Abstract
The goal of this work was to explore the most effective miRNAs affecting glioblastoma multiforme (GBM) phenotype transition and malignant progression. We annotated 491 TCGA samples’ miRNA expression profiles according to their mRNA-based subtypes and found that the mesenchymal tumors had significantly decreased miR-181 family expression compared with the other three subtypes while the proneural subtype harbored extremely high miR-181 family expression. Patients with high miR-181 family expression had longer overall survival (p = 0.0031). We also confirmed that NF-κB-targeting genes and the EMT (epithelial-mesenchymal transition) pathway were inversely correlated with miR-181 family expression and that the entire miR-181 family inhibited glioma cell invasion and proliferation; of these, miR-181b was the most effective suppressor. Furthermore, miR-181b was validated to suppress EMT by targeting KPNA4 and was associated with survival outcome in the TCGA and CGGA datasets and in another independent cohort. The EMT-inhibitory effect of miR-181b was lost after KPNA4 expression was restored. We also identified the antitumorigenic activity of miR-181b in vitro and in vivo. Our results showed that miR-181 family expression was closely correlated with TCGA subtypes and patients’ overall survival, indicating that miR-181b, a tumor-suppressive miRNA, could be a novel therapeutic candidate for treating gliomas.
Similar content being viewed by others
Introduction
Recently, several advantages have been discovered for the use of miRNAs instead of mRNAs as biomarkers1,2,3. There are nearly 1,000 miRNAs and more than 40,000 protein-coding genes in the human genome. Consequently, genome-wide gene expression data are much more extensive than miRNA expression data. It is more feasible to explore reliable miRNA biomarkers from genome-wide miRNA expression data than from genome-wide gene expression data. Furthermore, miRNAs are relatively stable and are therefore more amenable to analysis in formalin-fixed, paraffin-embedded tissues.
Glioblastoma multiforme (GBM) is the most common form of primary brain cancer. The median survival of primary GBM patients is approximately 1 year, but survival varies widely from <1 week to >3 years after diagnosis4. GBM has been classified into four subtypes—proneural, neural, classical and mesenchymal—by Verhaak et al5. These four subtypes have different biological behaviors and distinct markers. The contribution of miRNAs to molecular subtype has not been clarified previously. We annotated the miRNA microarrays of 491 TCGA samples by considering the four mRNA subtypes of GBM and analyzed two databases from America (TCGA) and China (CGGA), which together include nearly 700 samples. We found that extremely low miR-181 family expression was a hallmark for the mesenchymal subtype of glioblastomas.
miR-181 family members have been observed in several primary human cancers, including gastric, lung and prostate cancer, astrocytic tumors, acute myeloid leukemia and chronic lymphocytic leukemia6,7,8,9,10,11. Low expression of miRNA-181 family members is associated with a poor prognosis in several cancer subtypes9,11,12,13,14, consistent with our results in both TCGA and CGGA databases. Sun et al. demonstrated that miR-181b regulated NF-κB–mediated endothelial cell (EC) activation and vascular inflammation in response to proinflammatory stimuli by targeting KPNA4 and that the rescue of miR-181b expression could provide a new target for anti-inflammatory therapy15. Base on the inhibitory effect of miR-181b on NF-κB, we found that miR-181b overexpression could reverse the epithelial-mesenchymal transition (EMT) and inhibit tumor growth in vitro and in vivo.
Results
Aberrant expression of the miR-181 family correlated with TCGA subtypes and prognosis of patients with GBMs
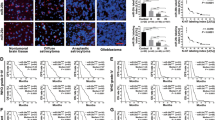
We annotated miRNA microarrays (Supplementary Figure 1) by considering the four mRNA-based subtypes of glioblastoma according to the TCGA (Supplementary Figure 2). By analyzing the miRNA expression in the four subtypes, we determined that the miR-181 family (miR-181a, miR-181b, miR-181c and miR-181d) was significantly down-regulated in the mesenchymal subtype compared with the other three subtypes, especially, the proneural subtype (Fig. 1A,B). The expression of the miR-181 family divided patients into two groups according to the expression of miR-181s. Patients in the miR-181 low-expression group had significantly shorter overall survival than those in the miR-181 high-expression group (p = 0.0031) (Fig. 1C). These data supported the hypothesis that miR-181 family members are tumor suppressors in glioma.
miR-181 family was preferentially expressed in proneural glioma.
