Abstract
Non-alcoholic fatty liver disease (NAFLD) is a common, progressive liver disease strongly associated with the metabolic syndrome. It is unclear how progression of NAFLD towards cirrhosis translates into systematic changes in circulating proteins. Here, we provide a detailed proteo-transcriptomic map of steatohepatitis and fibrosis during progressive NAFLD. In this multicentre proteomic study, we characterize 4,730 circulating proteins in 306 patients with histologically characterized NAFLD and integrate this with transcriptomic analysis in paired liver tissue. We identify circulating proteomic signatures for active steatohepatitis and advanced fibrosis, and correlate these with hepatic transcriptomics to develop a proteo-transcriptomic signature of 31 markers. Deconvolution of this signature by single-cell RNA sequencing reveals the hepatic cell types likely to contribute to proteomic changes with disease progression. As an exemplar of use as a non-invasive diagnostic, logistic regression establishes a composite model comprising four proteins (ADAMTSL2, AKR1B10, CFHR4 and TREM2), body mass index and type 2 diabetes mellitus status, to identify at-risk steatohepatitis.
Similar content being viewed by others
Main
Non-alcoholic fatty liver disease (NAFLD) is a chronic, progressive condition affecting about 25% of the global population that is strongly associated with features of the metabolic syndrome, including obesity and type 2 diabetes mellitus (T2DM)1. NAFLD is characterized by excessive accumulation of hepatic triglyceride and encompasses a range of disease states: from steatosis (non-alcoholic fatty liver, NAFL) through non-alcoholic steatohepatitis (NASH), defined by the presence of hepatocyte ballooning and lobular inflammation with increasing fibrosis stage, to cirrhosis and hepatocellular carcinoma1. Not every patient diagnosed with NAFL will develop NASH or progress to cirrhosis and end-stage liver disease, meaning that there is substantial interindividual variation in disease severity. Patients with greater steatohepatitic disease activity, defined by a histological NAFLD Activity Score (NAS, the sum of steatosis, hepatocyte ballooning and lobular inflammation) more than or equal to 4 with fibrosis stage of 2 or more (F ≥ 2) are considered to show ‘at-risk NASH’ that indicates a high likelihood of progressive disease2,3.
Several non-invasive tests have been developed to identify patients with advanced liver fibrosis. These include use of indirect markers reflecting liver function and biochemical changes, such as the NAFLD Fibrosis Score (NFS) or the FIB-4 (ref. 4), and biomarkers that directly measure collagen turnover, including cleaved pro-collagen type 3 peptide5 or thrombospondin-2 (ref. 6). The FibroScan-AST (FAST) score based on imaging assessment has proved to be an efficient way to identify NASH patients considered to be at risk of progressive disease7. More recently, proteomic approaches have been used to identify classifiers that differentiate advanced from early fibrosis8,9. In contrast, effective biomarkers that identify steatohepatitis and grade activity remain elusive, the field therefore relies on histological assessment that is invasive and has considerable interobserver variability.
