Abstract
Inhibition of bacterial growth under aerobic conditions by elevated levels of cyclic adenosine 3′,5′-monophosphate (cAMP), first revealed more than 50 years ago, was attributed to accumulation of toxic methylglyoxal (MG). Here, we report a Crp-dependent mechanism rather than MG accumulation that accounts for the phenotype in Shewanella oneidensis, an emerging research model for the bacterial physiology. We show that a similar phenotype can be obtained by removing CpdA, a cAMP phosphodiesterase that appears more effective than its Escherichia coli counterpart. Although production of heme c and cytochromes c is correlated well with cAMP levels, neither is sufficient for the retarded growth. Quantities of overall cytochromes c increased substantially in the presence of elevated cAMP, a phenomenon resembling cells respiring on non-oxygen electron acceptors. In contrast, transcription of Crp-dependent genes encoding both cytochromes bd and cbb3 oxidases is substantially repressed under the same condition. Overall, our results suggest that cAMP of elevated levels drives cells into a low-energetic status, under which aerobic respiration is inhibited.
Similar content being viewed by others
Introduction
Among living organisms, prokaryotes thrive in every potential habitat on the Earth suitable for life because of their unparallel metabolic diversity. In many bacteria, central to regulation of metabolism is the cAMP (cyclic adenosine 3′,5′-monophosphate)-Crp (cAMP receptor protein) regulatory system, as clearly illustrated in many bacteria, Escherichia coli in particular1,2. The primary role of the canonical cAMP-Crp system, revealed mostly by early studies on Escherichia coli, is to regulate uptake of preferred carbon sources and repression of genes required for utilization of less preferred ones, a process called carbon catabolite repression (CCR)3. However, this turns to be only the tip of the iceberg as more and more biological processes are reported to be regulated by the system in diverse bacteria1. By using a robust top-down physiological approach, You et al. recently demonstrated that the physiological function of the cAMP-Crp system is to coordinate the allocation of proteomic resources with different metabolic demands in different nutrient environments4. Although environmental cues that modulate cAMP signals vary depending on species or even strains (for example, several α-ketoacids in E. coli), there is a possibility that most, if not all, of relevant bacteria use the cAMP-Crp system the same way that E. coli does. That is, the transcription of Crp-dependent genes could be differently regulated by altered cAMP levels in response to environmental changes.
Shewanella, a genus of Gram-negative γ-proteobacteria thriving in diverse environments, possess highly adaptable metabolism, a quality that could be exploited for potential applications in bioremediation of heavy metals and energy generation via fuel cells5,6. While this subject has been a focus for more than two decades, the genus is now emerging as an important research model for general bacterial physiology. Many physiological traits displayed by shewanellae, mostly based on studies of the genus representative Shewanella oneidensis, are distinct, not found in Escherichia coli and other well-characterized model microorganisms. In addition, shewanellae are regarded as a reservoir for antibiotic resistance and the number of Shewanella species identified as pathogenic to animals including human being has been increasing with time7,8.
S. oneidensis is a strictly respiratory organism because the gene encoding 6-phosphofructokinase (PFK), an essential enzyme of glycolysis, is missing9. Moreover, the ability of S. oneidensis to utilize five- and six-carbon carbohydrates is rather poor because of the scarcity of enzymes for such sugars and their transport10. Despite this, the bacterium, probably all of shewanellae, is regarded respiratory versatile because it derives energy by coupling organic matter oxidation to the respiration of an array of terminal electron acceptors (EAs), such as oxygen, fumarate, nitrate and metal oxides5. To date, how this bacterium adopts different metabolic modes in response to the availability of different EAs has been intensively studied and some progresses have been made. First, Fnr (fumarate/nitrate regulator), whose E. coli courterpart is the major player in respiration, has no significant role in bacterial physiology11. Second, S. oneidensis uses the Arc (aerobic respiration control) system for regulating aerobic respiration without affecting genes in the tricarboxylic acid (TCA) cycle12. Third, it is evident that Crp is crucial in respiration because crp mutants are defective in utilizing several EAs, including oxygen, Fe3+, Mn4+, nitrate, nitrite, fumarate and dimethyl sulfoxide (DMSO)13,14,15,16,17,18,19.
During aerobiosis, the primary targets of the cAMP-Crp regulatory system are genes encoding terminal reductases, including those reducing oxygen but traditionally called as cytochrome oxidases16,17. Cytochrome oxidases generate energy by coupling the oxidation of a respiratory substrate such as a c-type cytochrome or quinol to the reduction of oxygen to water20. Like in most bacteria, there are multiple cytochrome oxidases in S. oneidensis, two cytochrome c oxidases (a caa3-type and a cbb3-type) and a bd-type quinol oxidase17. For respiration of oxygen, cytochrome cbb3 is the predominant system whereas cytochrome caa3 is not of significance17. Cytochrome bd, on the other hand, appears to mainly facilitate adaptation to a variety of stress conditions, especially nitrite, although it is able to support growth when cytochrome cbb3 is absent16,21.
Initially observed in E. coli and later in other bacteria, aerobic growth is impeded when cAMP is present at concentrations of 0.5 mM or higher with certain sugars as carbon sources22,23. This effect of cAMP is attributed to accumulation of methylglyoxal (MG), which is a toxic intermediate produced from dihydroxyacetone phosphate (DHAP) by MG synthase (MGS)22,24.
We have found by chance, in the course of studies on the cAMP-CRP regulation of genes for nitrate and nitrite reductases15, that cAMP at 2 mM also retarded aerobic growth in S. oneidensis. However, a gene encoding an E. coli MGS homologue is missing in the S. oneidensis genome9. Thus, possibilities for the growth defect associated with cAMP include i) another protein functioning as MGS if MG is responsible and (ii) a different mechanism. In this report, we describe the investigation of the subject. Our results demonstrate that cAMP at elevated levels retards growth mainly by compromising transcription of Crp-dependent genes for both the cytochrome cbb3-type and bd oxygen oxidases.
