Abstract
Terrestrial hot springs harbor diverse microbial communities whose compositions are shaped by the wide-ranging physico-chemistries of individual springs. The effect of enormous physico-chemical differences on bacterial and archaeal distributions and population structures is little understood. We therefore analysed the prevalence and relative abundance of bacteria and archaea in the sediments (n = 76) of hot spring features, in the Taupō Volcanic Zone (New Zealand), spanning large differences in major anion water chemistry, pH (2.0–7.5), and temperature (17.5–92.9 °C). Community composition, based on 16S rRNA amplicon sequence variants (ASVs) was strongly influenced by both temperature and pH. However, certain lineages characterized diverse hot springs. At the domain level, bacteria and archaea shared broadly equivalent community abundances across physico-chemically diverse springs, despite slightly lower bacteria-to-archaea ratios and microbial 16S rRNA gene concentrations at higher temperatures. Communities were almost exclusively dominated by Proteobacteria, Euryarchaeota or Crenarchaeota. Eight archaeal and bacterial ASVs from Thermoplasmatales, Desulfurellaceae, Mesoaciditogaceae and Acidithiobacillaceae were unusually prevalent (present in 57.9–84.2% of samples) and abundant (1.7–12.0% sample relative abundance), and together comprised 44% of overall community abundance. Metagenomic analyses generated multiple populations associated with dominant ASVs, and showed characteristic traits of each lineage for sulfur, nitrogen and hydrogen metabolism. Differences in metabolic gene composition and genome-specific metabolism delineated populations from relatives. Genome coverage calculations showed that populations associated with each lineage were distributed across a physicochemically broad range of hot springs. Results imply that certain bacterial and archaeal lineages harbor different population structures and metabolic potentials for colonizing diverse hot spring environments.
Similar content being viewed by others
Introduction
Globally, the environmental factor found to affect microbial community composition the most is salinity [1]. However, at smaller geographical scales or within single biomes where salinity is stable (e.g., freshwater environments), other physicochemical factors, such as temperature and pH become significant [2]. These factors are expected to have an even greater effect in hot springs, where temperature and pH ranges are extreme compared to many other environments, e.g., pH ranges of −0.8 to 10.5 and temperature ranges of <10 to >100°C [3,4,5,6]. The extremes of these ranges are almost at the boundaries of conditions able to support life [7,8,9,10,11], and are inhabited by acidophilic, alkaliphilic, and (hyper)thermophilic microorganisms [6, 12,13,14,15].
Phylogenetically diverse microorganisms are found across the physico-chemical ranges of hot springs. For example, the bacterial phyla Proteobacteria and Aquificae are common inhabitants of various hot springs such as Taupō Volcanic Zone (TVZ, New Zealand), Yellowstone National Park (YNP, USA), and Hveragerði (Iceland) [6, 12, 15,16,17]. In addition, various archaea from Euryarchaeota (e.g., Thermoplasma) and Crenarchaeota (e.g., Sulfolobus) have been isolated or molecularly characterized from hot springs globally [18, 19]. Likewise, several thermophilic cyanobacterial genera, including Synechococcus, Leptolyngbya, and Calothrix, have been reported across alkaline hot springs worldwide [20]. While both hot spring bacteria and archaea have been determined via 16S rRNA characterization [12, 21,22,23], most of these studies utilized different primer sets for bacteria and archaea, making archaeal and bacterial relative abundances incomparable. Moreover, of the studies that have addressed both domains together using a single universal prokaryotic primer set [6, 24], some were potentially affected by known primer biases against archaea [25], suggesting that further research is needed to understand the relative contributions of archaea and bacteria in various hot spring settings [6].
High phylogenetic diversity among hot spring microorganisms may be expected given the distinct adaptations required to survive across large ranges in temperature and pH [26, 27]. For example, genome streamlining is a characteristic of microorganisms adapted to high temperatures, allowing low costs of energy maintenance and increased fitness [28]. Thermophiles may also possess higher concentrations of saturated lipids compared to non-thermophiles to increase their membrane integrity [29]. To maintain pH homeostasis of their intracellular regions in highly acidic environments, acidophiles possess mechanisms to pump out excess intracellular protons, whereas alkaliphiles increase proton influx into cells to maintain intracellular charge balance [30, 31]. Accordingly, previous studies have shown that temperature and pH are strong drivers of differences in diversity and composition among hot spring microbial communities [6, 12,13,14,15]. Despite the selective pressures exerted by extreme physicochemical differences among hot springs, some groups of bacteria and archaea in hot spring communities, such as Acidithiobacillus, Venevibrio and some Thermoplasmatales, are present in hot springs spanning large physicochemical ranges [6, 15]. For example, we previously observed the same cosmopolitan, genome-streamlined, Acidithiobacillus species (particularly the TVZ_G3 group) in hot spring sediments spanning temperatures of 17.5–92.9°C and pHs of 1.0–7.5 [17]. Nevertheless, the cosmopolitanism of other dominant hot spring microorganisms across varying hot spring physicochemistries is yet to be determined.
Here, we evaluated the relative abundances of bacteria and archaea, and their 16S rRNA gene copy numbers, across a wide range of hot spring physicochemistries (i.e., temperature, pH, and major anions), and assessed cosmopolitanism among dominant hot spring microorganisms. To do this, we sampled 76 subaqueous sediments from four geothermal areas (up to 65 km apart) across the TVZ, New Zealand. We then examined the composition of 16S rRNA gene amplicon sequence variants (ASVs), and the abundance and prevalence of microbial ASVs/variants. The relationship of dominant and prevalent variants – those with >1% relative abundance in the overall microbial community and >50% prevalence across samples – to temperature and pH was determined. In addition, using metagenomics, genomes of abundant and prevalent variants were examined to evaluate population structures underpinning cosmopolitanism and the traits of these populations related to sulfur, nitrogen and hydrogen metabolism. Results provide insights into phylogenetic distributions, cosmopolitanism and niche differentiation among hot spring adapted microorganisms.
Materials and methods
Sample collection and physicochemical measurements
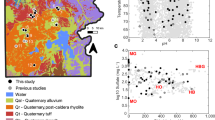
Seventy-six sediment samples were collected from four geothermal areas in the TVZ located 1–65 km apart – Parariki thermal stream, the Sinter Flats lagoon area at the Rotokawa geothermal field, Tikitere geothermal field, and Waiotapu Scenic Reserve located 1–65 km apart, in February and November 2019 (Table S1; Fig. 1 from [15]). Across these geothermal areas, eight sites with multiple co-located hot spring features, including 38 hot spring vents, 27 hot spring outflows, and 11 geothermally-influenced streams were sampled. Samples included 3–5 spatial replicates (<0.5 m apart) per feature, except for at Waiotapu A, which comprised a complex of numerous small vents with replicated chemistries, and Waiotapu B, where its vent and outflow were relatively small. Sediment was collected from a few millimeters to centimeters below the water surface into sterile 50 mL centrifuge tubes, transported on dry ice, and stored at -80°C. The pHs and temperatures of the hot spring fluids were measured in situ using a WTW 330i handheld meter (WTW GmbH, Germany) (Table S1). Major water anion data for Parariki stream, Rotokawa, and Tikitere were derived from prior studies [32,33,34], and Waiotapu water chemistry data were from the 1000springs project (http://1000springs.org.nz).
DNA extraction, amplicon sequencing and quantification, and metagenomic sequencing
DNA extractions, 16S rRNA gene V4-V5 amplicon sequencing from all 76 sediment samples, and metagenome sequencing from 18 samples, along with genome assembly, binning and annotation were undertaken as previously described [17] and are summarized in Supplementary Information. Droplet Digital PCR (ddPCR) was used to quantify the concentration of 16S rRNA genes in each sample using the same primers (without Illumina adapters) and PCR conditions with additional signal stabilizing steps [15].
