Abstract
Lactobacillus delbrueckii subsp. bulgaricus is one of the most widely used starter culture strains in industrial fermented dairy manufacture. It is also common in naturally fermented dairy foods made using traditional methods. The subsp. bulgaricus strains found in naturally fermented foods may be useful for improving current industrial starter cultures; however, little is known regarding its genetic diversity and population structure. Here, a collection of 298 L. delbrueckii strains from naturally fermented products in Mongolia, Russia and West China was analyzed by multi-locus sequence typing based on eight conserved genes. The 251 confirmed subsp. bulgaricus strains produced 106 unique sequence types, the majority of which were assigned to five clonal complexes (CCs). The geographical distribution of CCs was uneven, with CC1 dominated by Mongolian and Russian isolates and CC2–CC5 isolates exclusively from Xinjiang, China. Population structure analysis suggested six lineages, L1–L6, with various homologous recombination rates. Although L2–L5 were mainly restricted within specific regions, strains belonging to L1 and L6 were observed in diverse regions, suggesting historical transmission events. These results greatly enhance our knowledge of the population diversity of subsp. bulgaricus strains and suggest that strains from CC1 and L4 may be useful as starter strains in industrial fermentation.
Similar content being viewed by others
Introduction
Lactobacillus delbrueckii subsp. bulgaricus, a Gram-positive, rod-shaped, non-motile and non-spore-forming lactic acid bacterium (LAB), is one of the most common starter bacterial species in industrial fermentation of dairy products. Globally, almost all types of yogurt and a considerable proportion of other types of dairy products contain this bacterium. It is also an important component in the traditional production of naturally fermented dairy products, which generally use bacteria obtained from the local environment1. Previous reports have shown that many different LAB coexist during fermentation of traditional dairy products, contributing to their special flavor and texture1,2,3,4. Although the dominant species of LAB varies in different dairy products (e.g. Lactobacillus helveticus, Lactobacillus kefiranofaciens and L. delbrueckii were the predominant LAB species in airag, khoormog and tarag, respectively4), L. delbrueckii subsp. bulgaricus was identified as the predominant bacterial strain in certain types of naturally fermented products in China and Mongolia1,2.
Naturally fermented dairy products have a long history in China, Mongolia and Russia. A book titled A New Account of Tales of the World (Shi Shuo Xin Yu) was published in the Eastern Jin Dynasty in China (AD 317–420), documenting the production of a fermented dairy product named “lao”. Nomadic people across large rural areas of these countries still use traditional methods to produce fermented dairy products. These products have tremendous potential as sources of novel subsp. bulgaricus strains for improving industrial starter cultures. Improving our understanding of the diversity and phylogenetic relationships of subsp. bulgaricus isolates from naturally fermented products would provide insights into the evolutionary patterns of this subspecies and would support more effective selection of the next generation of starter strains for industrial applications.
Different molecular methods have been reported for the identification and typing of L. delbrueckii, including randomly amplified polymorphic DNA analysis5, sodium dodecyl sulfate polyacrylamide gel electrophoresis6, restriction fragment length polymorphism analysis7, amplified fragment length polymorphism analysis8 and multi-locus sequence typing (MLST)9,10,11. While all of these methods can distinguish subsp. bulgaricus from other subspecies, including subsp. delbrueckii, subsp. lactis12, subsp. indicus8, subsp. sunkii10 and subsp. jakobsenii11, MLST is generally the preferred method because of its convenience and higher resolution and the ease of data interchange between global laboratories13. However, only three MLST studies associated with subsp. bulgaricus have been published and all employ a limited number of strains. Therefore, our knowledge of the genetic diversity and population structure of subsp. bulgaricus strains still remains limited.
In this study, we collected 298 L. delbrueckii strains (287 strains identified by 16S rRNA sequencing obtained from naturally fermented dairy foods in traditional pasturing areas and 11 strains retrieved from the NCBI GenBank database), with subsp. bulgaricus accounting for 84% of isolates. We calculated the diversity and predicted the population structure of the strains based on a newly developed MLST scheme. From these results, we inferred the geographical distribution and recombination characteristics of different lineages in an attempt to provide information that will be useful in starter strain selection in the fermentation industry.
Results
MLST assay for identifying L. delbrueckii subsp. bulgaricus
A total of 298 strains were analyzed using the amplified fragments of eight housekeeping genes (clpX, dnaA, groEL, murE, pheS, pyrG, recA, and rpoB) (Supplementary Table S1). All gene fragments were successfully amplified from all strains tested. Our eight-gene assay shared a single locus, pheS, with a previously reported three-gene MLST assay9 and shared three loci (groEL/hsp60, pyrG, recA) with a seven-locus scheme10.