Methods
Bacterial strains, plasmids and culture conditions
The bacterial strains and plasmids used in this study are listed in Table 1. Sequences of the primers used in this study are available upon request. All chemicals are from Sigma-Aldrich Co. unless otherwise noted. E. coli and S. oneidensis were grown aerobically in Lysogeny broth (LB, Difco, Detroit, MI) at 37 and 30 °C for genetic manipulation. When appropriate, the growth medium was supplemented with the following: 2,6-diaminopimelic acid (DAP), 0.3 mM; ampicillin, 50 μg/ml; kanamycin, 50 μg/ml; gentamycin, 15 μg/ml.
Growth of S. oneidensis strains under aerobic conditions was measured at 600 nm (OD600) in either LB or MS defined medium, which contains 30 mM lactate as electron donor used as previously described25. For aerobic growth, mid-log phase cultures were inoculated into fresh media to an OD600 of ∼0.02 and shaken at 200 rpm at 30 °C.
In-frame mutant construction and complementation
In-frame deletion strains were constructed using the att-based fusion PCR method as described previously26. In brief, two fragments flanking the genes of interest were amplified by PCR and then linked by a second round of PCR. The fused fragments were introduced into plasmid pHGM01 using the Gateway BP clonase II enzyme mix (Invitrogen) according to the manufacturer’s instruction. Vectors carrying mutational constructs in E. coli WM3064, were subsequently transferred into S. oneidensis via conjugation. Integration of the mutagenized constructs into the chromosome was selected by resistance to gentamycin and confirmed by PCR. These transconjugants were grown in LB broth in the absence of NaCl and plated on LB supplemented with 10% sucrose. Gentamycin-sensitive and sucrose-resistant colonies were screened by PCR for deletions of the target genes. Mutants were verified by sequencing the region containing the intended mutations.
Plasmids pHG101 were used for genetic complementation of the mutants27. Wild-type genes and their adjacent promoters, were generated by PCR and cloned into pHG101. For inducible gene expression, genes of interest generated by PCR were introduced into pHGE-Ptac under the control IPTG-inducible promoter Ptac28. After verification by sequencing, the vectors were transferred into the relevant strains via conjugation for complementation and/or expression.
Chemical assays
Cultures of 3 ml grown to an OD600 of ∼0.2 were subjected to filtering through a 0.22 μm nylon membrane for separation of cells and cell-free filtrate. The filtrate was immediately for cAMP assay, which was performed by using a commercially available kit (cAMP direct immunoassay kit, BioVision, http://www.biovision.com/camp-direct-immunoassay-kit-colorimetric-2862.html) according to the manufacturer’s instructions. The external cAMP levels were used to estimate the cAMP excretion rate by multiplying the specific growth rate and normalizing to OD600 values as described elsewhere4. The relative cAMP excretion rate for each mutant strain was given by comparing to that of the wild-type, representing the relative internal cAMP level because it is proportional to the cAMP excretion rate29. Amounts of MG and heme c from cells were measured following the procedures described elsewhere22,30. Standard curves were made with commercial agents each time.
Viability assay
S. oneidensis strains grown to an OD600 of ∼0.2 were incubated with 0.4 mM MG or 4 mM cAMP for half an hour, then adjusted to approximately 107 CFUs/ml and followed by 10-fold serial dilutions. Ten microliters of each dilution was spotted onto LB plates. For nitrite susceptibility assay, ten microliters of each dilution of the untreated was spotted onto LB plates containing 5 mM nitrite. The plates were incubated at 30 °C before being read.
Cytochromes cbb3 activity assay
Visual analysis of cbb3 activity was done by staining colonies with the agents for the Nadi Assay. Nadi reactions were carried out by the addition of a-naphthol and N′,N′-dimethyl-p-phenylenediamine (DMPD) on LB agar plates31. Colonies were timed for formation of the indophenol blue.
SDS-PAGE and heme-staining
Unless otherwise noted, mid-log phase cells were harvested, washed with phosphate buffered saline (PBS), resuspended in the same buffer and sonicated. Protein concentrations of the cell lysates were determined by the bicinchoninic acid assay (Pierce Chemical). The cell lysates were resolved by SDS-PAGE using 12% polyacrylamide gels and stained with 3,3′,5,5′-tetramethylbenzidine (TMBZ) as described elsewhere32.
Promoter activity assay
The activity of various promoters was assessed using a single-copy integrative lacZ reporter system as described previously33. A fragment containing the sequence upstream of each operon from −300 to +1 (relative to the translation start codon) was amplified and cloned into the reporter vector pHGEI01 and verified by sequencing, These plasmids were then transferred by conjugation into relevant S. oneidensis strains. Plasmid pHGEI01 containing promoters of interest integrates into the chromosome and the antibiotic marker is then removed by an established approach16,33. Cells grown to the mid-log phase were collected and β-galactosidase activity assays were performed with an assay kit as described previously27.
Other analyses
Student’s t test was performed for pairwise comparisons. Values are presented as means +/− standard deviation (SD) in the relevant figures.
Results
Growth inhibition by cAMP is not due to accumulation of methylglyoxal in S. oneidensis
This investigation began with the chance observation that cAMP at 2 mM significantly retards aerobic growth of S. oneidensis in LB broth. To further assess the effect of cAMP on growth, we added cAMP of varying concentrations into liquid cultures (∼0.05 of OD600) prepared from the mid-log phase cells and monitored the consequences (Fig. 1A). While the addition of 1 mM cAMP hardly affected growth, the molecule at higher concentrations (2 and 4 mM) inhibited growth significantly and inhibition increased with cAMP levels. A similar trend was observed from the MS defined medium, but inhibition appeared more severe, with no visible growth in the presence of 4 mM cAMP (Fig. S1). Nevertheless, in both cases cell densities increased constantly when growth was not completely prohibited, a phenomenon not observed in E. coli, whose growth is completely arrested by much less cAMP (0.5 mM)22. In addition, we examined the effect of cAMP on viability. Cells of the mid-log phase (∼0.2 of OD600) were incubated with 4 mM cAMP for half an hour, properly diluted and dropped on LB plates. As shown in Fig. 1B, cell viability of S. oneidensis appeared to be slightly reduced. But this was due to the growth defect because there was no difference in the number of viable cells between samples treated by cAMP and not from viable-cell counting (data not shown). Thus we concluded that cell viability is not affected significantly by cAMP, distinct from the fact that E. coli cells die rapidly because of the MG accumulation22. These contrasting phenotypes suggest that the growth defect of S. oneidensis resulting from exogenous cAMP may not be due to a toxic metabolite.