Amplicon data analyses
QIIME2 (version 2019.10) was used to process demultiplexed sequence reads by read joining, quality filtering (Q score cutoff of 25), and denoising (with singletons removed) [35]. Tables of OTUs (clustered at 99% identity) and ASVs (sequences 100% identical) were generated using VSEARCH and deblur plug-ins, respectively [36, 37]. Taxonomy was assigned using the SILVA database version 132 [38] and q2-feature-classifier plug-in [39]. Rarefaction curves were generated using R (version 4.0.2) with the R package vegan (version 2.5-6). Alpha and beta-diversity were determined using vegan and visualized using ggplot2 (version 3.3.2). Statistical correlations were generated using R package ggpubr (version 0.4.0). Correlations and statistical significances of correlations were determined using Pearson’s correlation coefficients and t-distribution tables (df = n–1), respectively.
Metagenomic data analyses
Metagenome-assembled genomes (MAGs) shared across up to 18 sediment sample assemblies were grouped by 98% and 99% similarity threshold using dRep version 1.4.3 [40] and are referred to as equivalent populations. Barrnap version 0.9 [41] was used to extract 16S rRNA genes from MAGs. FastANI version 1.33 was used to calculate pairwise average nucleotide identities (ANI) [42]. DRAM version 1.4.6 was used for annotation against KEGG-based KOfam, UniRef and Pfam databases downloaded 26-May-2023 [43,44,45,46]. Single-nucleotide polymorphisms (SNPs) detection was analyzed using Snippy version 4.6.0 [47]. For full metagenomic methodology, refer to Supplementary Information.
Results and discussion
Physicochemical diversity of studied hot spring settings
The four geothermal areas sampled differed based on fluid pH and major anions, ranging from acid-sulfate-chloride to acid-sulfate-bicarbonate [48, 49] (Table S1). Because temperature is a major controller of microbial composition and diversity in hot springs [6, 12, 15, 23, 50], we also sampled multiple features at each site, including vents for high temperatures (38.0–92.9 °C), outflows for moderate temperatures (17.5–69.1 °C), and geothermally-influenced streams for low temperatures (18.3–33.4 °C) (Figure S1). Site temperatures and pHs differed by up to 57.4 °C and 3.5 units, with overall temperatures and pHs being 17.5–92.9 °C (average 47.7 °C) and 2.0–7.5 (average 4.0) (Figure S1). In the TVZ, most hot springs are highly acidic, in contrast, for example, to the YNP, which includes more circumneutral pH springs [51].
Bacteria and archaea were relatively abundant across acidic to circumneutral hot spring sediments
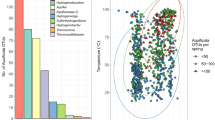
A total of 23,225 ASVs, comprising 47 phyla including unclassified archaea and bacteria, were obtained based on 16S rRNA amplicon sequences from hot spring and hot spring-influenced sediments. Of these, 55.3% of the ASVs, in terms of richness, belonged to two archaeal phyla, Euryarchaeota and Crenarchaeota, and one bacterial phylum, Proteobacteria (Fig. 1a). These three phyla were also dominant, and together comprised 72.1% of the overall hot spring microbial community abundance. While specific hot spring communities tended to be dominated by either bacteria or archaea, overall, bacterial and archaeal abundances were roughly similar (56.3% and 43.7%, respectively) and both exhibited broadly similar distribution patterns with respect to temperature and pH (Fig. 1b). However, we observed significant and opposing correlations between temperature and the relative abundances of archaea (R = 0.42, p = 0.00014, Pearson’s correlation coefficient) and bacteria (R = −0.42, p = 0.00014) (Figure S2). No correlations were observed with pH. These trends are consistent with archaeal preferences for high-temperature niches and the early characterization of archaea solely as extremophiles [52]. Such associations are supported by archaeal adaptations conferring high-temperature tolerance, such as the presence of tetraether lipids in cell membranes [26]. Despite this, bacteria still comprised, on average, over 40% of communities from hot springs with temperatures over 70 °C (45.4% on average) and also over 80 °C (43.2% on average). In fact, some of the most well-studied (hyper)thermophiles from hot springs are bacteria (e.g., Aquificales and Thermus spp.) [53], and studies of other TVZ hot spring features (water and siliceous deposits) also showed that bacterial taxa, such as Hydrogenobaculum (Aquificae), Venevibrio (Aquificae), and Acidithiobacillus (Proteobacteria) were common in high-temperature springs [6, 15, 50].
a Relative abundance of phyla based on 16S rRNA gene amplicons for prokaryotic communities associated with hot spring sediments across all samples. Each sample contained between 24 and 2,046 ASVs (or 16 to 274 after rarefying to the minimum number of sequences in a sample, 673, Table S1). Symbols in parentheses after the sample names refer to the local hot spring feature sampled: V = Vent, O = Outflow, and S = Geothermally-influenced stream. b Plots showing the relative abundances of bacteria and archaea across all samples (left), and their percent difference across hot springs with varying temperatures (middle) and pHs (right). Percent difference was calculated as archaeal abundance per sample - bacterial abundance per sample. The gap between pH 4.1 and 5.3 reflects the lack of hot springs with these pHs in the TVZ.
Euryarchaeota and Crenarchaeota, which dominated the archaeal fraction of hot spring communities in this study and were major constituents of the combined bacterial and archaeal communities, comprise a high proportion of the archaeal communities of many hot springs globally [18, 22, 23, 54, 55]. Other commonly observed archaeal phyla in this study (Fig. 1a) are also common inhabitants of hot springs elsewhere. For example, Thaumarchaeota, Diapherotrites, Hadesarchaeaeota, Korarchaeota, and Nanoarchaeaeota have been reported from hot springs in Kamchatka, YNP, and Iceland [56,57,58,59]. Likewise, the dominant and commonly observed bacterial phyla detected in our study (e.g., Proteobacteria, Thermotogae, Epsilonbacteraeota, Aquificae, and Planctomycetes) are found in geographically diverse hot springs [12,13,14, 24]. Taken together, these data suggest widespread occurrences of the same bacterial and archaeal lineages across hot springs globally. While our results show that archaea are at least as abundant as bacteria in acidic to circumneutral pH TVZ hot spring sediments, an extensive study of hot spring water samples in the TVZ showed that the abundance of archaea (6.4%) was much lower than bacteria (93.6%) [6], either reflecting underestimation of archaea due to a V4 primer bias [6] (Supplementary Information), or substantial differences between water and sediment community compositions [24].
Microbial composition, biomass, and diversity differed strongly with changes in temperature and/or pH
Beta-diversity analysis indicated that microbial community composition in hot spring and hot spring-associated sediments was shaped by temperature and pH (and major anions, which are pH-associated) (Figure S3), comparable to previous studies [6, 12,13,14, 24]. Higher temperature vent communities were differentiated from cooler outflow or geothermally-influenced stream communities in the constructed ordination. Similarly, greater diversity was previously reported at a distal area than at the spring vents [60]. Although the streams were cooler on average than hot spring outflows (26.4 ± 6.7°C versus 36.1 ± 14.1°C), half of the geothermally-influenced stream sediment communities were indistinguishable from outflow communities, reflecting their acidity and the large geothermal inputs received.
