Abstract
Bacteria have evolved many strategies to spare energy when nutrients become scarce. One widespread such strategy is facultative phototrophy, which helps heterotrophs supplement their energy supply using light. Our knowledge of the impact that such behaviors have on bacterial fitness and physiology is, however, still limited. Here, we study how a representative of the genus Porphyrobacter, in which aerobic anoxygenic phototrophy is ancestral, responds to different light regimes under nutrient limitation. We show that bacterial survival in stationary phase relies on functional reaction centers and varies depending on the light regime. Under dark-light alternance, our bacterial model presents a diphasic life history dependent on phototrophy: during dark phases, the cells inhibit DNA replication and part of the population lyses and releases nutrients, while subsequent light phases allow for the recovery and renewed growth of the surviving cells. We correlate these cyclic variations with a pervasive pattern of rhythmic transcription which reflects global changes in diurnal metabolic activity. Finally, we demonstrate that, compared to either a phototrophy mutant or a bacteriochlorophyll a overproducer, the wild type strain is better adapted to natural environments, where regular dark-light cycles are interspersed with additional accidental dark episodes. Overall, our results highlight the importance of light-induced biological rhythms in a new model of aerobic anoxygenic phototroph representative of an ecologically important group of environmental bacteria.
Similar content being viewed by others
Introduction
In natural environments, free-living bacteria face, between short intervals of fast multiplication, long periods of stasis or very slow growth [1,2,3]. To survive such periods, they have evolved diverse strategies to optimize resource utilization and spare energy [4,5,6,7]. Facultative photoheterotrophy is such a strategy that a subset of environmental bacteria use to supplement their mostly heterotrophic growth. It can be based on two main molecular mechanisms: retinalophototrophy, which uses rhodopsins to pump ions through the membrane, and chlorophototrophy, which is based on photochemical reaction centers containing chlorophylls or bacteriochlorophylls [8]. Aerobic Anoxygenic Phototrophy, hereafter AAnP, is a common chlorophototrophic pathway, whereby usable energy is produced from light under aerobic conditions in the absence of H2O dissociation and O2 release [9, 10].
Aerobic anoxygenic phototrophic bacteria include, among others, members of the alpha-, beta- and gammaproteobacteria [11]. These represent an important fraction of the prokaryotes present in the upper layers of seas and lakes, but are also widespread in environments as diverse as the phyllosphere, soil crusts, or thermal springs [12,13,14]. These bacteria encode a set of about 45 genes required to produce bacteriochlorophyll a, the main pigment needed for phototrophy, and the components of their light-harvesting complexes, including carotenoids [15]. While these bacteria are in general not able to grow photoautotrophically, with the notable exception of Dinoroseobacter shibae which can use the ethylmalonyl-CoA pathway [16], they can benefit from light exposure to maintain higher concentrations of intracellular ATP, incorporate limited amounts of CO2 via substrate co-assimilation or anaplerotic reactions, and spare organic carbon through downregulation of respiration [17,18,19,20,21]. However, bacteriochlorophyll a biosynthesis and AAnP represent a significant energy investment and are known to generate toxic byproducts, especially singlet oxygen and other reactive oxygen species, in the presence of oxygen and light [11, 22, 23]. That is why many aerobic anoxygenic phototrophs tend to regulate bacteriochlorophyll a production strictly and limit its synthesis to dark phases [24, 25].
The life history of aerobic anoxygenic phototrophs is likely to be deeply affected by diel cycles, both because these bacteria can actively utilize light when needed and because they tend to associate and cooperate with obligate photoautotrophs [23, 26,27,28,29]. While we know the preferences of some species in terms of light intensity and adaptations to non-optimal light conditions [30, 31], our understanding of the rhythmic physiology of these bacteria in response to diurnal cycles and more generally of the adaptive value of AAnP under different light regimes is still limited [32]. Indications can be found scattered in literature pertaining to quite diverse microbiological fields. Field studies in microbial ecology suggest that the transcription of AAnP genes may vary throughout the day, being maximal at night, and that cell division and cell activity are also driven by diurnal cycles in AAnP-competent bacteria [33,34,35]; in freshwater natural communities grown ex situ, respiration rates and organic substrate assimilation can vary in response to periodic infrared light exposure, suggesting that these variations are driven by AAnP [36]. Laboratory based studies of AAnP models show clear differences in global transcription patterns between cells cultured in the dark and in the light, some of these differences being directly connected to light-stress responses [37, 38]. Finally, studies on pure cultures of Roseobacter sp. OCH 114 and Dinoroseobacter shibae DFL12 under controlled conditions showed that these bacteria benefit from exposure to continuous light or dark-light cycles for their survival, although this survival benefit was never genetically linked to phototrophy [17, 39]. Despite these convergent indications, consistent evidence on a single experimental model, ranging from genetics to fitness measurements, is actually missing. Moreover, it is generally unclear how AAnP affects bacterial fitness under diel cycles and how facultative aerobic anoxygenic phototrophs navigate the tradeoff between costs and benefits of phototrophy.
Here, we investigate how a new freshwater model of AAnP-competent bacteria, Porphyrobacter sp. ULC335, responds to and benefits from diel dark-light cycles. When grown to stationary phase under dark-light alternance, Porphyrobacter sp. ULC335 presents rhythmic patterns of density variation that were not reported in most other species capable of AAnP. We explain these patterns by connecting them to variations in mortality and reproduction rates due to phototrophy, study their transcriptional correlates, and reveal their impact on fitness. In so doing, we contribute to a better understanding of the rhythmic patterns governing the physiology and lifestyle of bacteria prevalent in photic aquatic layers.
Methods
Detailed methods are available in a “Supplementary Methods” file.
Standard growth conditions
Porphyrobacter sp. ULC335 was grown in BG11 media [40] supplemented with 0.5 g/L tryptone, 0.25 g/L yeast extract, and vitamins. Standard batch growth took place at 30 °C with agitation (180 rpm). Growth in the Photon Systems Instruments MC1000 took place at 28 °C. Batch cultures for competition and survival experiments were done in 24-well plates at 28 °C with agitation.
Pigment characterization
Cell pellets were extracted with 7:2 acetone-methanol for 5 min; the absorbance of the supernatant was measured in quartz cuvettes (Genesys 10 s, Thermo Scientific) or in 96-well plates (SpectraMax i3x, Molecular Devices). For Thin Layer Chromatography, 100 µl of extract were spotted onto a Silica 60 F254 plates (Merk, art. 5554). The plate was migrated in 8:0.75:0.2:0.8:0.25 petroleum ether:hexane:isopropanol:acetone:methanol.