(A) After annotating the miRNA microarrays according to the TCGA mRNA microarray subtypes, we identified the differentially expressed miRNAs and observed that miR-181 family expression was dominant in the proneural subtype. (B) There was a significant difference in miR-181 family expression among the four TCGA subtypes. miR-181 family expression in the proneural subtype was significantly higher than in the other three subtypes, especially the mesenchymal subtype. (C) When the entire miR-181 family was considered, patients in the miR-181 family low-expression group had a significantly shorter survival time (p = 0.0031).
miR-181 family overexpression inhibited cell invasion and proliferation
We performed Gene set variation analysis (GSVA) of miR-181 expression to determine the biological functions of the miR-181 family in glioma (Fig. 2A). NF-κB targets and an EMT gene signature list were significantly inversely correlated with miR-181 family expression and the NF-κB targets and the EMT gene signature list were surprisingly similar. To further determine whether the miR-181 family was associated with glioma invasion, we used two common glioma cell lines U87 and LN229 and observed that invasion and proliferation decreased in cells transfected with miR-181 family members compared with control cells, especially in those transfected with miR-181b (Fig. 2B,C). These results indicated that miR-181 family overexpression participated in the regulation of glioma cell motility and proliferation in vitro and that miR-181b was the most effective member.
miR-181 family overexpression inhibited cell invasion and proliferation.
(A) Gene Set Variation Analysis indicated that NF-κB targets and the EMT gene signature were significantly enriched in the tumors with low expression of the miR-181 family. (B/C) U87 and LN229 cells were transfected with hsa-miR-181a, hsa-miR-181b, hsa-miR-181c, or hsa-miR-181d expression vectors. Transwell invasion assays revealed that cells transfected with miR-181 family member mimics all had weaker invasion capacity (*p < 0.05). Colony formation assays showed that treated cells exhibited a significant reduction of colony formation after 2 weeks of miR-181 family mimic treatment (*p < 0.05). miR-181b had a trend of more effective suppression on tumor cells invasion and colony formation.
Patients with elevated miR-181b expression had a better prognosis because of KPNA4 inhibition and the reversal of EMT
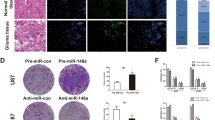
A survival analysis of 491 GBM miRNA microarray samples from the TCGA indicated that high-grade glioma patients with higher miR-181b expression had a significantly longer OS (p = 0.0080) than those with low expression (Fig. 3A). Similar results were found in the CGGA dataset (82 samples, p = 0.0230) and in another independent cohort (107 samples, p < 0.0001). We performed a bioinformatics search for potential targets of miR-181b (http://www.microrna.org/) to better understand its mechanism in glioma. There are two predicted binding sites for miR-181b in the KPNA4 DNA sequence (Fig. 3B). A luciferase assay was performed in a pMIR-REPORT miRNA reporter vector containing the putative wild-type (WT) and mutant (Mut) KPNA4 3′UTR binding sites. The overexpression of miR-181b inhibited wild-type, but not mutant, KPNA4 reporter activity, suggesting that miR-181b specifically targeted the 3′ UTR of KPNA4 (Fig. 3C).
miR-181b could reverse EMT by targeting KPNA4 and its expression correlates with better prognosis.
(A) In the TCGA and CGGA cohorts and another independent cohort, patients with elevated miR-181b expression had a better prognosis (p = 0.008, p = 0.023 and p < 0.0001, respectively). (B) Diagram of the seed sequence of miR-181b matching the 3′UTRs of the KPNA4 gene and the design of wild-type or mutant KPNA4 3′UTR-containing reporter constructs. (C) Luciferase reporter assays in glioma cells after co-transfection of cells with wild-type or mutant 3′UTR KPNA and miR-181b. The data represent the fold change in the expression (mean ± SE) of 3 replicates. *p < 0.05. (D) Western blot for KPNA4, N-cadherin, E-cadherin and Vimentin expression 48 hours after transfection with miR-NC or miR-181b. (E) Immunohistochemisty revealed that tumors in the miR-181b low expression group had lower KPNA4, N-cadherin and Vimentin and higher E-cadherin expression.
