Abstract
Cucumber is vulnerable to many foliage diseases. Recent studies reported cloning of candidate genes for several diseases in cucumber; however, the exact defence mechanisms remain unclear. Dof genes have been shown to play significant roles in plant growth, development and responses to biotic and abiotic stresses. Dof genes coding for plant-specific transcription factors can promote large-scale expression of defence-related genes at whole genome level. The genes in the family have been identified and characterized in several plant species, but not in cucumber. In the present study, we identified 36 CsDof members from the cucumber draft genomes which could be classified into eight groups. The proportions of the CsDof family genes, duplication events, chromosomal locations, cis-elements and miRNA target sites were comprehensively investigated. Consequently, we analysed the expression patterns of CsDof genes in specific tissues and their response to two biotic stresses (watermelon mosaic virus and downy mildew). These results indicated that CsDof may be involved in resistance to biotic stresses in cucumber.
Similar content being viewed by others
Introduction
Cucumber is a major vegetable crop worldwide1,2,3, which belongs to the Cucurbitaceae family and is vulnerable to infection or infestation by many bacterial, fungal and viral pathogens or insects such as Fusarium oxysporum (Fusarium wilt, FW)4,5, Podosphaera fusca (powdery mildew, PM)6, Pseudoperonospora cubensis (downy mildew, DM)7, Zucchini yellow mosaic virus (ZYMV)8, Corynespora cassiicola (target leaf spot, TLS)9 and Cladosporium cucumerinum (scab)10,11. These biotic stresses pose a substantial threat to yield and food security. In recent years, several resistance genes or resistance gene candidates have been cloned in cucumbers; however, the defence mechanisms remain unknown. Cucumber was the first vegetable crop with a draft genome sequence1,2. The publicly available whole-genome and many transcriptome sequences of cucumber provide a good opportunity to study disease resistance mechanisms, especially from a whole-genome perspective.
The Dof (DNA-binding with one finger) proteins are a group of plant-specific transcription factors (TFs) that have typically 200–400 amino acids with a highly conserved 52 amino acid Dof DNA-binding domain at the N-terminal. The Dof domain has a Cys2/Cys2 Zn2+ finger structure that recognises the specific cis-element of (AT)/AAAG in the promoter region of their target genes12,13,14. The C-terminal of the Dof proteins contains a transcriptional regulation domain with diverse functions including interaction with various regulatory proteins and activation of gene expression15. The N- and C- terminal regions of Dof protein may interact with various regulatory proteins or intercept signals to mediate the activation or repression of the target genes16.
The Dof TFs have been shown to play important roles in many biological processes, such as synthesis of seed storage protein, seed development and germination, photosynthetic processes and flowering17. Dof TFs participate in various physiological processes such as the cell cycle, adaxial-abaxial polarity, plant hormonal signalling, phytochrome and cryptochrome signalling, nitrogen use, biotic and abiotic stress tolerance18. The Dof members could play roles in the regulation of secondary metabolic processes, such as the biosynthesis of glucosinolates and flavonoids19,20 and cell cycle regulation21. In Arabidopsis, the Dof genes DAG1 and DAG2 are associated with seed germination22,23 and CDF1, CDF2 and CDF3 control photoperiodic flowering24. Some other Dof genes (for example, AtDof2.4, AtDof5.8 and AtDof5.6/HCA2) are specifically expressed in the early stage of vascular tissue development25,26. In tomato, StDof1 is specifically expressed in epidermal fragments of guard cells; StDof1 can interact with the promoter of the KST1 gene13. Rice OsDof3 was reported to be involved in gibberellins-regulated expression27. Maize Dof1 and Dof2 were confirmed to be activators of genes associated with carbohydrate metabolism and contained the C4 key photosynthetic gene phosphoenolpyruvate carboxylase (PEPC)28.
Dof transcription factors have been functionally characterised in Arabidopsis22,23 and a number of crops such as tomato18, rice27 and soybean29. However, no such information is available in cucumber. The objective of the present study was to conduct a genome-wide characterisation of Dof gene families in the cucumber genome. We identified 36 CsDof members in the cucumber draft genome, which were classified into eight subgroups. The proportions of the CsDof family genes, duplication events and chromosomal locations were investigated. We also analysed the expression patterns of CsDof members in various tissues and the response to inoculation of the watermelon mosaic virus (WMV) as well as downy mildew (DM) pathogens. Our study provided novel insights into the stress responses of CsDof genes to these two important cucumber pathogens and deepens our understanding of the structure and function of CsDof genes in the cucumber genome.
Results
Identification of CsDof homologues in the cucumber genome
To identify the Dof transcription factor coding genes in the cucumber genome, we used the HHM profile of the Dof domain (PF02701) as a query to perform an HMMER search (http://hmmer.janelia.org/) against the 9930 and Gy14 cucumber draft genomes. Thirty-six Dof gene sequences were identified from the HMMER database (http://www.ebi.ac.uk/Tools/hmmer/search/hmmsearch). We then performed SMART search (http://smart.emblheidelberg.de/) to confirm the presence of the conserved Dof domain which confirmed the existence of 36 putative CsDof genes in the cucumber genome. For convenience, the 36 CsDof genes were assigned names from CsDof01 to CsDof36 based on their chromosomal locations (Table 1). These CsDof genes encoded predicted peptides ranging from 150 to 503 aa with the pI value from 4.8 to 9.6 and the molecular weight from 16.9 to 55.2 kD. All CsDof proteins were predicted to be localised in the nucleus except for CsDof30 which was extracellular (Table 1). In addition, using BLASTN, we found nine CsDof genes (CsDof02, CsDof04, CsDof10, CsDof11, CsDof13, CsDof16, CsDof18, CsDof28 and CsDof36) showed polymorphisms between 9930 and Gy14. The SNPs and their locations within each gene are presented in Supplementary Table S1.