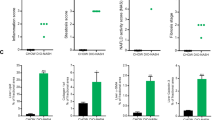
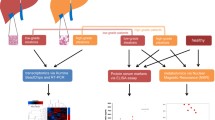
In this study, we integrate proteomics and RNA sequencing (RNA-seq) approaches to understand pathophysiological changes associated with NAFLD in humans and establish whether candidate circulating biomarkers might originate from the liver (Fig. 1a); a similar approach to that used recently in human alcoholic liver disease10 and NAFLD animal models11. Our study included 336 samples from patients with histologically characterized NAFLD derived from the European NAFLD Registry12. The discovery cohort comprised 191 plasma samples and the independent validation cohorts included 115 serum samples together with 30 liver biopsies. Within the discovery cohort, 38.4% were female, the average age was 55.2 (±11) years, average body mass index (BMI) was 33.5 (±6.7) and 60.7% had type 2 diabetes (Table 1 and Supplementary Table 1). Samples were processed for proteomics using the SomaScan v.4.0 platform, measuring 4,730 unique proteins and reads were corrected for sex, centre and T2DM (Extended Data Fig. 1). When stratifying patients on the basis of fibrosis stage (F), ranging from 0 to 4 and comparing advanced (F3–4) with mild (F0–2), we found 117 unique proteins (121 probes) to be differentially expressed (Fig. 1b and Supplementary Table 2). Functional annotation enrichment clustered proteins correlating to pathways such as ‘cell adhesion’, ‘inflammatory response’ and ‘carbohydrate metabolism’ (Fig. 1c). When stratifying patients on the basis of a high disease activity using NAS ≥ 4, we found 52 differentially expressed proteins (53 probes) (Fig. 1d and Supplementary Table 3). Enrichment analysis grouped proteins relating to ‘lipid metabolism’, ‘amino-acid biosynthesis’ or ‘bile acid catabolism’ (Fig. 1e). The two comparisons, advanced fibrosis and NAS ≥ 4, shared 30 differentially expressed proteins (Fig. 1f). When looking at the top 50 most significant differentially expressed proteins for each of these two comparisons, different dynamic expression patterns were observed as NAFLD progressed (Fig. 1g). Clear differences were seen in proteins associated with fibrogenesis and steatohepatitis during the pathogenesis of NAFLD: proteins associated with steatohepatitic activity (NAS ≥ 4) tended to peak in NASH F2–3 and then fall with progression to cirrhosis. By contrast, proteins purely associated with fibrosis increased steadily, peaking in cirrhosis (F4) (Fig. 1g).
a, Schematic overview of the study. b,c, Differentially expressed proteins in the discovery cohort of 191 patients stratified on the basis of fibrosis stage F3–4 versus baseline F0–2 (Benjamini–Hochberg false discovery rate) (b) and functional annotation enrichment analysis (c). d,e, Differentially expressed proteins in 191 patients stratified on the basis of a high disease activity score NAS ≥ 4 (Benjamini–Hochberg false discovery rate) (d) and enrichment analysis (e). f, Venn diagram showing overlap between the two different analyses. g, Heatmap showing expression of top 50 most significant proteins associated with fibrosis and NAS ≥ 4. FC, fold change.
To establish that the circulating proteins were of hepatic origin, and to further characterize their cellular origins within the liver, we conducted a two-stage analysis. First, a proteo-transcriptomic comparison in a cohort of matching plasma-liver biopsy samples that were a subset of the discovery cohort, and second, an integrated single-cell RNA-seq and tissue expression analysis using publicly available data13.
We performed linear correlations between circulating proteins on the basis of the SomaScan read-out and hepatic messenger RNA obtained from RNA-seq analysis in a subset of 52 cases from the discovery cohort with matching plasma-liver biopsy samples. Here, 4,584 protein probes, matching 4,292 proteins and/or genes, were identified in the RNA-seq data, of which 194 significantly correlated with each other (Fig. 2a and Supplementary Table 4). Within these 194 correlations, 31 proteins had been identified in the two previous comparisons described above (Fig. 2a). Eight of these 31 signals were associated with both NAS ≥ 4 and advanced fibrosis (F3–4), including THBS2, APOF, ADAMTSL2, CFHR4, TREM2, AKR1B10, SULT2A1 and PTGR1. In addition, 21 positive correlations were uniquely identified in the advanced fibrosis comparison, including GDF15, IGFBP7 and SHBG, while two correlations were from the NAS ≥ 4 comparison (ADSSL1 and ENO3) (Fig. 2a).
a, Pearson correlation analysis between plasma proteome (4,584 protein probes, matching 4,292 proteins and/or genes) and matching hepatic mRNA in a subset of the discovery cohort (n = 52). b, Integrated single-cell RNA-seq analysis to deconvolute the 31 signatures of proteins associated with hepatic mRNA.
GTex tissue expression analysis indicated that several of the 31 proteins in the signature are enriched in normal human liver, including the markers APOF, CFHR4, PTGR1, SULT2A1 and SHBG (Extended Data Fig. 2). Additionally, supervised analysis using bulk RNA-seq data from a large cohort of 206 patients with NAFLD14, showed that most of our signature changes occur in patients with advanced fibrosis and/or NAS ≥ 4 (Supplementary Table 5). Integrated single-cell RNA-seq analysis showed that the 31 signature proteins can be found in different hepatic cell populations (Fig. 2b). Of the 31 markers, 18 were enriched in epithelial cells, hepatocytes or cholangiocytes (including AKR1B10, CFHR4 and PTGR1) compared to other hepatic cells, while other markers were primarily restricted to fibroblasts (ADAMTSL2, THBS2) or macrophages (CXCL8 and TREM2) (Fig. 2b).
