Abstract
Background
Combination therapy can offer greater efficacy on medical treatments. However, the discovery of synergistic drug combinations is challenging. We propose a novel computational method, SyndrumNET, to predict synergistic drug combinations by network propagation with trans-omics analyses.
Methods
The prediction is based on the topological relationship, network-based proximity, and transcriptional correlation between diseases and drugs. SyndrumNET was applied to analyzing six diseases including asthma, diabetes, hypertension, colorectal cancer, acute myeloid leukemia (AML), and chronic myeloid leukemia (CML).
Results
Here we show that SyndrumNET outperforms the previous methods in terms of high accuracy. We perform in vitro cell survival assays to validate our prediction for CML. Of the top 17 predicted drug pairs, 14 drug pairs successfully exhibits synergistic anticancer effects. Our mode-of-action analysis also reveals that the drug synergy of the top predicted combination of capsaicin and mitoxantrone is due to the complementary regulation of 12 pathways, including the Rap1 signaling pathway.
Conclusions
The proposed method is expected to be useful for discovering synergistic drug combinations for various complex diseases.
Plain Language Summary
Adding drug treatments together can sometimes produce better results for patients. We introduced a new computer-based method called SyndrumNET, designed to identify effective drug combinations for treating diseases. The method uses data about how diseases and drugs interact at a molecular level to predict which drugs work well together. Tested on six different diseases, such as asthma and different types of cancer, SyndrumNET proved to be more accurate than previous approaches. For example, most of the drug combinations predicted by SyndrumNET to rank highly have shown better combination effects on leukemia cells. This method also helped understand why certain drug combinations work better by analyzing their effects on cellular pathways. The findings suggest that SyndrumNET could be a valuable tool in developing more effective treatment for various complex diseases.
Similar content being viewed by others
Introduction
Combination therapy, which is a treatment modality that combines two or more drugs, can offer greater efficacy or lower individual drug dosages compared with monotherapy1. Its effectiveness has been recognized for various complex diseases, such as cancers, hypertension, cardiovascular, neurological, and autoimmune disorders2,3. The number of drug combinations approved by the US Food and Drug Administration (FDA) has continuously increased since the first approval of co-administered drugs in the 1940s4; however, determining synergistic drug combinations is very challenging, particularly in heterogenous diseases. There are more than 13,000 drugs approved for human use by FDA5; thus, the number of possible drug pairs is approximately 85 million. Conducting clinical trials for all possible drug combinations is impractical. Thus, there is a strong need for methods to facilitate the identification of synergistic drug combinations for various diseases.
A variety of computational methods have been developed for predicting synergistic drug combinations6. A popular approach is to use supervised learning. For example, pharmacological features (e.g., target proteins, efficacy classes) enriched in approved drug combinations were extracted and new drug combinations associated with the pharmacological features were searched7. A sparsity-induced classifier with tensor-based representations of pharmacological features was proposed8. A machine learning model using an ensemble of weak predictive models was applied for the dataset in the AstraZeneca-Sanger Drug Combination Prediction DREAM Challenge9. A deep learning-based method with structural features of compounds was proposed for COVID-19 and the importance of the structural characteristics of the drugs was determined10. However, these supervised learning methods require the prior information on known synergistic drug combinations as a learning dataset to construct predictive models and their performance depends heavily on the quality and quantity of the learning dataset. The number of diseases for which sufficient information on synergistic drug combinations is very limited. For most diseases, synergistic drug combinations remain unknown.
Unsupervised learning is a more practical and realistic approach for predicting synergistic drug combinations because they can be applied to any disease without prior knowledge of synergistic drug combinations11,12,13. For example, a transcriptome-based approach was proposed to predict synergistic drug combinations for glioblastoma (GBM), where the drug was assumed to restore the disease-specific gene expression pattern. For a fixed drug, other drug partners were searched using the inverse correlation between the disease-specific transcriptional expression signatures and drug response gene expression signatures11; however, a general framework for any drug pair would be more practical. In addition, the gene expression signatures of diseases and its approved drugs are not always inversely correlated14. A network-based approach was proposed to predict synergistic drug combinations for hypertension and cancers based on the relationship between drug target genes and disease genes in the comprehensive molecular interaction network13. However, this approach is only applicable to drugs with known targets and it does not take into account the dynamic changes in the cells or organisms associated with drug treatment15,16. The therapeutic effect is not only determined by the network-based relationships between diseases and drugs. These previous unsupervised approaches are based on single omics data representing a few biological aspects. Diseases result from the disruption of many biological processes; thus, the integrative use of multi-omics data should contribute to the enhancement of the prediction accuracy of synergistic drug combination.
In this study, we propose a novel computational method, which we call SyndrumNET, to predict synergistic drug combinations by network propagation with trans-omics analyses. The prediction is based on multi-omics data such as genome, transcriptome, interactome, and diseasome data. We demonstrated the usefulness of the proposed method on the prediction of synergistic drug combinations for six diseases: acute myeloid leukemia (AML), chronic myeloid leukemia (CML), colorectal cancer, asthma, type II diabetes, and hypertension. We validated the prediction result for CML through in vitro experiments and identified the underlying mode-of-action of the synergistic effects of the drug combination at the pathway level by microarray analysis.
Methods
Construction of the human molecular interaction network
Human molecular interactions were constructed from seven databases (Supplementary Data S1): (i) Yeast-two-hybrid high-throughput datasets were retrieved from the yeast two-hybrid database (HuRI)17 (accessed on March 2, 2020), (ii) Protein complexes were retrieved from the CORUM database18 (accessed on September 3, 2018), (iii) Kinase–substrate pairs were retrieved from the PhosphositePlus database19 (accessed on September 7, 2018), (iv) Metabolic enzyme-coupled interactions were retrieved from the KEGG Rpair database20 (accessed on March 12, 2016), (v) Signaling interactions were retrieved from the Signalink v.2.0 database21 (accessed on December 3, 2018), (vi) Innate immune response interactions were retrieved from the InnateDB database22 (accessed on June 2, 2018), (vii) 3D structurally resolved protein-protein interactions were retrieved from the Instruct database23 (accessed on March 3, 2020). We used molecular interactions with biological annotations. We did not include interactions extracted from gene expression data or evolutionary considerations. All interactions from these databases were combined, and the union yielded a network of 13,524 proteins and 311,888 interactions (Supplementary Data S1). Duplicated interactions were excluded using the simplify function in the igraph library (1.2.6) of R. The giant component was used as a human molecular interaction network, and it consisted of 235,123 interactions involving 13,377 proteins (Supplementary Data S2).
This newly established human molecule interaction network offers two advantages. First, it facilitates easy comparison with prior work on network-based drug combination prediction methods. Certain steps in our proposed method align with the findings made by Cheng et al.13. To enhance comparability in predictive performance, we adhered to the network creation procedure outlined in the previous work. Second, the network allows for the selection of experimentally validated interaction types. In this study, our focus was solely on molecular interactions with biological annotations (e.g., physical interactions, phosphorylation and substrate–enzyme associations). This emphasis has the potential to enhance the reliability of the network.
Construction of disease-specific gene expression profiles
The CREEDS database provides gene expression signatures for 79 diseases with 14,804 genes24, derived from transcriptome data registered in Gene Expression Omnibus (GEO)25. We retrieved disease-specific gene expression signatures of AML, CML, colorectal cancer, asthma, and type 2 diabetes from CREEDS24 (accessed on April 22, 2020).
CREEDS lacked the gene expression signature for hypertension. Thus, we constructed the gene expression signature for hypertension according to the procedure in CREEDS24. The detail is written in Supplementary Note 1 (Construction of disease-specific gene expression profiles). Gene expression data for hypertension were retrieved from GEO (GSE24752, and GSE75360 (accessed on November 9, 2022). The disease-specific gene expression levels were determined relative to a healthy cohort. Finally, we obtained the disease-specific gene expression profiles with the same gene set in CREEDS (14,804 genes) for hypertension.
Identification of disease-specific genes from disease-specific gene expression profiles
We identified disease-specific genes for 79 diseases registered in CREEDS. Genes in CREEDS have nonzero scores indicating disease specificity. Therefore, disease-specific genes were defined as those with expression levels (not zeros) for each disease. For hypertension, we selected the top 5% of genes with positive fold change values and the top 5% of genes with negative fold change values compared to healthy control as disease-specific genes. Disease-specific genes were used to calculate network-based distance between diseases.
Construction of disease modules using disease susceptibility genes
A set of disease susceptibility genes on the human molecular interaction network were referred to as “disease modules”13. To investigate the susceptibility genes of our target diseases (AML, CML, colorectal cancer, asthma, type 2 diabetes, and hypertension), we sourced relevant genes from six databases as delineated in a prior study13, including (i) the Online Mendelian Inheritance in Man (OMIM) database26, (ii) ClinVar database27, (iii) the genome-wide association studies (GWAS) database28, (iv) the phenome-wide association study database (PheWAS)29, (v) the GWASdb database30, and (vi) the DisGeNET database31. The accessed date and specific search keys are detailed in Supplementary Data S1. The gene symbols (HGNC symbols) were converted into Entrez IDs using the biomaRt library32 in R (version 4.0.3). The number of the genes in the disease module of AML, CML, colorectal cancer, asthma, type 2 diabetes, and hypertension are 1075, 51, 408, 929, 1173, and 909, respectively. A comprehensive list of genes in the modules can be found in Supplementary Data S3.