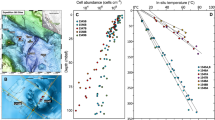
Biomass inferred from 16S rRNA gene concentrations was significantly and negatively correlated with temperature (R = −0.34, p = 0.0025; Fig. 2c), but not pH (R = 0.043, p = 0.71; Fig. 2d). Studies of creek sediments and soils have also reported a lack of correlation with pH [61, 62]. Correlated temperature and 16S rRNA gene concentrations across wide-ranging temperatures (17.5–92.9 °C) and eight sites, together with similar trends observed elsewhere, suggests inferred biomass is consistently reduced in higher temperature springs (e.g., 39.3–74.1°C across a single site [63]; 57–100 °C across three sites [12]). The trend identified here remained significantly negative after excluding geothermally-influenced streams (R = −0.3, p = 0.016; Figure S4). At high-temperature sites (≥70 °C), we detected almost two-fold lower copy number concentrations on average (1,329,131 ± 1,284,851 copies/gram of sediment) compared to low temperatures sites (≤40 °C, 2,571,607 ± 894,876 copies/gram of sediment). Moreover, at some extremely high-temperature sites (>80 °C), copy numbers were 1000-fold lower than at most low and moderate temperature hot spring sites (Figure S5a), highlighting the negative effect of temperature on inferred microbial biomass (Fig. 2c). Overall, concentration ranges were similar to those reported from hot springs elsewhere [12, 24]. However, copy numbers were 10 to 10,000-fold lower than those reported from freshwater and marine sediments [64, 65], potentially owing to the higher temperatures of hot springs in general. It is worth noting that although ddPCR quantifies gene copy numbers and not actual biomass or cell numbers, concentrations in this study were similar to those obtained using direct cell counts to estimate cell concentration ranges of 106 to 108 cells/ml in hot spring environments elsewhere [66].
Scatter plots showing (a) a non-significant correlation between Shannon’s indices and temperature (°C) (R = −0.17, p = 0.13); (b) a significantly positive correlation between Shannon’s indices and pH (R = 0.28, p = 0.013); (c) a significantly negative correlation between 16S rRNA gene copies and temperature (°C) (R = −0.34, p = 0.0025); (d) a non-significant correlation between 16S rRNA gene copies and pH (R = 0.043, p = 0.71). Correlations and statistical significances of correlations were determined using Pearson’s correlation coefficients and t-distribution tables (df = n–1), respectively. Lines represent linear regressions and shaded areas represent 95% confidence intervals.
Several studies have identified significant negative correlations between hot spring temperature and microbial alpha diversity (using Shannon indices) [12,13,14,15]. We instead identified a weak, non-significant, negative correlation (R = −0.17, p = 0.13; Fig. 2a and S5b), despite significant changes in beta-diversity and inferred biomass. Analysis of TVZ hot spring water (n = 925) also showed no association between temperature and alpha diversity, except at >70 °C [6]. However, we found that among features associated with individual hot springs, microbial diversity of lower temperature geothermally-influenced streams (18.3–33.4°C) was significantly higher than in vents and outflows (Figure S6). While the effect of temperature cannot be excluded, this difference could be due to higher reported nutrient (i.e., ammonia, nitrate, and phosphorus) concentrations in the streams [67], supporting a greater diversity of microorganisms. In contrast, results overall indicated a significant positive correlation between pH (range 2.0–7.5) and Shannon diversity (R = 0.28, p = 0.01; Fig. 2b), consistent with other TVZ hot spring studies [6, 15]. Although these studies show diversity increases with pH (up to at least pH 9.5), the growth of individual isolates from geothermal environments has been demonstrated across wide pH ranges (e.g., pHs of 1.0–6.0 for Acidianus brierleyi and 5.6–10.0 for Anoxybacillus kamchatkensis) [68, 69], indicating broad tolerances.
Predominance of a few microbial taxa across physico-chemically different hot springs
While microbial alpha diversity and community composition are influenced by pH (Figure S3), some taxa inhabit hot springs with wide-ranging pHs, for example, Sharp et al. [14] reported consistent relative abundances of Proteobacteria, Acidobacteria, Crenarchaeota, and Bacteroidetes across pH gradients. We previously identified a wide occurrence of Acidithiobacillus spp. across hot spring sediment and sinter pHs of at least 2.0 to 7.5 [15, 17], likely supported by multiple genes encoding amino acid decarboxylases, K+ transporters, Na+/H+ antiporters, and plasma-membrane proton efflux ATPases [17]. Features such as streamlined genomes and higher predicted proline contents also potentially facilitate the prevalence of TVZ Acidithiobacillus across broad hot spring temperature ranges [17]. The wide occurrence of Acidithiobacillus across physico-chemically different hot spring sediments and siliceous sinters was achieved by only a few dominant and cosmopolitan species or sub-species, based on an analysis of ASVs and MAGs [17]. To determine how widespread this trend was among other hot spring microorganisms, we examined all bacterial and archaeal variants. Most ASVs were detected at single or narrow ranges of temperature and pH. Specifically, 97.7% of ASVs displayed a temperature range of <10°C, and 98.2% of ASVs had a pH range of <1 pH unit. Nevertheless, some were distributed across large differences, and were found in up to 84% of samples (Fig. 3a–d). We also identified a strong relationship between prevalence and community abundance (R = 0.64, p = 2.2 × 10−16), with eight of the ten most abundant ASVs overall (comprising up to 12% of the overall community) being the most prevalent (Fig. 4a).
a Scatter plot illustrating the overall relative abundance (summed across samples) and prevalence of all ASVs. Colored dots represent eight ASVs that are highly abundant (>1% abundance in the community) and prevalent (>50% of samples). b, c Temperature and pH ranges where these eight prevalent ASVs were detected. Boxes, internal horizontal lines and whiskers represent upper/lower quartiles, median and minimum/maximum ranges, respectively. The gap between pH 4.1 and 5.3 reflects the lack of hot springs with these pHs in the TVZ, as shown in Fig. 1b.
A small fraction of ASVs comprised most of the overall community. In total, 280 ASVs (1.2%) comprised 90% of community abundance. Of these, the eight most prevalent (in >50% of samples) were distributed across temperatures from 17.5 °C and up to 92.9 °C and pHs from 2.0 and up to 7.5 (Fig. 4a–c and S7a). These eight ASVs accounted for 44% of hot spring community abundance and spanned four phyla (pairwise sequence identities 67.3% to 99.7%, Fig. 5a) that are commonly detected in hot spring environments in New Zealand [6, 15] and elsewhere [12, 14]. Three ASVs were Euryarchaeota/Thermoplasmatota (two A10 and one BSLdp215), two each were Proteobacteria (Acidithiobacillus) and Thermotogae/Thermotogota (Mesoaciditoga), and one was from Epsilonbacteraeota/Campylobacterota (Desulfurella). Consistent with their wide hot spring distributions, none showed strong correlations between their relative abundances and either temperature or pH (rs ≤ 0.4, p < 0.05), and three (Thermoplasmatales ASV2 and Mesoaciditoga ASVs 5 and 8) showed no significant correlations with either variable (Fig. 5b). All four lineages are well-documented in hot spring settings. For example, Acidithiobacillus is also prevalent in TVZ hot spring fluids [6], and has been found in hot spring fluids [46, 70] and sediment [71] elsewhere. At least two species of Desulfurella were first discovered from a sulfuric hot spring environment (Kamchatka, Russia) [72, 73]. Similarly, Mesoaciditogaceae has been isolated from acidic hot spring water, whereas members of Thermoplasmatota were among the first taxa recovered from hot spring environments [19, 74].
a Pairwise 16S rRNA gene sequence identities (%) among the eight abundant and prevalent ASVs. b Heatmap of Spearman’s correlations between ASV relative abundance and hot spring temperature or pH. Asterisks indicate significant correlations (p < 0.05). c Plot comparing ASV numbers and OTU relative abundance. The linear trend line shows a significantly positive correlation between the top 20 most abundant OTUs in the total community and number of ASVs observed per OTU. Test = Pearson’s correlation coefficients. Shaded areas represent 95% confidence intervals.
Environments tend to select for phylogenetically related microorganisms based on shared traits [75, 76]. Accordingly, six out of eight of the prevalent and abundant hot spring ASVs we identified were represented by just three taxonomic families or genera (Thermoplasmatales family A10, Mesoaciditoga, Acidithiobacillus; Fig. 5a) - i.e., there were two-to-three ASVs from each lineage. Results also indicated that rarer close relatives of these variants (those sharing >99% 16S rRNA gene amplicon similarity) were numerous (Supplementary Information and Tables S2-3), likely reflecting environmental selection based on shared traits [75, 76]. ASV numbers were significantly positively correlated with the summed relative abundance of close relatives (OTUs, Fig. 5c and S7b), indicating higher strain or species level diversity among successful hot spring taxa. A similar trend has also been found in communities along oxic-hypoxic gradients of deep lakes [77].