To demonstrate the potential power of our proteo-transcriptomics signature strategy to support development of new non-invasive diagnostics to detect fibrosing-steatohepatitis, we performed logistic regression analysis to identify patients with ‘at-risk NASH’, defined as NAS ≥ 4 (with at least one point deriving from each NAS component) plus F ≥ 2 fibrosis. Backward elimination of variables identified a composite model in the discovery cohort (n = 191) that could classify patients with at-risk NASH with an area under the curve (AUC) of 0.878 (±0.025) including the variables BMI, T2DM and circulating ADAMTSL2, AKR1B10, CFHR4 and TREM2 (Fig. 3a), independent from any other clinical variables. The classification model had a positive predictive value of 0.79 and a negative predictive value of 0.85 (Supplementary Table 6). It significantly outperformed established non-invasive tests including the FIB-4, NFS and aspartate transaminase (AST) to alanine transaminase (ALT) ratio scores in the entire discovery cohort of 191 patients, and had a higher AUC compared to the FAST score, which was available for a subset of 62 patients (Fig. 3b, Extended Data Fig. 3 and Supplementary Table 6). These findings were validated in an independent cohort of 115 samples where the model had an AUC of 0.80 (±0.04) (Fig. 3c, Extended Data Fig. 3 and Supplementary Table 6).
a, Binary logistic regression modelling identified a composite model in the discovery cohort (n = 191 patients) that could classify patients with at-risk NASH on the basis of the variables BMI, T2DM and circulating ADAMTSL2, AKR1B10, CFHR4 and TREM2. b, Performance of the classification model in two independent cohorts (discovery n = 191 patients, validation n = 115 patients) in comparison with NFS, Fibrosis-4 (FIB-4) and AST:ALT ratio. Bar charts present AUC for each score with the corresponding standard error of area, as calculated by ROC analysis. Paired-sample area difference under the ROC curve test was used to compare the classification model with the other scores (discovery cohort NFS P = 0.000001, FIB-4 P = 7.8667 × 10−7 and AST:ALT ratio P = 6.85 × 10−10; validation cohort NFS P = 0.000429, FIB-4 P = 0.000949 and AST:ALT ratio P = 0.000036) (****P < 0.0001, ***P < 0.001). c, Representative images of immunohistochemistry and quantification for AKR1B10 in human liver biopsies (n = 30 biologically independent patient samples). Scale bars, 100 μm. Data are presented as mean ± s.e.m. (Kruskal–Wallis with Bonferroni correction and Mann–Whitney U-test). Arrows indicate necro-inflammatory region with ballooned hepatocytes.
In this study, we have identified proteo-transcriptomic connections associated with features of progressive NAFLD. While only CFHR4 is uniquely expressed in healthy liver (Extended Data Fig. 2), ADAMTSL2, AKR1B10 and TREM2 have been previously been reported to play a role in the progression of liver diseases and NAFLD. Single-cell RNA-seq has showed that TREM2-positive macrophages are associated with hepatic portal fibrosis, while ADAMTSL2 reflects a zonal activation of hepatic stellate cells15,16. Soluble ADAMTSL2 proved to be a good biomarker to identify significant and advanced fibrosis in patients with NAFLD, while circulating TREM2 levels have proved to stratify patients with NASH9,17. Soluble levels of TREM2 are believed to reflect the recruitment and expansion of TREM2-positive macrophages localizing to fibrotic areas in the liver, in a response to resolve steatohepatitis18. Using a high-throughput RNA-seq approach in a cohort of 206 NAFLD biopsies to understand the pathogenesis disease progression, we recently showed that changes in transcription of the epithelial markers AKR1B10 and GDF15 can also lead to altered circulating concentrations of these proteins, serving as putative biomarkers for fibrosing-steatohepatitis14. To support these findings, we performed immunohistochemical stainings on series of 30 NAFLD biopsies. AKR1B10 positivity was more prominent in advanced NAFLD, and was observed in ballooned hepatocytes and hepatocytes neighbouring necro-inflammatory foci and periportal/periseptal areas (Fig. 2d).