Compared with the stable expression of miR-NC in U87 and LN229 cells, the stable expression of miR-181b reduced KPNA4 protein expression (Fig. 3D), significantly decreased the expression of the mesenchymal markers (N-cadherin and Vimentin) and increased the expression of the epithelial marker, E-cadherin. These data suggested that the tumor suppressor activity of miR-181b in glioma cells regulated EMT pathways. KPNA4 protein expression in glioma tissues was analyzed by immunohistochemistry (IHC) to determine whether it correlated with reduced miR-181b expression. miR-181b expression was negatively correlated with mesenchymal markers, such as N-cadherin and Vimentin; however, miR-181b expression was positively correlated with the epithelial marker E-cadherin (Table 1) (Fig. 3E). We performed a univariate Cox regression analysis of the independent cohorts using clinical and genetic variables and demonstrated that miR-181b expression, the extent of tumor resection, the preoperative KPS score and the IDH1 mutation status were significantly associated with OS; however, the variables sex, age and MGMT promoter methylation status were not associated with OS (Table 2). A multivariate Cox regression analysis of the independent cohorts indicated that miR-181b was an independent prognostic factor (OS: HR, 0.264; 95% CI, 0.144–0.484; p < 0.001).
Restoring KPNA4 expression counteracted miR-181b overexpression
We tested whether KPNA4 was an important target of miR-181b in controlling cell proliferation and invasion and discovered that cell proliferation and invasion were increased in U87/miR-181b and LN229/miR-181b cells transfected with KPNA4 (Fig. 4A). As we confirmed, invasion and proliferation were inhibited by overexpressing miR-181b, which targeted KPNA4. To determine whether miR-181b could directly down-regulate KPNA4 expression and its downstream pathways, we transfected pReceiver-Lv105-KPNA4 into U87 and LN229 cells stably expressing miR-181b or miR-NC. KPNA4 downregulation by miR-181b was counteracted by the overexpression of KPNA4 as evidenced by Western blotting (Fig. 4B). Interestingly, the downregulation of N-cadherin and Vimentin, which was thought to be an indirect result of miR-181b overexpression, was rescued by the upregulation of KPNA4. Similarly, the overexpression of KPNA4 in miR-181b-treated cells down-regulated E-cadherin expression (Fig. 4B).
KPNA4 expression overcomes miR-181b-mediated EMT repression.
(A) Cell invasion and colony formation were inhibited by pReceiver-Lv105-KPNA4, which reversed the effects of miR-181b. (B) Western blotting indicated that glioma cells transfected with KPNA4 expressed higher N-cadherin and Vimentin. GAPDH was used as a control. *p < 0.05.
miR-181b suppressed glioma tumor growth in orthotopic models
An in vivo model was established to investigate the relationship between miR-181b and glioma growth. LN229 cells stably expressing miR-181b or miR-NC were injected stereotactically into the brains of nude mice. miR-181b-treated LN229 cells displayed a marked reduction of tumor volume (Fig. 5A). The KPNA4 protein levels were significantly lower in xenografts from miR-181b-expressing cells than in those from miR-NC control cells (Fig. 5B), confirming that miR-181b overexpression suppressed KPNA4 expression in vivo. Moreover, the expression levels of Ki-67, N-cadherin, Vimentin and MMP-9 decreased, whereas E-cadherin expression increased in the miR-181b group relative to the miR-NC group (Fig. 5C). To further evaluate the therapeutic effect of miR-181b on nude mice, the survival period of each group was analyzed by a Kaplan–Meier curve. The miR-181b mimic-treated group showed a significant improvement in survival compared with the control group until the end of the observation period. Only three mice in the miR-181b treatment group died by day 49, whereas no mice were alive in the miR-NC treatment group at this time point (p = 0.013) (Fig. 5D).
miR-181b overexpression suppressed tumor growth in vivo.