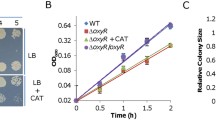
Effect of cAMP and MG on S. oneidensis physiology.
(A) LB broth containing cAMP (0–4 mM) or MG (0–0.4 mM) was inoculated with mid-log phase S. oneidensis cultures (~0.2 of OD600), incubated (200 rpm) under aerobic conditions. (B) Viability assessment. Mid-log phase S. oneidensis cultures were incubated with 4 mM cAMP or 0.4 mM MG, serially diluted and plated on LB plates. Photos were taken after 24 h. All experiments were performed at least three times with standard deviations presented as error bars in (A) and representative results presented in (B).
To rule out the possibility that MG underlies the growth defect in the presence of cAMP, we assessed impact of MG on growth. As shown in Fig. 1A, influences of cAMP and MG on growth were clearly different. We then examined MG on viability with cells prepared the same as above. Cells incubated with 0.4 mM MG for half an hour exhibited significantly reduced viability (Fig. 1B). Furthermore, despite of the lack of an E. coli MGS homologue, we examined levels of MG produced endogenously. In either rich or defined medium containing cAMP at concentrations that displays the strongest inhibition, MG was below the detection limit (data not shown). These data collectively conclude that the growth defect resulting from high concentrations of cAMP is not due to MG.
An S. oneidensis cpdA mutant is defective in aerobic growth
Given the data presented above, we reasoned that mutants lacking enzymes that catalyze cAMP degradation are likely more sensitive to the molecule. To date, such enzymes for cAMP hydrolysis have not been characterized in S. oneidensis. However, CpdA (SO_3901) appears to be an E. coli cAMP phosphodiesterase homologue encoded in the S. oneidensis genome, with 45% identity in amino acid sequence and an E value of 2e-76 in a BLASTp analysis. To confirm that S. oneidensis CpdA functions as a cAMP phosphodiesterase, we constructed a cpdA in-frame deletion strain (ΔcpdA). It is immediately evident that growth of ΔcpdA was significantly impaired (Fig. 2A). To validate this phenotype, a copy of the S. oneidensis cpdA gene under the control of the IPTG-inducible promoter (Ptac) was introduced into the mutant (Fig. 2A). Growth defect was partially corrected in the absence of IPTG because the promoter is slightly leaky34,35. With IPTG ranging from 0.01 to 0.2 mM, complementation was successful (Fig. 2A), indicating that the growth defect of the ΔcpdA strain was due to the intended mutation per se. More importantly, E. coli cpdA was also able to complement the growth defect, albeit not as effectively as its S. oneidensis counterpart (Fig. 2A).
S. oneidensis CpdA is a cAMP phosphodiesterase.
(A) Complementation of growth defect of the cpdA mutant in LB broth with the wild-type as control (WT). S. oneidensis cpdA (SocpdA) and E. coli cpdA (EccpdA) were placed behind the IPTG-inducible Ptac promoter as described in the experimental procedures. The cpdA mutants carrying empty vector (vec), S. oneidensis cpdA (SocpdA) and E. coli cpdA (EccpdA) were examined without IPTG or with 0.2 mM IPTG. (B) cAMP levels in cultures in (A) The averaged cAMP level in WT was set to 100%, to which cAMP levels in other strains were normalized. Experiments were performed at least three times with error bars representing the standard deviation.
To further provide evidence for the role of S. oneidensis CpdA as a cAMP phosphodiesterase, we assayed cAMP levels in relevant strains. Consistent with a previous report about an E. coli ∆cpdA strain36, intracellular levels of cAMP in an S. oneidensis ∆cpdA strain increased by over 2.5-fold (Fig. 2B). When either the S. oneidensis or E. coli cpdA gene was expressed, cAMP levels reduced greatly. Notably, with IPTG at 0.2 mM, cAMP levels between cells expressing the S. oneidensis cpdA and E. coli cpdA gene differed markedly, implying a difference in the efficacy of these two enzymes. Together with functional prediction based on the sequence, these data manifest that S. oneidensis CpdA functions to decompose cAMP.
Growth inhibition by cAMP is dependent on CRP in S. oneidensis
To unravel the mechanism responsible for the cAMP inhibition in S. oneidensis, we first examined whether such effect of cAMP requires Crp. A crp deletion strain (∆crp), whose aerobic growth is only slightly impaired11, was subject to the analysis of cAMP effect. In contrast to the wild-type, the Δcrp strain was resistant to exogenous cAMP with respect to growth (Fig. 3A). As this observation was confirmed by genetic complementation with an integrative system described in our previous study16, it supports that Crp is essential for cAMP-induced growth deficiency.
Growth inhibition by cAMP is dependent on Crp in S. oneidensis.
(A) Growth of a crp mutant in the presence of 4 mM cAMP (marked with an asterisk). ∆crp/crp represents the crp mutant carrying a single copy of crp for complementation, which was successful8. (B) Growth of ∆cpdA, ∆cya, ∆cpdA∆crp and ∆cpdA∆cya strains in LB broth. In both A and B, error bars representing standard deviations from at least three independent experiments.
In bacteria, cAMP is synthesized by adenylate cyclases (ACs)3,36. The S. oneidensis genome encodes three functional ACs, CyaA (SO_4312), CyaB (SO_3778) and CyaC (SO_1329), which have been characterized with respect to cAMP synthesis. Among them, CyaC is the major AC for cAMP production; the loss of all three ACs results in a phenotype similar to that of a crp mutant, in line with that both cAMP and Crp are essential to the physiological role of the cAMP-Crp complex14. To confirm that the growth defect requires the cAMP-Crp complex, we removed crp and cya (all three genes for ACs) from the ΔcpdA strain. In contrast to the ΔcpdA strain, the newly constructed ΔcpdAΔcrp and ΔcpdAΔcya strains displayed normal growth, comparable to that of the Δcrp strain (Fig. 3B). Based on these results, we conclude that the growth defect, resulting from either addition of exogenous cAMP or the cpdA mutation, is dependent on the cAMP-Crp complex.
