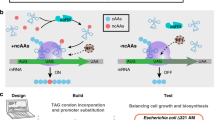
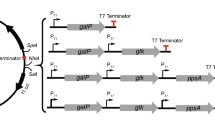
Abstract
Genome engineering has become a powerful tool for creating useful strains in research and industry. In this study, we applied singleplex and multiplex genome engineering approaches to construct an E. coli strain for the production of L-DOPA from glucose. We first used the singleplex genome engineering approach to create an L-DOPA-producing strain, E. coli DOPA-1, by deleting transcriptional regulators (tyrosine repressor tyrR and carbon storage regulator A csrA), altering glucose transport from the phosphotransferase system (PTS) to ATP-dependent uptake and the phosphorylation system overexpressing galactose permease gene (galP) and glucokinase gene (glk), knocking out glucose-6-phosphate dehydrogenase gene (zwf) and prephenate dehydratase and its leader peptide genes (pheLA) and integrating the fusion protein chimera of the downstream pathway of chorismate. Then, multiplex automated genome engineering (MAGE) based on 23 targets was used to further improve L-DOPA production. The resulting strain, E. coli DOPA-30N, produced 8.67 g/L of L-DOPA in 60 h in a 5 L fed-batch fermentation. This titer is the highest achieved in metabolically engineered E. coli having PHAH activity from glucose.
Similar content being viewed by others
Introduction
L-DOPA (3,4-dihydroxyphenyl-L-alanine) is an aromatic compound that is derived from L-tyrosine (Fig. 1). L-DOPA has been used to treat Parkinson’s disease, which is caused by deficiency of the neurotransmitter dopamine. Since Monsanto developed a commercial process for L-DOPA synthesis by asymmetric hydrogenation, L-DOPA has been produced by asymmetric, enzymatic and microbial synthesis1. However, the asymmetric synthesis has major disadvantages such as a poor conversion rate and low enantioselectivity. Thus, biotechnology approaches using microorganisms or enzymes have been explored as alternatives. Microorganisms with tyrosinase2,3,4,5,6,7,8, tyrosine phenol-lase (Tpl)9,10,11,12,13 and p-hydroxyphenylacetate 3-hydroxylase (PHAH)14 activity have been used to produce L-DOPA. However, the microbial fermentations require tyrosine or catechol/pyruvate as substrates, leading to high production costs. Nakagawa et al. constructed an E. coli expressing Streptomyces castaneoglobisporus tyrosinase gene, which can produce 293 mg/L of L-DOPA from glucose15. Muñoz et al. reported an engineered E. coli having PHAH activity, which can produce 1.5 g/L of L-DOPA from glucose16. However, the titer of L-DOPA in the engineered E. coli is lower than that of the microbial fermentation from tyrosine or catechol/pyruvate. Thus, further work must be carried out to increase L-DOPA production from glucose in E. coli.
Schematic representation of metabolic pathways involved in L-DOPA biosynthesis and regulation in E. coli.