(A) Luminescence imaging was performed on miR-181b-treated LN229 tumors and scramble-treated controls. miR-181b mimic-treated cells displayed a marked reduction in tumor size. (B) Kaplan-Meier survival curves indicating that mice transfected with miR-181b showed a significantly better outcome than the miR-NC-treated group (p = 0.0138). (C) Real-time PCR revealed that miR-181b expression in the miR-181b-treated group was higher than that in the scramble-treated group. (D) Immunohistochemistry for Ki-67, KPNA4, E-cadherin, N-cadherin, Vimentin and MMP-9 was performed on orthotopic tumor sections. The error bars represent the means ± SD for three independent experiments. *p < 0.05.
Discussion
miR-181 family members have been reported to be tumor suppressors in glioma16,17,18,19,20, but there have been no reports on the role of the miR-181 family as a whole. The aberrant expression of the entire miR-181 family is highly correlated with the prognosis and progression of glioma. In the present study, we investigated the expression of the miR-181 family in two independent cohorts comprising nearly 700 glioblastoma patients. We determined that the expression of the miR-181 family was down-regulated in the mesenchymal subtype of GBM compared with other subtypes, suggesting that the miR-181 family exhibited subtype preferences. The increased expression of the miR-181 family conferred a better prognosis and could be utilized as a prognostic indicator. miR-181b was the most effective member of the miR-181 family according to the functional assays in U87 and LN229 cells. The pathway analyses indicated that miR-181b inhibited EMT by blocking KPNA4 expression, which limited glioma growth in vitro and in vivo.
From a biomarker perspective, our findings suggested novel discovery strategies. First, it is known that each miRNA modulates the expression of hundreds of gene transcripts21. The resulting phenotype is likely the aggregate effect of numerous altered transcripts. In this context, we postulated that the miR-181 family collectively regulated other gene transcripts that contributed to the prognostic or predictive value of the miR-181 family; it would be of great interest to identify these transcripts. It is interesting that KPNA4 was the common target of all four members of the miR-181 family according to the bioinformatic prediction and literature. This finding suggested a cooperative effect of the miR-181 family on the malignant progression of glioma. Sun et al. reported that miR-181b served as a potent regulator of downstream NF-κB signaling in the vascular endothelium by targeting KPNA4 and they identified the correlated pathways in glioma cell lines via functional assays15. However, the expression levels of the miR-181 family members in gliomas are not reported and their underlying contribution to survival remains to be further investigated.
KPNA4 (importin α3) and importin α4 are the primary importin α isoforms that mediate NF-κB p50/p65 heterodimer translocation into the nucleus22. KPNA4 directly binds to the nuclear localization sequences (NLSs) in NF-κB p50 and p65. The NF-κB dimer is held in an inactive state in the cytoplasm by an inhibitory protein (IKB) that masks the NLSs in the subunits. The NLSs in p50 and p65 must be unmasked to enable transport into the nucleus by KPNA4 and the dimers can then activate NF-κB-responsive genes. NF-κB is essential for the epithelial-mesenchymal transition (EMT)23; in many cancer types, the loss of E-cadherin coincides with a gain in the expression of the mesenchymal cadherin, N-cadherin. This “cadherin switch” is a hallmark of EMT24, which is considered to be the most important malignant process in glioma. The suppression of metastasis via the inhibition of NF-κB activity has been reported for numerous human cancers25,26,27. NF-κB activity is necessary for cells to remain in the mesenchymal state, whereas inhibiting NF-κB reverses EMT, resulting in viable and healthy epithelial cells. These findings provide mechanistic insights into the role of NF-κB in late-stage mammary tumorigenesis and metastasis. Several NF-κB target genes have been induced during EMT, such as those encoding various MMPs; in the present study, we determined that miR-181b overexpression reduced MMP-9 expression (Fig. 5C).