Phylogenetic analysis
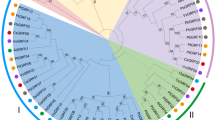
To clarify the phylogenetic relationships among 36 CsDof genes, a phylogenetic tree was constructed based on the alignment of the CsDof domain sequences (Supplementary Fig. S1). The neighbour joining (NJ) phylogenetic tree divided the CsDof homologs into four groups (A to D), which could be further classified into 8 subgroups: A, B1, B2, C1, C2.1, C2.2, D1 and D2. The bootstrap values are shown in Fig. 1. Subgroup B1 contained the most CsDof family members (9 genes, or 25%); B2 and C2.2 constituted the smallest clade with only 2 members each. Groups C2.1 and C2.2 contained 5 CsDof genes (13.9%); group D1 showed 7 members representing 19.4% of the total Dof genes. All 36 CsDof genes showed the conserved Cys2/Cys2 Zn2+ finger in the Dof domain (Supplementary Fig. S1).
Chromosomal locations and duplications of Dof members
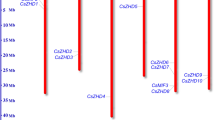
As shown in Fig. 2, the 36 CsDof genes were dispersed on 6 of the 7 cucumber chromosomes (none in chromosome 7). Chromosome 6 harboured the most (11 of 36) Dof genes, whereas 10, 8, 5, 1 and 1 were mapped on chromosomes 1, 3, 5, 2 and 4, respectively (Fig. 2).
Distribution of 36 CsDof genes onto seven cucumber chromosomes.
Graphical representation of physical locations for each CsDof gene on cucumber chromosomes. The CsDof duplications genes were connected with a line. The CsDof06/CsDof07 and CsDof34/CsDof35 were tandem duplicated genes and the CsDof01/CsDof03, CsDof02/CsDof04, CsDof03/CsDof25, CsDof07/CsDof35, CsDof09/CsDof36 and CsDof20/CsDof24 were undergoing segmental duplication events.
In this study, the CsDof genes duplications were analysed using the CoGe website. Two pairs of tandem duplicated genes were identified (CsDof06/CsDof07 and CsDof34/CsDof35) and 6 pairs of segmental duplication genes were observed (CsDof01/CsDof03, CsDof02/CsDof04, CsDof03/CsDof25, CsDof07/CsDof35, CsDof09/CsDof36 and CsDof20/CsDof24) (Fig. 2). To trace the dates of the duplication blocks, we estimated the Ks/Ka distances and ratios. The segmental duplications of the CsDof genes in cucumber originated from approximately 4.639 Mya (million years ago, Ks = 0.6031) to 11.66 Mya (Ks = 1.5161) with an average of 9.079 Mya (Ks = 1.180). The Ks of the tandem duplication of CsDof06 and CsDof07 was 1.8735, which dated the duplication event at 14.41 Mya; that the tandem duplication of CsDof34 and CsDof35 was 1.3398, dating the duplication event at 10.31 Mya (Supplementary Table S2).
Gene structure and conserved motifs analysis
In order to investigate the characterization of exon-intron structure, all of the CsDof genes were analyzed by using the Gene Structure Display Server. As shown in Fig. 3, the predicted number of exons among the 36 CsDof genes were relatively fewer and they varied from one to three with 17 members having one and 14 with two. Five (CsDof 05, CsDof15, CsDof 20, CsDof 27 and CsDof 35) had three exons. Furthermore, some CsDof genes within the same subgroup demonstrated similar exon/intron structure patterns in terms of exon number. For instance, all CsDof genes in subgroup A had no introns. The two exons CsDof gene were mainly in group C (four in C1, C2.1 and C2.2) and D (five in D1 and one in D2). The majority three excons were belonging in group B (B1 and B2). These similar structure feature may be related to their functions in cucumber genome (Fig. 3).
To obtain insights into the diversity of motif compositions in CsDof proteins, these proteins were assessed using the MEME programme. A total of 15 conserved motifs were identified (Supplementary Table S3; Fig. S2). These motifs are represented in their relative location within the protein. Motif 1 was uniformly observed in all Dof proteins and was confirmed to be the conserved Dof domain. Moreover, the CsDofs in each subgroup had several special motifs at C-terminal regions, suggesting that they had a similar function in the CsDof members within the same subgroups (Supplementary Fig. S2). Groups A, C2.2 and D2 showed one conserved motif (motif 1). Group D1 members contained motif 4 and motif 12 and Group B1 contained motif 2 and motif 7.