The dN/dS ratio is a common indicator of selection pressure, with dN/dS > 1 indicating positive selection and dN/dS < 1 indicating purifying selection for the gene sequence tested14. For each of the eight loci, the calculated dN/dS ratios were all less than one, especially for recA, where dN/dS = 0 (no nonsynonymous single nucleotide polymorphisms (SNPs) were found) (Table 1). This indicated that all loci were under strong purifying selection, which is optimal for MLST loci15.
Definition of L. delbrueckii subsp. bulgaricus
All isolates included in this study were previously identified as L. delbrueckii by 16S RNA gene sequencing (data not shown). However, with poor distinguishing power, 16S RNA-based typing methods cannot accurately identify strains to the subspecies level. Therefore, we included the sequences of type strains of each subspecies (subsp. bulgaricus: ATCC 11842T, subsp. delbrueckii: JCM 1012T, subsp. lactis: DSM20072T, subsp. indicus : DSM15966T, subsp. jakobsenii: ZN7a-9T, subsp. sunkii: DSM24966T, Table S2)8,11,16,17 in the analysis as reference strains to more accurately identify isolates belonging to subsp. bulgaricus.
The maximum likelihood tree based on the concatenated sequences of the eight loci showed that the six type strains formed distinct lineages (Supplementary Fig. S1). Most of the tested strains (249 of 298) grouped together with the subsp. bulgaricus type strain (ATCC 11842T) to form a distinct branch. Forty-five strains were genetically distant from subsp. bulgaricus and were more closely related to the other five type strains. Three of the strains (IMAU90010, IMAU90013 and IMAU94251) were located at an intermediate position between the subsp. bulgaricus lineage and the other lineages. A neighbor-joining tree was then generated for each locus separately and again, 249 of the strains clustered with the subsp. bulgaricus type strain (Supplementary Fig. S2). Only six, four and three loci supported the grouping of the three intermediate strains, IMAU90010, IMAU90013 and IMAU94251, respectively, with subsp. bulgaricus. The sequences of the remaining loci were more closely related to other subspecies, which may indicate inter-lineage homologous recombination within the species. As supported by the majority of the loci, we included the strain IMAU90010 as subsp. bulgaricus in further analyses.
Therefore, 251 strains were identified as subsp. bulgaricus in this study. Among the 246 subsp. bulgaricus isolates obtained from dairy products, 190 (77.2%) were from fermented cow milk and 45 (18.3%) were from fermented yak milk (Supplementary Table S2). Most of these strains were collected from Asia (China: 107, Mongolia: 117, Middle-South Russia: 19, Japan: 1), with the remaining strains originating from Denmark (NBIMCC1381), France (ATCC BAA-365), North America (CNCM1519, CNCM1632) and Bulgaria (NBIMCC1273, NBIMCC285 and ATCC11842T) (Supplementary Table S2).
Genetic diversity of subsp. bulgaricus
We aligned the sequences for each strain at each MLST locus and confirmed that neither indels nor premature stop codons were present in any of the sequences. The DNA fragments ranged from 470 bp (groEL) to 603 bp (rpoB) and the concatenated sequences produced a 4,261-bp fragment. Amongst the 298 L. delbrueckii strains, 20 (dnaA) to 27 (groEL and pyrG) alleles per locus were identified, corresponding to 119 STs. For the 251 subsp. bulgaricus strains, the alleles numbers varied from 12 (dnaA) to 20 (groEL) and 106 STs were assigned (Table 1). There were 113 polymorphic sites amongst the subsp. bulgaricus sequences, accounting for only half of the variation (224 sites) observed for the species L. delbrueckii. The nucleotide diversity (π, the average number of nucleotide differences per site between two randomly selected isolates) was also calculated. Within subsp. bulgaricus, the π values ranged from 0.0038 (dnaA) to 0.0069 (pyrG), while at the species level, the range was from 0.0056 (rpoB) to 0.0109 (clpX). Taking the eight loci together, the π value was 0.0030 for the subsp. bulgaricus lineage and 0.0042 for the whole species. Overall, although lower than for the whole species, subsp. bulgaricus still displayed a high level of nucleotide and allelic diversity.