Construction of drug response gene expression profiles
Drug-induced gene expression profiles were obtained from the LINCS Program L1000 mRNA profiling assay (http://www.lincsproject.org), where the gene expression levels for 978 landmark genes, termed L1000 genes. L1000 is highly reproducible, comparable to RNA sequencing, and suitable for computational inference of the expression levels of 81% of non-measured transcripts33. The gene expression profiles of L1000 were measured at various post-treatment intervals—3, 6, 24, 48, and 144 h—and across a range of concentrations using diverse human cell lines. Within the level 5 dataset, we extracted the gene expression profiles of 1488 drugs, and averaged the gene expression profiles of the same drug across experimental conditions, such as post-treatment intervals, the concentration of the drug, and cell lines. The details on drug names, efficacies, and procedures are provided in Supplementary Data S4 and Supplementary Note 1 (Construction of drug response gene expression profiles).
Construction of drug modules using drug response genes
The top 5% of genes with positive fold change values and the top 5% of genes with negative fold change values in the drug-induced gene expression profiles were considered drug response genes and were used to construct the drug module for each drug (Supplementary Data S4).
Evaluation of network-based distance between disease modules and drug modules
The network-based proximity between a query disease module and a drug module was evaluated using a network-based distance measure. The path length between genes constituting a disease module and drug response genes constituting a drug module was calculated. Q represents the set of genes \({q}_{1},\cdots ,{q}_{\left|Q\right|}\) in the query disease module, A represents the set of genes \({a}_{1},\cdots ,{a}_{\left|A\right|}\) in the drug A module, and \(d(q,a)\) represents the shortest path length between nodes q and a in the human molecular interaction network15 and was defined as:
To determine the significance of the network-based proximity measure between query disease Q and drug A, a reference distance distribution was created. First, a set of genes with the same size and degree of the query disease module was randomly selected. Second, a set of genes with the same size and degree of the drug module was randomly selected. Then, the proximity between the two sets of genes in the human molecular interaction network was calculated15. After 100 repetitions, the mean \({\mu }_{d(Q,A)}\) and the standard deviation \({\sigma }_{d(Q,A)}\) were calculated. The normalized network-based proximity measure was defined as
For a more efficient calculation of the permutation process, parallel computing using the PAR function was performed34. Finally, the \(z\left(Q,A\right)\) sign was inverted, scaled in the range of 0 to 1, and defined as PQA. The code for calculation of network-based distance between disease modules and drug modules is deposited in figshare35.
Evaluation of the network-based distance between diseases
The network-based proximity between query disease Q and disease α was evaluated using a network-based distance measure. The shortest path lengths between susceptibility genes and disease-specific genes were calculated according to formula (1). The normalized network-based proximity measure z(Q, α) was calculated according to formula (2). Although Sections 7 and 8 share similarities, they differ in focus. Section 7 elucidates the calculation of distance between a disease and a drug, while section 8 delves into the calculation of distance between different diseases. The code for calculation of the network-based distance between diseases is deposited in figshare35.
Evaluation of network-based distance between drug modules
To evaluate the network-based distance between drug A module and drug B module, the network-based separation measure of the drug response genes between drug A module and drug B module was calculated13. Briefly, the network-based separation measure 〈sAB〉 was defined as follows:
〈dAA〉 is the mean shortest distance between the response genes of drug A in a human molecular interaction network, 〈dBB〉 is the mean shortest distance between the response genes of drug B in a human molecular interaction network, and 〈dAB〉 is the mean shortest distance between the response genes of drug A and drug B.
If drug A had only a single response gene, the average shortest distance between the response genes of drug A (denoted as 〈dAA〉) would be 0. The computation of the distance between two drug modules was based on the response genes of drugs A and drug B. When both drugs A and drug B possessed only one response gene, and if that gene was identical, the distance between the drug modules (denoted as 〈sAB〉) would be 0. In this study, drug response genes were identified as the top 5% of genes exhibiting the positive or negative expression changes in the drug-induced gene expression profiles. Since the number of response genes for drug A (or drug B) ranged from 198 to 200, the associated distance was not 0.
The formulas for calculating the drug–disease distance (S-score using formula [1]) and the network-based separation measure of the drug response genes between drug A module and drug B share similarities, but they exhibit a distinction. Therefore, we used different notations. The determination of the distance between a drug module and a disease module relies on generating a random distribution of S-scores. In contrast, the distance between drugs is determined by the topological distance between one drug and another on the molecular network. This choice is motivated by the substantial number of drug pairs, where the computational expense of generating a random distribution is notably high. The code for calculation of network-based distance between drug modules is deposited in figshare35.
Evaluation of the network-based disease similarity based on disease-specific gene expression profiles
The similarity between query disease module Q and the other disease a was evaluated using the network-based proximity measure R(Q, a)36. The normalized network-based proximity measure between query disease module Q and the other disease a was evaluated as z(Q, a). The network-based similarity was calculated by sign inversion and scaling of the proximity z(Q, a) in the range of 0 to 1. The similarity of 79 diseases in the CREEDS database and six diseases (AML, CML, colorectal cancer, asthma, type II diabetes, and hypertension) was calculated.
Evaluation of drug-similarity based on the chemical structures
The structure-based similarity between drug A and drug B was evaluated as R(A,B) based on the chemical structures. Chemical structures (MOLfiles) for 8,287 drugs were retrieved from the KEGG DRUG37. KCF-S fingerprints38 for each drug were computed using kcfconvoy (https://github.com/KCF-Convoy/kcfconvoy). The generalized Jaccard similarity coefficient between drugs was calculated based on the fingerprints39.
Evaluation of transcriptional correlations between a disease module and drug modules
The transcriptional correlation of the gene expression profiles between the query disease Q module and the drug A module was evaluated by the cosine correlation coefficient represented as CQA. Similarly, the transcriptional correlation between the query disease Q module and the drug B module was evaluated as CQB. The cosine coefficient was calculated using the cosine function in the lsa library (version 0.73.2) in R. The code for calculation of transcriptional correlations between a disease module and drug modules is deposited in figshare35.
Amplification of component genes in the disease and drug modules by network propagation
The size of the query disease module and drug modules was increased by network propagation. A network propagation method with prior knowledge, called PRINCE, was leveraged40. The algorithm is implemented in https://github.com/kztakemoto/network_propagation.
The network propagation with prior knowledge calculates the probability of genes belonging to the query disease module or the drug module. Neighbor nodes of a query disease module were identified as candidates for new genes of the query disease module. The network-based similarity between diseases was used as prior knowledge in the network propagation procedure. Neighbor nodes for a drug module were identified as candidates for new genes of the drug module. The structure-based similarity between drugs was used as prior knowledge for the network propagation procedure.
Network propagation without prior knowledge was also performed as follows: Neighbor nodes of a query disease module were identified as candidates for new genes of the query disease module. Similarly, neighbor nodes of a drug module were identified as candidates for new genes of the drug module. The neighbors function in the igraph library (v.1.2.6) was used to identify the neighbor nodes41. The parameters of network propagation are summarized in Supplementary Supplementary Data S5.
Design of the prediction score for synergistic drug combinations
The prediction score of the synergistic effect of drug A and drug B for a query disease was designed using three components (Fig. 1). The first component was the network-based localization relationship score between a query disease module, drug A module, and drug B module, which is referred to as (TQAB). The second component was the network-based proximity score between a query disease module, drug A module, and drug B module, which is referred to as (PQAB). The third component was the transcriptional correlation coefficient between a query disease module, drug A module, and drug B module (CQAB).
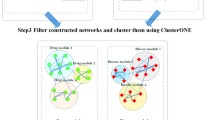
A comprehensive human molecular interaction network was constructed. Susceptibility genes and drug response genes were mapped onto the network. The network-based relationships between a query disease module and drug modules were calculated. The network-based proximities between a query disease module and drug modules were calculated. For the method without network propagation (Syndrum), the transcriptional correlations between a query disease module, and drug modules were calculated using the overlapping genes between a query disease module and the drug modules. The network-based similarities of diseases are calculated based on network-based proximity between diseases, referred to as disease similarity. The structural similarity of drugs was calculated based on the chemical structure, referred to as drug similarity. Network propagation using the similarities was performed to identify overlap genes between a query disease module and disease modules. For the method with network propagation (SyndrumNET), the transcriptional correlations between a query disease module, and drug modules were calculated using the newly identified overlapping genes. Finally, the network-based localization relationships score, the network-based proximities, and the transcriptional correlation coefficients between a query disease module and the drug modules were integrated. Q indicates ‘Query disease’, A indicates ‘Drug A’, and B indicates ‘Drug B’.