Cis-element analysis of CsDof genes
In the CsDof genes promoter region, many key defence cis-elements (such as ACGTABOX, ASF1MOTIFCAMV and WBOXATNPR1) were identified. Other key elements included those in hormone signalling, such as ARFAT (auxin), ASF1MOTIFCAMVT (auxin), ERELEE4 (ethylene), GAREAT (gibberellin), LECPLEACS2 (ethylene), MYBGAHV (gibberellin), NTBBF1ARROLB (auxin) and TATCCAOSAMY (gibberellin). Some CsDof genes harboured additional transcription factor cis-regulatory elements, e.g., MYB1AT, MYB1AT, MYB1AT, MYBGAHV, MYBCOREATCYCB1 and WRKY71OS (Supplementary Table S4). All CsDof gene promoters contained the conserved elements DOFCOREZM, suggesting that these CsDof genes could be regulated by themselves. Additionally, an in silico analysis showed that the CsDof gene promoters maintained several tissue-specific elements. Some examples included root-specific (ROOTMOTIFTAPOX1, RAV1AAT and SP8BFIBSP8BIB), leaf-specific (CCA1ATLHCB1, DOFCOREZM, GATABOX, GT1CONSENSUS, IBOXCORE and RAV1AAT) and flower-specific (CARGCW8GAT) responsive elements (Supplementary Table S4).
Regulated miRNAs
Among the 36 CsDof genes, 15 were targeted by 20 miRNAs. CsDof21 had the most abundant transcript, which was targeted by 6 plant miRNAs (miR5021, miR5658, miR2673a, miR2673b, miR2673 and miR7494b) (Supplementary Table S5). A sequence analysis of CsDof26 implied that the 188–288 bp region may be the target site of miR319a. The miR5658 targeted CsDof05, CsDof08, CsDof20 and CsDof21. The CsDof28 was targeted by 5 plant miRNAs (miR831, miR831-5p, miR393b-3p, miR393b-3p and miR3434-3p) (Supplementary Table S5).
Tissue-specific expression of CsDof genes
From analysis of publicly available transcriptome data, we observed 17 of 36 CsDofs that had a relatively high expression in root tissues including CsDof04, CsDof05, CsDof06, CsDof07, CsDof08, CsDof11, CsDof13, CsDof14, CsDof15, CsDof17, CsDof21, CsDof23, CsDof24, CsDof26, CsDof27, CsDof28 and CsDof32. Among the 17, CsDof genes, CsDof06, CsDof13, CsDof14, CsDof17, CsDof21 and CsDof23 were only expressed in the root, suggesting that these six genes may play specific roles in root development. Moreover, CsDof04, CsDof10, CsDof11, CsDof15, CsDof20, CsDof24, CsDof31 and CsDof32 were specifically expressed in the stems (Fig. 4). The expressions of five genes, CsDof01, CsDof02, CsDof03, CsDof18 and CsDof25, were specific to male flowers, whereas CsDof12, CsDof33 and CsDof35 were specifically expressed in female flowers. Finally, 3 (CsDof11, CsDof30 and CsDof34) and 3 (CsDof09, CsDof19 and CsDof29) genes were specifically expressed in the leaves and tendrils, respectively (Fig. 4).
Tissue-specific digital expression profiles of CsDof transcription factors.
The CsDof genes were listed on the right of the expression array and the colour scale was shown at the top. The higher expression for each gene was presented in red; otherwise, green was used. The genes with an RPKM equal to 0 were not used in this array. The capital letters indicated the following: R = root; S = stem; L = leaf; T = tendril; M = male flower; F = female flower.
Responses to DM and WMV pathogen inoculations
Of 36 CsDof genes, 22 were responsive to DM pathogen inoculation (Fig. 5). Of these, four CsDof genes (CsDof07, CsDof17, CsDof19 and CsDof34) had lower expression as compared with the control, whereas two genes (CsDof27 and CsDof29) exhibited higher expression than in the control after inoculation. Additionally, five members (CsDof03, CsDof18, CsDof28, CsDof35 and CsDof36) were first up- and then down- regulated. Moreover, four genes (CsDof10, CsDof12, CsDof19 and CsDof31) were the highest in expression on the first day after DM inoculation, whereas the expression of seven genes (CsDof02, CsDof04, CsDof18, CsDof25, CsDof28, CsDof35 and CsDof36) peaked on the second day after inoculation.
Expression profile of CsDof transcription factors under downy mildew.
The expression colour scale was shown at the top. The 22 CsDof genes were observed in transcriptome data with DM inoculation and the sparkline presented the expression variation. Higher expression for each gene was presented in red; otherwise, green was used. The genes with an RPKM equal to 0 were not used in this array. The columns indicated the changes in the CsDof gene expression levels at various time points (1, 2, 3, 4, 6 and 8 dpi; Ck was treated with MOCK) under DM stress.
Quantitative real-time PCR was conducted for these 36 CsDof genes to study their expression upon inoculation of the WMV viral pathogen (Fig. 6). Among them, 34 were up- or down-regulated after WMV inoculation and no changes were observed for CsDof08 or CsDof16. Three genes (CsDof02, CsDof14 and CsDof35) were initially up- regulated but down- regulated after 24 d. The expressions of ten genes (CsDof02, CsDof09, CsDof12, CsDof14, CsDof15, CsDof19, CsDof20, CsDof22, CsDof26 and CsDof32) increased on the first day after inoculation. Overall, 20 CsDof genes exhibited higher expression on the third day after WMV inoculation. CsDof28, CsDof30 and CsDof33 were highly expressed at 3 d, 9 d and 24 d after inoculation, respectively, whereas the expression of CsDof04 and CsDof13 peaked at 18 d after inoculation.