Materials and Methods
Patients and samples
Paired miRNA and mRNA profiling data were downloaded from the TCGA data portal (http://cancergenome.nih.gov) and utilized as a discovery set. A total of 491 TCGA glioblastoma samples were included in our study28. This method has been used previously to identify the integrated miR-181 family expression5. Gene expression subtypes were assigned using the Verhaak et al. 840-gene classifier. The dataset from the Chinese Glioma Genome Atlas (CGGA) (http://www.cgga.org.cn), which includes 189 glioblastoma samples (82 samples were used for miRNA microarrays and 107 samples were used for checking miR-181b expression), was obtained as a validation set. The microarray data set was deposited in the Gene Expression Omnibus (GEO) (accession number GSE25632) according to “minimum information about a microarray experiment” (MIAME) guidelines. Written informed consent was obtained from the patients for the publication of this report and any accompanying images. This study was performed with the approval of the Ethics Committee of Capital Medical University and was in compliance with the Helsinki Declaration. Methods were performed in accordance with the approved guidelines.
GSVA of miR-181 family expression
GSVA of miR-181 family expression was performed using the GSVA package in R. A list of NF-κB target genes was obtained from a published article (Supplementary Table 1)29. The EMT gene signature list was obtained from another article (Supplementary Table 2)30.
Real-time PCR and Western blotting
Real-time PCR and Western blotting were performed according to the manufacturer’s instructions as previously described31,32. The following primers were used: miR-181b sense, 5′-ACACTCCAGCTGGGAACATTCATTGCTGTCGG-3′; miR-181b anti-sense, 5′-TGGTGTCGTG GAGTCG-3′; U6 RT, 5′-TGGTGTCGTGGAGTCG-3′; U6 sense, 5′-CTCGCTTCGGCAGCACA-3′; U6 anti-sense, 5′-AACGCTTCACGAATTTGCGT-3′. qRT-PCR was performed using SYBR Premix DimerEraser (Takara, Dalian, China) on a 7900HT system. U6 levels were used as an internal control and fold changes were calculated by relative quantification (2−ΔΔCt). For Western blotting, the density of specific protein bands was quantified after normalization to the density of the GAPDH band in the same sample.
Fluorescence in situ hybridization, immunohistochemistry and cell biological assays
Fluorescence in situ hybridization (FISH) was utilized to detect miR-181b as previously described33. The colony formation and transwell invasion assays have been previously described34. Immunohistochemistry was performed using antibodies against E-cadherin (1:100 dilution; Cell Signaling Technology), N-cadherin (1:100 dilution; Cell Signaling Technology), Vimentin (1:250 dilution; Abcam), Ki67 (1:200 dilution; Abcam) and MMP9 (1:100 dilution; Abcam). The degree of immunostaining of sections was viewed and scored separately by two independent investigators. The scores were determined by combining the proportion of positively stained tumor cells and the intensity of staining. The proportion of positively stained tumor cells was graded as follows: 0, no positive tumor cells; 1, < 5% positive tumor cells; 2, 5–20% positive tumor cells; and 3, > 20% positive tumor cells. The intensity of staining was recorded on a scale of 0 (no staining), 1 (weak staining, light yellow), 2 (moderate staining, yellowish brown), or 3 (strong staining, brown). The staining index was calculated as follows: staining index = staining intensity x proportion of positively stained tumor cells. High expression was defined as a staining index score ≥4 and low expression was defined as a staining index <4.
Luciferase reporter assay
For the luciferase reporter assay, the 3′ UTR of KPNA4 was PCR-amplified from human cDNA. To create a mutant version, the sequence complementary to binding site 1 in the 3′ UTR of miR-181b (AATGAATGA) was replaced with ACGCGGACA and site 2 (ATGAATGT) was replaced with GACGGACA. The PCR products were digested with SacI and HindIII and inserted into the pMIR-REPORTER vector. The constructs were validated by sequencing. Cells were seeded into a 24-well plate and co-transfected with the wild-type or mutated KPNA4 3′UTR reporter plasmid, pRL-TK and miR-181b or miR-NC. Luciferase assays were performed 24 h after transfection using the Dual Luciferase Reporter Assay System (Promega, Madison, WI, USA).