Population snapshot
The 251 subsp. bulgaricus strains were assigned 106 different STs, with the allele profiles listed in Supplementary Table S2. According to eBURST analysis, the 251 strains could be assigned into five clonal complexes (CC1–CC5, 168 strains), seven doubletons (16 strains) and 53 singletons (67 strains) (Fig. 1). CC1, containing almost half of all strains (n = 123), was the largest CC and consisted of 19 STs. ST10, with eight single-locus variants (SLVs), was defined as the group founder. ST7 (accounting for the largest number of strains among the 106 STs) and ST13 were predicted to be subgroup founders of CC1. CC2 consisted of 18 strains within nine STs. The group founder, ST41, had five SLVs. CC3 included six strains with five STs (Supplementary Table S2). CC4 and CC5 contained 14 and seven strains, respectively, each with three STs. About one-third of strains (33%, singletons and doubletons) appeared to be less related to each other and the rest of the population.
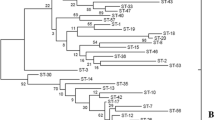
Population snapshot generated by eBURST based on the allelic profiles of 251 subsp. bulgaricus isolates.
Each dot indicates an individual sequence type (ST), with the size of the dot representing the number of strains. The numbers associated with the dots represent the assigned STs. Solid lines link single locus variants, i.e. STs identical at seven of the eight loci, to constitute the clonal complex.
The whole population contains six lineages
As shown in Supplementary Fig. S1, low bootstrap values on many branches of the maximum likelihood tree indicated uncertain phylogenetic relationships and the possible existence of homologous recombination. To identify sub-populations of subsp. bulgaricus that may have been affected by recombination, a linkage model generated by Structure analysis was applied to the concatenated sequences. The results of triple runs with K values from 3–20 were compared and K = 6 was selected for use in further analyses. This suggested that the collected subsp. bulgaricus strains were descendants of six distinct ancestor populations. The rebuilding of population structure showed that the 106 STs fell into six distinct lineages (L1–L6), with little intersection of ancestral sources. The STs within each lineage tended to be highly homogenous (Fig. 2a).
Population structure of subsp. bulgaricus based on concatenated sequences of the eight MLST loci.
(a) Population structure inferred by Structure analysis. Each bar represents one of 106 sequence types (STs). Different colors indicate the predicted ancestor populations of subsp. bulgaricus as inferred by Structure analysis (assuming K = 6 and applying the linkage model). The proportion of the colors for each ST represents the ancestors’ contribution to the ST. Based on this, six lineages (L1–L6) are predicted and given boundaries by the vertical gray lines. (b) Split network inferred by SplitsTree. Except for four STs, the six branches indicated by the black arcs are consistent with L1–L6 defined according to Structure analysis.
To verify the population structure of the six lineages, we generated a split graph using SplitsTree based on the concatenated sequences of the 106 STs. Except for four STs belonging to L6 (ST43, ST48, ST57, ST67) that were closer to L5 in the split graph, the network-like structure with six branches was largely consistent with the six lineages defined by Structure analysis (Fig. 2b). Therefore, although a minority of intermediate types might exist, both Structure and SplitsTree analysis results supported the hypothesis that subsp. bulgaricus strains could be separated into six major lineages.
The lineages show various recombination levels
The phi-test was used to give a statistical view of recombination18. The P value generated for all 106 STs was 0.0018 (Table 2), indicating significant evidence of recombination across the whole subspecies. However, P values for the six individual lineages varied, ranging from 0.0048–1 (Table 2). Only L3 and L6 showed significant intra-lineage recombination (both P values were <0.05). Correspondingly, higher ρ/θ ratios (the number of recombination events versus mutation events) were observed in L3 and L6 (39.969 and 17.587, Table 2), indicating recombination events occurred more frequently within these lineages.
The standardized index of association (IAS)19 was also calculated to measure the homologous recombination by estimating the linkage disequilibrium between the eight loci of each lineage (Table 2). In addition to lineages L3 and L6, the IAS value for L4 also suggested a tendency towards frequent recombination between alleles in this lineage. The IAS values for L1, L2 and L5 indicated significant linkage disequilibrium among the alleles, suggesting clonal relationships might dominate within these lineages.