Expression profiling of CsDof genes under WMV stress.
The 34 CsDof genes were determined to have varied expression after WMV inoculation via qRT-PCR and the sparkline presents the expression variation. The expression colour scale was shown at the top. A higher expression for each gene was presented in red; otherwise, green was used. The columns indicate the changes in CsDof gene expression levels at various time points (1 d, 3 d, 6 d, 9 d, 12 d, 18 d and 24 d under WMV inoculation; Ck was treated with MOCK).
Discussion
In recent years, gene family analysis has become an important approach to understand gene structure, function and evolution. The Dof genes are plant-specific transcription factors that are involved in various biological processes and are ubiquitous in many plant species. The function and evolution of Dof genes have been thoroughly studied in Arabidopsis (36 AtDof genes), rice (30 OsDof genes)30, soybean29 (78 GmDof genes), Chinese cabbage15 (76 BraDof genes), potato17 (35 StDof genes), pigeonpea31 (38 CcDof genes) and tomato18 (34 SlDof genes). Here, we conducted a comprehensive analysis of the CsDof family in cucumber to determine their potential functions in response to biotic stresses.
We identified 36 putative CsDof genes in the cucumber genome; the number of CsDof homologues was largely similar to that found in Arabidopsis or rice30. The 36 CsDof genes were classified into four groups (A to D) and eight subgroups (A, B1, B2, C1, C2.1, C2.2, D1 and D2) (Fig. 1). In Arabidopsis, all 36 AtDof genes were divided into four groups and nine subgroups (A, B1, B2, C1, C2.1, C2.2, C3, D1 and D2)30; the CsDof genes were not found in subgroup C3. In cucumber, subgroup B1 contained nine CsDof genes; only five were noted in Arabidopsis. Subgroup B2 had two CsDof members but three AtDof genes. These results indicated that CsDof and AtDof genes underwent different duplication events.
Similar to that found in rice and Arabidopsis30, the cucumber Dof genes had few introns (0–2) in each gene (Fig. 3). A motif analysis indicated that motif 1 was uniformly observed in all Dof proteins (Supplementary Fig. S2). Similar to the results from Arabidopsis, rice30 and tomato18, this phenomenon suggested that CsDof transcription factors were evolutionarily conserved in plants.
The whole-genome duplications, segmental/tandem duplication and transposition events were crucial for gene family expansion. Cucumber did not experience a recent whole-genome duplication1; thus, the remainder duplications would play crucial roles in gene expansion. In this study, we found 8 pairs of CsDof genes that had undergone duplications (Fig. 2). Intriguingly, we observed six pairs of segmental duplication and two pairs of tandem duplication events in CsDof genes. This indicated that CsDof gene segmental duplication is predominant in the evolution of cucumber and that tandem duplication is involved18. The duplication divergence of monocots and dicots occurred approximately 170–235 Mya32. This study showed that the mean date of CsDof genes tandem duplication events was at 12.36 Mya, whereas the mean date of segmental duplication events was at 9.079, which showed that the segmental/tandem duplication occurred after the divergence of the monocot-dicot split and that the tandem duplication occurred prior to the segmental duplication in cucumber CsDof genes.
Gene expression patterns provided important clues for gene function; thus, we conducted a digital gene expression analysis for duplicated CsDof genes in the root, leaf, stem, tendril and flower tissues using public RNA-seq data33. The Ka/Ks ratio analysis indicated that the CsDof genes had divergent functions after the duplication events. All tested 8 pairs of genes showed distinct expression patterns both in a tissue-specific and biotic responsive manner. For example, in one pair of segmental duplication CsDof genes, CsDof07 was highly expressed in the root and CsDof35 was highly expressed in female flowers (Fig. 4). The expression of CsDof07 was relatively low and reached maximum at the third day after WMV inoculation; however, CsDof35 was expressed at a higher level and peaked at the sixth day after WMV inoculation (Fig. 6). The tandemly duplicated CsDof genes were very differently expressed. CsDof34 and CsDof35 had specific expressions in the leaves and female flowers, respectively and CsDof34 had the highest expression at day zero (Mock treatment) under DM inoculations; however, CsDof35 reached its peak expression at 2 d after DM inoculation (Fig. 5). These results indicated that the duplicated CsDof genes may play crucial and diversifying roles in plant development.
Dof TFs have been shown to play crucial roles in the regulatory networks of plant defence, including responses to diverse biotic and abiotic stresses15,34,35. In tobacco, the Sar8.2b gene can be promoted by the Dof transcription factor, which is involved in systemic acquired resistance36. In tomato, the Dof transcription factor was found to regulate fungus resistance through the ACBP3 gene, which promoted autophagy-mediated leaf senescence and conferred resistance to Pseudomonas syringae pv. tomato DC300037. In barley seeds, the Dof transcription factors play a role in biotic stress tolerance based on their association with the cystatin gene38. The Dof gene may play an indirect role in responding to biotic stresses and we speculated that the CsDof transcript factor is involved in resistance through a direct or indirect manner; these may thus promote different target resistant genes in cucumber. In this study, we investigated the response patterns of CsDof genes to DM and WMV inoculations. We found that 19 and 34 Dof genes were up regulated after DM and WMV inoculations, respectively, suggesting that most of these Dof genes may play positive roles in host defence responses; however, additional work is needed to confirm their roles.