Endemism is suggested to be a feature of hot springs separated by large-scale geographic distances (e.g., different continents) due to dispersal limitations [78, 79]. In this study, the wide distribution of eight abundant microbial variants, up to 65 kilometers apart, and spread across a broad spectrum of temperatures and pHs, suggests cosmopolitanism and not niche differentiation, at least at the 16S rRNA amplicon level of resolution. However, it remains to be determined whether these variants are abundant and prevalent in hot springs outside of the TVZ. Cosmopolitanism of variants has been observed in various other environments. For example, some Vibrio oligotypes have both host-associated and free-living lifestyles [80]. ASVs of certain methane-oxidizing bacteria in eutrophic lakes also have been observed across both oxygen-deficient and methane-deficient conditions [81]. Different taxonomic resolution thresholds have been defined based on 16S rRNA genes to determine the cosmopolitanism of closely-related microbial taxa. For example, Ward et al. (2017) allowed 4-nucleotide variation in single ‘sub-OTUs’ [82], while a minute gap of dissimilarity is allowed in single oligotypes (99.2–99.8% identity or 0.2–0.8% sequence variation) [83, 84], and ASVs employ 100% identity [85]. Regardless, observations of dominant sequence variants based on full or partial 16S rRNA gene sequences do not preclude strain-level variation not captured by differences in 16S rRNA genes, which more typically enables genus and, in some cases, species level discrimination based on the full gene [86,87,88].
Population structure underpinning the distribution of cosmopolitan variants
To explore the population structures underpinning variant cosmopolitanism, we selected MAGs associated with the eight ASV lineages. Overall, there were 198 MAGs (75-100% complete with <5% contamination) that spanned 35 phyla from 18 hot spring samples (17.5–92.9°C and pH 2.0–7.5). Of these, 50 MAGs were classified as Mesoaciditogaceae, Desulfurellaceae, Acidithiobacillaceae and three Thermoplasmata families (Table S4), and are proposed here to represent 17 unique species based on average nucleotide identities (ANI’s) of ≥96.5% (alignable fractions of 43-96%) [89] (Table S5). Almost half contained 16S rRNA gene sequences, enabling sequence-based comparisons between cosmopolitan ASVs and 20 Thermoplasmata and three Acidithiobacillaceae MAGs (Fig. 6a and Table S4). ASVs 1 and 2 were 100% identical to 16S gene sequences from eight Thermoplasmata family ARK-15 MAGs (NCBI strain identifier A10-Griffin-MG), while ASV3 was identical to six GCA-001856825 family sequences. ASVs 6 and 7 were identical to 16S genes from two MAGs designated as Acidithiobacillaceae (ASV 6) and one Acidithiobacillaceae UBA2486 sp002341825 (ASV 7). Genomes sharing as little as 85% ANI or less can share identical 16S rRNA gene sequences [90], and microorganisms with identical 16S rRNA gene sequences, but divergent genomes, are known to occupy distinct niches [91]. In this study, ASVs 1 and 3 matched with the V4-V5 hypervariable regions of six MAGs each that shared strain-level similarity (≥99% and >98% ANI, respectively). In contrast, ASVs 2 and 6 matched to regions from MAGs with ANIs below the species delineation threshold (91% and 95% ANI), suggesting genus level resolution. Results therefore indicated that these ASVs encompassed collections of strains and distinct species.
a Heat maps showing the log relative genome abundance per site based on read mapping. Abundances were normalized to library size, and are included where the summed length of mapped reads equated to least 5% of each genome (72 Kbp to 6 Gbp). Samples are ordered by temperature (left plot), or pH (right plot), and sample conditions and locations are indicated below the x-axes. White dashed boxes indicate samples from which a genome was recovered. MAGs shown are representatives following dereplication at 98% ANI, and MAG cluster sizes based on 98% and 95% ANI thresholds are shown on the right, along with GTDB based taxonomy. Asterisks indicate references for ≥95% ANI clusters. Cosmopolitan ASV sequence matches are shown where 100% identical to a MAG-derived 16S rRNA gene sequence. b Heat maps showing gene copy numbers present (maximum = four) per MAG associated with sulfur metabolism (oxidation/reduction), energy-generating nitrogen-cycling processes (only nitrate reduction identified for the MAGs shown), and hydrogen metabolism (production/consumption). hydr hydrogenase, cyt cytochrome, aux auxiliary, fhl formate hydrogenlyase.
To determine the spatial distributions of the 50 MAGs, including those matched to cosmopolitan ASVs, the MAGs were clustered at 98% ANI. Dereplicating populations at this similarity yields a negligible 0.6% chance of indiscriminate mapping (ANI proportion to the power of read length, 0.98250) [92]. The resulting 18 representative MAGs had cluster sizes of one to nine, reflecting the recovery of highly similar genomes from multiple hot spring locations (Fig. 6a). Genomes with such high similarity have been termed ‘sequence-discrete’ populations [93,94,95]. Sequence-discrete populations share >95% nucleotide similarity, are separated from other populations by a genetic discontinuity, and are by inference conspecific [89]. They are more likely to share environmental niches than distinct populations (or species) [94, 95]. Here, genome coverages of representative MAG populations were determined by read mapping. Results indicated the presence of sub-species populations, associated with each of the four archaeal and bacterial lineages, that had broad physicochemical distributions, extending results reported previously for Acidithiobacillus [17]. At least one population from each phyla was present in the majority of sites from which metagenomes were derived (16 to 17 sites), which spanned temperatures from 23 to 74 °C or 80 °C and pHs from 2.5 to 7.3 (Fig. 6a, Table S6). Related populations, within and the same families, tended to share broadly similar spatial distribution patterns. However, some differentiation in spatial niches was evident, such as the relatively high abundances of ASV2-associated ARK-15 MAGs 224-226 at 69–80 °C. Striking differences were also evident between Thermoplasmata families - Thermoplasmataceae were all absent (or lower in abundance) at the higher pHs.
Metabolic traits of cosmopolitan lineages
Differentiation in functional genes as part of the accessory genome is believed to drive niche differentiation among closely-related taxa [96]. For example, key genes for osmolyte uptake are present in the marine subclades/ecotypes of the well-documented alphaproteobacterial bacterioplankton SAR11, but absent in freshwater SAR11 [97]. Similarly, the different responses to light intensity of Prochlorococcus ecotypes are likely due to differences in number of putative high-light-inducible proteins [98]. To determine differences in metabolic potential within and between populations we examined Mesoaciditogaceae, Desulfurellaceae, Acidithiobacillaceae, and Thermoplasmata genome annotations for genes involved in sulfur, nitrogen and hydrogen metabolism. Taxa within the four phyla are known for distinct metabolisms. Desulfurella and Acidithiobacillus are known for sulfur metabolism (reduction and oxidation, respectively), along with iron oxidation by Acidithiobacillus species [72], which might explain the prevalence of both genera in the sulfur-rich TVZ hot springs (http://1000springs.org.nz). Thermotoga are known as strictly anaerobic fermenters and Mesoaciditogaceae have been shown to reduce thiosulfate and iron [74, 99], while species of Thermoplasmata engage in heterotrophy, methanogenesis or iron oxidation [100, 101]. However, of the TVZ hot spring taxa, all had genes with similarity to those encoding hydrogenases - either FeFe (the Mesoaciditogaceae) or NiFe (Fig. 6b). Mechanisms for sulfur metabolism were ubiquitous, and predictably most numerous in Acidithiobacillaceae and scarcest in Mesoaciditogaceae.