Oligonucleotides and cell transfection
miR-181a, miR-181b, miR-181c and miR-181d mimics and KPNA4 wild-type and mutant plasmids were chemically synthesized by GenePharma Co., Ltd. (Shanghai, China). Cells at 50–70% confluence were transfected using Lipofectamine (Invitrogen, Carlsbad, CA, USA). Oligonucleotides were transfected into U87 and LN229 glioma cells (purchased from the Institute of Biochemistry and Cell Biology, Chinese Academy of Science, Shanghai, China) at a final concentration of 50 nM as previously described35.
Lentivirus packaging and establishing stable cell lines
A lentiviral packaging kit was purchased from Open Biosystems (Huntsville, AL, USA). Lentiviruses carrying hsa-miR-181b or hsa-miR-negative control (miR-NC) were packaged according to the manufacturer’s protocol. Stable U87 and LN229 cell lines were established by lentiviral infection and puromycin selection. The packaging steps were performed as previously described31.
Orthotopic nude mouse models and treatment
A nude mouse tumor xenograft model was established as previously described36,37. BALB/c-A nude mice (4 weeks old) were purchased from the Animal Center at the Cancer Institute of the Chinese Academy of Medical Science. Twelve mice were randomly divided into two groups. To establish intracranial gliomas, 0.5 × 105 LN229 glioblastoma cells transduced with luciferase lentivirus and stably expressing miR-181b or miR-NC were implanted stereotactically38. Overall survival time of the mice was monitored. After being observed for 50 days, the mice bearing xenograft tumors were sacrificed; the tumor tissues were removed and fixed in formalin and paraffin-embedded sections were prepared for immunohistochemical analysis.
Statistical analysis
We applied prediction analysis of microarrays (PAM) as previously reported for the molecular subtype annotation of the four datasets39. The integrated miR-181 family expression for predicting survival was developed based on a linear combination of miRNA expression weighted by the regression coefficient derived from the univariate Cox regression analysis. Quantitative results are presented as the means ± standard deviation. The differences in miR-181 family expression in the microarray data were analyzed by Student’s t-test. The survival curves for patients with high or low expression of the miR-181 family were calculated using the Kaplan-Meier method and the differences were analyzed via the two-sided log-rank test. A p value <0.05 was considered to be statistically significant. All the data analyses were performed in GraphPad Prism and R.
Additional Information
How to cite this article: Wang, H. et al. Upregulation of miR-181s reverses mesenchymal transition by targeting KPNA4 in glioblastoma. Sci. Rep. 5, 13072; doi: 10.1038/srep13072 (2015).
References
Calin, G. A. & Croce, C. M. MicroRNA signatures in human cancers. J. Nat Rev Cancer. 6, 857–866 (2006).
Volinia, S. et al. A microRNA expression signature of human solid tumors defines cancer gene targets. J. Proc Natl Acad Sci USA. 103, 2257–2261 (2006).
Lim, L. P. et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. J. Nature. 433, 769–773 (2005).
Ohgaki, H. et al. Genetic pathways to glioblastoma: a population-based study. J. Cancer Res. 64, 6892–6899 (2004).
Verhaak, R. G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR and NF1. J. Cancer Cell. 17, 98–110 (2010).
Marcucci, G. et al. Prognostic significance of and gene and microRNA expression signatures associated with, CEBPA mutations in cytogenetically normal acute myeloid leukemia with high-risk molecular features: a Cancer and Leukemia Group B Study. J. J Clin Oncol. 26, 5078–5087 (2008).
Shi, L. et al. hsa-mir-181a and hsa-mir-181b function as tumor suppressors in human glioma cells. J. Brain Res. 1236, 185–193 (2008).
Chen, G. et al. MicroRNA-181a sensitizes human malignant glioma U87MG cells to radiation by targeting Bcl-2. J. Oncol Rep. 23, 997–1003 (2010).
Visone, R. et al. miR-181b is a biomarker of disease progression in chronic lymphocytic leukemia. J. Blood. 118, 3072–3079 (2011).