Transposon library
The plasmid pRL27 [41] was conjugated into Porphyrobacter sp. ULC335 and several mutants whose color differed from the wild type were isolated. The insertion sites were identified using the method previously published [41]. Four mutants are central to our study: mutant B+C− (insertion in crtI, coding for the phytoene synthase, required for carotenoid production), mutant B−C+ (insertion in bchH, coding for a subunit of the magnesium chelatase, required for Bchla synthesis), mutant B−C− (insertion in ispA, coding for the farnesyl diphosphate synthase, required to synthesize a precursor of both Bchla and carotenoids), mutant B++C++ (insertion in the ppaA gene coding for an AAnP regulator).
Genome sequencing and annotation
The genomic DNA was purified using the Qiagen Genomic-tip 20/g kit and sequenced on PacBio Sequel instruments. The genome was assembled using PacBio CCS 4.1.0 for the read correction step and Flye 2.6 [42] for the assembly. The functional classification of the genome content was done using the online blastKOALA service with the genus Porphyrobacter as a reference.
RNA-sequencing
RNA was extracted from 1 ml culture using TRIzol (Invitrogen) followed by purification on RNeasy mini-kit columns (Qiagen). RNA-sequencing was carried out at Novogene on Illumina NovaSeq systems after ribosomal depletion and in-house library construction. Reads were trimmed and filtered using fastP [43], mapped using bowtie2 [44], and counted using featureCounts [45]. Comparison between wildtype and B−C+ strains was done using the DESeq2 package [46]. Temporal analysis of individual genes was done using the ImpulseDE2 package [47]. Each temporally regulated gene was assigned to a cluster using hclust (Euclidian algorithm, 15 clusters) on the Z-scores of the rlog-transformed count data for all four timepoints. The ggVennDiagram [48] and ggplot [49] libraries were used to plot the summary figures.
Phylogenetic analysis
Fourteen protein sequences characteristic of the AAnP system from complete alphaproteobacterial genomes were concatenated and aligned using muscle 3.8.31 [50]; a phylogenetic tree was constructed using iqtree 2.1.1 [51] with default parameters (evolution model chosen automatically). In parallel, a set of conserved ribosomal proteins, elongation factors, and gyrases were retrieved and a phylogenetic tree was constructed following the same procedure. Strains were considered to encode the anoxygenic phototrophy gene cluster if they gave positive blastp matches with at least 13 out of the 14 proteins.
Flow cytometry and fluorescence microscopy
Cells stained with LIVE/DEAD BacLight (Invitrogen) were analyzed by flow cytometry (BD Accuri C6 Plus). The zones corresponding to the dead, compromised, and live subpopulations were determined empirically based on the density distribution of live and 2.5% glutaraldehyde-fixed cells. Visualization and quantification were done with R [52] using flowCore [53]. The same cells were analyzed using fluorescence microscopy, where only dead and live subpopulations could be distinguished.
Cells stained with DyeCycle Orange (Invitrogen) were analyzed by flow cytometry. The coordinates of the highest density in the regions of the 1 N and 2 N peaks (green fluorescence) were determined empirically. The lowest density point between the 1 N and the 2 N peaks on the empirical density plot was used as a proxy for the “actively replicating” population.
Survival and competition experiments
For survival experiments, late exponential phase cultures of wild type, B−C+, and B++C++ were diluted and grown under continuous light, continuous dark, or 12 h/12 h dark-light cycles in triplicates. At the beginning of stationary phase (48 h) and after four additional days (144 h), CFUs were determined and the survival rate (144 h vs 48 h) was calculated.
For competition experiments, exponential phase cultures of the two mutant strains were mixed with the wild type strain in equal proportions and grown under continuous light, continuous dark, or 12 h/12 h dark-light cycles in triplicates. The dual cultures were plated at 48 h and 144 h like for the survival experiment, and the ratios of the mutant to the wild type were calculated for both timepoints based on CFUs.
In survival and competition experiments, irradiance during light periods was 80 µmol/s/m2.
Results
Strain ULC335 is a representative of the Porphyrobacter genus, in which AAnP is ancestral
In the course of a study on a bacterial community associated with a single cyanobacterial species (ULC335) [54] (Suplementary File 1), we isolated a nonmotile dark-red colony-forming bacterial strain which, based on its 16S ribosomal RNA sequence, was assigned to the Porphyrobacter-Erythrobacter group (Erythrobacteraceae genus I–VI) [55]. The strain encoded an allele of the pufM gene, coding for a light-harvesting complex subunit, and readily produced, under standard aerobic culture conditions, bacteriochlorophyll a (hereafter “Bchla”) (Suplementary Figs. 1 and 2), suggesting that the strain was capable of AAnP [56, 57].
Bacteria belonging to the Porphyrobacter-Erythrobacter group are ubiquitous in aquatic environments and often found associated with Cyanobacteria [58,59,60,61]. While many genera among alphaproteobacteria contain members capable of anoxygenic phototrophy (Suplementary Fig. 3), the Porphyrobacter-Erythrobacter group is one of the few taxonomic units in which the AAnP gene cluster is most likely ancestral and present in the vast majority of sequenced strains (Figs. 1A, SFig. 4). Moreover, very few members of the Porphyrobacter-Erythrobacter have lost the gene cluster, which indicates that it consistently provides a benefit. Noteworthy, the dominant mode of transmission of the genes coding for anoxygenic phototrophy within alphaproteobacteria, including within the Porphyrobacter-Erythrobacter group, is vertical, suggesting that the gene cluster has extensively co-evolved and co-adapted with the core genome (Suplementary Figs. 5 and 6). On this basis, we consider the Porphyrobacter-Erythrobacter group as an excellent model to study the effects of long-term adaptation to AAnP.
A Simplified phylogenetic tree made from an alignment of concatenated core-genome proteins of Erythrobacteraceae rooted on Caulobacter crescentus (Caulobacterales); a tree comprising more strains can be found in Suplementary Fig. 4. Porphyrobacter sp. ULC335 branches within the Porphyrobacter-Erythrobacter clade, in which the anoxygenic phototrophy gene cluster is quasi-ubiquitous (branches in green). Scale bar: average number of substitutions per site. Numbers at the nodes are support values (percent of generated trees supporting the branch). B The aerobic anoxygenic phototrophy (AAnP)-associated genes found in Porphyrobacter sp. ULC335. The AAnP-associated genes are scattered in six loci, one major locus more than 40 kb in length, and five minor loci, comprising two or three genes. The color codes indicate the predicted functional categories of the products: dark green for proteins involved in bacteriochlorophyll a synthesis, yellow for structural proteins of the reaction centers, orange for proteins involved in carotenoid synthesis, light green for regulatory genes, gray for genes associated with the AAnP genes but whose function is not clear. C Schematic overview of the genome of Porphyrobacter sp. ULC335. The concentric circles represent, from inside out, the GC skew (yellow and gold), the GC content (light and dark brown), RNAs (red), genes encoded on the reverse and forward strand (gray and black), AAnP-related loci (same color code as in B), genome coordinates using the coding sequence of dnaA as a reference.