Population structure in relation to geographical distribution
An unbalanced distribution of the six lineages of subsp. bulgaricus was observed among the sampling regions, with three lineages associated with a specific region, i.e. L2 in QSG of China, L3 in Tibet and L5 in Xinjiang (Fig. 3). This geographical clustering may suggest possible adaptive advantages for these lineages in the local region. Comparatively, except for three STs isolated from QSG of China, approximately half of STs of L1 were isolated from Mongolia, while the other half of the lineage was from Tibet. A similar composition was also observed for L6, with half of the STs from Mongolia and the other half from Xinjiang, China. Therefore, transfer of the strains over long distances may have occurred during the evolutionary history of subsp. bulgaricus.
Geographical distribution of clonal complexes (CCs) and lineages of subsp. bulgaricus.
Each bar represents one of the 106 sequence types (STs), of which the colors and height indicate the region of isolation and numbers of isolates, respectively. ST type and CC distribution (red stars for doubletons and blanks for singletons) are indicated below the bars. The gray shades indicate the six lineages defined by Structure analysis. “QSG of China” refers to Qinghai, Sichuan and Gansu provinces of China and “Other location” indicates Japan (ST1), Denmark and France (ST3), North America (ST4, ST5) and Bulgaria (ST2, ST110, ST111).
Interestingly, the distribution of the CCs also showed geographical dependency (Supplementary Fig. S3). Strains of the largest clonal complex, CC1, were all collected from Mongolia and Russia, except for two strains each from Xinjiang and QSG of China. The remaining four CCs only contained strains isolated from Xinjiang, China. Except for one strain from CC1, the strains from QSG of China formed four doubletons and seven singletons. Interestingly, all 28 STs (31 strains) from Tibet were singletons, although they were collected from a limited geographical area (Supplementary Table S2).
Discussion
This study has greatly enhanced our knowledge of the genetic diversity of L. delbrueckii subsp. bulgaricus. We identified 106 STs and six lineages amongst the strains examined, while a previous study identified 15 STs and two lineages among 25 strains9. Our results also expand the known nucleotide diversity of the species L. delbrueckii, with increased π values compared with previous research observed for three loci (pyrG, 0.0090 vs. 0.0068; groEL/hsp60, 0.0081 vs. 0.0066; recA, 0.0079 vs. 0.0066)10. The MLST assay developed here may be useful in further determining the population structure of subsp. bulgaricus, but also in distinguishing subsp. bulgaricus from the other subspecies of L. delbrueckii. However, as a limited number of strains from non-bulgaricus subspecies were examined in this study, the typing efficiency of this assay for other subspecies of L. delbrueckii should be examined in further work.
Mongolia has one of the largest livestock husbandry industries in the world and is famous for its dairy products. In this research, we collected subsp. bulgaricus strains from 13 of the 22 Mongolian provinces. The majority of the strains collected from Mongolia (101 of 117, 86%) belonged to CC1 (Fig. 3), suggesting that strains of this clonal complex had selective advantages in producing naturally fermented dairy foods in Mongolia. CC1 also had the richest diversity, with 19 STs and 123 strains and was found across a much wider area, including South-west Russia (regions adjoining Mongolia), Xinjiang and QSG of China, implying strong adaptability in different niches by these strains. Although any genetic mechanisms accounting for this selective advantage still need to be illuminated, probably by whole genome sequencing, strains of CC1 could potentially be used for improving current industrial starter cultures. Additionally, all strains from industrial dairy products or their descendants (ATCC BAA-365, 2038, CNCM I-1519, CNCM I-1632, NBIMCC1273, NBIMCC1381)20,21,22 were grouped in L4. Any of the other isolates in this lineage would likely display similar characters in fermentation of dairy products and could therefore be another potential source of new starter culture strains. It is notable that many lactobacilli coexist in the natural fermented dairy foods and both the species composition and their relative abundance contribute to the special flavor and texture. Therefore, although CC1 and L4 might be used as potential references for improving starter cultures, it should be considering the impact of the other co-existed species and the relative abundance, which could be delineated by meta-genomics analysis on the bacteria community in natural fermented dairy foods.