This study has some limitations as we assessed linear associations between protein and hepatic mRNA in a European White cohort only, which does not exclude the potential contribution of other organs to the expression of the circulating proteins or that other factors contribute in different ethnic groups. We were also limited in our ability to confirm some proteomic findings in hepatic tissue due to limited availability of appropriate antibodies. Nevertheless, we have highlighted the complexity of the different liver cell populations and showed that circulating proteins correlating with hepatic mRNA can be used to identify patients with at-risk NASH.
Methods
Patient selection
A total of 336 histologically characterized cases were derived from the European NAFLD Registry (NCT04442334); samples were collected as previously described12. European White patients have been treated and diagnosed for NAFLD on the basis of histology at specialized centres including Angers and Paris (France), Mainz (Germany), Turin (Italy), Linköping (Sweden) and Newcastle upon Tyne (UK). The discovery cohort comprised 191 plasma samples and the independent validation cohorts included 115 serum samples and 30 paraffin liver biopsies sections (Table 1 and Supplementary Table 1). A subset of the discovery cohort, comprising 52 of these cases had frozen liver tissue available for RNA extraction. All liver samples were centrally scored according to the semiquantitative NASH-CRN Scoring System by an expert liver pathologist. Fibrosis stage ranged from F0 to F4 (cirrhosis) and the NAS was defined as the sum of the scores for steatosis, hepatocyte ballooning and lobular inflammation3. Alternative diagnoses and aetiologies such as excessive alcohol intake, autoimmune liver diseases, viral hepatitis and steatogenic medication use were excluded. Sex and/or gender of participants was determined on the basis of self-report. This study has been approved by the relevant Ethical Committees in the participating centres and all patients having provided informed consent.
Proteomics
The proteomic aptamer-based SomaScan Platform (SomaLogic) was used to process 191 plasma and 115 serum human samples (20 μl, 1 in 20 dilution)19. To each sample slow off-rate modified labelled aptamers were added to form SOMAmer–protein bead complexes. The beads were captured, and non-specifically bound reagents were subsequently removed. SOMAmers were quantified by hybridization to DNA microarrays. Relative quantity of SOMAmer reagents measured by the SomaScan assay reflecting original protein concentrations (that is, relative fluorescent units, RFUs). Counts were analysed for differential expression using linear models as implemented by the R package limma (https://www.bioconductor.org/) and correcting for centre, sex and T2DM. Statistical significance was determined by a corrected P value less than 0.05 (Benjamini–Hochberg false discovery rate) and a fold change of more than 1.25.
RNA-seq
As previously described, mRNA was extracted from frozen liver biopsy samples and processed with the TruSeq RNA Library Prep Kit v.2 and sequenced on the NextSeq 550 System (Illumina)14. Data are available on the NCBI GEO repository (GSE135251). Raw sequencing quality assessment and alignment to the reference genome (GRCh38, Ensembl release 76) was done using Fastqc (v.0.11.5) and MultiQC (v.1.2dev), and gene count tables were produced with HT-Seq. Counts were normalized using the trimmed mean of M values method and transformed using limma’s voom methodology. A correction for centre, sex and batch was implemented. Pearson correlation was used to investigate linearity between hepatic mRNA and circulating proteins. A P < 0.01 was considered significant. Tissue expression analysis was conducted using GTEx (https://gtexportal.org/). Supervised analysis was done as previously described14. Deconvolution to identify cell of origin was done using publicly available single-cell RNA-seq data (GSE115469) from liver samples obtained from neurological deceased individuals13. The transformed normalized and cluster identifiers were obtained from the Human Protein Atlas (https://www.proteinatlas.org/). For each marker of interest, the Z score was calculated to visualize expression per cell cluster. The DAVID annotation tool was used for functional protein pathway enrichment on the basis of UniprotKB Keywords and Homo Sapiens background20.