Genomes within each of the four phyla shared characteristic traits (Fig. 6b). For example, all 12 Acidithiobacillus MAGs contained soxABXYZ, sulfide-quinone reductase (sqr), and sulfite oxidase genes. Almost all Thermoplasmata MAGs (24 of 25) contained genes homologous to asrAB anaerobic sulfite reductase (analogous to sulfhydrogenase, cytochrome c3 hydrogenase). These were co-located with an archaeal-type formate dehydrogenase alpha subunit with a molybdopterin oxidoreductase 4Fe-4S domain (Table S7), which potentially substitutes for siroheme-binding AsrC oxidoreductase. Asr couples sulfite reduction to hydrogen sulfide with electron acceptors, including NADH, H2, and formate [102]. Energy yielding nitrogen cycling mechanisms were largely absent. However, all four Desulfurellaceae MAGs contained nar respiratory nitrate reductase genes, and other spatially-localized Campylobacterota relatives present contained periplasmic nap nitrate reductase genes (Table S7) [103], indicating that diverse members of this phylum conserve energy via nitrate reduction, albeit via different mechanisms.
Despite shared traits overall, some populations could be delineated from relatives by consistent differences in gene presence or copy numbers. For example, all five Acidithiobacillaceae UBA2486 sp002341825 MAGs were distinguished from other Acidithiobacillaceae by the presence of dsrE genes (involved in sulfur transfer) [104], and only a single copy of the soxABX sulfur oxidation genes [105]. Comparably, among the Thermoplasmata, only the nine ARK-15 genomes conspecific with MAG 214 had syntenous cysAT sulfur transport genes annotated (Fig. 6b) [106], suggesting other Thermoplasmata relied on an alternative mechanism for importing thiosulfate/sulfate. The sulfate adenylyltransferase gene, sat, involved in converting sulfur to adenosine 5′-phosphosulfate in the Dsr pathway [107], was instead almost exclusive to other Thermoplasmata families (GCA-001856825 and Thermoplasmataceae).
Some traits appeared to be MAG-specific. In particular, complete sets of adenylyl-sulfate reductase aprAB and dissimilatory sulfite reductase dsrABC genes were found in a handful of distantly related Thermoplasmata MAGs across all three families. Likewise, syntenous clusters of formate hydrogenlyase (fhl subunits 1–5) genes that annotations suggest encode a bidirectional NiFe hydrogenase [108], were present in a small number of ARK-15 MAGs. Although genome incompleteness likely accounts for some differences observed, the complete absence of these genes in a several MAGs, estimated to be 95–99% complete (Table S4) points to strain-level differences in sulfur and hydrogen metabolism.
Single-nucleotide polymorphisms differentiate populations of Thermoplasmatales
To further explore the genomic diversity among Thermoplasmatales populations, which contained the largest number of genomes and the greatest number of matches to cosmopolitan ASVs, we performed SNP analysis. Results show that SNPs — including indels, non/synonymous point mutations, and complex or multiple/combined point mutations and indels — within a population accounted for an average of 0.2 ± 0.1% of the genomes, whereas the rates between populations rose to 2.6 ± 0.9% (Fig. 7a). Of these SNPs, more than 80% were located in CDS regions (1730 ± 1145 SNPs per Mbp for intra-population and 19,884 ± 12,918 SNPs per Mbp for inter-population). In contrast, only 0–0.03% of SNPs were detected in non-coding rRNA gene regions (0–0.8 SNPs per Mbp for both intra/inter-populations) (Fig. 7b), as expected for the highly conserved nature of 16S rRNA genes. Similarly, Leptospirillum and Ferroplasma populations, isolated from acid mine drainage, showed no SNPs in their 16S or 23S rRNA gene sequences, while the average SNP rates of their whole genomes were 0.08% and 2.2%, respectively [109]. In addition, we found that about half of SNPs (47.8–66.2%) were synonymous and are not predicted to alter encoded amino acids (Fig. 7c), whereas approximately a quarter were missense SNPs, along with a small number of frameshifts and stop/start codon disruptions (25.1–34.5%) that encode distinct amino acids and may represent phenotypic differences. For example, functional shifts caused by SNPs, which are common among pathogenic microorganisms (e.g., Staphylococcus aureus), have been associated with increases in virulence and antibiotic resistance [110]. Comparable to our results, one-third of SNPs detected in Leptospirillum and Ferroplasma populations in the acid mine drainage study were suggested to cause changes at the protein-coding level [109]. Taken together, our results imply that the cosmopolitanism observed among hot spring ASVs encompasses genomic variation resulting from intra- and inter-population point mutations (as illustrated with Thermoplasmatales), along with differences in metabolic gene composition.
a Types of SNP point mutations including substitutions, insertions, and deletions, and complex (i.e., multiple points/combined mutations of substitutions and indels). b Genomic regions where SNPs were detected. SNPs identified as ‘unspecified’ were excluded, specifically 200–700 SNPs for intrapopulation and 100-3,200 SNPs for interpopulation MAGs. c The effects to CDS by SNPs.
Conclusions
This study reports the influence of temperature and pH on the relative abundance, diversity, and prevalence of bacteria and archaea in hot spring sediments. We found that bacteria and archaea shared similar abundances in the overall hot spring microbial communities, despite a slight proportional increase in archaea with higher temperatures. The dominant phyla overall were a mixture of bacteria (Proteobacteria) and archaea (Euryarchaeota and Crenarchaeota). Higher microbial diversity was associated with an increase in pH from 2.0 to 7.5, while microbial cell concentrations, as inferred from 16S rRNA gene copies, were primarily influenced by and positively correlated with temperature. Although differences in microbial composition were driven by temperature and pH, we identified eight phylogenetically diverse bacterial and archaeal variants (based on ASVs) that were found in up to 84% of the hot spring communities and accounted for 44% of the relative abundance. These variants, belonging to Thermoplasmatales, Desulfurella, Mesoaciditoga, and Acidithiobacillus, were present across geographically distant hot spring sites with wide-ranging temperatures and pHs, illustrating the lack of constraint of temperature and pH on their distributions. Amplicon results suggest bacterial and archaeal cosmopolitanism may be a common feature of hot spring environments. Metagenomic results indicated the presence of strain-level (sub-species) populations associated with each of these four lineages that had broad spatial and physicochemical ranges. In general, diverse members of each lineage (different families or genera) shared common metabolic traits, although variations in mechanisms for sulfur, nitrogen and hydrogen metabolism were evident both between and within predicted species. Accordingly, when inspecting nucleotide-level difference among the numerous Thermoplasmata MAGs recovered, we found that both intra-species, and to a greater extent, inter-species populations were differentiated by a mixture of synonymous and non-synonymous SNPs, indicating differences in amino acid coding and potential differences in protein function. Results suggest that multiple prokaryotic lineages, including both bacteria and archaea, are successful in colonizing a range of hot spring conditions by harboring diverse population structures and genome-specific metabolic traits.
Data availability
Sequence data can be accessed through NCBI BioProject PRJNA644733.
References
Lozupone CA, Knight R. Global patterns in bacterial diversity. Proc Natl Acad Sci. 2007;104:11436–40.
Kent AD, Yannarell AC, Rusak JA, Triplett EW, McMahon KD. Synchrony in aquatic microbial community dynamics. ISME J. 2007;1:38–47.
Bodvarsson G. Physical characteristics of natural heat resources in Iceland. Joekull. 1961;11:29–38.
Brock TD. Bimodal distribution of pH Values of thermal springs of the world. GSA Bull. 1971;82:1393–4.
Jones B, Renaut RW, Rosen MR. Microbial construction of siliceous stalactites at geysers and hot springs: examples from the Whakarewarewa Geothermal Area, North Island, New Zealand. PALAIOS. 2001;16:73–94.