Xinjiang appears to have a particular richness of subsp. bulgaricus strains, as all five CCs were found there and CC2–5 were exclusively isolated from this region. It was reported recently that the oldest known pieces of cheese were discovered in the tombs of an early Bronze Age cemetery in Xinjiang23,24. Accordingly, the production of fermented foods in Xinjiang can be traced back at least 3800 years. Xinjiang is also an important hub that has bridged human commercial activities between Eastern and Western societies for several thousands of years. Ancient traders may have brought fermented dairy products and hence subsp. bulgaricus strains, from distant regions to Xinjiang, which promoted the spread and enriched the population diversity of the strains in this region. Therefore, the long history of fermented dairy food production, along with ancient human activity, might contribute to the higher number of subsp. bulgaricus CCs in Xinjiang, China. Comparatively, only 31 isolates consisting of 28 singletons and no clonal complexes were identified in Tibet. This restriction of clonal expansion might be attributed to the harsh environment of the plateau, which would suppress the growth and transmission of subsp. bulgaricus strains. Therefore, it would be interesting to explore the mechanisms of adaptive evolution of subsp. bulgaricus in different regions by deciphering the association between specific genomic features and environmental factors in their sampling regions.
Conclusion
In this study, we designed an eight-locus MLST scheme and identified 251 subsp. bulgaricus isolates from a collection of 298 L. delbrueckii strains. In total, 106 STs, five CCs and six lineages were identified amongst the subsp. bulgaricus strains, with different recombination rates and geographical distribution patterns observed across the lineages. This study improved our understanding of the genetic diversity of this important industrial bacterium. Further work based on whole genome sequence analysis would provide more subtle evolutionary details on subsp. bulgaricus and provide insight into genetic mechanisms involved in the natural fermentation of dairy products. Together, this information will assist in improving bacterial starter cultures used in industrial production of dairy products.
Materials and Methods
Bacterial strains and DNA extraction
A total of 298 L. delbrueckii strains were analyzed in this study, including six type strains. Except for the L. delbrueckii subsp. sunkii type strain DSM24966T (obtained from the Deutsche Sammlung von Mikroorganismen und Zellkulturen), completed whole genome sequences are available from the NCBI website (http://www.ncbi.nlm.nih.gov/genome/genomes/514) for all type strains, as well as for five other subspecies of L. delbrueckii. Details are listed in Supplementary Table S2. The strains included 284 isolates from the Collection Centre of Lactic Acid Bacteria (LABCC) of the Inner Mongolia Agriculture University, China and three isolates derived from the National Bank for Industrial Microorganisms and Cell Cultures (NBIMCC) in Bulgaria. The study included strains isolated from naturally fermented dairy products made by nomads in the autonomous regions of Xinjiang and Tibet and from the Qinghai, Sichuan and Gansu provinces of China (the latter three provinces were adjacent to each other and designated “QSG of China” in this paper). Strains were also isolated from 13 provinces in Mongolia and from Respublika Buryatiya, Kalmykia and Tuwa, Russia (Supplementary Table S2), from 2005–2013. The three NBIMCC strains were isolated from yogurt starters used in Bulgaria and Denmark. The 284 LABCC strains were analyzed by 16S rRNA gene sequencing and identified as L. delbrueckii species. Detailed strain information is provided in Supplementary Table S2.
All 287 L. delbrueckii strains (LABCC: 284, NBIMCC: 3), as well as DSM24966T, were maintained in MRS broth (OXOID, Germany) at 37 °C for 18–24 h. Cells were harvested by centrifugation at 3000 rpm for 5 min and cell pellets were used for total genomic DNA extraction as described previously25. Purified DNA was diluted to a final concentration of 100 ng/μL for further applications.
Selection of MLST loci and primer design
Selection of the target housekeeping genes was based on the presence of SNPs between L. delbrueckii subsp. bulgaricus ATCC 11842T (GenBank accession no. NC_008054) and L. delbrueckii ND02 (NC_014727) (Data not shown). We initially selected 12 loci for the MLST scheme, but four loci were abandoned because of amplification difficulties or low sequence quality. As a result, eight housekeeping genes encoding the following proteins were chosen for analysis: ATP-dependent protease ATP-binding subunit ClpX (clpX), chromosomal replication initiation protein (dnaA), CTP synthetase (pyrG), chaperonin GroEL (groEL), UDP-N-acetylmuramoylalanyl-D-glutamate-L-lysine ligase (murE), phenylalanyl-tRNA synthetase subunit alpha (pheS), recombinase A (recA) and DNA-directed RNA polymerase subunit beta (rpoB). All loci were present as a single copy, shared a conserved sequence, were widely distributed across the chromosome and were mutually unlinked in location26. Primers to amplify the eight loci were designed using Primer Premier 5.0 (PREMIER Biosoft) based on the known genome sequence of L. delbrueckii subsp. bulgaricus ATCC 11842 T. Primer sequences are listed in Supplementary Table S1.