The strategies for constructing a genetically defined strain for L-DOPA overproduction are also shown. The ×’s indicate that the genes are deleted. Encircled − or + symbols indicate inhibition or activation, respectively. The genes targeted by MAGE are underlined. PTS: phosphotransferase system; TCA: tricarboxylic acid cycle; G6P: glucose 6-phosphate; 6PBL: 6-phospho D-glucono-1,5-lactone; Ribu5P: D-ribulose 5-phosphate; X5P: D-xylulose 5-phosphate; R5P: D-ribose 5-phosphate; S7P: D-sedoheptulose 7-phosphate; F6P: fructose 6-phosphate; GAP: glyceraldehyde 3-phosphate; E4P: D-erythrose 4-phosphate; PEP: phosphoenolpyruvate; Pyr: pyruvate; Ac-CoA: acetyl-CoA; OAA: oxaloacetate; CIT: citrate; DAHP: 3-Deoxy-arabino-heptulonate 7-phosphate; DHQ: 3-Dehydroquinate; DHSH:3-Dehydroshikimate; SHK: shikimate; S3P: shikimate 3-phosphate; EPSP: 5-enolpyruvyl-shikimate 3-phosphate; CHA: Chorismate; PRE: prephenate; HPPH: 4-hydroxyphenylpyruvate. galP: galactose permease gene; glk: glucokinase gene; zwf: glucose-6-phosphate dehydrogenase gene; tktA: transketolase I gene; pckA: PEP carboxykinase gene; ppc: PEP carboxylase gene; ppsA: PEP synthase gene; pykFA: pyruvate kinase I/II gene; aroF, aroG and aroH: DAHP synthase gene; aroB: DHQ synthase gene; aroD: DHQ dehydratase; aroE/ydiB: shikimate/quinate dehydrogenase gene; aroA: 3-phosphoshikimate-1-carboxyvinyltransferase gene; aroC: CHA synthase; tyrA: CHA mutase/prephenate dehydrogenase gene; tyrB: tyrosine aminotransferase gene; trpED: anthranilate synthetase gene; pheA: prephenate dehydratase gene; hpaBC: E. coli W p-hydroxyphenylacetate 3-hydroxylase gene. nadK: NAD kinase gene; rpoD: sigma 70 factor gene; rpoA: a subunit of RNA polymerase gene; csrA: carbon storage regulator A; tyrR: tyrosine repressor.
Genome engineering is a powerful technique to manipulate entire genomes for obtaining desired phenotypes. The singleplex and multiplex genome engineering approaches have been successfully used for strain development17,18,19,20,21,22. Thus, we first focus on increasing the supply of the precursor, tyrosine, by using a singleplex genome engineering approach. We then apply multiplex automated genome engineering (MAGE) to develop an E. coli strain overproducing L-DOPA.
Results and Discussion
E. coli W hpaBC has been successfully introduced into E. coli to produce L-DOPA from glucose16. Figure 1 shows that tyrosine availability should first be increased to improve L-DOPA production from glucose. Successful strategies for engineering E. coli strains that can overproduce tyrosine include: (i) improving the carbon flow through the biosynthetic pathway of interest by removing transcriptional and allosteric regulation; (ii) increasing the availability of the direct precursors phosphoenolpyruvate (PEP) and erythrose-4-phosphate (E4P); (iii) preventing loss of carbon to competing pathways; (iv) enhancing the first enzymatic reaction of the shikimate pathway to yield 3-deoxy-D-arabino-heptulosonate-7-phosphate (DAHP); (v) and identifying and relieving rate-limiting enzymatic reactions. Thus, we first used singleplex genome engineering to increase the supply of tyrosine.
Removal of transcriptional regulators
Tyrosine repressor (TyrR) is a transcriptional dual regulator that represses the transcription of several genes encoding enzymes involved in aromatic acid biosynthesis23. Carbon storage regulator A (CsrA) is a regulator of carbohydrate metabolism. CsrA regulates the levels of three enzymes that participate directly in phosphoenolpyruvate (PEP) metabolism. It activates pyruvate kinase (PykF) and represses PEP carboxykinase (PckA) and PEP synthase (PpsA) in E. coli24. It has been reported that the inactivation of tyrR and csrA improves aromatic compound production25,26,27,28. Thus, we first deleted tyrR and csrA to obtain E. coli AROM-1 (Fig. 1), resulting in a slight increase in L-DOPA production from 138.7 ± 4.9 mg/L to 148.3 ± 11.7 (Table 1). Munoz et al. also reported that knocking out tyrR enhanced L-DOPA production in E. coli16.