Intracellular cAMP influences quantities of cytochromes c
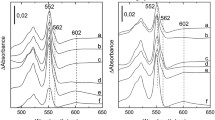
S. oneidensis colonies are brown-red on plates, largely because of more than 40 c-type cytochromes11,37,38. Previously, we reported that the loss of Crp decreases the levels of c-type cytochromes approximately by 60%11. During this investigation, we noticed that the color of Δcya colonies (or cell pellets) was similar (Fig. 4A). In contrast, the color of ΔcpdA colonies was much deeper, so was the wild-type with 4 mM cAMP. We therefore hypothesized that the levels of c-type cytochromes increase with intracellular cAMP. To test this, heme c levels in relevant strains were determined with a ΔccmF mutant used as negative control (Fig. 4A). The ccmF gene encodes a cytochrome c heme lyase, which is essential to c-type cytochrome maturation in S. oneidensis28,39. Compared to the wild-type, deletion of cya significantly lowered levels of heme c, which was comparable to that of the Δcrp strain and could be recovered by exogenous cAMP. In contrast, inactivation of cpdA elevated levels of heme c. Moreover, levels of heme c in the ΔcpdAΔcrp and ΔcpdAΔcya strains were similar to those of the Δcrp and Δcya strains. This observation was further confirmed by the profile of c-type cytochromes revealed by heme-staining (Fig. 4B). In a word, these data clearly show that the levels of c-type cytochromes in S. oneidensis increase with cAMP.
cAMP influences quantities of cytochromes c.
(A) cAMP influences heme c levels. Mid-log phase cultures (~0.2 of OD600) of indicated strains were pelletted and photographed, then were lysed for quantition of heme c levels. The average amount of heme c from the wild-type strain was set to 100%. ∆ccmF, which could not produce cytochromes c, was used as negative control. WT/cAMP represents WT grown with 2 mM cAMP.(B) Proteins (10 μg per lane) extracted from the indicated samples were resolved by SDS-PAGE and analyzed by heme staining. All experiments were performed at least three times with standard deviations presented as error bars or similar results were obtained.
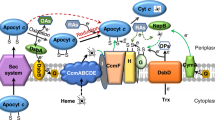
cAMP-CRP regulates heme biosynthesis and cytochrome c maturation
To elucidate the mechanism underlying growth defect and/or increased production of c-type cytochromes caused by the cpdA mutation, we focused on the heme synthetic pathway and the cytochrome c maturation system. S. oneidensis possesses the most common pathway for heme synthesis (Fig. 5A), as illustrated in E. coli, which entails nine reactions that converts glutamyl-tRNA to protoporphyrin IX40,41. Interestingly, there are multiple candidates for HemB, HemG and HemH. To determine which of heme synthetic genes are affected by cAMP, we monitored abundance of the transcript of these hem genes by qRT-PCR provided that they are not organized into operons except hemC and hemD. In the ∆cya and ∆cpdA strains, the hemA gene was repressed and induced approximately 2-fold respectively, whereas the other hem genes were affected insignificantly (Fig. 5A). To confirm this observation, we used a lacZ-reporter to assay β-galactosidase activities driven by hemA, hemG2 and hemC promoters. Although robustness of these promoters differed substantially, they showed the same trend as observed from qRT-PCR (Fig. 5B).
cAMP-CRP regulates heme biosynthesis and cytochrome c maturation.
(A) Expression of the hem and ccm genes in ∆cya and ∆cpdA analyzed by qRT-PCR. Enzymes for heme biosynthesis are shown above: multiple candidates for HemB, HemG and HemH are present; HemN and HemF HemN and HemF are coproporphyrinogen III oxidases, catalyzing the same reaction under different conditions. Cells of mid-log phase were prepared as described in the experimental procedures. The averaged expression level of each gene in mutants was compared to that in the wild-type. (B) Five genes were further analyzed by lacZ-reporter for confirmation. (C) Heme c levels in the wild-type overproducing HemA or CcmF. The average amount of heme c from the wild-type strain containing the empty vector was set to 100%. (D) Growth of the wild-type under indicated conditions as in C. All experiments were performed in triplicate and error bars indicate the standard error.
HemA (glutamyl-tRNA reductase) catalyzes the first dedicated, rating-limiting step in heme synthesis40. To test whether HemA accounts for the phenotype of the ∆cya and ∆cpdA strains, we placed the hemA gene under the control of IPTG-inducible promoter Ptac to examine effects of HemA of varying quantities on heme c levels and growth of the wild-type. Surprisingly, HemA influenced the heme c levels in a dose-dependent way (Fig. 5C). With IPTG at no more than 0.05 mM, the heme c levels increased with HemA but further enhanced production of HemA by IPTG at 0.1 mM and above played an inhibitory role, resulting in significant reduction in heme c levels. Altered production of HemA also had an apparent impact on growth (Fig. 5D). When IPTG was added to levels more than 0.1 mM, growth was significantly retarded. In contrast, HemA induced by IPTG at 0.05 mM or lower did not exert any negative effect on growth.
We then examined whether the cytochrome c maturation system may be the cause for growth defect and increased heme c levels of the cpdA mutant. In contrast to the hem genes, the ccm genes are organized into three operons, ccmABCDE, ccmI and ccmFGH39. qRT-PCR analysis of the transcript of the ccmA, ccmI and ccmF genes revealed that the ccmF operon but not others was affected by the both cya and cpdA mutations (Fig. 5A). This observation was then confirmed by using the lacZ-reporter (Fig. 5B). Interestingly, forced production of CcmF by IPTG displayed an effect on heme c levels similar to that observed from HemA, although it appeared milder in the overproduction end (Fig. 5C). Consistently, growth was also similarly impacted (Fig. 5D). All together, these data suggest that HemA and CcmF, when present in certain range, can modestly affect quantities of c-type cytochromes, but in large excess exert a significant negative impact on c-type cytochrome production. Despite this, it is clear that neither of these two proteins appears to be critical for the growth defect of the cpdA mutant because their overproduction compromises quantities of c-type cytochromes.