Immunohistochemistry
Human formalin-fixed paraffin-embedded liver biopsies (n = 30) were immunostained with antibody directed against human AKR1B10 (ab232623, Abcam; EDTA, 1/500). Immunostains were performed manually at room temperature using Envision Flex+ reagent (Dako) as secondary antibody with 3′,3′-diaminobenzidine visualization. Immunopositive cells were quantified in three different high power fields (magnification ×400) using a bright field microscope.
Statistics
The Kolmogorov–Smirnov and Shapiro–Wilk normality tests, one-way analysis of variance with Dunnett’s test, Chi-square, Mann–Whitney U-test and Kruskal–Wallis test with Bonferroni correction were performed in IBM SPSS v.s27 or GraphPad Prism 9. Binary logistic regression analysis was carried out in IBM SPSS v.27 using Backward Stepwise Likelihood Ratio model including clinical parameters sex, age, BMI, ALT, AST, albumin, platelet count and T2DM, and the uncorrected values of the circulating proteins as measured by SomaScan identified as hepatic markers associated with F3–4 and NAS ≥ 4. The model identifying patients with NASH + F ≥ 2 + NAS ≥ 4 with at least one point deriving from each NAS component, and the FIB-4, NFS and FAST scores were calculated as follows:
-
(1)
Classification model = −6.236112 + (0.082163 × BMI) + (1.110341 × T2DM) + (0.001084 × ADAMTSL2) − (0.000031 × CFHR4) + (0.000060 × TREM2) + (0.000048 × AKR1B10)
-
(2)
FIB-44 = (age (years) × AST (U l−1))/((platelets (109 per l)) × √ALT (U l−1))
-
(3)
NFS4 = 1.675 + 0.037 × age (years) + 0.094 × BMI (kg m−2) + 1.13 × T2DM + (0.99 × AST:ALT ratio) (0.013 × platelet (109 per l)) (0.66 × albumin (g dl−1))
-
(4)
FAST7 = (e \({}^{(-1.65+1.07\times {\mathrm{In(LSM)}}+2.66\times 10^{-8}\, {\mathrm{CAP}}3-63.3\times {\mathrm{AST}}^{-1})}\))/(1 + e\({}^{(-1.65+1.07\times {\mathrm{In(LSM)}}+2.66\times 10^{-8}\, {\mathrm{CAP}}3-63.3\times {\mathrm{AST}}^{-1})}\))
Receiver operating characteristic (ROC) analyses and AUC calculations were performed with IBM SPSS v.27. Paired-Sample Area Difference under the ROC curve was used as statistical test. The binary cut-off for the classification model was set at greater than −0.4491 to rule in patients with NASH + F ≥ 2 + NAS ≥ 4, the FIB-4 score was set at more than 1.3, the FAST score at more than 0.67 to rule in and equal to or less than 0.35 to rule out. Graphs have been generated using R ggplot2, R pheatmap and GraphPad Prism 9. Illustrations within Fig. 1a were created with BioRender.com.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
RNA-seq data are available on the NCBI GEO repository (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE135251). Source data are provided with this paper.
Code availability
The analysis investigating the differentially expressed proteins and transcripts was carried out following the vignette for the R package limma (https://bioconductor.org/packages/release/bioc/html/limma.html).
References
Anstee, Q. M., Reeves, H. L., Kotsiliti, E., Govaere, O. & Heikenwalder, M. From NASH to HCC: current concepts and future challenges. Nat. Rev. Gastroenterol. Hepatol. 16, 411–428 (2019).
Dyson, J. K., McPherson, S. & Anstee, Q. M. Non-alcoholic fatty liver disease: non-invasive investigation and risk stratification. J. Clin. Pathol. 66, 1033–1045 (2013).
Kleiner, D. E. et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 41, 1313–1321 (2005).
Younes, R. et al. Long-term outcomes and predictive ability of non-invasive scoring systems in patients with non-alcoholic fatty liver disease. J. Hepatol. 75, 786–794 (2021).
Boyle, M. et al. Performance of the PRO-C3 collagen neo-epitope biomarker in non-alcoholic fatty liver disease. JHEP Rep. 1, 188–198 (2019).