To summarise, in this study, we reported a comprehensive analysis of CsDof transcription factor genes in the cucumber genome. The 36 CsDof genes were categorised into eight subgroups and the structural and functional properties of each CsDof member were characterised. Most of the CsDof genes were induced by biotic stresses. Our work will assist in understanding the roles of these CsDof transcription factors in response to biotic stresses and their potential interactions with defence-related genes in the disease resistance network.
Methods
Genome-wide identification of CsDof homologue sequences
The conserved Dof domain based on a Hidden Markov Model (HMM) (PF02701) was downloaded from the Pfam protein family database (http://pfam.sanger.ac.uk/). To identify the Dof transcription factor coding genes of Cucumis sativus, we used the HHM profile of the Dof domain as a query to perform a HMMER search (http://hmmer.janelia.org/) against the cucumber genome databases (http://www.icugi.org/; http://cucumber.genomics.org.cn/; http://wenglab.horticulture.wisc.edu/). All non-redundant sequences encoding complete Dof domains were considered to be putative Dof genes. Each non-redundant sequence was double checked for the presence of the conserved Dof domain using a SMART search (http://smart.emblheidelberg.de/). The candidate cucumber CsDof genes were named based on their distribution on the seven cucumber chromosomes. The ExPASy server (http://web.expasy.org/compute_pi/)39 was used to compute the pI and molecular weight of the identified CsDof proteins. We performed a nuclear localisation signal (NLS) analysis prediction of the Dof protein on the website (http://cello.life.nctu.edu.tw/).
Phylogenetic characterisation of CsDof homologs
For a phylogenetic analysis of the plant Dof gene family, nucleotide or protein sequences from Arabidopsis were obtained from previous studies30. The information of Dof genes in Arabidopsis are presented in Table S6. Multiple sequence alignments were conducted on the amino acid sequences of Dof proteins from cucumber and Arabidopsis using ClusterW with default settings. Subsequently, MEGA 6.0 software was employed to construct an unrooted phylogenetic tree based on alignments using the Neighbour-Joining (NJ) method with the following parameters: JTTmodel, pairwise gap deletion and 1000 bootstraps40. Furthermore, maximum likelihood, minimal evolution and PhyML methods were also applied for the tree construction to validate the results of the NJ method.
Chromosomal locations and duplications
Chromosomal locations for each CsDof gene were determined via BLASTP search against the cucumber genome databases with default settings. Tandem duplications and segmental duplications in the cucumber genome were analysed using the website tool CoGe (https://genomevolution.org/CoGe/)41 and the duplicated genes were linked with coloured lines which created by using the Circos software42 (http://circos.ca/).
To estimate the synonymous and non-synonymous substitution rates, we used the software DnaSp43. The time (million years ago, Mya) of duplication and divergence of each CsDof genes were estimated using a synonymous mutation rate of λ substitutions per synonymous site per year, T = Ks/2λ (λ = 6.5610 e-9)33.
Gene structure analysis and conserved motif identification
The exon-intron organisations of the genes were determined using the Gene Structure Display Server (http://gsds.cbi.pku.edu.cn) through a comparison of their full-length cDNA or predicted coding sequence (CDS)44. The motifs of the Dof protein sequences were statistically identified using the MEME programme (http://meme-suite.org/tools/meme) with the motif length set to 6–100 and motif sites to 2–120. The maximum number of motifs was set to 15, the distribution of one single motif was “any number of repetitions” and the other parameter was “search given strand only”.
Cis-element and miRNA target analysis
The mature miRNAs sequences were downloaded from miRBase v20.0 (http://www.mirbase.org) and PMRD (http://bioinformatics.cau.edu.cn/PMRD). Known cucumber miRNAs were used to identify the miRNAs target genes in the CsDof families. The prediction was conducted using the Plant Small RNA Target Analysis Server (psRNA Target: http://plantgrn.noble.org/psRNATarget) with default parameters. Alignment between all known plant miRNA and their potential CsDofs targets were evaluated using previously described parameters45. To identify the putative cis-acting regulatory elements presented in the promoter regions of CsDof genes, nucleotide sequences of 2000 bp upstream regions from the translational start codon (ATG) were retrieved from the PGSC database. An in silico promoter analysis was carried out using the PLACE database (http://www.dna.affrc.go.jp/PLACE/signalscan.html).
Tissue-specific expression
The cucumber genome-wide digital gene expression was assessed as described by Baloglu et al.33. Illumina sequencing reads from RNA-Seq studies were retrieved from a public repository database (SRA, Sequence Read Archive) with the following accession numbers: SRR351499 (cucumber root tissue), SRR351905 (cucumber stem tissue), SRR351906 (cucumber leaf tissue), SRR351908 (cucumber male flower tissue), SRR351912 (cucumber female flower tissue) and SRR351910 (cucumber tendril tissue). All transcript data were analysed using Gene-E 3.0.240 (www.broadinstitute.org/cancer/software/GENE-E).