Increasing the availabilities of the precursor PEP by altering glucose transport
Increasing PEP availability is a common strategy for engineering E. coli strains for the overproduction of aromatic compounds. In E. coli, glucose is mainly transported and phosphorylated by the phosphotransferase system (PTS). Under standard growth conditions, 50% of the glycolytic intermediate PEP resulting from the catabolism of glucose is used as the phosphate donor for phosphorylation and translocation by the PTS. The properties of the PTS limit the production of compounds that have PEP as a precursor. Carmona et al. suggested that inactivation of the PTS is the primary strategy for engineering E. coli to overproduce aromatic metabolites29. Thus, we deleted the PTS (ptsHIcrr) to further improve L-DOPA production. The inactivation of the PTS increased the L-DOPA titer to 176 ± 3.6 mg/L (Table 1). Non-PEP-mediated glucose transport and phosphorylation systems have successfully been used for the replacement of the PTS to increase PEP availability30,31,32. Thus, we integrated the galP and glk under the control of the P37 promoter into the E. coli knockout strain AROM-2 to obtain E. coli AROM-3. The titer of L-DOPA and growth of E. coli AROM-3 harboring pQE30-hpaBC showed no significant difference compared to E. coli AROM-2 (p < 0.05, Table 1).
Knockout of Glucose-6-phosphate dehydrogenase gene
Glucose-6-phosphate dehydrogenase (encoded by zwf) catalyzes the oxidization of glucose-6-phosphate to gluconate-6-phosphate. It has been reported that knocking out zwf drives more carbon flux into the Embden-Meyerhof-Parnas (EMP) pathway and tricarboxylic acid (TCA) cycle33. They also found that the zwf mutant is able to synthesize pentose phosphate (PP) pathway-derived compounds independently from the oxidative part of the PP pathway by directing its carbon flow from the EMP pathway directly into the non-oxidative part of the PP pathway. Thus, we disrupted zwf in E. coli AROM-3 to obtain E. coli AROM-4. E. coli AROM-4 (pQE30-hpaBC) produced L-DOPA at 205.3 ± 2.5 mg/L, which was greater than E. coli AROM-3 (pQE30-hpaBC) (Table 1). The stoichiometric analysis demonstrated that the yield of the aromatic compound DAHP approaches the theoretical maximum when E4P is provided by the nonoxidative part of the PP pathway and pyruvate is recycled to PEP by PpsA34. The improvement of L-DOPA titer after zwf deletion was experimentally demonstrated for the first time.
Removal of competing pathway
Prephenate can be converted into either tyrosine or phenylalanine. To eliminate the loss of prephenate to the competing reaction (phenylalanine biosynthesis), we deleted prephenate dehydratase and its leader peptide genes (pheLA) in E. coli AROM-4 to obtain E. coli TYR-1. The pheLA deletion slightly increased the L-DOPA titer to 209.2 ± 0.9 mg/L (Table 1). Some other groups have previously reported that the pheLA deletion increases L-tyrosine production35,36.
Coordinating expression of the downstream pathway of chorismate
The bifunctional enzyme Chorismate (CHA) mutase/prephenate dehydrogenase, TyrA, catalyzes the first and second step of L-tyrosine biosynthesis (Fig. 1). TyrA catalyzes both reactions in separate domains of the protein and the CHA mutase/prephenate hydrogenase is feedback-inhibited by L-tyrosine (up to 95% inhibition of the prephenate dehydrogenase and 45% of the CHA mutase activity28. Feedback-resistant mutants of the TyrA E. coli enzyme have been used for L-tyrosine overproduction35,36. Thus, TyrAfbr [M53I/A354V] was used to deregulate the feedback inhibition by tyrosine. Substrate channeling is a powerful tool for balancing the expression of genes. It can increase the catalytic efficiency of the sequential reactions in a biosynthetic pathway37,38. To increase the rate of CHA conversion to L-DOPA, we first fused the tyrAfbr, tyrB and hpaBC genes with a (G4S)3 linker, then integrated the fusion protein chimera under the control of the 7P37 promoter into the chromosome of E. coli TYR-1 to obtain E. coli DOPA-1. E. coli DOPA-1 produced 307.4 ± 3.7 mg/L of L-DOPA.