To further characterize Porphyrobacter sp. ULC335, we sequenced its genome, obtaining a single closed circular sequence of 3.6 Mbps. Besides the typical genetic complement of free-living alphaproteobacteria (Suplementary Fig. 7A, B), it encoded a complete AAnP gene cluster comparable to the one found in sequenced members of the Porphyrobacter-Erythrobacter group (Fig. 1B) [15] and no less than five blue-light responsive proteins including a BLUF domain [26, 62] (Suplementary Fig. 7C). Some of the genes coding for the carotenoid synthesis pathway are found in the AAnP gene cluster (crtC, crtD, crtF), others in secondary loci (crtZ, crtW; crtB, crtI, crtY; crtG) (Fig. 1B, C). Thin layer chromatography revealed that, besides Bchla and its intermediates, Porphyrobacter sp. ULC335 synthesizes a number of red and yellow pigments (Suplementary Fig. 8). Porphyrobacter sp. ULC335 does not encode a full carbon fixation pathway and is unlikely to grow autotrophically. However, like Erythrobacter sp. NAP1, it produces key enzymes of the 3-hydroxypropionate bicycle pathway for carbon fixation, which would give it the capacity to co-assimilate CO2 while using glycolate, glyoxylate, or acetate, among others, as carbon sources (Suplementary Fig. 9) [19, 63]. Altogether, these data suggest that Porphyrobacter sp. ULC335 could make use of light to spare organic substrates under nutrient limitation [20].
In stationary phase, Porphyrobacter sp. ULC335 presents light-induced rhythmic growth patterns dependent on a functional photosystem
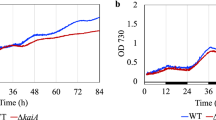
When monitored under 12 h/12 h dark-light cycles, Porphyrobacter sp. ULC335 presented, during stationary phase, strong variations in optical density in response to light in a range of light intensities (Fig. 2A, Suplementary Fig. 10). The bacterial density increased during the light phase and decreased during the dark phase; the rhythms strictly followed the photoperiod and did not persist under constant light or darkness (Fig. 2A, Suplementary Fig. 11), which suggests that they were not driven by a circadian clock but by external light variations. We hypothesized that such variations in optical density could be a consequence of the phototrophic activity of the organism, although, to our knowledge, such conspicuous rhythms have not been reported so far in bacteria capable of AAnP in the absence of additional phototrophic systems. To test this hypothesis, we generated a Tn5 transposon mutant library of the strain and screened for color variations which would betray an alteration of Bchla or carotenoid content. We obtained four mutants of interest (Fig. 2B): strain B+C− produced Bchla but no carotenoids (insertion in crtI, coding for the phytoene synthase), strain B−C+ produced carotenoids, although less than the wild type, but no Bchla (insertion in bchH, coding for a subunit of the magnesium chelatase required for Bchla synthesis), strain B−C− produced neither Bchla nor carotenoids (insertion in ispA, coding for the farnesyl diphosphate synthase, required to synthesize a precursor of both Bchla and carotenoids), strain B++C++ overproduced both Bchla and carotenoids, especially in rich medium and under continuous light (insertion in the ppaA gene coding for an AAnP regulator) (Fig. 2C; Suplementary Fig. 12). Of the four mutants, only the B++C++ strain presented rhythmic growth similar to the wild type strain (Fig. 2D). By contrast, strains B−C− and B−C+ did not present any periodic optical density variation (Fig. 2E). The strain B+C− was very sensitive to light: as soon as light was switched on the density started decreasing and continued during the subsequent cycles (Fig. 2E); upon light exposure, the viability of cells, as quantified by Life/Dead staining and CFUs, decreased by at least two orders of magnitude (Suplementary Fig. 13). This suggests that, in the absence of carotenoids, Bchla production might be toxic to the cells, especially in the presence of light. Overall, these data indicate that the rhythmic density variations in response to light depend on a functional reaction center including both Bchla and carotenoids.
A Rhythmic variations in optical density (measured at 680 nm) of three replicate cultures of Porphyrobacter sp. ULC335 under dark-light alternance (light at 180 µmol/s/cm2). Light intensity, culture temperature (water bath), and room temperature are shown as evidence that density changes are from biological and not technical origin. Cultures were maintained in 80 ml tubes in a device adapted for the culture of cyanobacteria or algae. The rhythms do not present persistence under continuous light (120 h–156 h) or continuous darkness (204 h–240 h), and follow light variation seamlessly when the period is shorter than 24 h (240 h–288 h). B Colony phenotype of Porphyrobacter sp. ULC335 (B+C+) and four transposon mutants grown on BG11-P-agar in the dark. The color of colonies are in agreement with the absorbance profiles of liquid cultures (Suplementary Fig. 12). C Simplified pathway of carotenoid and bacteriochlorophyll a biosynthesis. In red, the genes in which the transposon is inserted for each of the mutant strains. Ticked lines represent multiple synthesis steps (enzymatic reactions) or multiple convergent effects (regulators). D Rhythmic variations in optical density (measured at 680 nm) of the wild type compared with the bacteriochlorophyll a and carotenoid overproducer strain. No significant difference in pattern is apparent between the two strains. The culture conditions are similar to (A). E Variations in optical density (OD, measured at 680 nm) of the wild type compared with the three bacteriochlorophyll a and/or carotenoid mutant strains. B−C+ and B−C− did not present any rhythmic growth pattern; B+C− presented slight variations in slope between dark and light phases, the speed of the OD decline increasing during light phases. Because B+C− was strongly inhibited by light, the cultures were maintained under continuous darkness until stationary phase, when a 12 h/12 h dark-light alternance illumination regime was started.
The optical density variations in response to light correlate with changes in cell viability and DNA replication
To further investigate the origin of these rhythms, we sought to determine what happens to Porphyrobacter sp. ULC335 cells when they are exposed to darkness. To do so, we sampled six parallel cultures of Porphyrobacter sp. ULC335 grown to stationary phase and maintained under continuous light for two days; three of the cultures were switched to the dark for 12 h before sampling (Time Point 1 [TP1]) and switched back to light immediately after sampling (dark treatment); a second sample of all six tubes was taken eight hours after the switch back to light (TP2) (continuous light treatment) (Fig. 3A, Suplementary Fig. 14). Using spectrophotometry and flow cytometry, we did not find any difference in pigment content, cell morphology or aggregation between the treatments (Suplementary Fig. 15). However, using Live/Dead staining, we observed a clear difference in the proportion of living and compromised (permeable) cells between the two treatments: while 70% cells cultured under continuous light were viable and non-compromised, based on fluorescence microscopy, this proportion dropped to 40% after 12 h in the darkness; interestingly, it was enough for the latter cultures to be exposed back to light for 8 h to restore the proportion of viable cells to original levels (Fig. 3B). Colony forming units confirmed this trend (Suplementary Fig. 16). Flow cytometry revealed that three subpopulations coexist in the cultures, independently of the treatment: viable noncompromised cells, compromised (permeable) cells, and dead cells (Fig. 3C; controls: Suplementary Fig. 17). Like the “dead” subpopulation, the “compromised” subpopulation was significantly larger in the cultures exposed to 12 h of darkness than in the cultures exposed to continuous light (Suplementary Fig. 18). Moreover, by analyzing the DNA content of the cells, we found that the frequency of cells undergoing DNA replication was significantly lower in the cultures exposed to darkness than in the cultures kept under continuous light; upon reexposure to light, the two populations were indistinguishable again (Fig. 3D; Suplementary Fig. 19). Taken together, these results suggest that the life history of Porphyrobacter sp. ULC335 follows alternate phases in response to illumination conditions: during dark-phases, cells slow down replication and a significant proportion of them dies; during light-phases, part of the surviving cells replicate and divide, and this, possibly along with the partial recovery of some compromised cells, leads to the restoration of previous viability levels in the population.