Power JF, Carere CR, Lee CK, Wakerley GLJ, Evans DW, Button M, et al. Microbial biogeography of 925 geothermal springs in New Zealand. Nat Commun. 2018;9:2876.
Brock TD. Lower pH limit for the existence of blue-green algae: evolutionary and ecological implications. Science. 1973;179:480–3.
Brock TD. Life at high temperatures. Science. 1985;230:132–8.
Brock TD, Darland GK. Limits of microbial existence: temperature and pH. Science. 1970;169:1316–8.
Kashefi K, Lovley DR. Extending the upper temperature limit for life. Science. 2003;301:934.
Stetter KO. Hyperthermophilic procaryotes. FEMS Microbiol Rev. 1996;18:149–58.
Krebs JE, Vaishampayan P, Probst AJ, Tom LM, Marteinsson VT, Andersen GL, et al. Microbial community structures of novel Icelandic hot spring systems revealed by PhyloChip G3 analysis. Astrobiology. 2014;14:229–40.
Miller SR, Strong AL, Jones KL, Ungerer MC. Bar-coded pyrosequencing reveals shared bacterial community properties along the temperature gradients of two alkaline hot springs in Yellowstone National Park. Appl Environ Microbiol. 2009;75:4565–72.
Sharp CE, Brady AL, Sharp GH, Grasby SE, Stott MB, Dunfield PF. Humboldt’s spa: microbial diversity is controlled by temperature in geothermal environments. ISME J. 2014;8:1166–74.
Sriaporn C, Campbell KA, Millan M, Ruff SW, Van Kranendonk MJ, Handley KM. Stromatolitic digitate sinters form under wide-ranging physicochemical conditions with diverse hot spring microbial communities. Geobiology. 2020;18:619–40.
Hugenholtz P, Pitulle C, Hershberger KL, Pace NR. Novel division level bacterial diversity in a Yellowstone Hot Spring. J Bacteriol. 1998;180:366–76.
Sriaporn C, Campbell KA, Van Kranendonk MJ, Handley KM. Genomic adaptations enabling Acidithiobacillus distribution across wide-ranging hot spring temperatures and pHs. Microbiome. 2021;9:135.
Barns SM, Fundyga RE, Jeffries MW, Pace NR. Remarkable archaeal diversity detected in a Yellowstone National Park hot spring environment. Proc Natl Acad Sci. 1994;91:1609–13.
Segerer A, Langworthy TA, Stetter KO. Thermoplasma acidophilum and Thermoplasma volcanium sp. nov. from Solfatara Fields. Syst Appl Microbiol. 1988;10:161–71.
Ward DM, Castenholz RW, Miller SR. Cyanobacteria in geothermal habitats. In: Whitton BA, editor. Ecology of Cyanobacteria II: Their diversity in space and time. Dordrecht: Springer Netherlands; 2012. p. 39-63.
Meyer-Dombard DR, Shock EL, Amend JP. Archaeal and bacterial communities in geochemically diverse hot springs of Yellowstone National Park, USA. Geobiology. 2005;3:211–27.
Pagaling E, Grant WD, Cowan DA, Jones BE, Ma Y, Ventosa A, et al. Bacterial and archaeal diversity in two hot spring microbial mats from the geothermal region of Tengchong, China. Extremophiles. 2012;16:607–18.
Purcell D, Sompong U, Yim LC, Barraclough TG, Peerapornpisal Y, Pointing SB. The effects of temperature, pH and sulphide on the community structure of hyperthermophilic streamers in hot springs of northern Thailand. FEMS Microbiol Ecol. 2007;60:456–66.
Colman DR, Feyhl-Buska J, Robinson KJ, Fecteau KM, Xu H, Shock EL, et al. Ecological differentiation in planktonic and sediment-associated chemotrophic microbial populations in Yellowstone hot springs. FEMS Microbiol Ecol. 2016;92:fiw137.
Parada AE, Needham DM, Fuhrman JA. Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ Microbiol. 2016;18:1403–14.
Konings WN, Albers S-V, Koning S, Driessen AJM. The cell membrane plays a crucial role in survival of bacteria and archaea in extreme environments. Antonie van Leeuwenhoek. 2002;81:61–72.
Woese CR. Bacterial evolution. Microbiol Rev. 1987;51:221–71.
Sabath N, Ferrada E, Barve A, Wagner A. Growth temperature and genome size in bacteria are negatively correlated, suggesting genomic streamlining during thermal adaptation. Genome Biol Evol. 2013;5:966–77.
Oshima M, Miyagawa A. Comparative studies on the fatty acid composition of moderately and extremely thermophilic bacteria. Lipids. 1974;9:476–80.
Baker-Austin C, Dopson M. Life in acid: pH homeostasis in acidophiles. Trends Microbiol. 2007;15:165–71.
Padan E, Bibi E, Ito M, Krulwich TA. Alkaline pH homeostasis in bacteria: new insights. Biochimica et Biophysica Acta (BBA) - Biomembranes. 2005;1717:67–88.
Dobson M. Facies Mapping and Analysis of Diverse Hydrothermal Sedimentary Facies and Siliceous Spicular Sinter at Hell’s Gate, Tikitere Geothermal Field, Taupō Volcanic Zone, New Zealand [University of Auckland Honours Dissertation]. Auckland, New Zealand: University of Auckland; 2018.
Schinteie R, Campbell KA, Browne PRL. Microfacies of stromatolitic sinter from acid-sulphate-chloride springs at Parariki Stream, Rotokawa Geothermal Field, New Zealand. Palaeontol Electron. 2007;10:1–33.
Camp SL. Hydrothermal Sedimentary Facies and Environmental Controls on Their Formation in The Sinter Flat Lagoon area, Rotokawa Geothermal Field, Taupo Volcanic Zone, New Zealand [University of Auckland Honours Dissertation]. Auckland, New Zealand: University of Auckland; 2018.
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol. 2019;37:852–7.
Amir A, McDonald D, Navas-Molina JA, Kopylova E, Morton JT, Zech XZ, et al. Deblur rapidly resolves single-nucleotide community sequence patterns. mSystems. 2017;2:e00191–16.
Rognes T, Flouri T, Nichols B, Quince C, Mahé F. VSEARCH: a versatile open source tool for metagenomics. PeerJ. 2016;4:e2584.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41:D590–D6.
Bokulich NA, Kaehler BD, Rideout JR, Dillon M, Bolyen E, Knight R, et al. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome. 2018;6:90.
Olm MR, Brown CT, Brooks B, Banfield JF. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 2017;11:2864–8.
Seemann T. Barrnap 0.9: rapid ribosomal RNA prediction. 2018. Available from https://github.com/tseemann/barrnap.
Jain C, Rodriguez-R LM, Phillippy AM, Konstantinidis KT, Aluru S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat Commun. 2018;9:5114.
Shaffer M, Borton MA, McGivern BB, Zayed AA, La Rosa SL, Solden LM, et al. DRAM for distilling microbial metabolism to automate the curation of microbiome function. Nucleic Acids Res. 2020;48:8883–900.
Aramaki T, Blanc-Mathieu R, Endo H, Ohkubo K, Kanehisa M, Goto S, et al. KofamKOALA: KEGG Ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics. 2020;36:2251–2.
Wang Y, Wang Q, Huang H, Huang W, Chen Y, McGarvey PB, et al. UniProt Consortium. A crowdsourcing open platform for literature curation in UniProt. PLOS Biol. 2021;19:e3001464.
Mistry J, Chuguransky S, Williams L, Qureshi M, Salazar GA, Sonnhammer EL, et al. Pfam: The protein families database in 2021. Nucl Acids Res. 2021;49:D412–9.
Seemann T. Snippy: fast bacterial variant calling from NGS reads. 2015. Available from https://github.com/tseemann/snippy.
White DE. Thermal waters of volcanic origin. GSA Bull. 1957;68:1637–58.