PCR amplification and sequencing
Genomic DNA from each strain was used as a template for amplification of MLST loci using a Veriti 96-Well Thermal Cycler (Applied Biosystems). For each target, a 50-μL PCR mixture was prepared, containing 150 ng of genomic DNA, 10 mM of each dNTP, 10 pmol of each primer, 2.5 U of Taq polymerase and 1 × PCR buffer (with Mg2+). Parameters for the thermal cycler were: 94 °C for 4 min; 30 cycles of 94 °C for 1 min, 55–60 °C (optimal annealing temperatures for each locus are listed in Supplementary Table S1) for 45 s and 72 °C for 1 min; and a final extension of 72 °C for 7 min. PCR products were electrophoresed in a 1.0% agarose gel. PCR products were sequenced by the Shanghai Majorbio Bio-pharm Technology Corporation using the same primers as for PCR.
Diversity analyses
Both forward and reverse sequences for each strain were trimmed, aligned and analyzed using BioNumerics software (version 6.6, Applied Maths). Allele definition was also performed with it. For each MLST locus, the sequences obtained for all isolates were compared and each unique sequence was assigned with an allele number. Unique sequences were defined as those differing from the others at one or more nucleotides. Each isolate was unambiguously defined by an allele profile or sequence type (ST) derived from the combination of numbers corresponding to the alleles at each of the loci analyzed. The same ST was assigned to multiple strains when they shared the same allelic profile. Allelic profiles were used for subsequent analysis.
The mean GC content of the DNA along with the dN/dS ratios (where dS is the number of synonymous substitutions per synonymous site and dN is the number of nonsynonymous substitutions per nonsynonymous site) were calculated using START v2.027. Nucleotide diversity per site (π) was estimated using DnaSP version 5.028.
Recombination analyses
The phi-test for recombination, based on individual loci from the whole strain collection, was performed using SplitsTree v4.12.618. The P-value (a value of P < 0.05 was considered significant herein) indicated the DNA regions exhibiting the strongest evidence of mosaicism. The LDhat program29,30 of the RDP4 program31 was used to calculate per-site ρ/θ ratios based on concatenated sequences of the eight loci with 1,000,000 Markov Chain Monte Carlo (MCMC) updates. The parameters ρ and θ represented the rates of recombination and mutation respectively. The standardized index of association (IAS) was calculated using the lian program v3.619.
Phylogenetic analyses
Phylogenetic trees were constructed from the concatenated sequences (4261 bp) using the maximum likelihood method and the HKY97 model in PhyML32 with 1000 replications.
Population structure analyses
eBURST33 was used to analyze population structure. STs were clustered into CCs based on the similarity of their allelic profiles. According to the grouping criterion, only one of the eight loci analyzed (SLVs) was allowed for intra-CC variation. The split decomposition method was used to assess the degree of tree-like structure for alleles of each locus and all STs using SplitsTree v4.12.618,34. According to the algorithm, a tree-like structure indicates a clonal descent, while an interconnecting network or a parallelogram indicates recombination. Structure v2.3 with linkage model35 was used to infer the lineage ancestry of the unique STs, assuming that the ancestry of each ST was derived from K ancestral subpopulations. Three individual runs per value of K (chosen between 3 and 20) were performed using 500,000 MCMC iterations, consisting of 200,000 burn-in iterations and 300,000 sampling iterations. The K-value that generated the highest median posterior probability was used as the probable number of ancestral populations.
Nucleotide sequence accession numbers
The sequences of the eight MLST loci have been deposited in the GenBank database under accession numbers KM227013–KM229308 and KP057215–KP057222.
Additional Information
How to cite this article: Song, Y. et al. Genetic diversity and population structure of Lactobacillus delbrueckii subspecies bulgaricus isolated from naturally fermented dairy foods. Sci. Rep. 6, 22704; doi: 10.1038/srep22704 (2016).
References
Sun, Z. et al. Identification and characterization of the dominant lactic acid bacteria from kurut: the naturally fermented yak milk in Qinghai, China. J Gen Appl Microbiol 56, 1–10 (2010).
Yu, J. et al. Diversity of lactic acid bacteria associated with traditional fermented dairy products in Mongolia. J Dairy Sci 94, 3229–3241, doi: 10.3168/jds.2010-3727 (2011).
Bao, Q. et al. Isolation and identification of cultivable lactic acid bacteria in traditional yak milk products of Gansu Province in China. J Gen Appl Microbiol 58, 95–105 (2012).