A Experimental design. Six Porphyrobacter sp. ULC335 cultures were maintained in parallel under dark-light alternance up to stationary phase to allow for carotenoid and bacteriochlorophyll a accumulation; then, the cultures were exposed for 48 h to continuous light, before three of the six cultures were switched to the dark for 12 h and back again to light for another 12 h. Two series of samples were taken, one at the end of the dark period and one 8 h after return to light, and analyzed using fluorescence microscopy and flow cytometry. B Living cells as determined using fluorescence microscopy on Live/Dead stained cells. The proportion of living cells dropped after the dark phase and returned to baseline after exposure to light. At least 500 cells were counted per replicate. C Dead, compromised, and alive subpopulations as observed using flow cytometry after Live/Dead staining. The arrows indicate the probable path from alive to compromised to dead cells, as membrane permeability increases; lysed cells have released their cellular content, including DNA, and cannot be observed using flow cytometry. D The proportion of replicating cells, as determined by flow cytometry using DyeCycle Orange stain, decreased during the dark phase and returned to baseline after reexposure to light. A total of 40'000 cells was analyzed. The calculation of the replication index is explained in the methods; see Suplementary Fig. 19 for a more detailed representation of flow cytometry data. Amount of light during light periods: 150 µmol/s/m2. Asterisks indicate statistically significant differences (bilateral Student’s t-test, p-value < 0.05).
In Porphyrobacter sp. ULC335, direct responses to light and responses to electron flow underlie rhythmic physiology
To understand the physiological correlates of this behavior, we sequenced the full transcriptome of stationary phase cultures under 12 h/12 h darklight cycles. We were particularly interested in transcriptional responses upon light phase transition, so we sampled one hour before (Late Night, “LN”) and after (Early Day, “ED”) the switch to light, one hour before (Late Day, “LD”) and after (Early Night, ”EN”) the switch to darkness. In addition, we treated in parallel the wild type strain and the arrhythmic phototrophy mutant B−C+ in order to be able to discriminate between direct responses to light (independent of Bchla) and responses due to the electron flow generated by phototroph.
Disrupting photosynthesis had massive consequences at the transcriptional level. Among protein-coding genes, 1533 out of 3418 were differentially regulated more than two-fold at least at one of the four timepoints in the phototrophy mutant, about half being downregulated and half being upregulated (Fig. 4A; Suplementary File 2). The time of day had a strong influence on the divergence of the transcriptional profile between the two strains. The number of differentially regulated genes between the two strains differed the most at the EN time point and differed the least at the LN time point, suggesting that, in the prolonged absence of light exposure, the physiologies of the two strains tended to converge. Globally, the mutant and the wild type seem to time transcription differently. Genes more transcribed in the mutant compared to the wild type were most abundant during the day (and EN), while genes less transcribed in the mutant compared to the wild type were most abundant during the night. Among genes downregulated in the mutant, functions linked to photosynthesis, including porphyrin metabolism, and ribosome biogenesis were enriched, especially during the day; genes upregulated in the mutant included categories linked to DNA repair, translation, and energy metabolism (Fig. 4B; Suplementary File 3). This indicates that the transcriptional investment into energy metabolism, especially photosynthesis, and translation machinery is modified, both in terms of timing and intensity, in the phototrophy mutant compared to the wild type.
A Summary of the experiment: number of genes upregulated (right) or downregulated (left) in the Bchla null mutant compared to the wild type at different timepoints (ED: early day; LD: late day; EN: early night; LN: late night). B KEGG categories enriched among differentially regulated genes in the Bchla null mutant compared to the wild type; ED and LD on one side and EN and LN on the other side are considered together in this analysis. C The figure summarizes information about the temporal regulation of genes in the wild type strain and its Bchla null mutant. The Venn diagram indicates the number of temporally regulated genes in the two strains. The major clusters of genes showing a consistent light-response pattern in either of the two strains are presented based on their temporal profile (upper plot; the datapoints represent the relative transcription rates [the Z-scores of the rlog-transformed count data]) and the fraction they represent among temporally regulated genes in both strains (lower plot; light colors represent genes falling in the same cluster in the wild type and the Bchla null mutant). The same data split by strain is presented in Suplementary Fig. 20. D Heatmap of gene expression for the AAnP-associated genes. The first two columns show the relative expression of each gene in the two strains; the next eight columns represent the variation over time of the two strains. The gene number and name are shown in cyan for Bchla biosynthesis genes, gold for reaction center genes, orange for carotenoid biosynthesis genes, and red for regulators; gray is used for genes whose function is dubious.
We then identified genes differentially regulated over time (Impulse DE algorithm, p-value < 0.01) in either the wild type or the B−C+ strain, clustered them by temporal profile and compared the different sets of genes between the two strains (Suplementary File 4; Suplementary Fig. 20). Again, photosynthesis disruption had a huge impact: only 283 genes (8% of the genome) varied over time in the B−C+ strain, compared to 1933 genes (56% of the genome) in the wild type strain. This indicates that functional AAnP not only provides energy to the cells, but also drives a pervasive rhythmic physiology that is entirely absent in the phototrophy mutant. More than half of the genes temporally regulated in the wild type strain show opposite expression trends during daytime and nighttime. Four gene clusters (655 genes in total) are consistently upregulated during the day and downregulated during the night (Fig. 4C, lower part), while two clusters (444 genes in total) are downregulated during the day and upregulated during the night (Fig. 4C, upper part). Overall, this suggests that the transcriptional profile, and consequently the physiology, of Porphyrobacter sp. ULCC335 presents a marked contrast between day and night involving opposed regulation of more than 1000 genes (31% of the genes). Genes downregulated during the day and upregulated during the night are enriched for the AAnP pathway, ribosome biosynthesis, and oxidative phosphorylation, including the electron transfer chain; genes upregulated during the day and downregulated during the night include transporters, energy metabolism and especially genes involved in the 3-hydroxypropionate bicycle and possibly implicated in CO2 co-assimilation (Suplementary File 5). This suggests that the night is dedicated to replenish Bchla and the ribosomal content of the cells, while nutrient assimilation might be privileged during the daytime.