Network-based localization relationship score (TQAB) was determined based on the topological classes of the query disease module, drug A module, and drug B module. Six types of topological classes (Class I ~ Class VI) were defined based on a previous study13. Within the six classes, class II is designated as Complementary Exposure and it tends to have synergistic effects. This class represents situations where two separated drug modules that overlap individually with a query disease module (\(\normalsize z\left(Q,A\right) \, < \, 0, z \left(Q,B\right) \, < \, 0 \, {and} \; {s}_{{AB}} \, < \, 0\)). Based on the topological classes, the score of the network-based localization relationship was assigned as follows:
The network-based proximity score (PQAB) was calculated by averaging the network-based proximity between the query disease Q module and drug A module (PQA) and the network-based proximity between the query disease Q module and drug B module (PQB), as follows:
The transcriptional correlation score (CQAB) was calculated by averaging the absolute value of the transcriptional correlation coefficient between the query disease Q module and drug A module (CQA) and the transcriptional correlation coefficient between the query disease Q module and drug B module (CQB), as follows:
Finally, the prediction score was calculated by adding the network-based localization relationship score (TQAB), network-based proximity score (PQAB), and transcriptional correlation score (CQAB) as follows35:
Collection of drug combinations with known synergistic effects for various diseases
The known synergistic drug pairs for AML, CML, colorectal cancer, asthma type 2 diabetes, and hypertension were obtained from the PubMed database. The keywords “synergy,” “synergic,” “synergistic,” “synergism,” “interaction,” and “combination” along with disease names were used as keywords for the search procedure. For AML and CML, known synergistic drug pairs were retrieved from DrugCombDB (2019.05.31 release version)42. DrugCombDB contains drug combinations for human cancer cell lines. We linked the drugs and diseases according to the cell line. For hypertension, known synergistic drug pairs were retrieved from a previous paper13. The curated known synergistic drug pairs are summarized in Supplementary Data S6.
Performance evaluation protocol
The area under the receiver operating characteristic curve (AUC) was calculated using the performance function in the ROCR library (v.1.0-11) in R. The code for the performance evaluation is deposited in figshare35.
Chemicals used for the cell survival assay
Capsaicin was purchased from FUJIFILM Wako Pure Chemical Industries, Ltd. (Osaka, Japan). Daunorubicin hydrochloride, idarubicin, and topotecan were purchased from Cayman Chemicals (Ann Arbor, Michigan, USA). Mitoxantrone, fasudil were purchased from Tokyo Chemical Industries Co., Ltd. (Tokyo, Japan). Cell Counting Kit-8 used for the WST-8 assay was purchased from DOJINDO (Tokyo, Japan).
Cell culture and reagents
The K562 human CML cell line was obtained from the RIKEN BioResource Center (Tokyo, Japan) and grown in RPMI 1640 (NACALAI TESQUE, INC., JAPAN) medium supplemented with 10% fetal bovine serum (Funakoshi Co., Ltd., JAPAN). All cells were incubated at 37 °C in a humidified atmosphere containing 5% (v/v) CO2.
Cell survival assay
In vitro growth inhibition was evaluated according to the manufacturer’s standard protocol using the Cell Counting Kit-8. Cells were seeded in 96-well plates at a density of 5000 cells/well in a total volume of 100 µL and exposed to various drugs for 72 h at 37 °C in a 5% CO2 atmosphere. WST-8 was added, and after 3 h, the absorbance was measured at a wavelength of 450 nm (reference 630 nm) using a microplate reader (Bio-Rad Laboratories, Inc., Hercules, CA). The results are expressed as percentages [i.e., as the ratio of the absorbance of treated cells to that of the control (drug untreated group, 100%)]. Percent survival was calculated using the following formula: percentage survival = (absorbance of treated wells − absorbance of blank wells)/(absorbance of untreated wells − absorbance of blank wells) × 100. The number of biological replicates is three.
Statistical evaluations of the significance of combinatory effects for drug synergy
Two essential models were used to evaluate the significance of combinatory effects for drug synergism: Bliss’s IA43 model and Loewer’s additivity (CA) model44.
For Bliss’s IA model, it is assumed that the effects of drugs are stochastic events under the non-interaction assumption between drugs. The effects of a drug combination are calculated as the joint probability of each effect. Drug A causes v% effects and drug B causes w% effects at a given combination of concentrations. The total effect rate of the combination can be computed as \({{CI}}_{{mix}}=1-\left(1-v\right)\left(1-w\right)\) under the additive assumption. Thus, if \({{CI}}_{{mix}} \, > \, 1\), a given drug combination is considered to have synergistic effect based on Bliss’s IA model. \({{CI}}_{{mix}}\) is denoted as the IA score in this study.
For Loewer’s additivity model, it is assumed that the toxic unit equals one under the non-interaction assumption between drugs. The concentrations weighted by the effect of each drug (toxic unit [TU]) are added together to yield the TU of a given drug combination.
For a given drug combination, Ca and Cb stand for the concentrations of drug A and drug B, respectively. \({{EC}}_{u}^{A}\) and \({{EC}}_{u}^{B}\) are the concentrations of the drugs causing u% effect by drug A and drug B, respectively. If TU = 1, the effect rate of the drug combination remains at u% under the additive assumption. Thus, if TU < 1, a given drug combination is assumed to have synergistic effect based on Loewer’s additivity (CA) model, when u% is selected as the results of the growth inhibition assay. \({{\mbox{TU}}}\) is denoted as the CA score in this study.
Sample preparation for microarray analysis
Cells were seeded at a density of 50,000 cells/mL in a 10-cm diameter dish and exposed to various drugs (capsaicin 50 μM, mitoxantrone 30 nM) for 24 h at 37 °C in a 5% (v/v) CO2 atmosphere. Experiments were conducted in three independent wells for each group. Total RNA was extracted using the RNeasy Mini Kit (QIAGEN, Valencia, CA) according to the manufacturer’s protocol and used for microarray experiments by the well. The extracted total RNA from these wells was combined by exposure condition.
Mode-of-action of drug combinations by microarray analysis
Cyanine-3 (Cy3) labeled cRNA was prepared from 150 ng RNA using the One-Color Low Input Quick Amp Labeling kit (Agilent) according to the manufacturer’s instructions, followed by RNAeasy column purification (QIAGEN, Valencia, CA). Dye incorporation and cRNA yield were assessed using a NanoDrop ND-1000 Spectrophotometer.
Cy3-labeled cRNA (600 ng, specific activity >6 pmol Cy3/µg cRNA) was fragmented at 60 °C for 30 min in a reaction volume of 25 µl containing 25× Agilent fragmentation buffer and 10× Agilent blocking agent following the manufacturer’s instructions. Upon completion of the fragmentation reaction, 25 µl of 2× Agilent hybridization buffer was added to the fragmentation mixture and hybridized to Agilent SurePrint GE Unrestricted Microarrays (G2519F) for 17 h at 65 °C in a rotating Agilent hybridization oven. After hybridization, microarrays were washed for 1 min at room temperature with GE Wash Buffer 1 (Agilent) and 1 min at 37 °C with GE Wash Buffer 2 (Agilent), then dried immediately.
The slides were scanned immediately after washing using an Agilent DNA Microarray Scanner (G2505C) with a one-color scan setting for 8 × 60 K array slides (Scan Area 61 × 21.6 mm, Scan resolution 3 µm, Dye channel was set to Green and Green PMT was set to 100%).
The scanned images were analyzed with Feature Extraction Software 10.7.1.1 (Agilent) using default parameters (protocol GE1_107_Sep09 and Grid: 028282_D_F_20110531) to obtain background subtracted and spatially detrended processed signal intensities. Features flagged in Feature Extraction as Feature Non-uniform outliers were excluded. Our microarray data have been registered and are publicly available in the Gene Expression Omnibus (GEO) database (GSE254052). We calculated the log2 fold change (log2FC) for each probe when comparing the control and exposed groups. Then, we averaged the log2FC for genes. We defined differential expression genes (DEGs) as genes with absolute value of log2FC greater than one (\(\left|{\log }_{2}{FC}\right|\ge \,1\)) in the exposed group compared to the control group.
Enrichment analysis of transcription factors
Gene set enrichment analysis was performed using DAVID45 and “clusterProfiler” (v4.4.4) library in R46. The transcription factor enrichment analysis was performed using ChEA47. The results were considered statistically significant at p < 0.05.
Statistics and reproducibility
We performed two-sided t-test using “stats” (v4.3.1) library in R and Fisher’s exact test using DAVID45 and “clusterProfiler” (v4.4.4) library in R46. We examined whether the mean distance from a disease to a drug with a known effect differs from that to a drug with an unknown effect by two-sided t-test. For t-test, the sample size is 1,106,328 drug pairs for each query disease. For Fisher’s exact test using DAVID45 and “clusterProfiler”, the sample size is the number of genes of functional pathways in the KEGG database, totaling 8,156 and 8,772 genes, respectively. We conducted cell survivable assay following microarray analysis. For cell survivable assay, we preformed three biological replicates. The mRNA from these replicates were mixed and used for microarray analysis.