Renaut RW, Jones B. Hydrothermal environments, terrestrial. In: Thiel JRV, editor. Encyclopedia of Geobiology Amsterdam, The Netherlands: Springer; 2011. p. 467-79.
Ward L, Taylor MW, Power JF, Scott BJ, McDonald IR, Stott MB. Microbial community dynamics in Inferno Crater Lake, a thermally fluctuating geothermal spring. The. ISME Journal. 2017;11:1158–67.
Rowe JJ, Fournier RO, Morey GW. Chemical analysis of thermal waters in Yellowstone National Park, Wyoming, 1960-65. Geol Surv Bull 1303. US Government Printing Office; 1973. p. 31.
Woese CR, Magrum LJ, Fox GE. Archaebacteria. J Mol Evol. 1978;11:245–52.
Brock TD, Freeze H. Thermus aquaticus gen. n. and sp. n., a nonsporulating extreme thermophile. J Bacteriol. 1969;98:289–97.
Childs AM, Mountain BW, O’Toole R, Stott MB. Relating microbial community and physicochemical parameters of a hot spring: Champagne Pool, Wai-o-tapu, New Zealand. Geomicrobiol J. 2008;25:441–53.
Mondal N, Peketi A, Mapder T, Roy C, Mazumdar A, Chakraborty R, et al. Indus and Nubra Valley hot springs affirm the geomicrobiological specialties of Trans-Himalayan hydrothermal systems. J Earth Syst Sci. 2022;131:12.
Eme L, Reigstad LJ, Spang A, Lanzén A, Weinmaier T, Rattei T, et al. Metagenomics of Kamchatkan hot spring filaments reveal two new major (hyper)thermophilic lineages related to Thaumarchaeota. Res Microbiol. 2013;164:425–38.
Merkel AY, Podosokorskaya OA, Sokolova TG, Bonch-Osmolovskaya EA. Diversity of methanogenic archaea from the 2012 terrestrial hot spring (Valley of Geysers, Kamchatka). Microbiology. 2016;85:342–9.
Miller-Coleman RL, Dodsworth JA, Ross CA, Shock EL, Williams AJ, Hartnett HE, et al. Korarchaeota diversity, biogeography, and abundance in Yellowstone and Great Basin hot springs and ecological niche modeling based on machine learning. PLOS One. 2012;7:e35964.
Reigstad LJ, Jorgensen SL, Schleper C. Diversity and abundance of Korarchaeota in terrestrial hot springs of Iceland and Kamchatka. ISME J. 2010;4:346–56.
Roy C, Rameez MJ, Haldar PK, Peketi A, Mondal N, Bakshi U, et al. Microbiome and ecology of a hot spring-microbialite system on the Trans-Himalayan Plateau. Sci Rep. 2020;10:5917.
Fabisch M, Beulig F, Akob D, Küsel K. Surprising abundance of Gallionella-related iron oxidizers in creek sediments at pH 4.4 or at high heavy metal concentrations. Front Microbiol. 2013;4:390.
Wu Y, Zeng J, Zhu Q, Zhang Z, Lin X. pH is the primary determinant of the bacterial community structure in agricultural soils impacted by polycyclic aromatic hydrocarbon pollution. Sci Rep. 2017;7:40093.
Hall JR, Mitchel KR, Jackson-Weaver O, Kooser AS, Cron BR, Crossey LJ, et al. Molecular characterization of the diversity and distribution of a thermal spring microbial community by using rRNA and metabolic genes. Appl Environ Microbiol. 2008;74:4910–22.
Bai Y, Shi Q, Wen D, Li Z, Jefferson WA, Feng C, et al. Bacterial communities in the sediments of Dianchi Lake, a partitioned eutrophic waterbody in China. PLOS One. 2012;7:e37796.
Schippers A, Neretin LN. Quantification of microbial communities in near-surface and deeply buried marine sediments on the Peru continental margin using real-time PCR. Environ Microbiol. 2006;8:1251–60.
Siering PL, Clarke JM, Wilson MS. Geochemical and biological diversity of acidic, hot springs in Lassen Volcanic National Park. Geomicrobiol J. 2006;23:129–41.
Penrose L. A Terrestrial Analogue for the Origin of Life? : The Role of Mixing Zones in Modern Hot Springs in Rotorua, New Zealand [University of Auckland MSc Theses]: Univerisity of Auckland; 2020.
Kevbrin VV, Zengler K, Lysenko AM, Wiegel J. Anoxybacillus kamchatkensis sp. nov., a novel thermophilic facultative aerobic bacterium with a broad pH optimum from the Geyser valley, Kamchatka. Extremophiles. 2005;9:391–8.
Segerer A, Neuner A, Kristjansson JK, Stetter KO. Acidianus infernus gen. nov., sp. nov., and Acidianus brierleyi Comb. nov.: facultatively aerobic, extremely acidophilic thermophilic sulfur-metabolizing archaebacteria. Int J Syst Evol Microbiol. 1986;36:559–64.
Bohorquez LC, Delgado-Serrano L, López G, Osorio-Forero C, Klepac-Ceraj V, Kolter R, et al. In-depth characterization via complementing culture-independent approaches of the microbial community in an acidic hot spring of the Colombian Andes. Microb Ecol. 2012;63:103–15.
Wemheuer B, Taube R, Akyol P, Wemheuer F, Daniel R. Microbial diversity and biochemical potential encoded by thermal spring metagenomes derived from the Kamchatka Peninsula. Archaea. 2013;2013:136714.
Bonch-Osmolovskaya EA, Sokolova TG, Kostrikina NA, Zavarzin GA. Desulfurella acetivorans gen. nov. and sp. nov. —a new thermophilic sulfur-reducing eubacterium. Arch Microbiol. 1990;153:151–5.
Miroshnichenko ML, Rainey FA, Hippe H, Chernyh NA, Kostrikina NA, Bonch-Osmolovskaya EA. Desulfurella kamchatkensis sp. nov. and Desulfurella propionica sp. nov., new sulfur-respiring thermophilic bacteria from Kamchatka thermal environments. Int J Syst Evol Microbiol. 1998;48:475–9.
Itoh T, Onishi M, Kato S, Iino T, Sakamoto M, Kudo T, et al. Athalassotoga saccharophila gen. nov., sp. nov., isolated from an acidic terrestrial hot spring, and proposal of Mesoaciditogales ord. nov. and Mesoaciditogaceae fam. nov. in the phylum Thermotogae. Int J Syst Evol Microbiol. 2016;66:1045–51.
Amend AS, Martiny AC, Allison SD, Berlemont R, Goulden ML, Lu Y, et al. Microbial response to simulated global change is phylogenetically conserved and linked with functional potential. ISME J. 2016;10:109–18.
Handley KM, Piceno YM, Hu P, Tom LM, Mason OU, Andersen GL, et al. Metabolic and spatio-taxonomic response of uncultivated seafloor bacteria following the Deepwater Horizon oil spill. ISME J. 2017;11:2569–83.
Salmaso N. Effects of habitat partitioning on the distribution of bacterioplankton in deep lakes. Front Microbiol. 2019;10:2257.
Papke RT, Ramsing NB, Bateson MM, Ward DM. Geographical isolation in hot spring cyanobacteria. Environ Microbiol. 2003;5:650–9.
Podar PT, Yang Z, Björnsdóttir SH, Podar M. Comparative analysis of microbial diversity across temperature gradients in hot springs From Yellowstone and Iceland. Front Microbio. 2020;11:1625.
Schmidt VT, Reveillaud J, Zettler E, Mincer TJ, Murphy L, Amaral-Zettler LA. Oligotyping reveals community level habitat selection within the genus Vibrio. Front Microbiol. 2014;5:563.
Mayr MJ, Zimmermann M, Guggenheim C, Brand A, Bürgmann H. Niche partitioning of methane-oxidizing bacteria along the oxygen–methane counter gradient of stratified lakes. ISME J. 2020;14:274–87.