cAMP in excess inhibits activity of both cytochrome bd and cbb3 oxidases
To look further for answers addressing the growth defect of the cpdA mutant, we turned to cytochrome oxidases because these enzymes provide proton motive force for energy under aerobic conditions. Moreover, prior studies showed that the functional oxidases, bd-type (encoded by cydABX) and cbb3-type (encoded by ccoNOPQ), are under the direct control of the cAMP-Crp complex16,17,21. To test activities of the bd-type and cbb3-type oxidases in the ΔcpdA strain, we performed nitrite susceptibility assay and Nadi plate assay, respectively. Consistent with the notion that cytochrome bd confers resistance to nitrite in S. oneidensis16, a cyd null mutant (Δcyd) was hypersensitive to nitrite (Fig. 6A). Like the Δcrp and Δcya strains, the ΔcpdA strain displayed substantially increased susceptibility to nitrite and loss of both crp and cpdA did not further elevate susceptibility. Importantly, this increased susceptibility due to the CpdA loss was restored to the level of wild-type by its expression in trans, only in the presence of cytochrome bd. Moreover, the phenotype was also complemented by forced production of cytochrome bd.
cAMP in excess inhibits cytochromes cbb3 and bd.
(A) Effect of cAMP on susceptibility of indicated strains to 5 mM nitrite on LB plates. Cytochrome bd confers S. oneidensis resistance to nitrite. ∆cpdA/cpdA, ∆cpdA∆crp/cpdA and ∆cpdA∆crp/cyd represent indicated mutants expressing cpdA or cyd by IPTG at levels sufficiently high for successful complementation as described in the text or previous publications. (B) Effect of cAMP on cytochrome cbb3 activity of indicated strains on LB plates by the Nadi assay. The method is based on the rapid formation of indophenol blue from colorless a-naphthol catalyzed by cytochrome c oxidase, using N′,N′-dimethyl-p-phenylenediamine monohydrochloride as an exogenous electron donor. Photos were taken at indicated times after the reaction started. The wild-type and ΔccoN strains serve as positive and negative controls. (C) Growth of ∆cpdA∆cyd in LB. (D) Growth of ∆cpdA∆cco in LB. (E) Impacts of cAMP on expression of cyd and cco operons. Activities of the cyd and cco operon promoters from mid-log phase samples were assayed by a lacZ-reporter system used previously17. ∆cpdA/cpdA and ∆cpdA∆crp/cpdA represent indicated mutants expressing cpdA by IPTG at levels sufficiently high for successful complementation as described in the text or previous publications. All experiments were performed at least three times with standard deviations presented as error bars or similar results were obtained.
In the case of the cytochrome cbb3, similar results were obtained. With Nadi assay, which specifically detects cytochrome c oxidase-dependent respiration31, we visualized activities of the cytochrome cbb3 in relevant strains (Fig. 6B). As shown before17, loss of Crp compromised the cytochrome cbb3 activity. Surprisingly, the cytochrome cbb3 activity was most drastically reduced in the ΔcpdA strain, with the indophenol blue ring barely visible in one minute. This severe defect was dependent on Crp as the ΔcpdAΔcrp and Δcrp strains were indistinguishable.
Under standard conditions, the cytochrome bd is dispensable for aerobic growth of the S. oneidensis wild-type17. However, this was not observed with the cpdA mutation as the ΔcpdAΔcyd strain had growth defect more severe than the ΔcpdA strain (Fig. 6C), suggesting that the cytochrome bd is crucial for supporting growth when the cpdA gene is absent. Similar results were obtained from the ΔcpdAΔccoN strain (Fig. 6D). Notably, the ΔcpdAΔcco strain had the slowest growth rate when compared to the wild type, ΔcpdA and ΔcpdAΔcyd strains, suggesting that in the ΔcpdA strain the cytochrome cbb3 still plays a predominant role in supporting growth as in the wild-type. These data, collectively, indicate that activities of both cytochrome bd and cbb3 oxygen reductases are impaired in the ΔcpdA strain, leading to growth deficiency.
To unravel the mechanism for reduced activities of both oxidases, we examined their expression levels in the ΔcpdA strain. As shown in Fig. 6E, there was no difference in activities of the cyd promoter in strains lacking any of tested genes, crp, cya, cpdA, or even two of them combined, suggesting that cAMP in absent and in excess has a similar regulatory effect on cyd expression. On the contrary, loss of crp or cya resulted in a modest reduction in cco expression but cAMP in overabundance was more detrimental. Altogether, these results suggest that the growth defect of the cpdA mutant is due to reduced production of both cytochrome bd and cbb3.
Discussion
Cyclic nucleotides act as second messengers in diverse signaling cascades throughout all kingdoms of life, among which cAMP is first discovered and most extensively studied in bacteria1,42. The actions of cAMP are mediated by downstream cAMP-binding proteins, which are involved in diverse processes43,44,45,46. In bacteria, the central to cAMP-mediated regulation is formation of the cAMP-Crp complex, a transcriptional regulator of a number of metabolic operons, including those involved in the transport of substrates, glycolysis, the tricarboxylic acid cycle and aerobic respiration2,47. It has been proposed that a key physiological role of the cAMP-Crp complex is to ensure the proteomic resources to be spent on distinct metabolic sectors as needed in different nutrient environments4. As a consequence, some carbon sources are transported and utilized when cAMP levels are manipulated. One of such examples is xylose, which can be converted to be DHAP, leading to production of toxic MG and thereby growth defect22. However, this is not the case in S. oneidensis. As our data presented here eliminate the possibility that MG is accountable for the growth phenotype in S. oneidensis, a different mechanism must exist.