Inclusion & ethics statement
All members involved in this study have met the authorship criteria mandated by Nature Portfolio journals and have been listed as authors. Their contributions were vital to the study’s design and execution. The roles and responsibilities of each collaborator were clearly defined and mutually agreed upon before initiating the research. This research faced no severe restrictions or prohibitions within our operational environment and was conducted in a manner that avoids causing stigmatization, incrimination, discrimination, or personal risk to any parties involved. In preparing our manuscript, we diligently referenced research that aligns with our study, ensuring that our citations reflect the pertinent scientific context and contributions.
Results
Overview of the proposed trans-omics methods
We propose “SyndrumNET”, a network-based trans-omics approach, to predict synergistic drug combinations by integrating genome, transcriptome, interactome, and diseasome. An overview of the proposed method is shown in Fig. 1, and the detailed procedures are described in the Methods section. Disease susceptibility genes and drug target genes are not randomly dispersed throughout the human molecular interaction network. Instead, they form localized clusters, termed either disease modules or drug modules13,48. If two drug modules are close to a disease module but distant from each other, these two drugs tend to have synergistic effects on the disease13.
In the first step, we aimed to understand the relationships between a disease and drugs based on their localization in the human molecular interaction network. We constructed a comprehensive human molecular interaction network by integrating various types of molecular interactions (e.g., physiological protein-protein interactions) from multiple databases (Fig. 1, Supplementary Data S1, Supplementary Data S2 and see Methods). Then, we identified disease modules and drug modules using disease susceptibility genes and drug response genes (Supplementary Data S3, Supplementary Data S4 and see Methods). We measured the network-based proximity between a query disease module and drug modules and network-based separation between drug modules. We evaluated the relationships between a disease and drugs based on their localization in the network, which we term as the network-based localization relationships between a query disease module and the drug modules.
In the second step, we aimed to understand the relationships between a disease and drugs based on their proximity in the human molecular interaction network. The network-based drug-disease proximity can quantify the influence of the drug on the disease15. We averaged the network-based proximity between a query disease and each of the two drug modules (Fig. 1 and Methods).
In the third step, we evaluated the transcriptional correlations between a query disease module and the drug modules. The gene expression profiles for the diseases were constructed from Crowd Extracted Expression of Differential Signatures (CREEDS)24 and the gene expression profiles for the drugs were constructed from the Library of Integrated Network-Based Cellular Signatures (LINCS)49. There is a limitation to the number of genes that overlap between a query disease module and the drug modules. To overcome this problem, we amplified the number of overlapping genes between a query disease module and the drug modules by network propagation with the similarities of diseases and drugs (Fig. 1 and Methods).
Finally, we developed a scoring scheme by integrating the network-based proximity and the transcriptional correlations between a query disease module and the drug modules (see Methods). The proposed method without network propagation is referred to as Syndrum, while the proposed method with network propagation is referred to as SyndrumNET. We applied these methods to predicting synergistic drug combinations for six diseases including AML, CML, colorectal cancer, asthma, type 2 diabetes, and hypertension.
Synergistic drug combinations can be explained by the topological relationship between drug modules and disease modules in the human molecular interaction network
We examined the relationships between disease modules and drug modules for six diseases and 1488 drugs, including AML, CML, colorectal cancer, asthma, type 2 diabetes, and hypertension. We calculated the network-based proximity between a query disease module and its approved drug modules using the shortest path length (see Methods). Then, we compared the results between approved drugs and the other drugs with respect to the network-based proximity for a query disease. For cancers and asthma, a query disease and its approved drugs are closely located compared with the other drugs in terms of network-based proximity (Fig. S1). The results suggest that drug response genes tend to be close to susceptibility genes for cancers and asthma if the drugs are effective for the treatment of the disease.
Next, we examined the network-based proximity relationships between 1488 drug modules and six disease modules with respect to known drug synergy. For a query disease, we compared the averaged network-based proximity between drug pairs with known synergistic effects and the other drug pairs. This tends to be shorter compared with that of randomly selected drug module pairs (Fig. 2a). The results suggest that synergistic drug combinations can be explained by the distance between a disease module of interest and drug modules in the human molecular interaction network.
a Distribution of average network-based proximity (PQAB) between a query disease module (Q) and drug modules (A and B). Gray color indicates the network-based proximity between a query disease module and drug modules with unknown effects (AML: 1,106,100 pairs, CML: 1,106,103 pairs, colorectal cancer: 1,105,598 pairs, asthma: 1,106,327 pairs, type 2 Diabetes: 1,106,324 pairs, hypertension: 1,106,318 pairs). The orange color indicates the network-based proximity between a query disease module and drug modules with known synergistic effects (AML: 228 pairs, CML: 225 pairs, colorectal cancer: 730 pairs, asthma: 1 pair, type 2 Diabetes: 4 pairs, hypertension: 10 pairs). Upper and lower whiskers mean standard deviations, and points indicate mean. b Distribution of the number of overlapped genes between a query disease module (Q) and individual drug modules (A or B). Light green represents the number of overlapped genes in the method without network propagation (Syndrum). Green represents the number of overlapped genes in the method with network propagation (SyndrumNET). c Distribution of average absolute transcriptional correlation coefficients between a query disease module (Q) and drug modules (A and B). Gray color indicates the average absolute transcriptional correlation coefficients between a query disease module and drug modules with unknown synergistic effects. Green color indicates the average absolute transcriptional correlation coefficients between a query disease module and drug modules with known synergistic effects. The number of pairs of each group is consistent with Fig. 2a. Upper and lower whiskers mean standard deviations, and points indicate mean. The p-values were calculated using two-sided t-test.
Network propagation with disease similarities and drug similarities emphasizes the transcriptional correlation between a query disease module and drug modules
We examined the transcriptional correlations between each query disease module and individual drug modules using the overlapped genes in the gene expression profiles. We identified overlapping genes between 1,488 drug modules and each of the six diseases. There are less than ten overlapped genes for each query disease module with the drug modules for most disease–drug pairs (Fig. 2b and Fig. S2a).
We obtained the averaged transcriptional correlation coefficients for 1,106,328 drug pairs for the query disease and compared these coefficients between known synergistic drug pairs and the other drug pairs. We observed no significant difference between drug pairs with synergistic effects and those without synergistic effects (Fig. S2b). This observation may be due to the low number of overlapping genes between the query disease and drug modules.
We amplified the number of overlapping genes between the query disease module and drug modules by network propagation. We performed network propagation with prior knowledge on disease similarities and drug similarities (see Methods), which determines the prioritization of genes belonging to the query disease module or the drug module40. Based on the rank, we identified neighbor nodes for a query disease module as candidates for new genes. A summary of the parameters of network propagation are listed in Supplementary Data S5. The number of overlapped genes between the query module and drug modules is greater than ten for all disease–drug pairs (Fig. 2b and Fig. S2c). The network propagation process successfully increased the number of overlapped genes between the query module and drug modules.
Next, we examined the transcriptional correlation between each query disease and the individual drug modules using newly identified genes in the modules (Fig. 2c and Fig. S2d). We then averaged the transcriptional correlation coefficients of two drug modules for the query disease and obtained the averaged transcriptional correlation coefficients for 1,106,328 drug pairs for the query disease. We observed that the averaged transcriptional correlation coefficients of drug pairs with synergistic effects are significantly higher than those without synergistic effects (Fig. 2c) (p-value for CML < 0.001, colorectal cancer <0.001, and type 2 diabetes = 0.005 by two-sided t-test), except for AML and hypertension (p-value for AML = 0.301 and hypertension = 0.074 by two-sided t-test). These results suggest that network propagation emphasizes the strength of transcriptional correlations between a disease and drugs in the human molecular interaction network.
Performance evaluation of the proposed network-based trans-omics approach
We evaluated the performance of our proposed network-based trans-omics methods, Syndrum and SyndrumNET, to identify synergistic drug combinations among 1,488 drugs (Table 1). We focused on six diseases with at least one known synergistic drug combinations (Supplementary Data S6). We compared the prediction performance between the previous method, Syndrum, and SyndrumNET. Note that the previous method corresponds to the use of the network-based separation between drug modules13.
SyndrumNET works the best (Table 1). The prediction accuracy increased by 31.4% on average compared with the previous method. In particular, the AUC score for type 2 diabetes is increased 63.4% by SyndrumNET compared with the previous method (Table 1). These results suggest that it is important to consider various biological processes represented by multi-omics data for predicting synergistic drug combinations.
SyndrumNET is superior to Syndrum, which suggests that the enhancement of overlapped genes between the disease module and drug modules contributes to the enhancement of prediction accuracy (Table 1). The use of disease and drug similarities in the network propagation is more useful for predicting synergistic drug combinations.