Multiplex automated genome engineering
MAGE is an efficient and rapid tool for the genome engineering of bacterial strains. We selected aroF, aroG, aroB, aroD, ydiB, aroE, ppsA, tktA, nadK, aroL, aroK, aroA, tyrA, tyrB and tyrAfbr (M53I/A354V) as target sites to tune translation by ribosome binding site (RBS) replacement (Fig. 1). The RBS sequences were designed to be DDRRRRRDDDD (D = A, G, T; R = A, G) with a total pool complexity of 3.5 × 105 (36 × 25 × 15). Six genes (aroFP148L, aroGD146N, tyrAM53I, tyrAA354V, rpoDD521E and rpoAV257R) were targeted for amino acid mutations in their open reading frames (ORF). The introduction mutations in aroF, aroG and tyrA were used to remove product feedback inhibition23,26,27,28,35,36. The rpoD and rpoA mutants have been successfully used to increase tyrosine production39. Two genes (trpD and trpE) were targeted for inactivation by introducing a revertible premature stop codon into each ORF. To increase the MAGE allelic replacement frequency, the methyl-directed mismatch repair protein gene (mutS) of E. coli DOPA-1 was first deleted to obtain E. coli DOPA-2. E. coli DOPA-2 (pSIM6) was used as the starting strain for MAGE. After 30 cycles of MAGE, 1.3 × 1010 genetic variants (4.3 × 108 bp variations per cycle for 30 MAGE cycles19) were generated. According to an allelic replacement efficiency calculation22, 30 MAGE cycles generate 2.3% of genomes with at least 3 out of 23 targeted loci and 6.1 × 10−12 of genomes with all 23 targeted loci. One hundred clones from the 5th, 10th, 15th, 20th and 25th cycle and 1000 clones from the 30th cycle were screened in deep-well microplate culture. L-DOPA can be easily oxidized to dopachrome and then polymerized nonenzymatically to form the black pigment melanin40. Thus, we selected strains that produced darker cultures for further analysis. Darker cultures in the 48-well microplates were selected for HPLC analysis to determine L-DOPA concentration. Six MAGE strains from the 30th cycle showed higher L-DOPA concentrations in the deep-well microplate analysis and these were further analyzed in shake flasks. Of the six strains, strain 30-30 produced the highest level of L-DOPA, which was 34% higher than that of the starting strain E. coli DOPA-2 (Table 2). Table 2 also shows that all MAGE strains produced more tyrosine and total tyrosine plus L-DOPA than the starting strain. The reason may be because the above modification strategies were used to increase the availability of the precursor, tyrosine. Thus, we removed pSIM6 from MAGE strain 30-30 to obtain E. coli DOPA-30, which was used as the L-DOPA-producing strain in subsequent tests. After sequencing, we found that three genes have codon mutations in their ORFs (aroF: P148L; tyrA: M53I and rpoD: D521E, Supplementary Table 1). Only 3 modified loci out of 23 targets may be due to the low MAGE allelic replacement frequency (ARF) for multiple targeted loci. Only 2–4 modified targets were also observed in the MAGE lycopene-producer after 35 cycle MAGE based on 20 targets19. The ARF may be increased by increasing cycle numbers, Coselection MAGE (CosMAGE)21 or CRMAGE41. CosMAGE improves the ARF of each target site by around four-fold21. CRMAGE increases the efficiency from 6% of traditional MAGE to 66%41.