Zhi, F. et al. The use of hsa-miR-21, hsa-miR-181b and hsa-miR-106a as prognostic indicators of astrocytoma. J. Eur J Cancer. 46, 1640–1649 (2010).
Schwind, S. et al. Prognostic significance of expression of a single microRNA, miR-181a, in cytogenetically normal acute myeloid leukemia: a Cancer and Leukemia Group B study. J. J Clin Oncol. 28, 5257–5264 (2010).
Slaby, O. et al. MicroRNA-181 family predicts response to concomitant chemoradiotherapy with temozolomide in glioblastoma patients. J. Neoplasma. 57, 264–269 (2010).
Schaefer, A. et al. Diagnostic and prognostic implications of microRNA profiling in prostate carcinoma. J. Int J Cancer. 126, 1166–1176 (2010).
Marcucci, G. et al. MicroRNA expression in cytogenetically normal acute myeloid leukemia. J. N Engl J Med. 358, 1919–1928 (2008).
Sun, X. et al. MicroRNA-181b regulates NF-kappaB-mediated vascular inflammation. J. J Clin Invest. 122, 1973–1990 (2012).
Li, P. et al. MiR-181b suppresses proliferation of and reduces chemoresistance to temozolomide in U87 glioma stem cells. J. J Biomed Res. 24, 436–443 (2010).
Wang, J., Sai, K., Chen, F. R. & Chen, Z. P. miR-181b modulates glioma cell sensitivity to temozolomide by targeting MEK1. J. Cancer Chemother Pharmacol. 72, 147–158 (2013).
Lorenz, M. P. et al. Ethene adsorption and dehydrogenation on clean and oxygen precovered Ni(111) studied by high resolution x-ray photoelectron spectroscopy. J. J Chem Phys. 133, 014706 (2010).
Conti, A. et al. miR-21 and 221 upregulation and miR-181b downregulation in human grade II-IV astrocytic tumors. J. J Neurooncol. 93, 325–332 (2009).
Zhang, W. et al. miR-181d: a predictive glioblastoma biomarker that downregulates MGMT expression. J. Neuro Oncol. 14, 712–719 (2012).
Hendrickson, D. G. et al. Concordant regulation of translation and mRNA abundance for hundreds of targets of a human microRNA. J. PLoS Biol. 7, e1000238 (2009).
Fagerlund, R., Kinnunen, L., Kohler, M., Julkunen, I. & Melen, K. NF-{kappa}B is transported into the nucleus by importin {alpha}3 and importin {alpha}4. J. J Biol Chem. 280, 15942–15951 (2005).
Huber, M. A. et al. NF-kappaB is essential for epithelial-mesenchymal transition and metastasis in a model of breast cancer progression. J. J Clin Invest. 114, 569–581 (2004).
Lehembre, F. et al. NCAM-induced focal adhesion assembly: a functional switch upon loss of E-cadherin. J. EMBO J. 27, 2603–2615 (2008).
Huang, S., Pettaway, C. A., Uehara, H., Bucana, C. D. & Fidler, I. J. Blockade of NF-kappaB activity in human prostate cancer cells is associated with suppression of angiogenesis, invasion and metastasis. J. Oncogene. 20, 4188–4197 (2001).
Huang, S., DeGuzman, A., Bucana, C. D. & Fidler, I. J. Nuclear factor-kappaB activity correlates with growth, angiogenesis and metastasis of human melanoma cells in nude mice. J. Clin Cancer Res. 6, 2573–2581 (2000).
Andela, V. B., Schwarz, E. M., Puzas, J. E., O’Keefe, R. J. & Rosier, R. N. Tumor metastasis and the reciprocal regulation of prometastatic and antimetastatic factors by nuclear factor kappaB. J. Cancer Res. 60, 6557–6562 (2000).
Zhang, W. et al. Whole-genome microRNA expression profiling identifies a 5-microRNA signature as a prognostic biomarker in Chinese patients with primary glioblastoma multiforme. J. Cancer. 119, 814–824 (2013).