Comprehensive prediction of new drug combinations for six diseases
Using SyndrumNET, we predicted new synergistic drug combinations for six diseases (Supplementary Data S7). For AML, the top 20 drug pairs with antineoplastic and antihypertensive activity are predicted (Supplementary Data S7). For CML, analgesic, antibiotic, antineoplastic, and vasodilator drugs are among the top 20 (Table 2 and Supplementary Data S7). For colorectal cancer, most of the top 20 drug pairs are known antineoplastic medications (Supplementary Data S7). For asthma, anti-inflammatory drugs are among the predicted top 20 pairs (Supplementary Data S7). For type 2 diabetes, combinations of antihyperlipidemic drugs and a vitamin E supplement, tocopherol, are among the top 20 predicted drugs. In particular, the combination of an antihyperlipidemic drug (gemfibrozil) and supplement (tocopherol), which are commonly used for the treatment of diabetes separately50, are among the top five (Supplementary Data S7). For hypertension, many antineoplastic drug combinations are predicted in the top 20. An antihypertensive drug, zofenopril, and an antineoplastic drug is predicted as the fifth combination (Supplementary Data S7).
We examined the prediction results for CML in more detail. The combination of capsaicin and mitoxantrone is predicted to be the top-ranked pair. Mitoxantrone is an antitumor drug used for AML. Furthermore, the combination of topotecan and mitoxantrone is ranked fifth. A preclinical study of CML patients revealed that this combination exhibited modest activity in the accelerated phase of CML51. This suggests that SyndrumNET successfully reproduced a known synergistic drug combination with a high score.
Drug combinations may exert synergistic effects by targeting specific regions of functional pathways
We examined the relationships between the query disease module and two drug modules with respect to functional pathways in the case of CML, for example. First, we performed a functional pathway enrichment analysis of the genes associated with the CML, capsaicin, and mitoxantrone modules. Using the Syndrum method, 35 pathways including the leukemia-related pathway and the Ras1 signaling pathway are enriched in the CML module (Fig. S3 and Supplementary Data S7). Eleven pathways, such as the p53 signaling pathway, are enriched in the capsaicin module (Fig. S3 and Supplementary Data S7). Fifty-nine pathways, including human T-cell leukemia virus 1 infection, are enriched in the mitoxantrone module (Fig. S3 and Supplementary Data S7). Using the SyndrumNET method, the number of pathways identified in the CML, capsaicin, and mitoxantrone modules are 187, 138, and 145, respectively (Fig. 3a, Fig. S3 and Supplementary Data S7). These results suggest that network propagation increased the diversity of functional pathways enriched in the modules (Fig. 3a).
a Enriched functional pathways in the CML module, and the capsaicin and mitoxantrone modules in the method with network propagation (SyndrumNET). Purple color indicates CML. Light green color indicates capsaicin. Yellow color indicates mitoxantrone. The p-value calculated by Fisher exact test using clusterProfiler46. The sample number is 8,772 genes. The size of points reflects the p-value of the enrichment analysis. b Distribution of the coverage of enriched pathways between the CML module and drug modules. Vertical axis indicates coverage of the functional pathways by drug module pairs. Horizontal axis indicates rank of the drug module pairs when sorted in order of the coverage. Center line, median; box limits, upper and lower quartiles; whiskers, 1.5× interquartile range; white points, mean; black points, outliers. c Network-based relationship between the CML module and a synergistic drug combination, capsaicin and mitoxantrone, in the chronic myeloid leukemia pathway (hsa05220). Purple-colored nodes indicate CML module genes. Green-colored nodes indicate capsaicin module genes and orange-colored nodes represent mitoxantrone module genes. Aqua-colored nodes indicate clopamide module genes. The purple edged nodes indicate that the gene belongs to the CML module. The purple color edges show the interaction between the genes in the CML module. Green and orange color edges represent the interaction between the module genes of capsaicin and mitoxantrone, respectively.
Second, we examined the relationship of the functional pathways between the CML disease module and two drug modules by calculating the coverage of the enriched functional pathways (Fig. 3b). For example, 169 functional pathways are enriched in the capsaicin and mitoxantrone modules, of which 151 pathways are also enriched in the CML module. The functional pathway coverage of the CML module by capsaicin and mitoxantrone modules was 0.81. The functional pathway coverage was decreased along with the predicted rank (Fig. 3b). This suggests that the top predicted drug pairs tend to target functional pathways enriched in the CML module.
Next, we determined whether the genes in the query disease and drug modules were clustered into specific regions of a functional pathway. We measured the size and significance of the largest connected component (LCC) formed by the genes of the disease module and two drug modules in a functional pathway. The module genes of CML, capsaicin, and mitoxantrone formes a larger LCC than expected by chance (Z-score > 1.95) in 50 of the 187 CML-enriched pathways. For example, the genes associated with the capsaicin and mitoxantrone modules are in the neighborhood of genes in the CML module and are clustered in the chronic myeloid leukemia pathway (hsa05220) (Fig. 3c). On the other hand, the genes in the clopamide module are located aside from the CML module in the functional pathway (Fig. 3c). These results suggest that the genes in the capsaicin and mitoxantrone modules are specifically localized into the neighborhoods of the functional pathways enriched in the CML module.
In vitro experimental validation of the predicted drug combinations for CML
We validated the prediction results of SyndrumNET in vitro by conducting cell survival assays on CML cells (K562) for the top 20 predicted drug combinations (Table 2). Tiludronic acid was excluded from the validation list because it is not considered to act directly on cancer cells as it exhibits a preferential effect on skeletal muscle cells. Propranolol was also excluded from the validation list because it lowers blood pressure by blocking beta-adrenergic receptors in the heart and suppressing the heartbeat. In addition, cancer patients often experience cardiac dysfunction resulting from anticancer drug exposure and other pathological conditions, such as cancer cachexia. Among the top 20 predicted drug combinations, three pairs include either tiludronic acid or propranolol. The number of excluded combinations was 3, leaving 17 combinations for the final validation.
We exposed K562 cells to the drug pairs above for 72 h and measured cell survival using the WST assay. The survival ratio for each drug on K562 cells is shown in Table 2 as a ratio of drugs A or B. In addition, cell viability for the drug combination is shown as a survival ratio (Table 2). The statistical significance of synergistic effects for the top 17 drug combinations were evaluated using the CA model44,52 and the independent action (IA) model43,52, which are standard indicators of drug synergy. The output from the CA model is referred to as a CA score. If the CA score is lower than 1, the corresponding drug pair is considered to have a synergistic effect. The smaller the CA score, the higher the synergy. The output from the IA model is referred to as the IA score. If the IA score is higher than 1, the corresponding drug pair is considered to exhibit synergy. Synergistic effects were observed for 76.5% of the drug combinations in the CA model and 88.2% in the IA model. These results demonstrate that our prediction approach has high accuracy.
Elucidation of the mode-of-action of the synergistic drug combination for CML by microarray analysis
We examined the mode-of-action of synergistic drug combinations by microarray analysis. We focused on the drug combination of capsaicin and mitoxantrone, which were the top predicted drug pair identified by SyndrumNET. To determine the transcriptomic responses of CML cells to capsaicin, mitoxantrone, and the combination, we conducted a microarray analysis of CML cells exposed to the individual drugs and the combination. We identified 617 differentially expressed genes (DEGs) for capsaicin (Fig. 4a, b and Supplementary Data S8), 679 for mitoxantrone (Fig. 4a, b, and Supplementary Data S8), and 2,048 associated with the combination (Fig. 4a, b, and Supplementary Data S8). Moreover, 91 DEGs are common to the three groups (Fig. 4b), whereas 1,313 DEGs are detected only in the combined exposure group (Fig. 4b). The DEGs in the combined exposure group are different from those in the capsaicin and mitoxantrone exposure groups. These results suggest that the mechanism underlying transcriptional changes may be different between single and combined drug exposure.
a The heatmap of log2FC of capsaicin, mitoxantrone, and the combination exposure group. Yellow and green colors indicate up- and down-regulation, respectively. b The number of DEGs and overlapping genes between the three exposure groups. Cyan represents the combination exposure group. Green represents the capsaicin exposure group. Yellow represents the mitoxantrone exposure group. c The significance of the pathways enriched only in the combination exposure group. Cyan represents the combination exposure group. Green represents the capsaicin exposure group. Yellow represents the mitoxantrone exposure group. N.S. indicates not significant based on the Fisher exact test (p > 0.05) using DAVID45. The number of samples is 8,156 genes. d The log2FC of THBS1, RASGRP3, and PDGFB that belong to the RAP1 singling pathway and thought to be synergistically regulated by the combination exposure group. Green, yellow, and cyan represent capsaicin, mitoxantrone, and the combination exposure groups, respectively. Exposure experiments were conducted in three independent wells for each exposure condition. The extracted total RNA from these wells was combined by exposure condition for microarray analysis. Thus, the sample number for this figure is one. e The mode-of-action of the effects of combination exposure. The red solid lines represent the connections predicted by the transcription factor enrichment assay. The black solid lines indicate the known interactions between the proteins. The green- and orange-colored lines represent the assumed connection between the transcription factor and capsaicin and mitoxantrone, respectively.