Ward CS, Yung C-M, Davis KM, Blinebry SK, Williams TC, Johnson ZI, et al. Annual community patterns are driven by seasonal switching between closely related marine bacteria. ISME J. 2017;11:1412–22.
Eren AM, Morrison HG, Lescault PJ, Reveillaud J, Vineis JH, Sogin ML. Minimum entropy decomposition: unsupervised oligotyping for sensitive partitioning of high-throughput marker gene sequences. ISME J. 2015;9:968–79.
Kleindienst S, Grim S, Sogin M, Bracco A, Crespo-Medina M, Joye SBDiverse. rare microbial taxa responded to the Deepwater Horizon deep-sea hydrocarbon plume. ISME J. 2016;10:400–15.
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP. DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods. 2016;13:581–3.
Benítez-Páez A, Portune KJ, Sanz Y. Species-level resolution of 16S rRNA gene amplicons sequenced through the MinION™ portable nanopore sequencer. GigaScience. 2016;5:s13742–016-0111-z.
Olsen GJ, Woese CR. Ribosomal RNA: a key to phylogeny. FASEB J. 1993;7:113–23.
Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, Schleifer K-H, et al. Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol. 2014;12:635–45.
Konstantinidis KT, Rosselló-Móra R, Amann R. Uncultivated microbes in need of their own taxonomy. ISME J. 2017;11:2399–406.
Kim M, Oh H-S, Park S-C, Chun J. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol. 2014;64:346–51.
Jaspers E, Overmann J. Ecological significance of microdiversity: identical 16S rRNA gene sequences can be found in bacteria with highly divergent genomes and ecophysiologies. Appl Environ Microbiol. 2004;70:4831–9.
Olm MR, Crits-Christoph A, Bouma-Gregson K, Firek BA, Morowitz MJ, Banfield JF. inStrain profiles population microdiversity from metagenomic data and sensitively detects shared microbial strains. Nat Biotechnol. 2021;39:727–36.
Bendall ML, Stevens SLR, Chan L-K, Malfatti S, Schwientek P, Tremblay J, et al. Genome-wide selective sweeps and gene-specific sweeps in natural bacterial populations. ISME J. 2016;10:1589–601.
Caro-Quintero A, Konstantinidis KT. Bacterial species may exist, metagenomics reveal. Environ Microbiol. 2012;14:347–55.
Konstantinidis KT, DeLong EF. Genomic patterns of recombination, clonal divergence and environment in marine microbial populations. ISME J. 2008;2:1052–65.
Ward DM, Cohan FM, Bhaya D, Heidelberg JF, Kühl M, Grossman A. Genomics, environmental genomics and the issue of microbial species. Heredity. 2008;100:207–19.
Henson MW, Lanclos VC, Faircloth BC, Thrash JC. Cultivation and genomics of the first freshwater SAR11 (LD12) isolate. ISME J. 2018;12:1846–60.
Rocap G, Larimer FW, Lamerdin J, Malfatti S, Chain P, Ahlgren NA, et al. Genome divergence in two Prochlorococcus ecotypes reflects oceanic niche differentiation. Nature. 2003;424:1042–7.
Huber R, Langworthy TA, König H, Thomm M, Woese CR, Sleytr UB, et al. Thermotoga maritima sp. nov. represents a new genus of unique extremely thermophilic eubacteria growing up to 90 C. Arch Microbiol. 1986;144:324–33.
Iino T, Tamaki H, Tamazawa S, Ueno Y, Ohkuma M, Suzuki KI, et al. Candidatus Methanogranum caenicola: a novel methanogen from the anaerobic digested sludge, and proposal of Methanomassiliicoccaceae fam. nov. and Methanomassiliicoccales ord. nov., for a methanogenic lineage of the class Thermoplasmata. Microbes Environ. 2013;28:244–50.
Golyshina OV, Pivovarova TA, Karavaiko GI, Kondratéva TF, Moore ER, Abraham WR, et al. Ferroplasma acidiphilum gen. nov., sp. nov., an acidophilic, autotrophic, ferrous-iron-oxidizing, cell-wall-lacking, mesophilic member of the Ferroplasmaceae fam. nov., comprising a distinct lineage of the Archaea. Int J Syst Evo Microbiol. 2000;50:997–1006.
Hallenbeck PC, Clark MA, Barrett EL. Characterization of anaerobic sulfite reduction by Salmonella typhimurium and purification of the anaerobically induced sulfite reductase. J Bacteriol. 1989;171:3008–15.
Sparacino-Watkins C, Stolz JF, Basu P. Nitrate and periplasmic nitrate reductases. Chem Soc Re. 2014;43:676–706.
Dahl C. Cytoplasmic sulfur trafficking in sulfur‐oxidizing prokaryotes. IUBMB Life. 2015;67:268–74.
Wang R, Lin JQ, Liu XM, Pang X, Zhang CJ, Yang CL, et al. Sulfur oxidation in the acidophilic autotrophic Acidithiobacillus spp. Front Microbiol. 2019;9:3290.
Sirko A, Hryniewicz M, Hulanicka D, Böck A. Sulfate and thiosulfate transport in Escherichia coli K-12: nucleotide sequence and expression of the cysTWAM gene cluster. J Bacteriol. 1990;172:3351–7.
Kappler U, Dahl C. Enzymology and molecular biology of prokaryotic sulfite oxidation. FEMS Microbiol Lett. 2001;203:1–9.
Maeda T, Sanchez-Torres V, Wood TK. Escherichia coli hydrogenase 3 is a reversible enzyme possessing hydrogen uptake and synthesis activities. Appl Microbiol Biotechnol. 2007;76:1035–42.
Tyson GW, Chapman J, Hugenholtz P, Allen EE, Ram RJ, Richardson PM, et al. Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature. 2004;428:37–43.
Gill SR, Fouts DE, Archer GL, Mongodin EF, DeBoy RT, Ravel J, et al. Insights on evolution of virulence and resistance from the complete genome analysis of an early Methicillin-resistant Staphylococcus aureus Strain and a biofilm-producing Methicillin-resistant Staphylococcus epidermidis Strain. J Bacteriol. 2005;187:2426–38.
Acknowledgements
We thank B. Drake, M. Rowe, L. Steller, L. Penrose, T. Hamilton, J. Havig, M. Dobson, and A. Hamilton for helping with field work and geochemistry data collection, and H. Sze, J. Boey, and E. Gios, for bioinformatics support. We acknowledge Tauhara North No 2 Trust, Mercury Energy (Rotorua), and the New Zealand Department of Conservation (DOC) for Parariki Stream and Lake Rotokawa site access, and the Ngati Tahu-Ngati Whaoa Runanga Trust and the staff of Wai-O-Tapu Thermal Wonderland for Waiotapu site access. We are also grateful for site access provided by Hell’s Gate Geothermal Park (P. Rayner, Operations Manager), and acknowledge the Whakapoungakau 24 Ahu Whenua trust as kaitiakitanga of the Tikitere geothermal area. The authors acknowledge the use of New Zealand eScience Infrastructure (NeSI) high performance computing facilities. This study was funded by Australian Research Council discovery project DP180103204 and Genomics Aotearoa project 1806. Research was also supported by a Royal Society Te Apārangi Rutherford Discovery Fellowship awarded to KMH.
Author information
Authors and Affiliations
Contributions
KMH, MVK, KAC and CS conceived of the study. KAC and MVK led the field trips, and CS and KMH collected samples. CS conducted the laboratory work. CS and KMH analyzed and interpreted data and wrote the first draft of the manuscript with input from KAC and MVK. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sriaporn, C., Campbell, K.A., Van Kranendonk, M.J. et al. Bacterial and archaeal community distributions and cosmopolitanism across physicochemically diverse hot springs. ISME COMMUN. 3, 80 (2023). https://doi.org/10.1038/s43705-023-00291-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43705-023-00291-z
- Springer Nature Limited