The purpose of this study was to unravel the mechanism. The study was facilitated by an S. oneidensis cpdA mutant that stably maintains intracellular cAMP at levels sufficiently high to elicit a similar growth defect as the wild-type with 2 ~ 4 mM cAMP. S. oneidensis CpdA, A homolog of the E. coli counterpart, is verified to be a cAMP phosphodiesterase by cross-complementation. It should be noted that the E. coli CpdA could not fully complement the phenotype of the S. oneidensis cpdA mutant. With IPTG at 2 mM, forced production of S. oneidensis CpdA reduces the cAMP concentration below the wild-type level whereas cells producing E. coli CpdA exhibit only a 2-fold decrease in cAMP concentration (Fig. 2). This may not be surprising as the cAMP phosphodiesterase activity of E. coli CpdA is poor, with a rather high Km for cAMP (~500 μM) relative to intracellular cAMP concentration11. Thus, at least in the context of in vivo data presented here, S. oneidensis CpdA functions more effectively than its E. coli counterpart.
As the growth defect of S. oneidensis caused by cAMP at elevated concentrations is dependent on Crp, we adopt the non-hypothesis-driven approach of testing whether some members of the cAMP-Crp regulon might be accountable for the defect when expressed differently. In addition to growth defect, the cpdA mutant, as well as strains lacking either ACs or Crp, differs from the wild-type in color of colony/pellet, which largely reflects the cellular amount of c-type cytochromes11. Apparently, cAMP levels correlate well with overall production of c-type cytochromes. Given that reduced quantities of c-type cytochromes, as due to loss of either ACs or Crp, do not significantly impede aerobic growth11,13,14, we tested whether c-type cytochromes in increased production could lead to retarded growth under aerobic conditions.
Amounts of c-type cytochromes are determined by two systems, responsible for the heme synthesis and cytochrome c maturation respectively. The heme synthesis is carried out by 10 enzymes (HemN and HemF for the same reaction under different conditions), of which only a few are found to be conditionally inducible48. In the present study, we found that hemA, whose product catalyzes the rating-limiting step40, is induced about 2-fold in the cpdA mutant. Previous studies have shown that HemA in Salmonella typhimurium responds to heme availability at the level of protein lifetime49,50: when heme is abundant, it binds to HemA to promote degradation of the latter. While whether the stabilization of S. oneidensis HemA is also an issue remains unknown, its induction by increased concentrations of cAMP, to our knowledge, is unprecedented. Seemingly, this regulation by cAMP-Crp is indirect because by prediction there is no Crp-binding site located upstream of the hemA gene11,51. However, given the negative effect of overproduced HemA on overall amounts of c-type cytochromes, the possibility that HemA plays an important role in the growth defect appears small. Intriguingly, this is also true of CcmF, the cytochrome c lyase. We therefore conclude that neither heme synthesis nor cytochrome c maturation is accountable for growth defect or increased level of c-type cytochromes observed from the cpdA mutant.
Rather, increased levels of heme c may be a result of concerted upregulation of many cytochrome c genes because more than two thirds of them are predicted to be under the direct control of the cAMP-Crp complex11,13,14,33,51. This surely gains support from heme-staining analysis (Fig. 4). We have previously shown that anaerobic respiration of various EAs favors overall cytochrome c production11,14, a scenario resembling the cpdA mutant to some extent. It has been suggested that cAMP concentrations increase in response to the low internal energetic status in E. coli, promoting catabolism and inhibiting anabolism4,52. In S. oneidensis, a similar notion has been proposed17. Thus, it seems logic that elevated cAMP drives cells into a low-energy mode, favoring respiration of non-oxygen EAs. As a consequence, genes encoding proteins important for respiration of oxygen are repressed, such as those for cytochrome cbb3 and bd. We propose that this explains the growth defect.
According to previous reports, S. oneidensis cAMP-Crp binds to DNA motifs similar to its E. coli counterpart whereas Crp alone fails in binding11,15,16,17,18,19. Data presented here reveal that cAMP at varying levels impacts expression of Crp-regulon members differently: the cyd operon (bd) behaves the same in cAMP-deficient and -overproduction strains whereas the cco operon (cbb3) is further repressed by increased concentrations of cAMP. This is consistent with the finding that S. oneidensis Crp functions in a dose-dependent manner16. Coincidently, a study of E. coli demonstrates that cAMP-binding has a biphasic effect on site-specific DNA-binding by Crp53.
In recent years, cAMP-Crp complexes with distinct features have been found. In Mycobacterium, Crp can not only operate at extremely high levels of cAMP, based on the finding that the intracellular cAMP levels are as high as 3–4 mM54, but also bind to DNA in a specific manner and regulate transcription without cAMP55. In Pseudomonas putida, a bacterium that also has an incomplete glycolysis pathway (lacking PFK), Crp exhibits an affinity binding of cAMP approximately 1000 times higher than that of E. coli Crp56,57. A consequence of these differences is that the Crp regulons of these bacteria, including S. oneidensis, differ drastically from that of E. coli, as suggested in P. putida58.
Additional Information
How to cite this article: Yin, J. et al. Reduced expression of cytochrome oxidases largely explains cAMP inhibition of aerobic growth in Shewanella oneidensis. Sci. Rep. 6, 24449; doi: 10.1038/srep24449 (2016).
References
McDonough, K. A. & Rodriguez, A. The myriad roles of cyclic AMP in microbial pathogens: from signal to sword. Nat. Rev. Micro. 10, 27–38 (2012).
Green, J. et al. Cyclic-AMP and bacterial cyclic-AMP receptor proteins revisited: adaptation for different ecological niches. Curr. Opin. Microbiol. 18, 1–7 (2014).
Botsford, J. L. & Harman, J. G. Cyclic AMP in prokaryotes. Microbiol. Rev. 56, 100–122 (1992).
You, C. et al. Coordination of bacterial proteome with metabolism by cyclic AMP signalling. Nature 500, 301–306 (2013).
Fredrickson, J. K. et al. Towards environmental systems biology of Shewanella. Nat. Rev. Micro. 6, 592–603 (2008).
Malvankar, N. S. & Lovley, D. R. Microbial nanowires for bioenergy applications. Curr. Opin. Biotechnol. 27, 88–95 (2014).
Janda, J. M. & Abbott, S. L. The genus Shewanella: from the briny depths below to human pathogen. Crit. Rev. Microbiol. 40, 293–312 (2014).
Yin, J., Sun, Y., Mao, Y., Jin, M. & Gao, H. PBP1a/LpoA but not PBP1b/LpoB are involved in regulation of the major β-lactamase gene blaA in Shewanella oneidensis. Antimicrob. Agents Chemother. 59. 3357–3364 (2015).