Like in other AAnP-competent bacteria, genes involved in Bchla biosynthesis and components of the light-harvesting complexes tended to be downregulated during the day and upregulated at night in both the wild type and the Bchla null mutant strain (Fig. 4D; Suplementary Fig. 21). However, the amplitude of expression changes across the four timepoints was attenuated in the wild type strain compared to the Bchla null mutant. In B−C+, more than half of the expressed genes presented a four-fold difference in expression between late night (maximal expression) and late day (minimal expression), while a variety of attenuated and shifted profiles was represented in the wild type. These differences in the expression of AAnP-related genes suggest that genes which mainly respond to light cues in the B−C+ strain are tuned, in the wild type, via mechanisms sensitive to energy flow which are only active when AAnP is functional.
Overall, our transcriptome analysis revealed that diel light cycles have a major impact on the transcriptional program and physiology. The activity of the cells differs markedly between the daytime and the nighttime, and there is a clear physiological shift between early and late timepoints in either day or night. Consistent with our observations on optical density, inactivation of AAnP hampers rhythmic transcription and leads to a transcriptional profile that is most similar to the one presented by the AAnP-competent strain at the end of the night, when the benefits of phototrophy wear off.
Phototrophy provides a survival benefit in stationary phase that correlates with Bchla production
Porphyrobacter sp. ULC335 cultures present oscillations in density in stationary phase that correlate with changes in mortality and reproduction rates. However, it is unclear how these variations affect the overall survival of the bacteria, especially in view of the absence of any density variation in Bchla null strains (B−C+ and B−C−). We took advantage of the set of mutants we had generated (Fig. 2B, C) to ask whether AAnP actually provides some fitness advantage to Porphyrobacter sp. ULC335 depending on the illumination regime (DL: 12 h/12 h Dark-Light alternance, D: constant Dark, L: constant Light).
While AAnP did not provide a significant advantage in exponential up to early stationary phase (Suplementary Figs. 22 and 23), survival after 96 h spent in stationary phase was strongly affected by the genotype and the illumination regime. This is consistent with the accumulation pattern of Bchla and carotenoids, whose content increases a lot between 24 h and 48 h of growth, especially in the D and DL treatments, in the two strains capable of AAnP (Fig. 5A). When AAnP was impossible, either in the absence of light or in the absence of Bchla (B−C+ mutant), the CFUs decreased by about two orders of magnitude leading to survival rates around or below 5% (Fig. 5B). The capacity to produce energy through AAnP helped the wild type and B++C++ strains survive better under the DL regime, reaching 35% and 60% survival respectively. Under continuous light, again, the wild type and B++C++ strains survived better than the Bchla null mutant, but mortality was nonetheless much higher than under the DL regime, presumably because Bchla accumulation was impaired (Fig. 5A). These results demonstrate that AAnP provides a survival advantage to Porphyrobacter sp. ULC335 that depends on both growth phase and illumination regime, and partially correlates with Bchla accumulation. In our strain, AAnP is therefore not primarily used to supplement active heterotrophic growth, but to enhance survival during periods of starvation, especially under DL illumination.
A Bchla content at different timepoints during batch culture. The barplots represent the absorbance at 770 nm (Bchla) corrected by the optical density of the culture (average of two replicates), normalized so that the average of the first timepoint of the wild type strain equals 1. The wild type strain is shown in blue, the Bchla overproducer (B++C++) in yellow. The experiment was done under 12 h/12 h dark-light cycles, under continuous darkness, or under continuous light. B Percent survival, based on colony forming units, of the wild type, the Bchla null mutant (B−C+) and the Bchla overproducer (B++C++) after five days in stationary phase. The experiment was done under 12 h/12 h dark-light cycles, under continuous darkness, or under continuous light. The baseline is taken in early stationary phase (48 h of growth). C, D Outcome of competitions between the wild type and each of the two mutants after five days in stationary phase. The two competing strains were mixed in equal proportions after they had reached stationary phase (48 h of growth). E Figure summarizing the competitiveness of each strain under different illumination conditions. The inner color of the circles varies with relative Bchla/carotenoid content; the most competitive strains are outlined in red; the relative competitiveness is represented by arrows of three different thicknesses.
To complement these conclusions, we ran competition experiments in which the wild type strain and a competitor strain (either B−C+ or B++C++) were grown together in batch under a DL, D, or L regime. We sampled the cultures at 48 h (start of stationary phase) and at 144 h (late stationary phase), and compared the ratios between the strains. The competitiveness of the strains varied depending on the light regime in predictable ways (Fig. 5C, D). In the absence of light (D), the Bchla null mutant outcompeted both Bchla producers, and the wild type strain outcompeted the B++C++ strain. In the presence of light, under both the DL and L regimes, the B++C++ strain outcompeted the wild type strain which, in turn, outcompeted the Bchla null mutant. Altogether, these results indicate that, in Porphyrobacter sp. ULC335, AAnP has a significant cost, including in stationary phase, but that the fitness advantages it provides as soon as light is present—even when light is continuous and prevents the efficient accumulation of Bchla—warrants its maintenance in natural populations.
It stands out that, in our experimental conditions, the wild type strain is less fit than the Bchla and carotenoid overproducer B++C++ whenever light is present. This suggests that, in the natural environment of Porphyrobacter sp. ULC335, the photoperiod is on average lower than 12 h or that the regular diel dark-light alternance can often be interrupted by periods of continuous accidental darkness (Fig. 5E).
Discussion
In this work, we have isolated and described a new Porphyrobacter strain, which we found associated with a freshwater cyanobacterium (Snowella sp.). In stationary phase, this strain presented rhythmic variations in optical density in response to dark-light cycles, that we correlated with phases of high mortality and low reproduction in the darkness and phases of lower mortality and higher reproduction in the light. This behavior is dependent on Bchla production and a functional reaction center, including carotenoids contributing to photoprotection and light-harvesting. Underlying the cycles, we found pervasive rhythmic transcription patterns arising from direct responses to light, possibly driven by any of the five BLUF-domain containing proteins encoded in the genome, and indirect responses to increased electron flow and oxidative stress caused by phototrophic activity. Production of Bchla was found to be costly under continuous darkness and inefficient under continuous light; it provides maximal survival benefits in stationary phase under dark-light cycles, indicating that the strain is adapted to diurnal light cycles. So overall, in this study, we described a distinct bacterial rhythmic behavior, explained it in terms of fluctuations in death and reproduction rates at the cellular level, partially elucidated the transcriptional patterns that underlie it, and linked it to a survival benefit, providing a rationale for its evolution.
