A. Cycling up the epidermis: reconciling 100 years of debate. Exp. Dermatol 23, 87–91 (2014).
Larsen, S. B. et al. Establishment, maintenance, and recall of inflammatory memory. Cell Stem Cell 28, 1758–1774.e1758 (2021).
Ordovas-Montanes, J. et al. Allergic inflammatory memory in human respiratory epithelial progenitor cells. Nature 560, 649–654 (2018).
Gallo, R. L. Human skin is the largest epithelial surface for interaction with microbes. J. Investig. Dermatol 137, 1213–1214 (2017).
Harris-Tryon, T. A. & Grice, E. A. Microbiota and maintenance of skin barrier function. Science 376, 940–945 (2022).
Kerkhoff, C. et al. Novel insights into the role of S100A8/A9 in skin biology. Exp. Dermatol 21, 822–826 (2012).
Su, W., Wang, P., Dong, Q., Li, S. & Hu, S. S100A8 accelerates wound healing by promoting adipose stem cell proliferation and suppressing inflammation. Regen. Ther. 21, 166–174 (2022).
Pickert, G. et al. STAT3 links IL-22 signaling in intestinal epithelial cells to mucosal wound healing. J. Exp. Med. 206, 1465–1472 (2009).
Hsueh, Y. J. et al. STAT3 regulates the proliferation and differentiation of rabbit limbal epithelial cells via a ΔNp63-dependent mechanism. Investig. Ophthalmol. Vis. Sci. 52, 4685–4693 (2011).
Takahashi, M. et al. The antimicrobial peptide human β-Defensin-3 accelerates wound healing by promoting angiogenesis, cell migration, and proliferation through the FGFR/JAK2/STAT3 signaling pathway. Front. Immunol. 12, 712781 (2021).
Wang, X. et al. The proinflammatory cytokines IL-1β and TNF-α modulate corneal epithelial wound healing through p16(Ink4a) suppressing STAT3 activity. J. Cell Physiol. 235, 10081–10093 (2020).
Hu, Z. et al. Small proline-rich protein 2A is a gut bactericidal protein deployed during helminth infection. Science 374, eabe6723 (2021).
Li, S., Gallup, M., Chen, Y. T. & McNamara, N. A. Molecular mechanism of proinflammatory cytokine-mediated squamous metaplasia in human corneal epithelial cells. Investig. Ophthalmol. Vis. Sci. 51, 2466–2475 (2010).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Xu, Z. et al. Anatomically distinct fibroblast subsets determine skin autoimmune patterns. Nature 601, 118–124 (2022).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Cavalcante, R. G. & Sartor, M. A. annotatr: genomic regions in context. Bioinformatics 33, 2381–2383 (2017).
Ashburner, M. et al. Gene ontology: tool for the unification of biology.The gene ontology consortium. Nat. Genet. 25, 25–29 (2000).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics 16, 284–287 (2012).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Chin, C. H. et al. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 8, S11 (2014). Suppl 4.
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Doncheva, N. T., Morris, J. H., Gorodkin, J. & Jensen, L. J. Cytoscape StringApp: network analysis and visualization of proteomics data. J. Proteome Res 18, 623–632 (2019).
Acknowledgements
We thank Shufang He, Hong Lu and other members in the Bioimaging facility of Shanghai Institute of Precision Medicine for providing the technical support. We thank the support from Zeng Lab members at Shanghai Institute of Precision of Medicine for scientific discussion. This research was funded by: the National Natural Science Foundation of China (Grant No. 82204421, H.Z.); Innovative Research Team of High-Level Local Universities in Shanghai (Grant No. SHSMU-ZLCX20211700, H.Z. and M.L.); the Industry Support Foundation of Huangpu District, Shanghai (Grant No. XK2020007, H.X.).
Author information
Authors and Affiliations
Contributions
X.X., L.Z., and H.Z. conceived and designed the study. X.X. performed and L.Z. supervised the data acquisition and analysis. X.X. performed the mice experiments. R.L. and P.H. performed the in vivo experiments. Y.Z. provided clinical resources. Y.M., Y.Z., M.L., and R.J. provided insightful discussions. X.X. wrote the initial draft of the manuscript. H.Z., H.X., and L.Z. provided critical revisions to the manuscript. All authors have read and agreed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Eirini Trompouki and Joao Valente. [A peer review file is available.]
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xuanyuan, X., Zhang, L., Zheng, Y. et al. SPRR1B+ keratinocytes prime oral mucosa for rapid wound healing via STAT3 activation. Commun Biol 7, 1155 (2024). https://doi.org/10.1038/s42003-024-06864-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06864-5
- Springer Nature Limited