Heidelberg, J. F. et al. Genome sequence of the dissimilatory metal ion-reducing bacterium Shewanella oneidensis. Nat. Biotech. 20, 1118–1123 (2002).
Serres, M. H. & Riley, M. Genomic analysis of carbon source metabolism of Shewanella oneidensis MR-1: predictions versus experiments. J. Bacteriol. 188, 4601–4609 (2006).
Gao, H. et al. Physiological roles of ArcA, Crp and EtrA and their interactive control on aerobic and anaerobic respiration in Shewanella oneidensis. Plos ONE 5, e15295 (2010).
Gao, H., Wang, X., Yang, Z., Palzkill, T. & Zhou, J. Probing regulon of ArcA in Shewanella oneidensis MR-1 by integrated genomic analyses. BMC Genomics 9, 42 (2008).
Saffarini, D. A., Schultz, R. & Beliaev, A. Involvement of cyclic AMP (cAMP) and cAMP receptor protein in anaerobic respiration of Shewanella oneidensis. J. Bacteriol. 185, 3668–3671 (2003).
Charania, M. A. et al. Involvement of a membrane-bound class III adenylate cyclase in regulation of anaerobic respiration in Shewanella oneidensis MR-1. J. Bacteriol. 191, 4298–4306 (2009).
Dong, Y. et al. A Crp-dependent two-component system regulates nitrate and nitrite respiration in Shewanella oneidensis. PLos ONE 7, e51643 (2012).
Fu, H. et al. Crp-dependent cytochrome bd oxidase confers nitrite resistance to Shewanella oneidensis. Environ. Microbiol. 15, 2198–2212 (2013).
Zhou, G. et al. Combined effect of loss of the caa3 oxidase and Crp regulation drives Shewanella to thrive in redox-stratified environments. ISME J. 7, 1752–1763 (2013).
Wu, G., Li, N., Mao, Y., Zhou, G. & Gao, H. Endogenous generation of hydrogen sulfide and its regulation in Shewanella oneidensis. Front. Microbiol. 6, 374 (2015).
Gao, T., Ju, L., Yin, J. & Gao, H. Positive regulation of the Shewanella oneidensis OmpS38, a major porin facilitating anaerobic respiration, by Crp and Fur. Sci. Rep. 5, 14263 (2015).
Borisov, V. B., Gennis, R. B., Hemp, J. & Verkhovsky, M. I. The cytochrome bd respiratory oxygen reductases. Biochim. Biophys. Acta 1807, 1398–1413 (2011).
Chen, H., Luo, Q., Yin, J., Gao, T. & Gao, H. Evidence for the requirement of CydX in function but not assembly of the cytochrome bd oxidase in Shewanella oneidensis. Biochim. Biophys. Acta 1850, 318–328 (2015).
Ackerman, R. S., Cozzarelli, N. R. & Epstein, W. Accumulation of toxic concentrations of methylglyoxal by wild-type Escherichia coli K-12. J. Bacteriol. 119, 357–362 (1974).
Botsford, J. L. Cyclic AMP phosphodiesterase in Salmonella typhimurium: characteristics and physiological function. J. Bacteriol. 160, 826–830 (1984).
Ferguson, G. P., Tötemeyer, S., MacLean, M. J. & Booth, I. R. Methylglyoxal production in bacteria: suicide or survival? Arch. Microbiol. 170, 209–218 (1998).
Shi, M., Wan, F., Mao, Y. & Gao, H. Unraveling the mechanism for the viability deficiency of Shewanella oneidensis oxyR null mutant. J. Bacteriol. 197, 2179–2189 (2015).
Jin, M. et al. Unique organizational and functional features of the cytochrome c maturation system in Shewanella oneidensis. PLos ONE 8(9), e75610 (2013).
Wu, L., Wang, J., Tang, P., Chen, H. & Gao, H. Genetic and molecular characterization of flagellar assembly in Shewanella oneidensis. PLos ONE 6, e21479 (2011).
Luo, Q., Dong, Y., Chen, H. & Gao, H. Mislocalization of Rieske protein PetA predominantly accounts for the aerobic growth defect of tat mutants in Shewanella oneidensis. PLos ONE 8, e62064 (2013).
Epstein, W., Rothman Denes, L. B. & Hesse, J. Adenosine 3′:5′ cyclic monophosphate as mediator of catabolite repression in Escherichia coli. Proc. Natl. Acad. Sci. USA 72, 2300–2304 (1975).
Berry, E. A. & Trumpower, B. L. Simultaneous determination of hemes a, b and c from pyridine hemochrome spectra. Anal. Chem. 161, 1–15 (1987).
Marrs, B. & Gest, H. Genetic mutations affecting the respiratory electron-transport system of the photosynthetic bacterium Rhodopseudomonas capsulata. J. Bacteriol. 114, 1045–1051 (1973).
Thomas, P. E., Ryan, D. & Levin, W. An improved staining procedure for detection of peroxidase activity of cytochrome P-450 on sodium dodecyl sulphate polyacrylamide gels. Anal. Biochem. 75, 168–176 (1976).
Fu, H., Jin, M., Ju, L., Mao, Y. & Gao, H. Evidence for function overlapping of CymA and the cytochrome bc1 complex in the Shewanella oneidensis nitrate and nitrite respiration. Environ. Microbiol. 16, 3181–3195 (2014).
Shi, M., Gao, T., Ju, L., Yao, Y. & Gao, H. Effects of FlrBC on flagellar biosynthesis of Shewanella oneidensis. Mol. Microbiol. 93, 1269–1283 (2014).
Gao, T., Shi, M., Ju, L. & Gao, H. Investigation into FlhFG reveals distinct features of FlhF in regulating flagellum polarity in Shewanella oneidensis. Mol. Microbiol. 98, 571–585 (2015).
Imamura, R. et al. Identification of the cpdA gene encoding cyclic 3′,5′-adenosine monophosphate phosphodiesterase in Escherichia coli. J. Biol. Chem. 271, 25423–25429 (1996).