Based on average nucleotide identity with known genomes, Porphyrobacter sp. ULC335 is likely a new species in the Porphyrobacter-Erythrobacter clade (Suplementary Fig. 24). While Porphyrobacter sp. ULC335 probably obtains most of its energy, reductive power, and carbon from organic substrates, it can use phototrophy to supplement heterotrophic growth and to ensure survival when organic substrates become limiting like other AAnP-competent bacteria [11]. Because of the phylogenetic prevalence of AAnP in the clade, these organisms are expected to have evolved ways to tune their physiology to nutrient and light fluctuations, as evidenced by the high numbers of blue-light sensing proteins encoded in the genome of Porphyrobacter sp. ULC335 together with homologs for PpsR, AppA, and FnrL, known to integrate phototrophy with metabolism [64, 65]. Because Porphyrobacter strains are frequently associated with filamentous or colonial Cyanobacteria [58, 66, 67], their physiology could also indirectly follow the light and circadian clock-driven life cycle of their host like it has been suggested for other Cyanobacteria-associated heterotrophs [68]. This makes members of the Porphyrobacter genus excellent candidates for the study of bacterial rhythmic behaviors and interactions.
The conspicuous light-driven oscillations in optical density we detected in batch cultures of Porphyrobacter sp. ULC335 have never been reported or studied so far in AAnP-competent bacteria, with the exception of Sphingomonas glacialis AAP5, recently isolated from an alpine lake, which combines retinalophototrophy and chlorophotoptrophy [69]. In our case, the oscillations involve an increase in density during light phases and a decrease in density during dark phases. Interestingly, the net difference in optical density per 24-h period is almost null (Suplementary Fig. 10) suggesting that the biomass lost during dark phases is regained during light phases. The full oscillatory pattern is strictly dependent on both Bchla and carotenoids; however, when Bchla is produced in the absence of carotenoids, light phases trigger a sharp decrease in optical density that hardly slows down during dark phases (Fig. 2E, Suplementary Fig 13). This suggests that Bchla, or some derivative of it, might become toxic, for instance by generating reactive oxygen species, in the presence of light and cause cell lysis, including in a delayed way during the dark phase [22, 70, 71]. We showed that the decrease in optical density observed during dark phases correlates with an increase in the frequency of membrane-compromised and non-viable cells and a decrease in the frequency of replicating cells in the bacterial population. Based on the decrease in optical density, which corresponds to about 2 to 3% of the total biomass, it is likely that only a fraction of the 20% to 40% of compromised or dead cells (see Fig. 3B) actually lyse during the night. The decrease in compromised or dead cell proportion during the light phases is therefore most likely due to membrane repair and, to some extent, to continued lysis of affected cells, compensated by cell multiplication. Overall, these observations suggest that AAnP functions as a double edge sword: when exposed to light, AAnP photosystems sustain energy production and recycling of organic compounds; during dark phases, however, when energy becomes scarce, a reaction consecutive to Bchla exposure to light leads to membrane permeabilization and cell lysis; during the following light phase, the population recovers both in terms of membrane integrity and in terms of viability. It is however unclear whether this recovery is mostly due to compromised cells being able to repair their membrane or to the renewed growth of the surviving population based on nutrients released by lysing cells.
Physiological rhythms in bacteria and especially in heterotrophs remain understudied. Our RNA-sequencing data set reveals that, when AAnP is active, the alternance of dark and light phases affects the transcription of more than 50% of the genes. The transcripts responding to light in the absence of Bchla are only a minority of at most 10%. This indicates that physiological rhythms are pervasive in AAnP-competent bacteria and most likely generated through complex interactions between direct light responses (via light sensors), and indirect responses to light stress, redox and energy state, metabolism, cell-cycle, etc. About half of the genes are consistently upregulated or downregulated by light. This suggests that there is a degree of temporal separation of cellular activities between day and night in these bacteria. Night-activated genes include many genes connected to translation, especially components of the ribosome and translation complexes, as well as genes involved in phototrophy like in other AAnP-competent bacteria [37, 38]. This suggests that dark phases are dedicated to the replenishment of components essential for light-harvesting and cellular functioning; they tend to limit, however, active DNA replication as observed through flow cytometry. By contrast, light phases promote the transcription of genes involved in energy metabolism and protection against oxidative light stress, but also key genes in DNA replication (dnaA which falls in cluster 2, among genes maximally expressed during the day) and cell division (ftsZ, which falls in cluster 4, among proteins maximally expressed by the end of the day); these genes and functions would stimulate DNA replication and cell division, supporting our flow cytometry data (Fig. 3D).
Despite a few precursory studies, the evolutionary benefit of AAnP has not benefited from much attention. Previous research has found that exposure to light increases survival of Roseobacter and Dinoroseobacter strains in stationary phase, but without formal demonstration that this benefit can be traced back to AAnP [17, 39]. Here we provide, based on manipulation of genotypes and growth environments, an in-depth assessment of costs and benefits of AAnP in terms of its effect on survival and fitness in stationary phase. We find that Bchla production has a significant cost under constant darkness, where Bchla producers get outcompeted by nonproducers. By contrast, under continuous light or alternance, Bchla producers outcompete nonproducers. Under continuous light and, to a lesser extent, under light alternance, strains producing higher levels of Bchla and/or carotenoids get a strong selective advantage. This suggests that the regulation of AAnP in the wild type Porphyrobacter sp. ULC335 is adapted to dark-light cycles with a photoperiod lower than 12 h or to environments where cells can be accidentally cut off from light by opaque bodies or move out of the photic zone.
Data availability
The Porphyrobacter sp. ULC335 genome is publicly available on the NCBI repository (SAMN19986570, GCF_025917005.1). The RNA-sequencing data is publicly available on the NCBI Gene Expression Omnibus (GEO) repository (GSE245047).
References
Kolter R, Siegele DA, Tormo A. The stationary phase of the bacterial life cycle. Annu Rev Microbiol. 1993;47:855–74.
Hengge-Aronis R. Survival of hunger and stress: the role of rpoS in early stationary phase gene regulation in E. coli. Cell. 1993;72:165–8.
Himeoka Y, Mitarai N. Dynamics of bacterial populations under the feast-famine cycles. Phys Rev Res. 2020;2:013372.
Schink SJ, Biselli E, Ammar C, Gerland U. Death rate of E. coli during starvation is set by maintenance cost and biomass recycling. Cell Syst. 2019;9:64–73.e3.
López D, Vlamakis H, Losick R, Kolter R. Cannibalism enhances biofilm development in Bacillus subtilis: cannibalism and biofilm matrix. Mol Microbiol. 2009;74:609–18.
Lopez D, Vlamakis H, Kolter R. Generation of multiple cell types in Bacillus subtilis. FEMS Microbiol Rev. 2009;33:152–63.