Next, we investigated the functional pathways that were synergistically regulated by exposure to the drug combination (Table 3). Functional pathways enriched in the combined exposure group, but not in the single exposure group, are considered synergistically regulated. We identified 12 functional pathways that are synergistically affected in the combined exposure group. Interestingly, the Rap1 signaling pathway is identified only in the combined exposure group, but not in the single exposure groups (Table 3 and Fig. 4c). The Rap1 signaling pathway plays an essential role in the migration of leukocytes and lymphocytes and in the regulation of tumor progression53,54,55. These results suggest that the 12 functional pathways including the Rap1 signaling pathway are important to the synergistic effects of the drug combination in CML.
We examined the gene expression changes in the Rap1 signaling pathway and compared them with the exposure groups. The fold changes of RASGRP3, PDGFB, and THBS1 expression in the combined exposure group are more than twice higher compared with the total fold changes of capsaicin and mitoxantrone exposure groups (Fig. 4d, Fig. S4 and Fig. S5). These results suggest that the combination synergistically accelerated the expression of these genes.
Finally, we examined transcription factors (TFs) enriched in the three genes (e.g., RASGRP3, PDGFB, and THBS1). TFs are identified by Fisher’s exact test using the Enricher analysis tool56. We selected the ChIP-x Enrichment Analysis (ChEA) database as a gene set library. The ChEA database contains putative targets for TFs extracted from publications on experimental profiling of TFs binding to DNA in mammalian cells47. The stem cell leukemia gene (SCL), also known as Tal-157, is the most statistically enriched TF (Fig. S4). This suggests that capsaicin and mitoxantrone induce RASGRP3, PDGFB, and THBS1 expression through SCL to inhibit cell survival (Fig. 4e).
Discussion
We proposed a network-based trans-omics approach, SyndrumNET, to predict synergistic drug combinations for various human diseases. The originality of the method lies in its ability to identify drug combinations considering various biological processes, which was achieved by integrating multi-omics data representing different biological processes, such as static information on molecular interaction networks and dynamic information on drug transcriptomic responses. We emphasized transcriptional correlations between disease modules and drug modules using network propagation, to improve prediction accuracy. The method can be expanded to encompass additional diseases and tailored to specific gene signatures. By constructing gene expression profiles for novel drugs, our proposed method can be effectively employed for analyzing drug-specific response profiles. This method is also adaptable to any compounds with available response profiles. We predicted new drug combinations for CML using this method and validated the anticancer activity of predicted drug combination in vitro. We identified the underlying mode-of-action the drug synergy by microarray analysis at the pathway level. The proposed method will be useful for predicting synergistic drug combinations for various diseases.
We demonstrated that drug modules constructed by drug response genes tend to be in the neighborhood of associated diseases. Network-based methods have traditionally used drug target molecules to characterize drugs and construct drug modules13,15,58,59; however, few drugs have known target molecules. In addition, only a limited number of drugs have a sufficient number of target molecules to construct a drug module16. A recent study using a network-based method demonstrated that genes which respond to drugs and compounds are useful for predicting the efficacy of the drugs and compounds60. Therefore, it is reasonable to use drug response genes for constructing drug modules as an alternative to target molecules.
We used transcriptional correlations between disease modules and drug modules. This transcriptome-based method facilitates the systematic comparison of gene expression profiles that characterize the response to drugs and biological states of interest. The disadvantage of a transcriptome-based method is that correlation scores for disease–drug pairs tend to be low11. It is reported that most disease–drug pairs exhibit a weak transcriptional correlation when using gene expression of all protein-coding genes14. We found that modules were expanded by network propagation and the overlapping genes between expanded modules emphasized the transcriptomic correlation between the disease and drug module. This suggests that the expansion of modular genes using network propagation is effective for deterring transcriptional correlations between diseases and drugs. The network propagation method will contribute to the enhancement of prediction accuracy of the transcriptome-based method.
In this study, we focused on CML. Three discrete clinical stages are defined for CML: the chronic phase, the accelerated phase, and the blast crisis. Drugs known as tyrosine kinase inhibitors (TKIs) that target BCR-ABL are the standard treatment for CML61. Patients in the chronic phase achieve a 10-year survival rate of more than 90% with the TKIs. On the other hand, the prognosis of patients in the blast phase is poor, and treatments are limited. We used the K562 cell line derived from a patient in the blast phase for experimental validation to propose effective drug combinations for the blast phase of CML. We found that the combination of capsaicin and mitoxantrone exhibited synergistic effects on the CML cells. Neither capsaicin nor mitoxantrone targets the BCR-ABL fusion gene. In addition, neither drug module included BCR or ABL gene. The synergistic effects of the combination of capsaicin and mitoxantrone seems not be occurred through BCR-ABL inhibition and could be combined with standard therapy using TKIs. The synergistic effects of capsaicin and mitoxantrone seems to be occurred through the inhibition of DNA repair by mitoxantrone and the anticancer effect of capsaicin. Anticancer mechanisms are primarily related to induction of apoptosis and autophagy, reduced proliferation, as well as the inhibition of angiogenesis and metastasis62. Consistent with previous results, DEGs associated with capsaicin exposure were enriched in cancer pathways in our microarray analysis (Fig. 4). Mitoxantrone is known to be an inhibitor of topoisomerase II63. Thus, mitoxantrone may alter the expression of several genes, such as TFs and receptors, for capsaicin. The genes stimulated by capsaicin may alter the expression of downstream genes. Indeed, the expression of 1,313 genes were affected only by combination treatment (Fig. 4).
We detected the RAP1 signaling pathway as an important functional pathway exerting a synergistic effect in response to capsaicin and mitoxantrone. Our hypothesis is based on observations from our microarray analysis and previous observations about the function of genes in the RAP1 signaling pathway on cancer. RAP1, a member of the shelterin complex, has been implicated in cancer development64. For example, RAP1 activated by RasGRP3 increased cell migration and invasion in glioma cells65. From our microarray analysis, the expression of RAP1 was not altered by combination exposure. On the other hand, RASGRP3 was overexpressed by combination exposure (Fig. 4e and Fig. S8). Thus, it is possible that overexpression of RASGRP3 by combined drug exposure activates RAP1.
THBS1, as a tumor suppressor gene, influences the growth of tumors by inhibiting angiogenesis and activating the transforming growth factor. THBS1 is weakly expressed in AML patients66, which is associated with a shorter survival time66. From our microarray analysis, THBS1 expression was notably increased by combination exposure (Fig. 4e and Fig. S4). The results suggest that enhanced THBS1 expression by combination exposure suppresses the proliferation of CML cells. Future experimental validation is needed to confirm the mode-of-action of the drug combination through the RAP1 signaling pathway.
We examined the mode-of-action of the synergistic drug combination in the context of transcriptome factors. We found that SCL was enriched in the promoter region of overexpressed genes by combination treatment, such as RASGRP3, PDGFB, and THBS1. Increased expression of SCL is associated with leukemia and poor prognosis of T-cell acute lymphoblastic leukemia67. Interestingly, the expression of SCL was slightly decreased in the drug combination group (Fig. S6). This suggests that the suppressed expression of SCL may contribute to the synergistic effects of the combination.
We opted for microarray technology to understand the mode of action of the synergistic effect of capsaicin and mitoxantrone over RNA-seq analysis due to our emphasis on assessing the expression levels of coding regions in genes with known functions. Microarray technology offers advantages in terms of speed, simplicity, and affordability and requires minimal RNA input for the detection of the expression levels of known genes. While acknowledging the inherent differences between RNA-Seq and microarray technologies, it is worth noting that previous studies68,69 have reported a notable overlap (approximately 70%–80%) in differentially expressed genes identified by both methods, with a Spearman’s correlation ranging from 0.7 to 0.8. These findings imply that transitioning from microarray to RNA-Seq may not substantially alter the results. Furthermore, the primary distinction between microarray and RNA-Seq technologies lies in their gene detection capabilities: microarrays quantify a predetermined set of known genes (e.g., mRNA), whereas RNA-Seq can sequence all RNAs present, including those with unknown functions like miRNA and non-coding RNA. Our analysis of gene expression profiles aimed to uncover the mechanism behind the observed synergistic effect. KEGG pathway analysis excludes genes with unknown functions due to current limitations in pathway databases and analytical frameworks. Given these limitations, we argue that microarray data suffice for this study’s objectives because our focus is on elucidating mechanisms through known genes and their functional relationships in established pathways. However, we acknowledge the potential advantages of RNA-Seq, such as its broader dynamic range and ability to identify expressions of genes with unknown functions, including miRNA and non-coding RNA. The use of RNA-Seq offers significant analytical benefits, especially when the aim is to elucidate novel pathways or characterize genes with previously unknown functions.