Response to biotic stresses
To investigate the expression profiling of CsDof genes in response to downy mildew inoculation, the GEO data were downloaded from the NCBI PubMed database (http://www.ncbi.nlm.nih.gov/pubmed/) based on previous literature46. The data were used to draw heatmaps with Gene-E 3.0.240 (www.broadinstitute.org/cancer/software/GENE-E).
We also experimentally tested the responses of the CsDof genes to the inoculation of WMV with the cucumber line ‘Europe 8’, which is susceptible to WMV. The original WMV isolate was kindly provided by Prof. Rosario Provvidenti of Cornell University (Ithaca, New York, USA). The virus was maintained in plants of Cucurbita pepo. For inoculum preparation, diseased leaves were washed with ddH2O and the suspension was diluted in 0.2 mol • L−1 phosphoric acid buffer (pH 7.0) to a concentration of 1:3 (W/V). We used the leaf inoculation method of friction. Seeds of “Europe 8” were soaked in 55 °C warm water for 4 h for disinfection. Then, the seeds were germinated in an incubator (at 28 °C) for 18 h and planted in 4 × 8 plugs in a greenhouse. When the second true leaf was fully expanded, we sprinkled a small amount of 600 ~ 800 mesh emery after frictional artificial inoculation, flushed it with clear water and used the phosphoric acid buffer to inoculate the health leaves as a control. We set up 3 duplications and 10 cucumber seedlings for each. Both the inoculated and control seedlings were maintained in an insect-free growth chamber with 25 ~ 30 °C and at 100% RH (Relative Humidity). Leaf samples were harvested at 0, 1, 3, 6, 9, 12, 18 and 24 days post-inoculation (dpi), which were used to extract intercellular fluid or gene expression analysis.
To investigate the CsDof gene expression, total RNA was extracted using a Huayueyang Quick RNA isolation Kit (Cat. No.: ZH120, Huayueyang Biotechnology, Beijing, China) following the manufacturer’s procedure. To remove trace DNA contamination, DNase (Cat. No.: D2270A, TaKaRa Biotechnology, Dalian, China) was added to the total RNA samples. The qualities and quantities of RNA were determined using agarose gel electrophoresis and a NanoDrop ND-2000 Spectrophotometer (Thermo Fisher Scientific Inc., USA). For cDNA synthesis, 1 μg high-quality total RNA was reverse-transcribed with oligodT and random primers with Super Script III Reverse Transcriptase (TaKaRa, Dalian, China) according to the manufacturer’s instructions.
For qRT-PCR analysis, the specific primers for each CsDof gene were designed according to the Dof gene sequences using Primer 3 online (http://primer3.ut.ee/) (Supplementary Table S7). The cucumber ubiquitin extension protein gene (primer sequences: 5′-GGCAGTGGTGGTGAACATG-3′ and 5′-TTCTGGTGATGGTGTGAGTC-3′) was used as the reference gene47. qRT-PCR reactions were performed using the SYBR Premix Ex TaqTM kit (TaKaRa, Dalian, China) and a Roche LightCycler 480. All qRT-PCR experiments were performed with three biological and three technical replications. Relative gene expression was calculated using the 2−ΔΔCt method. Then, the data were compiled to make a heatmap via Gene-E 3.0.240 (www.broadinstitute.org/cancer/software/GENE-E).
Additional Information
How to cite this article: Wen, C.-l. et al. Identification and characterisation of Dof transcription factors in the cucumber genome. Sci. Rep. 6, 23072; doi: 10.1038/srep23072 (2016).
References
Huang, S. et al. The genome of the cucumber Cucumis sativus L. Nat Genet. 41, 1275–1281, doi: 10.1038/ng.475 (2009).
Qi, J. et al. A genomic variation map provides insights into the genetic basis of cucumber domestication and diversity. Nat Genet. 45, 1510–1515, doi: 10.1038/ng.2801 (2013).
Li, J. L. et al. Transcriptomic analysis reveals the roles of microtubule-related genes and transcription factors in fruit length regulation in cucumber (Cucumis sativus L.). Sci. Rep. 5, 8031, doi: 10.1038/srep08031 (2015).
Wang, M. et al. Water balance altered in cucumber plants infected with Fusarium oxysporum f. sp. Cucumerinum. Sci. Rep. 5, 7722, doi: 10.1038/srep07722 (2015).
Zhang, S. P. et al. A major quantitative trait locus conferring resistance to Fusarium wilt was detected in cucumber by using recombinant inbred lines. Mol Breeding 34, 1805–1815 (2014).
Nie, J. et al. Identification and fine mapping of pm5.1: a recessive gene for powdery mildew resistance in cucumber (Cucumis sativus L.). Mol. Breeding 35, 1–13 (2015).
Zhang, S. P. et al. Chromosomal mapping and QTL analysis of resistance to downy mildew in Cucumis sativus. Plant Dis. 97, 245–251 (2013).
Amano, M. et al. High-resolution mapping of zym, a recessive gene for zucchini yellow mosaic virus resistance in cucumber. Theor Appl Genet. 126, 2983–2993, doi: 10.1007/s00122-013-2187-5 (2013).
Wen, C. L. et al. Fine genetic mapping of target leaf spot resistance gene cca-3 in cucumber, Cucumis sativus L. Theor Appl Genet. 18, 1–12, doi: 10.1007/s00122-015-2604-z (2015).