Rosenthal AZ, Qi Y, Hormoz S, Park J, Li SH-J, Elowitz MB. Metabolic interactions between dynamic bacterial subpopulations. eLife. 2018;7:e33099.
Bryant DA, Frigaard N-U. Prokaryotic photosynthesis and phototrophy illuminated. Trends Microbiol. 2006;14:488–96.
Kirchman DL, Hanson TE. Bioenergetics of photoheterotrophic bacteria in the oceans: phototrophy bioenergetic. Environ Microbiol Rep. 2013;5:188–99.
Zubkov MV. Photoheterotrophy in marine prokaryotes. J Plankton Res. 2009;31:933–8.
Koblížek M. Ecology of aerobic anoxygenic phototrophs in aquatic environments. FEMS Microbiol Rev. 2015;39:854–70.
Tang K, Jia L, Yuan B, Yang S, Li H, Meng J, et al. Aerobic anoxygenic phototrophic bacteria promote the development of biological soil crusts. Front Microbiol. 2018;9:2715.
Hamilton TL, Bennett AC, Murugapiran SK, Havig JR. Anoxygenic phototrophs span geochemical gradients and diverse morphologies in terrestrial geothermal springs. mSystems. 2019;4:e00498–19.
Zervas A, Zeng Y, Madsen AM, Hansen LH. Genomics of aerobic photoheterotrophs in wheat phyllosphere reveals divergent evolutionary patterns of photosynthetic genes in methylobacterium spp. Genome Biol Evol. 2019;11:2895–908.
Zheng Q, Zhang R, Koblížek M, Boldareva EN, Yurkov V, Yan S, et al. Diverse arrangement of photosynthetic gene clusters in aerobic anoxygenic phototrophic bacteria. PLoS One. 2011;6:e25050.
Bill N, Tomasch J, Riemer A, Müller K, Kleist S, Schmidt-Hohagen K, et al. Fixation of CO2 using the ethylmalonyl-CoA pathway in the photoheterotrophic marine bacterium Dinoroseobacter shibae: CO2 fixation in Dinoroseobacter Shibae. Environ Microbiol. 2017;19:2645–60.
Shiba T. Utilization of light energy by the strictly aerobic bacterium Erythrobacter sp. OCH114. J Gen Appl Microbiol. 1984;30:239–44.
Tang K-H, Feng X, Tang YJ, Blankenship RE. Carbohydrate Metabolism and Carbon Fixation in Roseobacter denitrificans OCh114. PLoS ONE. 2009;4:e7233.
Zarzycki J, Fuchs G. Coassimilation of Organic Substrates via the Autotrophic 3-Hydroxypropionate Bi-Cycle in Chloroflexus aurantiacus. Appl Environ Microbiol. 2011;77:6181–8.
Hauruseu D, Koblížek M. Influence of Light on Carbon Utilization in Aerobic Anoxygenic Phototrophs. Appl Environ Microbiol. 2012;78:7414–9.
Schada von Borzyskowski L, Bernhardsgrütter I, Erb TJ. Biochemical unity revisited: microbial central carbon metabolism holds new discoveries, multi-tasking pathways, and redundancies with a reason. Biol Chem. 2020;401:1429–41.
Anthony JR, Warczak KL, Donohue TJ. A transcriptional response to singlet oxygen, a toxic byproduct of photosynthesis. Proc Natl Acad Sci. 2005;102:6502–7.
Ziegelhoffer EC, Donohue TJ. Bacterial responses to photo-oxidative stress. Nat Rev Microbiol. 2009;7:856–63.
Yurkov V, Hughes E. Genes associated with the peculiar phenotypes of the aerobic anoxygenic phototrophs. Adv Bot Res. 2013:66;327–58.
Yurkov V, Hughes E. Aerobic anoxygenic phototrophs: four decades of mystery. In: Hallenbeck PC, editors.. Modern Topics in the Phototrophic Prokaryotes. Springer International Publishing, Cham:2017. pp 193–214.
Gomelsky M, Hoff WD. Light helps bacteria make important lifestyle decisions. Trends Microbiol. 2011;19:441–8.
Staley C, Ferrieri AP, Tfaily MM, Cui Y, Chu RK, Wang P, et al. Diurnal cycling of rhizosphere bacterial communities is associated with shifts in carbon metabolism. Microbiome. 2017;5:65.
Losi A, Gärtner W. A light life together: photosensing in the plant microbiota. Photochem Photobiol Sci. 2021;20:451–73.
Newman A, Picot E, Davies S, Hilton S, Carré IA, Bending GD. Circadian rhythms in the plant host influence rhythmicity of rhizosphere microbiota. BMC Biol. 2022;20:235.
Piwosz K, Kaftan D, Dean J, Šetlík J, Koblížek M. Nonlinear effect of irradiance on photoheterotrophic activity and growth of the aerobic anoxygenic phototrophic bacterium Dinoroseobacter shibae: photoheterotrophic activity of Dinoroseobacter shibae. Environ Microbiol. 2018;20:724–33.
Koblížek M, Dachev M, Bína D, Nupur, Piwosz K, Kaftan D. Utilization of light energy in phototrophic Gemmatimonadetes. J Photochem Photobiol B. 2020;213:112085.
Bryant DA. Phototrophy and phototrophs. Reference Module in Life Sciences. Elsevier; 2019. p B9780128096338207000.
Liu R, Zhang Y, Jiao N. Diel variations in frequency of dividing cells and abundance of aerobic anoxygenic phototrophic bacteria in a coral reef system of the South China Sea. Aquat Microb Ecol. 2010;58:303–10.
Fecskeová LK, Piwosz K, Hanusová M, Nedoma J, Znachor P, Koblížek M. Diel changes and diversity of pufM expression in freshwater communities of anoxygenic phototrophic bacteria. Sci Rep. 2019;9:18766.
Garcia-Chaves MC, Cottrell MT, Kirchman DL, Ruiz-González C, del Giorgio PA. Single-cell activity of freshwater aerobic anoxygenic phototrophic bacteria and their contribution to biomass production. ISME J. 2016;10:1579–88.
Piwosz K, Villena-Alemany C, Mujakić I. Photoheterotrophy by aerobic anoxygenic bacteria modulates carbon fluxes in a freshwater lake. ISME J. 2022;16:1046–54.
Tomasch J, Gohl R, Bunk B, Diez MS, Wagner-Döbler I. Transcriptional response of the photoheterotrophic marine bacterium Dinoroseobacter shibae to changing light regimes. ISME J. 2011;5:1957–68.
Fiebig A, Varesio LM, Alejandro Navarreto X, Crosson S. Regulation of the Erythrobacter litoralis DSM 8509 general stress response by visible light. Mol Microbiol. 2019;112:442–60.
Soora M, Cypionka H. Light enhances survival of dinoroseobacter shibae during long-term starvation. PLoS One. 2013;8:e83960.