Feuerhake, F. et al. NFkappaB activity, function and target-gene signatures in primary mediastinal large B-cell lymphoma and diffuse large B-cell lymphoma subtypes. J. Blood. 106, 1392–1399 (2005).
Zarkoob, H., Taube, J. H., Singh, S. K., Mani, S. A. & Kohandel, M. Investigating the link between molecular subtypes of glioblastoma, epithelial-mesenchymal transition and CD133 cell surface protein. J. PLoS One. 8, e64169 (2013).
Shi, Z. M. et al. MiRNA-181b suppresses IGF-1R and functions as a tumor suppressor gene in gliomas. J. RNA. 19, 552–560 (2013).
Qian, X. et al. Sequence-dependent synergistic inhibition of human glioma cell lines by combined temozolomide and miR-21 inhibitor gene therapy. J. Mol Pharm. 9, 2636–2645 (2012).
Liu, Y. et al. MiR-218 reverses high invasiveness of glioblastoma cells by targeting the oncogenic transcription factor LEF1. J. Oncol Rep. 28, 1013–1021 (2012).
Wang, H., Wang, Y. & Jiang, C. Stromal protein periostin identified as a progression associated and prognostic biomarker in glioma via inducing an invasive and proliferative phenotype. J. Int J Oncol. 42, 1716–1724 (2013).
Tong, M. et al. Rab25 is a tumor suppressor gene with antiangiogenic and anti-invasive activities in esophageal squamous cell carcinoma. J. Cancer Res. 72, 6024–6035 (2012).
Zhang, C. Z. et al. MiR-221 and miR-222 target PUMA to induce cell survival in glioblastoma. J. Mol Cancer. 9, 229 (2010).
Zhang, J. et al. AKT2 expression is associated with glioma malignant progression and required for cell survival and invasion. J. Oncol Rep. 24, 65–72 (2010).
Shi, Z. et al. AC1MMYR2, an inhibitor of Dicer-mediated biogenesis of oncomir miR-21, reverses epithelial-mesenchymal transition and suppresses tumor growth and progression. J. Cancer Res. 73, 5519–5531(2013).
Yan, W. et al. Molecular classification of gliomas based on whole genome gene expression: a systematic report of 225 samples from the Chinese Glioma Cooperative Group. J. Neuro Oncol. 14, 1432–1440 (2012).
Acknowledgements
This work was supported by grants from the National High Technology Research and Development Program of China (863) (No. 2012AA02A508), the International Cooperation Program (No. 2012DFA30470), the National Natural Science Foundation of China (Nos. 81201993, 81402052 and 81101901).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: C.J. and T.J. Performed the experiments: H.W., T.T., Y.F., Y.W., J.C. and W.Y. Analyzed the data: H.W. Contributed reagents/materials/analysis tools: W.Y. and J.C. Wrote the paper: H.W., Y.W. and Y.P.Y. All authors contributed significantly and all authors are in agreement with the content of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wang, H., Tao, T., Yan, W. et al. Upregulation of miR-181s reverses mesenchymal transition by targeting KPNA4 in glioblastoma. Sci Rep 5, 13072 (2015). https://doi.org/10.1038/srep13072
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep13072
- Springer Nature Limited
This article is cited by
-
Ginsenoside Rg3 inhibits osteosarcoma progression by reducing circ_0003074 expression in a miR-516b-5p/KPNA4-dependent manner
Journal of Orthopaedic Surgery and Research (2021)
-
Potential role of microRNAs as biomarkers in human glioblastoma: a mini systematic review from 2015 to 2020
Molecular Biology Reports (2021)
-
LncRNA ST7-AS1, by regulating miR-181b-5p/KPNA4 axis, promotes the malignancy of lung adenocarcinoma
Cancer Cell International (2020)
-
MiR-596 activated by EP300 controls the tumorigenesis in epithelial ovarian cancer by declining BRD4 and KPNA4
Cancer Cell International (2020)
-
miR-504 suppresses mesenchymal phenotype of glioblastoma by directly targeting the FZD7-mediated Wnt–β-catenin pathway
Journal of Experimental & Clinical Cancer Research (2019)