Zhou, Q. et al. A sequencing-based linkage map of cucumber. Mol. Plant. 8(6), 961–3, doi: 10.1016/j.molp.2015.02.008 (2015).
Kang, H. X. et al. Fine genetic mapping localizes cucumber scab resistance gene Ccu into an R gene cluster. Theor Appl Genet. 122, 795–803, doi: 10.1007/s00122-010-1487-2 (2011).
Plesch, G., Ehrhardt, T. & Mueller-Roeber, B. Involvement of TAAAG elements suggests a role for Dof transcription factors in guard cellspecific gene expression. Plant J. 28, 455–464, doi: 10.1046/j.1365-313X.2001.01166.x (2001).
Yanagisawa, S. The Dof family of plant transcription factors. Trends Plant Sci. 7, 555–560, doi.org/ 10.1016/S1360-1385(02)02362-2 (2002).
Yanagisawa, S. Dof domain proteins: plant-specific transcription factors associated with diverse phenomena unique to plants. Plant Cell Physiol. 45, 386–391, doi: 10.1093/pcp/pch055 (2004).
Ma, J., Li, M. Y., Wang, F., Tang, J. & Xiong, A. S. Genome-wide analysis of Dof family transcription factors and their responses to abiotic stresses in Chinese cabbage. BMC Genomics. 16, 1–15, doi: 10.1186/s12864-015-1242-9 (2015).
Noguero, M., Atif, R. M., Ochatt, S. & Thompson, R. D. The role of the DNA-binding One Zinc Finger (DOF) transcription factor family in plants. Plant Sci. 209, 32–45, doi: 10.1016/j.plantsci.2013.03.016 (2013).
Venkatesh, J. & Park, S. W. Genome-wide Analysis and Expression Profiling of DNA-binding with One Zinc Finger (Dof) Transcription Factor Family in Potato. Plant Physiol. Bioch. 94, 73–85, doi: 10.1016/j.plaphy.2015.05.010 (2015).
Cai, X. et al. Genome-wide Analysis of Plant-specific Dof Transcription Factor Family in Tomato. J Integr. Plant Biol. 55, 552–566, doi: 10.1111/jipb.12043 (2013).
Skirycz, A. et al. DOF transcription factor AtDof1.1 (OBP2) is part of a regulatory network controlling glucosinolate biosynthesis in Arabidopsis. Plant J. 47, 10–24, doi: 10.1111/j.1365-313X.2006.02767.x (2006).
Skirycz, A. et al. Transcription factor AtDOF4,2 affects phenylpropanoid metabolism in Arabidopsis thaliana. New. Phytol. 175, 425–438, doi: 10.1111/j.1469-8137.2007.02129.x (2007).
Skirycz, A. et al. The DOF transcription factor OBP1 is involved in cell cycle regulation in Arabidopsis thaliana. Plant J. 56, 779–792, doi: 10.1111/j.1365-313X.2008.03641.x (2008).
Gualberti, G. et al. Mutations in the Dof zinc finger genes DAG2 and DAG1 influence with opposite effects the germination of Arabidopsis seeds. Plant Cell 14, 1253–1263, doi: 10.1105/tpc.010491 (2002).
Papi, M. et al. Identification and disruption of an Arabidopsis zinc finger gene controlling seed germination. Genes Dev. 14, 28–33 (2000).
Imaizumi, T., Schultz, T. F., Harmon, F. G., Ho, L. A. & Kay, S. A. FKF1 Fbox protein mediates cyclic degradation of a repressor of CONSTANS in Arabidopsis. Science. 309, 293–297, doi: 10.1126/science.1110586 (2005).
Guo, Y., Qin, G., Gu, H. & Qu, L. J. Dof5.6/HCA2, a Dof transcription factor gene, regulates interfascicular cambium formation and vascular tissue development in Arabidopsis. Plant Cell. 21, 3518–3534, doi: 10.1105/tpc.108.064139 (2009).
Konishi, M. & Yanagisawa, S. Sequential activation of two Dof transcription factor gene promoters during vascular development in Arabidopsis thaliana. Plant Physiol Biochem. 45, 623–629, doi: 10.1016/j.plaphy.2007.05.001 (2007).
Washio, K. Functional dissections between GAMYB and Dof transcription factors suggest a role for protein-protein associations in the gibberellin-mediated expression of the RAmy1A gene in the rice aleurone. Plant Physiol. 133, 850–863, doi: 10.1104/pp.103.027334 (2003).
Yanagisawa, S. Dof1 and Dof2 transcription factors are associated with expression of multiple genes involved in carbon metabolism in maize. Plant J. 21, 281–288, doi: 10.1046/j.1365-313x.2000.00685.x (2000).
Guo, Y. & Qiu, L. J. Genome-Wide Analysis of the Dof Transcription Factor Gene Family Reveals Soybean-Specific Duplicable and Functional Characteristics. Plos One. 8, e76809, doi: 10.1371/journal.pone.0076809 (2013).
Lijavetzky, D., Carbonero, P. & Vicente-Carbajosa, J. Genome-Wide Comparative Phylogenetic Analysis Of The Rice And Arabidopsis Dof Gene Families. BMC Evol. Biol. 3, 17, doi: 10.1186/1471-2148-3-17 (2003).