Andersen RA (ed). Algal culturing techniques. Elsevier/Academic Press, Burlington, Mass;2005.
Larsen RA, Wilson MM, Guss AM, Metcalf WW. Genetic analysis of pigment biosynthesis in Xanthobacter autotrophicus Py2 using a new, highly efficient transposon mutagenesis system that is functional in a wide variety of bacteria. Arch Microbiol. 2002;178:193–201.
Kolmogorov M, Bickhart DM, Behsaz B, Gurevich A, Rayko M, Shin SB, et al. metaFlye: scalable long-read metagenome assembly using repeat graphs. Nat Methods. 2020;17:1103–10.
Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–i890.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–9.
Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30:923–30.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Fischer DS ImpulseDE2: Differential expression analysis of longitudinal count data sets. (2019).
Gao C-H ggVennDiagram: A ‘ggplot2’ Implement of Venn Diagram. (2022).
Wickham H ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag New York (2016).
Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–7.
Nguyen L-T, Schmidt HA, von Haeseler A, Minh BQ. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol. 2015;32:268–74.
R Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria (2021).
Ellis B, Haaland P, Hahne F, Meur NL, Gopalakrishnan N, Spidlen J, et al. flowCore: flowCore: Basic structures for flow cytometry data. (2021).
Cornet L, Bertrand AR, Hanikenne M, Javaux EJ, Wilmotte A, Baurain D. Metagenomic assembly of new (sub)polar cyanobacteria and their associated microbiome from non-axenic cultures. Microb Genom. 2018;4:e000212.
Xu L, Sun C, Fang C, Oren A, Xu X-W. Genomic-based taxonomic classification of the family Erythrobacteraceae. Int J Syst Evol Microbiol. 2020;70:4470–95.
Achenbach LA, Carey J, Madigan MT. Photosynthetic and phylogenetic primers for detection of anoxygenic phototrophs in natural environments. Appl Environ Microbiol. 2001;67:2922–6.
Béjà O, Suzuki MT, Heidelberg JF, Nelson WC, Preston CM, Hamada T, et al. Unsuspected diversity among marine aerobic anoxygenic phototrophs. Nature. 2002;415:630–3.
Hughes R-A, Jin X, Zhang Y, Zhang R, Tran S, Williams PG, et al. Genome sequence, metabolic properties and cyanobacterial attachment of Porphyrobacter sp. HT-58-2 isolated from a filamentous cyanobacterium–microbial consortium. Microbiology. 2018;164:1229–39.
Kim M, Shin B, Lee J, Park HY, Park W. Culture-independent and culture-dependent analyses of the bacterial community in the phycosphere of cyanobloom-forming Microcystis aeruginosa. Sci Rep. 2019;9:20416.
Mironov KS, Leusenko PA, Ustinova VV, Bolatkhan K, Zayadan BK, Kupriyanova EV, et al. Draft genome sequences of a putative prokaryotic consortium (IPPAS B-1204) consisting of a cyanobacterium (Leptolyngbya sp.) and an alphaproteobacterium (Porphyrobacter sp.). Microbiol Resour Announc. 2019;8:e01637–18.
Hughes R-A, Zhang Y, Zhang R, Williams PG, Lindsey JS, Miller ES. Genome sequence and composition of a tolyporphin-producing cyanobacterium-microbial community. Appl Environ Microbiol. 2017;83:e01068–17.
Masuda S. Light detection and signal transduction in the BLUF photoreceptors. Plant Cell Physiol. 2013;54:171–9.
Zarzycki J, Brecht V, Müller M, Fuchs G. Identifying the missing steps of the autotrophic 3-hydroxypropionate CO 2 fixation cycle in Chloroflexus aurantiacus. Proc Natl Acad Sci. 2009;106:21317–22.
Kovács ÁT, Rákhely G, Kovács KL. The PpsR regulator family. Res Microbiol. 2005;156:619–25.
Ebert M, Laaß S, Thürmer A, Roselius L, Eckweiler D, Daniel R, et al. FnrL and three Dnr regulators are used for the metabolic adaptation to low oxygen tension in dinoroseobacter shibae. Front Microbiol. 2017;8:642.
Castro WDO, Lima ARJ, Moraes PHG, Siqueira AS, Aguiar DCF, Baraúna ARF, et al. Complete genome sequence of Porphyrobacter sp. Strain CACIAM 03H1, a proteobacterium obtained from a nonaxenic culture of microcystis aeruginosa. Genome Announc. 2017;5:e01069–17.
Li Q, Lin F, Yang C, Wang J, Lin Y, Shen M, et al. A large-scale comparative metagenomic study reveals the functional interactions in six bloom-forming microcystis-epibiont communities. Front Microbiol. 2018;9:746.
Biller SJ, Coe A, Chisholm SW. Torn apart and reunited: impact of a heterotroph on the transcriptome of Prochlorococcus. ISME J. 2016;10:2831–43.
Kopejtka K, Tomasch J, Kaftan D, Gardiner AT, Bína D, Gardian Z, et al. A bacterium from a mountain lake harvests light using both proton-pumping xanthorhodopsins and bacteriochlorophyll-based photosystems. Proc Natl Acad Sci. 2022;119:e2211018119.
Maisch T, Baier J, Franz B, Maier M, Landthaler M, Szeimies R-M, et al. The role of singlet oxygen and oxygen concentration in photodynamic inactivation of bacteria. Proc Natl Acad Sci. 2007;104:7223–8.
Wainwright M, Maisch T, Nonell S, Plaetzer K, Almeida A, Tegos GP, et al. Photoantimicrobials—are we afraid of the light? Lancet Infect Dis. 2017;17:e49–e55.
Acknowledgements
We thank Annick Wilmotte for providing the ULC335 community. We express our gratitude to Pilar Junier, Martina Valentini, and Despoina A. Mavridou for their constructive comments on earlier versions of this manuscript. This work was mainly funded by a Swiss National Foundation Ambizione grant to DG (PZ00P3_180142). The contribution of CTi was supported by a grant from the Fondation Mercier pour la Science to DG.
Author information
Authors and Affiliations
Contributions
CTi.: Investigation (supporting), Visualization (supporting), Formal analysis (supporting), Review and editing (equal). MP: Investigation (supporting), Review and editing (equal). CTe.: Investigation (supporting), Review and editing (equal). DG: Investigation (lead), Conceptualization (lead), Visualization (lead), Data curation (lead), Formal analysis (lead), Project administration (lead), Software (lead), Writing (lead), Review and editing (equal).
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tinguely, C., Paulméry, M., Terrettaz, C. et al. Diurnal cycles drive rhythmic physiology and promote survival in facultative phototrophic bacteria. ISME COMMUN. 3, 125 (2023). https://doi.org/10.1038/s43705-023-00334-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43705-023-00334-5
- Springer Nature Limited