In this study, we employed a human molecular interaction network created from various public databases. This network includes 13,377 proteins and 235,123 interactions, covering 65% of the genes responsible for human protein coding (Supplementary Data S2). It is worth noting that protein interaction databases undergo continuous updates. Since our methodology can be adjusted to accommodate new datasets, utilizing updated protein interaction network data holds promise for enhancing the accuracy and comprehensiveness of drug combination discovery. Developing predictive models based on the most recent protein interaction network data stands as a vital avenue for future research.
One limitation of the proposed method is that it does not consider disease subtypes. For example, the World Health Organization (WHO) classified AML into 25 subtypes, including two provisional entities, which differ in prognosis and treatment70; however, the genetic features of some subtypes remain unclear71. In addition, there may be few differences in genetic characteristics between the subtypes. According to the WHO classification, AML with RUNX1 mutation, AML with NPM1 mutation, and AML with biallelic CEBPA mutations are considered distinct categories70. In our proposed method, genetic features are used to define disease modules. A lack of variation in genetic features among subtypes results in reduced diversity in the predicted results and prediction accuracy. To predict effective drug combinations for each subtype, genetic data for the individual subtypes is needed. A recent study succeeded in predicting drug combinations for melanoma subtypes by considering transcriptome correlations and network centralities of genes between disease subtypes and drugs72. The incorporation of variation in genetic features could establish a prediction method for individual disease subtypes.
Data availability
We have only use existing publicly available datasets for analysis. The source databases for constructing human molecular interactions are summarized in from Supplementary Data S1. Human molecular interactions constructed in this study are available in Supplementary Data S2. A set of disease susceptibility genes and drug related genes are described in Supplementary Data S3 and S4. Gene expression signatures for 79 diseases with 14,804 genes are in the CREEDS database24. Gene expression signatures for hypertension can be retrieved from GEO (GSE24752, and GSE75360). Drug-induced gene expression profiles can be obtained from the LINCS Program L1000 mRNA profiling assay49. Drug combinations with synergistic effects are listed in Supplementary Data S6. The prediction results by SyndrumNET can be seen in Supplementary Data S7. Our microarray data can be found at GSE254052. The source data for Fig. 2a–c are in Supplementary Data Set 173, Supplementary Data Set 274, and Supplementary Data Set 375, respectively. The source data for Fig. 3a–c are in Supplementary Data Set 476, Supplementary Data Set 577, and Supplementary Data S2, S3, and S4, respectively. The source data of Figs. 4a, b, d are in GSE254052 and Supplementary Data S8, and for 4c is in Supplementary Data Set 678. All other data are available from the corresponding author on reasonable request.
Code availability
The functions used for our analyses are available at igraph (1.2.6), biomaRt (4.0.3), lsa (0.73.2), ROCR (v.1.0-11), and clusterProfiler (v4.4.4). The code for network propagation is available at: https://github.com/kztakemoto/network_propagation. The scripts for calculating prediction score and AUC are in Supplementary Data Set 735.
References
Mokhtari, R. B. et al. Combination therapy in combating cancer. Oncotarget 8, 38022–38043 (2017).
Doroshow, J. H. & Simon, R. M. On the design of combination cancer therapy. Cell 171, 1476–1478 (2017).
Gradman, A. H., Basile, J. N., Carter, B. L. & Bakris, G. L. Combination therapy in hypertension. J. Clin. Hypertens. 13, 146–154 (2011).
Das, P., Delost, M. D., Qureshi, M. H., Smith, D. T. & Njardarson, J. T. A survey of the structures of US FDA approved combination drugs. J. Med. Chem. 62, 4265–4311 (2019).
FDA. Fact Sheet: FDA at a Glance. U.S. FOOD & DRUG ADMINISTRATION From the OFFICE OF THE COMMISSIONER November (2021). Available at: https://www.fda.gov/about-fda/fda-basics/fact-sheet-fda-glance. (Accessed: 31st October 2022).
Kong, W. et al. Systematic review of computational methods for drug combination prediction. Comput. Struct. Biotechnol. J. 20, 2807–2814 (2022).
Zhao, X. M. et al. Prediction of drug combinations by integrating molecular and pharmacological data. PLOS Comput. Biol. 7, e1002323 (2011).
Iwata, H., Sawada, R., Mizutani, S., Kotera, M. & Yamanishi, Y. Large-scale prediction of beneficial drug combinations using drug efficacy and target profiles. J. Chem. Inf. Model. 55, 2705–2716 (2015).
Celebi, R. O., Bear Don’t Walk, R., Movva, S., Alpsoy, S. & Dumontier, M. In-silico prediction of synergistic anti-cancer drug combinations using multi-omics data. Sci. Rep. 9, 8949 (2019).
Jin, W. et al. Deep learning identifies synergistic drug combinations for treating COVID-19. Proc. Natl Acad. Sci. USA 118, e2105070118 (2021).
Stathias, V. et al. Drug and disease signature integration identifies synergistic combinations in glioblastoma. Nat. Commun. 9, 5315 (2018).
Li, X., Qin, G., Yang, Q., Chen, L. & Xie, L. Biomolecular Network-Based Synergistic Drug Combination Discovery. Biomed. Res. Int. 2016, 1–11 (2016).
Cheng, F., Kovács, A. & Barabási, A. Network-based prediction of drug combinations. Nat. Commun. 10, 1197 (2019).
Iwata, M. et al. Regulome-based characterization of drug activity across the human diseasome. npj Syst. Biol. Appl. 8, 44 (2022).
Guney, E., Menche, J., Vidal, M. & Barábasi, A.-L. Network-based in silico drug efficacy screening. Nat. Commun. 7, 10331 (2016).
Yıldırım, M. A., Goh, K.-I., Cusick, M. E., Barabási, A.-L. & Vidal, M. Drug—target network. Nat. Biotechnol. 25, 1119–1126 (2007).
Luck, K. et al. A reference map of the human binary protein interactome. Nature 580, 402–408 (2020).
Giurgiu, M. et al. CORUM: the comprehensive resource of mammalian protein complexes—2019. Nucleic Acids Res. 47, D559–D563 (2019).
Hornbeck, P. V. et al. PhosphoSitePlus: a comprehensive resource for investigating the structure and function of experimentally determined post-translational modifications in man and mouse. Nucleic Acids Res. 40, D261–70 (2012).
Shimizu, Y., Hattori, M., Goto, S. & Kanehisa, M. Generalized reaction patterns for prediction of unknown enzymatic reactions. Genome Inf. 20, 149–58 (2008).
Fazekas, D. et al. SignaLink 2 – a signaling pathway resource with multi-layered regulatory networks. BMC Syst. Biol. 7, 7 (2013).
Breuer, K. et al. InnateDB: Systems biology of innate immunity and beyond - Recent updates and continuing curation. Nucleic Acids Res. 41, D1228–D1233 (2013).
Meyer, M. J., Das, J., Wang, X. & Yu, H. INstruct: A database of high-quality 3D structurally resolved protein interactome networks. Bioinformatics 29, 1577–1579 (2013).
Wang, Z. et al. Extraction and analysis of signatures from the Gene Expression Omnibus by the crowd. Nat. Commun. 7, 12846 (2016).
Edgar, R. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Amberger, J. S., Bocchini, C. A., Schiettecatte, F., Scott, A. F. & Hamosh, A. OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an Online catalog of human genes and genetic disorders. Nucleic Acids Res. 43, D789–D798 (2015).
Landrum, M. J. et al. ClinVar: Public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–5 (2014).
Welter, D. et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 42, D1001–D1006 (2014).
Denny, J. C. et al. Systematic comparison of phenome-wide association study of electronic medical record data and genome-wide association study data. Nat. Biotechnol. 31, 1102–1111 (2013).
Li, M. J. et al. GWASdb: A database for human genetic variants identified by genome-wide association studies. Nucleic Acids Res. 40, D1047–D1054 (2012).
Pinero, J. et al. DisGeNET: A discovery platform for the dynamical exploration of human diseases and their genes. Database 2015, bav028 (2015).
Durinck, S. et al. BioMart and Bioconductor: A powerful link between biological databases and microarray data analysis. Bioinformatics 21, 3439–3440 (2005).
Subramanian, A. et al. A Next Generation Connectivity Map: L1000 Platform and the First 1,000,000 Profiles. Cell 171, 1437–1452.e17 (2017).
Berenger, F., Coti, C. & Zhang, K. Y. J. PAR: A PARallel And Distributed Job Crusher. Bioinformatics. https://doi.org/10.1093/bioinformatics/btq542 (2010)
Iida, M. Code for A network-based trans-omics approach for predicting synergistic drug combinations. https://doi.org/10.6084/m9.figshare.25735206. (2024)
Iida, M., Iwata, M. & Yamanishi, Y. Network-based characterization of disease-disease relationships in terms of drugs and therapeutic targets. Bioinformatics 36, i516–i524 (2020).