Meyer, T. E. et al. Identification of 42 possible cytochrome c genes in the Shewanella oneidensis genome and characterization of six soluble cytochromes. OMICS 81, 57–77 (2004).
Gao, H. et al. Impacts of Shewanella oneidensis c-type cytochromes on aerobic and anaerobic respiration. Microb. Biotechnol. 3, 455–466 (2010).
Fu, H., Jin, M., Wan, F. & Gao, H. Shewanella oneidensis cytochrome c maturation component CcmI is essential for heme attachment at the non-canonical motif of nitrite reductase NrfA. Mol. Microbiol. 95, 410–425 (2015).
Heinemann, I. U., Jahn, M. & Jahn, D. The biochemistry of heme biosynthesis. Arch. Biochem. Biophys. 474, 238–251 (2008).
Brennan, C. M. et al. Reduced heme levels underlie the exponential growth defect of the Shewanella oneidensis hfq Mutant. PLoS ONE 9, e109879 (2014).
Rehmann, H., Wittinghofer, A. & Bos, J. L. Capturing cyclic nucleotides in action: snapshots from crystallographic studies. Nat. Rev. Mol. Cell. Biol. 8, 63–73 (2007).
Endoh, T. & Engel, J. N. CbpA: a polarly localized novel cyclic AMP-binding protein in Pseudomonas aeruginosa. J. Bacteriol. 191, 7193–7205 (2009).
Schünke, S., Stoldt, M., Novak, K., Kaupp, U. B. & Willbold, D. Solution structure of the Mesorhizobium loti K1 channel cyclic nucleotide‐binding domain in complex with cAMP. EMBO rep. 10, 729–735 (2009).
Liu, W.-j., Dong, H., Peng, X.-w. & Wu, Q.-m. The Cyclic AMP-binding protein CbpB in Brucella melitensis and its role in cell envelope integrity, resistance to detergent and virulence. FEMS Microbiol. Lett. 356, 79–88 (2014).
Banerjee, A. et al. A universal stress protein (USP) in Mycobacteria binds cAMP. J. Biol. Chem. 290, 12731–12743 (2015).
Haverkorn van Rijsewijk, B. R. B., Nanchen, A., Nallet, S., Kleijn, R. J. & Sauer, U. Large-scale 13C-flux analysis reveals distinct transcriptional control of respiratory and fermentative metabolism in Escherichia coli. Mol. Syst. Biol. 7, 477 (2011).
Mancini, S. & Imlay, J. A. The induction of two biosynthetic enzymes helps Escherichia coli sustain heme synthesis and activate catalase during hydrogen peroxide stress. Mol. Microbiol. 96, 744–763 (2015).
Wang, L., Elliott, M. & Elliott, T. Conditional stability of the HemA protein (glutamyl-tRNA reductase) regulates heme biosynthesis in Salmonella typhimurium. J. Bacteriol. 181, 1211–1219 (1999).
Jones, A. M. & Elliott, T. A purified mutant HemA protein from Salmonella enterica serovar Typhimurium lacks bound heme and is defective for heme-mediated regulation in vivo. FEMS Microbiol. Lett. 307, 41–47 (2010).
Novichkov, P. S. et al. RegPrecise 3.0–A resource for genome-scale exploration of transcriptional regulation in bacteria. BMC Genomics 14, 745–745 (2013).
Narang, A. Quantitative effect and regulatory function of cyclic adenosine 5′-phosphate in Escherichia coli. J. Biosci. 34, 445–463 (2009).
Passner, J. M. & Steitz, T. A. The structure of a CAP-DNA complex having two cAMP molecules bound to each monomer. Proc. Natl. Acad. Sci. USA 94, 2843–2847 (1997).
Dass, B. K. M., Sharma, R., Shenoy, A. R., Mattoo, R. & Visweswariah, S. S. Cyclic AMP in Mycobacteria: characterization and functional role of the rv1647 ortholog in Mycobacterium smegmatis. J. Bacteriol. 190, 3824–3834 (2008).
Stapleton, M. et al. Mycobacterium tuberculosis cAMP receptor protein (Rv3676) differs from the Escherichia coli paradigm in its cAMP binding and DNA binding properties and transcription activation properties. J. Biol. Chem. 285, 7016–7027 (2010).
Arce-Rodríguez, A. et al. The Crp regulator of Pseudomonas putida: Evidence of an unusually high affinity for its physiological effector, cAMP. Environ. Microbiol. 14, 702–713 (2012).
Chavarría, M., Nikel, P. I., Pérez-Pantoja, D. & de Lorenzo, V. The Entner–Doudoroff pathway empowers Pseudomonas putida KT2440 with a high tolerance to oxidative stress. Environ Microbiol 15, 1772–1785 (2013).
Milanesio, P., Arce-Rodríguez, A., Muñoz, A., Calles, B. & de Lorenzo, V. Regulatory exaptation of the catabolite repression protein (Crp)–cAMP system in Pseudomonas putida. Environ Microbiol 13, 324–339 (2011).
Acknowledgements
This research was supported by National Natural Science Foundation of China (31270097, 41476105) and the Fundamental Research Funds for the central Universities (2015FZA6001).
Author information
Authors and Affiliations
Contributions
H.G. conceived the idea and designed the project. J.Y., M.Q. and H.F. carried out the experiments. J.Y., M.Q. and H.G. analyzed data. J.Y., M.Q. and H.G. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Yin, J., Meng, Q., Fu, H. et al. Reduced expression of cytochrome oxidases largely explains cAMP inhibition of aerobic growth in Shewanella oneidensis. Sci Rep 6, 24449 (2016). https://doi.org/10.1038/srep24449
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep24449
- Springer Nature Limited
This article is cited by
-
Understanding and engineering electrochemically active bacteria for sustainable biotechnology
Bioresources and Bioprocessing (2019)
-
Loss of OxyR reduces efficacy of oxygen respiration in Shewanella oneidensis
Scientific Reports (2017)
-
Investigation of a spontaneous mutant reveals novel features of iron uptake in Shewanella oneidensis
Scientific Reports (2017)
-
NapB in excess inhibits growth of Shewanella oneidensis by dissipating electrons of the quinol pool
Scientific Reports (2016)