Malviya, N. et al. Genome wide in silico characterization of Dof gene families of pigeonpea (Cajanus cajan (L) Millsp.). Mol. Biol. Rep. 42, 535–552, doi: 10.1007/s11033-014-3797-y (2015).
Blanc, G. & Wolfe, K. H. Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. Plant Cell. 16, 1667–1678, doi: 10.1105/tpc.021345 (2004).
Baloglu, M. C., Eldem, V., Hajyzadeh, M. & Unver, T. Genome-Wide Analysis of the bZIP Transcription Factors in Cucumber. Plos One 9, e96014, doi: 10.1371/journal.pone.0096014 (2014).
Gupta, S. et al. Insights into structural and functional diversity of Dof (DNA binding with one finger) transcription factor. Planta. 241, 549–562, doi: 10.1007/s00425-014-2239-3 (2015).
Corrales, A. R. et al. Characterization of tomato Cycling Dof Factors reveals conserved and new functions in the control of flowering time and abiotic stress responses. J. Exp. Bot. 65, 995–1012, doi: 10.1093/jxb/ert451 (2014).
Song, F. & Goodman, R. M. Cloning and identification of the promoter of the tobacco Sar8.2b gene, a gene involved in systemic acquired resistance. Gene. 290, 115–12, doi: 10.1016/S0378-1119(02)00584-X (2002).
Zheng, S. X., Xiao, S. & Chye, M. L. The gene encoding Arabidopsis acyl-CoA-binding protein 3 is pathogen inducible and subject to circadian regulation. J Exp Bot. 63, 2985–3000, doi: 10.1093/jxb/ers009 (2012).
Martinez, M. et al. The barley cystatin gene (Icy) is regulated by DOF transcription factors in aleurone cells upon germination. J Exp Bot. 56, 547–556, doi: 10.1093/jxb/eri033 (2005).
Gasteiger, E. Protein identification and Analysis Tools on the ExPASy Server in Proteomics Protocols Handbook (eds Walker, John M. ) pp. 571–607 (Humana, 2005).
Tamura, K., Stecher, G., Peterson, D., Filipsk, I. A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729, doi: 10.1093/molbev/mst197 (2013).
Tang H. et al. SynFind: Compiling Syntenic Regions across Any Set of Genomes on Demand. Genome Biol Evol. 11, 7(12), 3286–3298, doi: 10.1093/gbe/evv219 (2015).
Krzywinski M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645, doi: 10.1101/gr.092759.109 (2009).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452, doi: 10.1093/bioinformatics/btp187 (2009).
Guo, A. Y., Zhu, Q. H., Chen, X. & Luo, J. C. GSDS: a gene structure display server. Yi Chuan 29, 1023–1026 (2007).
Zhang, Y. miRU: an automated plant miRNA target prediction server. Nucleic Acids Res. 33 (Web Server issue), W701–4, doi: 10.1093/nar/gki383 (2005).
Li, Z. et al. Rna-seq improves annotation of protein-coding genes in the cucumber genome. BMC Genomics. 12, 540, doi: 10.1186/1471-2164-12-540 (2011).
Wang, H. Z., Li, S. J. & Guan W. The identification and genetic analysis of target spot in cucumber sources. China Vegetables 1, 24–25 (2010).
Acknowledgements
This work was supported in part by grants from the National Science & Technology Pillar Program (2015BAD02B00, 2014BAD01B09 and 2012BAD02B03), the National Natural Science Foundation of China (No. 31401873), the Technology Foundation for Selected Overseas Chinese Scholar, Ministry of Human Resources and Social Security of the People Republic of China, the Beijing Municipal Human Resource and Social Security Bureau and the Beijing Academy of Agricultural and Forestry Sciences (GJHZ201505/QNJJ201501).
Author information
Authors and Affiliations
Contributions
C.-l.W., Q.C., Y.W. and Y.X. designed the research. Q.C., A.M., S.Y. and J.Y. performed the research. C.-l.W., Q.C., L.Z. and Y.W. analysed the data and wrote the manuscript. All authors have read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wen, Cl., Cheng, Q., Zhao, L. et al. Identification and characterisation of Dof transcription factors in the cucumber genome. Sci Rep 6, 23072 (2016). https://doi.org/10.1038/srep23072
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep23072
- Springer Nature Limited
This article is cited by
-
Overexpression of BBF3 encoding a tobacco Dof transcription factor alters gene expression and cell death induction during N-mediated hypersensitivity to tobacco mosaic virus
Journal of General Plant Pathology (2024)
-
Morphological, physio-biochemical, and molecular indications of heat stress tolerance in cucumber
Scientific Reports (2023)
-
Analyses of Sequence Features and Expression Patterns of DOF Transcription Factor Genes in Pearl Millet (Pennisetum glaucum (L.) R. Br.)
Tropical Plant Biology (2023)
-
Genome-wide identification and characterization of a plant-specific Dof transcription factor gene family in olive (Olea europaea) and its comparison with Arabidopsis
Horticulture, Environment, and Biotechnology (2021)
-
Genome-wide survey of sugar beet (Beta vulgaris subsp. vulgaris) Dof transcription factors reveals structural diversity, evolutionary expansion and involvement in taproot development and biotic stress response
Biologia (2021)