Kanehisa, M., Goto, S., Furumichi, M., Tanabe, M. & Hirakawa, M. KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res. 38, D355–D360 (2010).
Kotera, M. et al. KCF-S: KEGG Chemical Function and Substructure for improved interpretability and prediction in chemical bioinformatics. BMC Syst. Biol. 7, S2 (2013).
Aoto, Y. et al. Time-series analysis of tumorigenesis in a murine skin carcinogenesis model. Sci. Rep. 8, 12994 (2018).
Vanunu, O., Magger, O., Ruppin, E., Shlomi, T. & Sharan, R. Associating genes and protein complexes with disease via network propagation. PLoS Comput. Biol. 6, e1000641 (2010).
Csardi, G. & Nepusz, T. The igraph software package for complex network research. InterJournal Complex Sy, 1695 (2006).
Liu, H. et al. DrugCombDB: a comprehensive database of drug combinations toward the discovery of combinatorial therapy. Nucleic Acids Res. 48, 871 (2020).
Bliss, C. I. The toxicity of poisons applied jointly 1. Ann. Appl. Biol. 26, 585–615 (1939).
Loewe, S. & Muischnek, H. Effect of combinations: Mathematical basis of the problem. Arch. Exp. Pathol. Pharmakol. 114, 313–326 (1926).
Sherman, B. T. et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50, W216–W221 (2022).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innov 2, 100141 (2021).
Lachmann, A. et al. ChEA: Transcription factor regulation inferred from integrating genome-wide ChIP-X experiments. Bioinformatics 26, 2438–2444 (2010).
Menche, J. et al. Disease networks. Uncovering disease-disease relationships through the incomplete interactome. Science 347, 1257601 (2015).
Koleti, A. et al. Data Portal for the Library of Integrated Network-based Cellular Signatures (LINCS) program: Integrated access to diverse large-scale cellular perturbation response data. Nucleic Acids Res. 46, D558–D566 (2018).
Choi, S. W. & Ho, C. K. Antioxidant properties of drugs used in Type 2 diabetes management: could they contribute to, confound or conceal effects of antioxidant therapy? Redox Rep. 23, 1–24 (2018).
Park, S. J. et al. Topotecan-based combination chemotherapy in patients with transformed chronic myelogenous leukemia and advanced myelodysplastic syndrome. Korean J. Intern. Med. 15, 122–126 (2000).
He, L. et al. Methods for high-throughput drug combination screening and synergy scoring. Methods Mol. Biol. 1711, 351–398 (2018).
Looi, C. K., Hii, L. W., Ngai, S. C., Leong, C. O. & Mai, C. W. The role of Ras-associated Protein 1 (Rap1) in cancer: bad actor or good player? Biomedicines 8, 334 (2020).
Wittchen, E. S. et al. Rap1 GTPase inhibits leukocyte transmigration by promoting endothelial barrier function. J. Biol. Chem. 280, 11675–11682 (2005).
Katagiri, K. et al. Rap1 is a potent activation signal for leukocyte function-associated antigen 1 distinct from protein kinase C and phosphatidylinositol-3-OH kinase. Mol. Cell. Biol. 20, 1956–1969 (2000).
Chen, E. Y. et al. Enrichr: Interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinf. 14, 128 (2013).
Barton, L. M. et al. Regulation of the stem cell leukemia (SCL) gene: A tale of two fishes. Proc. Natl Acad. Sci. USA 98, 6747–6752 (2001).
Barabási, A. L., Gulbahce, N. & Loscalzo, J. Network medicine: A network-based approach to human disease. Nat. Rev. Genet. 12, 56–68 (2011).
Cheng, F. et al. Network-based approach to prediction and population-based validation of in silico drug repurposing. Nat. Commun. 9, 2691 (2018).
do Valle, I. F. et al. Network medicine framework shows that proximity of polyphenol targets and disease proteins predicts therapeutic effects of polyphenols. Nat. Food 2, 143–155 (2021).
American Cancer Society. Targeted Therapies for Chronic Myeloid Leukemia. Available at: https://www.cancer.org/cancer/chronic-myeloid-leukemia/treating/targeted-therapies.html (2023). (Accessed: 1st March 2023).
Zhang, S., Wang, D., Huang, J., Hu, Y. & Xu, Y. Application of capsaicin as a potential new therapeutic drug in human cancers. J. Clin. Pharm. Ther. 45, 16–28 (2020).
Osheroff, N., Corbett, A. H. & Robinson, M. J. Mechanism of action of topoisomerase II-targeted antineoplastic drugs. Adv. Pharmacol. 29, 105–126 (1994).
Deregowska, A. & Wnuk, M. RAP1/TERF2IP-a multifunctional player in cancer development. Cancers (Basel) 13, 5970 (2021).
Lee, H. K. et al. RasGRP3 regulates the migration of glioma cells via interaction with Arp3. Oncotarget 6, 1850–1864 (2015).
Zhu, L. et al. THBS1 is a novel serum prognostic factors of acute myeloid leukemia. Front. Oncol. 9, 1567 (2020).
Porcher, C., Chagraoui, H. & Kristiansen, M. S. SCL/TAL1: A multifaceted regulator from blood development to disease. Blood 129, 2051–2060 (2017).
Xu, X. et al. Parallel comparison of Illumina RNA-Seq and Affymetrix microarray platforms on transcriptomic profiles generated from 5-aza-deoxy-cytidine treated HT-29 colon cancer cells and simulated datasets. BMC Bioinforma. 14, S1 (2013).
Rao, M. S. et al. Comparison of RNA-Seq and microarray gene expression platforms for the toxicogenomic evaluation of liver from short-term rat toxicity studies. Front. Genet. 9, 636 (2019).
Narayanan, D. & Weinberg, O. K. How I investigate acute myeloid leukemia. Int. J. Lab. Hematol. 42, 3–15 (2020).
Rose, D. et al. Subtype-specific patterns of molecular mutations in acute myeloid leukemia. Leukemia 31, 11–17 (2017).
Regan-Fendt, K. E. et al. Synergy from gene expression and network mining (SynGeNet) method predicts synergistic drug combinations for diverse melanoma genomic subtypes. npj Syst. Biol. Appl. 5, 6 (2019).
Iida, M. Figure 2a Distribution of average network-based proximity (PQAB) between a query disease module (Q) and drug modules (A and B). https://doi.org/10.6084/m9.figshare.25699275.v1.
Iida, M. Figure 2b Distribution of the number of overlapped genes between a query disease module (Q) and individual drug modules (A or B). https://doi.org/10.6084/m9.figshare.25699356.v1.
Iida, M. Figure 2c Distribution of average absolute transcriptional correlation coefficients between a query disease module (Q) and drug modules (A and B). https://doi.org/10.6084/m9.figshare.25699434.v1.
Iida, M. Figure 3a Enriched functional pathways in the CML module, and the capsaicin and mitoxantrone modules in the method with network propagation (SyndrumNET). https://doi.org/10.6084/m9.figshare.25699446.v1.
Iida, M. Figure 3b Distribution of the coverage of enriched pathways between the CML module and drug modules. https://doi.org/10.6084/m9.figshare.25705947.
Iida, M. Figure 4c The significance of the pathways enriched only in the combination exposure group. https://doi.org/10.6084/m9.figshare.25705959.v1.
Acknowledgements
This work was supported by the Japan Society for the Promotion of Science (JSPS), KAKENHI Grant-in-Aid for Transformative Research Areas (B) [grant number 20H05797], and Transformative Research Areas [Synergy pharmacology] for Y. Y. This work was also supported by the JSPS, KAKENHI Grant-in-Aid for Transformative Research Areas (B) [grant number 20H05799] for M. G. This study was also supported by the Naito foundation, an Intramural Research Grant, Grant-in-Aid for JSPS Fellows Restart Postdoctoral Fellow (RPD) [22J40019], and Grant-in-Aid for Scientific Research (C) [22K12265] to M.I.
Author information
Authors and Affiliations
Contributions
M.I.: Conceptualization, Formal analysis, Writing - Original Draft, Visualization, Funding acquisition. Y.K.: Methodology, Investigation. K.Y.: Methodology, Investigation. M.G.: Methodology, Investigation, Writing - Original Draft. S.N.: Data Curation. J.T.: Methodology, Writing - Original Draft. R.S.: Data Curation. Michio Iwata: Data Curation. Y.Z.: Methodology, Investigation, Writing - Review & Editing. K.I.: Methodology, Investigation. Y.Y.: Conceptualization, Resources, Writing - Review & Editing, Visualization, Supervision, Funding acquisition.
Corresponding author
Ethics declarations
Competing interests
Authors declare no competing interests.
Peer review
Peer review information
Communications Medicine thanks the anonymous reviewers for their contribution to the peer review of this work
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Iida, M., Kuniki, Y., Yagi, K. et al. A network-based trans-omics approach for predicting synergistic drug combinations. Commun Med 4, 154 (2024). https://doi.org/10.1038/s43856-024-00571-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43856-024-00571-2
- Springer Nature Limited