Kaihei, O., Dugersuren, J., Demberel, S. & Watanabe, K. Pyrosequencing analysis of the microbial diversity of airag, khoormog and tarag, traditional fermented dairy products of mongolia. Bioscience of microbiota, food and health 33, 53 (2014).
Torriani, S., Zapparoli, G. & Dellaglio, F. Use of PCR-based methods for rapid differentiation of Lactobacillus delbrueckii subsp. bulgaricus and L. delbrueckii subsp. lactis. Appl. Environ. Microbiol 65, 4351–4356 (1999).
Gatti, M., Fornasari, M. E. & Neviani, E. Differentiation of Lactobacillus delbrueckii subsp. bulgaricus and Lactobacillus delbrueckii subsp. lactis by SDS-PAGE of cell-wall proteins. Lett Appl Microbiol 32, 352–356 (2001).
Giraffa, G. et al. Molecular typing of Lactobacillus delbrueckii of dairy origin by PCR-RFLP of protein-coding genes. Int J Food Microbiol 82, 163–172 (2003).
Dellaglio, F., Felis, G. E., Castioni, A., Torriani, S. & Germond, J. E. Lactobacillus delbrueckii subsp. indicus subsp. nov., isolated from Indian dairy products. Int J Syst Evol Microbiol 55, 401–404, doi: 10.1099/ijs.0.63067-0 (2005).
Cebeci, A. & Gürakan, G. C. Comparative typing of L. delbrueckii subsp. bulgaricus strains using multilocus sequence typing and RAPD–PCR. Eur Food Res Technol 233, 377–385, doi: 10.1007/s00217-011-1526-5 (2011).
Tanigawa, K. & Watanabe, K. Multilocus sequence typing reveals a novel subspeciation of Lactobacillus delbrueckii. Microbiology 157, 727–738, doi: 10.1099/mic.0.043240-0 (2011).
Adimpong, D. B. et al. Lactobacillus delbrueckii subsp. jakobsenii subsp. nov., isolated from dolo wort, an alcoholic fermented beverage in Burkina Faso. Int J Syst Evol Microbiol 63, 3720–3726, doi: 10.1099/ijs.0.048769-0 (2013).
Weiss, N., Schillinger, U. & Kandler, O. Lactobacillus lactis, Lactobacillus leichmannii and Lactobacillus bulgaricus, Subjective Synonyms of Lactobacillus delbrueckii and Description of Lactobacillus delbrueckii subsp. lactis comb. nov. and Lactobacillus delbrueckii subsp. bulgaricus comb. nov. Syst Appl Microbiol 4, 552–557, doi: 10.1016/s0723-2020(83)80012-5 (1983).
Maiden, M. C. et al. MLST revisited: the gene-by-gene approach to bacterial genomics. Nat Rev Microbiol 11, 728–736, doi: 10.1038/nrmicro3093 (2013).
Jordan, I. K., Rogozin, I. B., Wolf, Y. I. & Koonin, E. V. Microevolutionary genomics of bacteria. Theor Popul Biol 61, 435–447 (2002).
Bilhere, E., Lucas, P. M., Claisse, O. & Lonvaud-Funel, A. Multilocus sequence typing of Oenococcus oeni: detection of two subpopulations shaped by intergenic recombination. Appl Environ Microbiol 75, 1291–1300, doi: 10.1128/AEM.02563-08 (2009).
van de Guchte, M. et al. The complete genome sequence of Lactobacillus bulgaricus reveals extensive and ongoing reductive evolution. Proc Natl Acad Sci USA 103, 9274–9279, doi: 10.1073/pnas.0603024103 (2006).
Kudo, Y., Oki, K. & Watanabe, K. Lactobacillus delbrueckii subsp. sunkii subsp. nov., isolated from sunki, a traditional Japanese pickle. Int J Syst Evol Microbiol 62, 2643–2649, doi: 10.1099/ijs.0.037051-0 (2012).
Huson, D. H. & Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23, 254–267, doi: 10.1093/molbev/msj030 (2006).
Haubold, B. & Hudson, R. R. LIAN 3.0: detecting linkage disequilibrium in multilocus data. Bioinformatics 16, 847–848 (2000).
Hao, P. et al. Complete sequencing and pan-genomic analysis of Lactobacillus delbrueckii subsp. bulgaricus reveal its genetic basis for industrial yogurt production. PLoS One 6, e15964, doi: 10.1371/journal.pone.0015964 (2011).