Kozumi, K. et al. Transcriptomics identify thrombospondin-2 as a biomarker for NASH and advanced liver fibrosis. Hepatology 74, 2452–2466 (2021).
Newsome, P. N. et al. FibroScan-AST (FAST) score for the non-invasive identification of patients with non-alcoholic steatohepatitis with significant activity and fibrosis: a prospective derivation and global validation study. Lancet Gastroenterol. Hepatol. 5, 362–373 (2020).
Luo, Y. et al. SOMAscan proteomics identifies serum biomarkers associated with liver fibrosis in patients with NASH. Hepatol. Commun. 5, 760–773 (2021).
Corey, K. E. et al. ADAMTSL2 protein and a soluble biomarker signature identify significant and advanced fibrosis in adults with NAFLD. J. Hepatol. 76, 25–33 (2022).
Niu, L. et al. Noninvasive proteomic biomarkers for alcohol-related liver disease. Nat. Med. 28, 1277–1287 (2022).
Veyel, D. et al. Biomarker discovery for chronic liver diseases by multi-omics – a preclinical case study. Sci. Rep. 10, 1314 (2020).
Hardy, T. et al. The European NAFLD Registry: a real-world longitudinal cohort study of nonalcoholic fatty liver disease. Contemp. Clin. Trials 98, 106175 (2020).
MacParland, S. A. et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 9, 4383 (2018).
Govaere, O. et al. Transcriptomic profiling across the nonalcoholic fatty liver disease spectrum reveals gene signatures for steatohepatitis and fibrosis. Sci. Transl. Med. 12, eaba4448 (2020).
Ramachandran, P. et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature 575, 512–518 (2019).
Dobie, R. et al. Single-cell transcriptomics uncovers zonation of function in the mesenchyme during liver fibrosis. Cell Rep. 29, 1832–1847.e8 (2019).
Indira Chandran, V. et al. Circulating TREM2 as a noninvasive diagnostic biomarker for NASH in patients with elevated liver stiffness. Hepatology 77, 558–572 (2023).
Hendrikx, T. et al. Soluble TREM2 levels reflect the recruitment and expansion of TREM2(+) macrophages that localize to fibrotic areas and limit NASH. J. Hepatol. 77, 1373–1385 (2022).
Williams, S. A. et al. Plasma protein patterns as comprehensive indicators of health. Nat. Med. 25, 1851–1857 (2019).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Acknowledgements
This study has been supported by the LITMUS (Liver Investigation: Testing Marker Utility in Steatohepatitis) consortium funded by the Innovative Medicines Initiative (IMI2) Program of the European Union under grant agreement no. 777377, which receives funding from the EU Horizon 2020 programme and EFPIA; the Elucidating Pathways of Steatohepatitis consortium funded by the Horizon 2020 Framework Program of the European Union under grant agreement no. 634413; the European NAFLD Registry and the Newcastle National Institute for Health and Care Research Biomedical Research Centre. Q.M.A. is a National Institute of Health and Care Research Senior Investigator.
Author information
Authors and Affiliations
Consortia
Contributions
Q.M.A. created the study concept. Q.M.A., O.G. and A.K.D. have designed the study and drafted the manuscript. L.A. generated the SomaLogic data. O.G., M.H. and S.C. performed the bioinformatics analysis and statistical analysis. D.T. and O.G. have done the immunohistochemistry and histopathology. O.G., M.H., L.A., S.C., D.T., M.E., J.M.S., J.B., E.B., V.R., A.K.D. and Q.M.A. contributed to data acquisition, analysis and interpretation, and critically revised the manuscript for intellectual content.