McNulty, N. P. et al. The impact of a consortium of fermented milk strains on the gut microbiome of gnotobiotic mice and monozygotic twins. Sci Transl Med 3, 106ra106, doi: 10.1126/scitranslmed.3002701 (2011).
Claesson, M. J., van Sinderen, D. & O’Toole, P. W. Lactobacillus phylogenomics–towards a reclassification of the genus. Int J Syst Evol Microbiol 58, 2945–2954, doi: 10.1099/ijs.0.65848-0 (2008).
Archaeology: Ancient cheese found with mummies. Nature 507, 10–10, doi: 10.1038/507010a (2014).
Yang, Y. et al. Proteomics evidence for kefir dairy in Early Bronze Age China. J Archaeol Sci 45, 178–186 (2014).
Yu, J. et al. Phylogenetic study of Lactobacillus acidophilus group, L. casei group and L. plantarum group based on partial hsp60, pheS and tuf gene sequences. Eur Food Res Technol 234, 927–934, doi: 10.1007/s00217-012-1712-0 (2012).
Xu, H. et al. Multilocus sequence typing of Lactococcus lactis from naturally fermented milk foods in ethnic minority areas of China. J Dairy Sci 97, 2633–2645, doi: 10.3168/jds.2013-7738 (2014).
Jolley, K. A., Feil, E. J., Chan, M. S. & Maiden, M. C. Sequence type analysis and recombinational tests (START). Bioinformatics 17, 1230–1231 (2001).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
McVean, G. A. et al. The fine-scale structure of recombination rate variation in the human genome. Science 304, 581–584, doi: 10.1126/science.1092500 (2004).
Auton, A. & McVean, G. Recombination rate estimation in the presence of hotspots. Genome Res 17, 1219–1227, doi: 10.1101/gr.6386707 (2007).
Martin, D. P. et al. RDP3: a flexible and fast computer program for analyzing recombination. Bioinformatics 26, 2462–2463, doi: 10.1093/bioinformatics/btq467 (2010).
Vilella, A. J. et al. EnsemblCompara GeneTrees: Complete, duplication-aware phylogenetic trees in vertebrates. Genome Res 19, 327–335, doi: 10.1101/gr.073585.107 (2009).
Feil, E. J., Li, B. C., Aanensen, D. M., Hanage, W. P. & Spratt, B. G. eBURST: inferring patterns of evolutionary descent among clusters of related bacterial genotypes from multilocus sequence typing data. J Bacteriol 186, 1518–1530 (2004).
Huson, D. H. SplitsTree: analyzing and visualizing evolutionary data. Bioinformatics 14, 68–73 (1998).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7, 574–578, doi: 10.1111/j.1471-8286.2007.01758.x (2007).
Acknowledgements
Financial support for this research was provided by the National Natural Science Foundation of China (Grant No. 31430066), the International S&T Cooperation Program of China (ISTCP, 2014DFR31150) and the China Agriculture Research System (Grant No. CARS-37).
Author information
Authors and Affiliations
Contributions
R.Y. and H.Z. designed this study. W.L. and B.M. collected the samples. J.Y. and Y.S. performed the experiments. Y.S., C.G. and Y.W. contributed to the data analyses. Y.S., Z.S., R.Y. and H.Z. wrote this manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Song, Y., Sun, Z., Guo, C. et al. Genetic diversity and population structure of Lactobacillus delbrueckii subspecies bulgaricus isolated from naturally fermented dairy foods. Sci Rep 6, 22704 (2016). https://doi.org/10.1038/srep22704
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep22704
- Springer Nature Limited
This article is cited by
-
Multilocus sequence typing of L. bulgaricus and S. thermophilus strains from Turkish traditional yoghurts and characterisation of their techno-functional roles
Food Science and Biotechnology (2024)
-
Distribution of Genes Related to Probiotic Effects Across Lacticaseibacillus rhamnosus Revealed by Population Structure
Probiotics and Antimicrobial Proteins (2023)
-
Safety Evaluation of Lactobacillus delbrueckii subsp. lactis CIDCA 133: a Health-Promoting Bacteria
Probiotics and Antimicrobial Proteins (2022)
-
Draft genome sequence of Lactobacillus delbrueckii subsp. bulgaricus LBP UFSC 2230: a tool for preliminary identification of enzymes involved in CLA metabolism
Brazilian Journal of Microbiology (2021)
-
Clustering of Pan- and Core-genome of Lactobacillus provides Novel Evolutionary Insights for Differentiation
BMC Genomics (2018)