Corresponding author
Ethics declarations
Competing interests
O.G. reports grants from NanoString Technologies. L.A. is employed by SomaLogic, Inc. D.T. reports consultation fees from Histoindex Pte Ltd. M.E. is a member of the advisory board at AMRA Medical AB. J.M.S. reports consultancy for Apollo Endosurgery, Albireo Pharma Inc, Bayer, BMS, Boehringer Ingelheim, Echosens, Genfit, Gilead Sciences, GSK, Heel GmbH, Intercept Pharmaceuticals, Ipsen, Inventiva Pharma, Julius Clinical, Madrigal, MSD, Nordic Bioscience, Novartis, Novo Nordisk, Pfizer, Roche, Sanofi, Shinogi, Siemens Healthcare GmbH and Summit Clinical Research; research funding from Gilead Sciences, Boehringer Ingelheim, Nordic Bioscience and Siemens Healthcare GmbH; and speaker honoraria from MedPublico GmbH and Boehringer Ingelheim, all outside of the submitted work. J.B. consults for and received grants from Echosens and Siemens. E.B. served as a consultant or advisory board member for Boehringer Ingelheim, Gilead Sciences, Intercept, Inventiva, Merck, Novo Nordisk, Pfizer, ProSciento and a speaker for Gilead Sciences, MSD and Novo Nordisk, outside of the submitted work. Q.M.A. reports grants from European Commission during the conduct of the study; other grants from AstraZeneca, Boehringer Ingelheim, Intercept; consultancy for Alimentiv, Akero, AstraZeneca, Axcella, 89Bio, Boehringer Ingelheim, Bristol Myers Squibb, Galmed, Genfit, Genentech, Gilead, GlaxoSmithKline, Hanmi, HistoIndex, Intercept, Inventiva, Ionis, IQVIA, Janssen, Madrigal, Medpace, Merck, NGMBio, Novartis, Novo Nordisk, PathAI, Pfizer, Prosciento, Poxel, Resolution Therapeutics, Roche, Ridgeline Therapeutics, RTI, Shionogi, Terns. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Isabella Samuelson, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Exploration of confounding effects on the expression of Soma probes.
Differentially expressed Soma probes after correction for sex and centre in the discovery cohort (n = 191) stratified based on fibrosis, NAS ≥ 4 and T2DM. The lower panel provides an overview showing the effect of corrections on the probe expression.
Extended Data Fig. 2 Tissue expression analysis of the 31 proteo-transcriptome signature based on GTEx portal.
Expression of the hepatic 31 proteo-transcriptome signature in different organs based on GTEx portal.
Extended Data Fig. 3 Performance of non-invasive scores to identify patients with NASH + F ≥ 2 + NAS ≥ 4.
Comparison of the classification model, FibroScan-AST (FAST), NAFLD fibrosis score (NFS), Fibrosis-4 (FIB-4) and AST/ALT ratio in a subset of the discovery, n = 62 (a), and validation, n = 28 (b), cohort. Bar charts present Area under the Curve (AUC) for each score with the corresponding standard error of area, as calculated by Receiver operating characteristic (ROC) analysis. Paired-Sample Area Difference under the ROC curve test was used to compare the classification model with other scores (discovery cohort FAST p = 0.111826, NFS p = 0.050406, FIB-4 p = 0.003247 and AST/ALT ratio p = 0.016038; validation cohort FAST p = 0.045552, NFS p = 0.038341, FIB-4 p = 0.036835 and AST/ALT ratio p = 0.005652) (**p < 0.01, *p < 0.05).
Supplementary information
Supplementary Information
Supplementary Tables 1–6, LITMUS consortium.
Source data
Source Data Fig. 1
Proteomics data.
Source Data Fig. 3
Overview scans of the immunohistochemical stainings against AKR1B10 on 30 NAFLD biopsies.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Govaere, O., Hasoon, M., Alexander, L. et al. A proteo-transcriptomic map of non-alcoholic fatty liver disease signatures. Nat Metab 5, 572–578 (2023). https://doi.org/10.1038/s42255-023-00775-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-023-00775-1
- Springer Nature Limited
This article is cited by
-
SomaLogic proteomics reveals new biomarkers and provides mechanistic, clinical insights into Acetyl coA Carboxylase (ACC) inhibition in Non-alcoholic Steatohepatitis (NASH)
Scientific Reports (2024)
-
Multi-omic prediction of incident type 2 diabetes
Diabetologia (2024)
-
Serum proteome signatures associated with liver steatosis in adolescents with obesity
Journal of Endocrinological Investigation (2024)
-
Plasma ALS and Gal-3BP differentiate early from advanced liver fibrosis in MASLD patients
Biomarker Research (2024)
-
Development of a novel non-invasive biomarker panel for hepatic fibrosis in MASLD
Nature Communications (2024)