As shown in Table 2, not all of the tyrosine was converted to L-DOPA in E. coli DOPA-30. In order to convert all L-tyrosine into L-DOPA, we added a single additional copy of the hpaBC into pQE30-hpaBC to obtain pQE30-2hpaBC and transformed the plasmid into E. coli DOPA-30. As shown in Table 3, overexpression of hpaBC in E. coli DOPA-30 indeed increased L-DOPA production, but this strain cannot also convert all the L-tyrosine into L-DOPA. However, the engineered E. coli with the hpaBC reported by Munoz et al. produced few L-tyrosine16. Comparing the sequence of the hpaBC in pQE30-2hpaBC with that reported by Munoz et al.16, the 5′-UTR sequence of the hpaC has been changed. The change may lead to the imbalanced expression between the hpaB and hpaC. Is this change resulted in the accumulation of L-tyrosine in the engineered strain? We re-amplified the hpaBC operon with the native 5-UTR sequence of the hpaC to obtain pQE30-hpaBCN. As shown in Table 3, E. coli DOPA-30 harboring pQE30-hpaBCN cannot produce L-tyrosine. Thus, the hpaBC in E. coli DOPA-30 was replaced with the hpaBCN to obtain E. coli DOPA-30N. As shown in Table 3, E. coli DOPA-30N cannot also produce L-tyrosine and produced 614.3 mg/L of L-DOPA.
Fed-batch fermentation
Fed-batch fermentation of E. coli DOPA-30N was performed in a 5 L bioreactor. As shown in Fig. 2, the strain produced 8.67 g/L of L-DOPA at 60 h. The OD600 of the culture reached 110. The L-DOPA productivity was 144.5 mg/L/h. The L-DOPA yield from glucose was 62.7 mg/g. The titer and yield were 5.7- and 1.2-fold higher than that reported by Muñoz et al.15, respectively. In addition, it was found that all the L-tyrosine was converted to L-DOPA after 40 h. The similar phenomenon was also observed by Muñoz et al.15. It indicates that the rate of hydroxylation of L-tyrosine by the HpaBC is slower than the rate of L-tyrosine synthesis. Therefore, the catalytic efficiency of the PHAH encoded by hpaBC should be improved.
Comparison with other microorganisms
L-DOPA production by microorganisms is summarized in Table 4. The L-DOPA titer obtained in this study is higher by a factor of 5.7 than the highest level previously reported using metabolically engineered E. coli strain that have PHAH activity from glucose16. The value is also higher than that obtained in microorganisms that have tyrosinase activity from tyrosine2,3,4,5,6,7,8. However, the value in this study is lower than that obtained in some microorganisms with Tpl activity from catechol and pyruvate9,10,12. It indicates that further works should be carried out for improving L-DOPA production.
Although the L-DOPA titer of our engineered E. coli is considerably higher than that previously reported, all of the tyrosine was converted to L-DOPA only after 40 h (Fig. 2). It indicates that PHAH is the rate-limited step for L-DOPA biosynthesis in this strain. The catalytic efficiency of the PHAH encoded by hpaBC should be improved. Directed evolution may be used to increase its catalytic efficiency. Because only three targets were found in the MAGE strain (Supplementary Table 1), we can apply other strategies to further enhance the availability of tyrosine, such as upregulating tktA, increasing NADPH availability and upregulating hpaBC.
In conclusion, we first constructed an L-DOPA-producing E. coli strain, DOPA-1, using a singleplex genome engineering approach based on knockouts of genes and integration of the tyrAfbr, tyrB and hpaBC fusion protein chimera. MAGE based on 23 targets was then used to further improve L-DOPA production, which yielded the strain E. coli DOPA-30N. E. coli DOPA-30N produced 8.67 g/L of L-DOPA in 60 h in a 5L fed-batch fermentation. This titer is the highest reported in metabolically engineered E. coli that has PHAH activity from glucose. This strain, E. coli DOPA-30N, can serve as a base strain for developing more efficient strains capable of producing L-DOPA or other aromatic compounds. The rapid and efficient markerless deletion approach using the IPTG-inducible ccdB as a counter-selectable marker will be generally useful for gene knockout of E. coli.
Methods
Strains, plasmids and primers
The strains and plasmids used in this study are listed in Table 5. The primers are listed in Supplementary Table 2.
Genetic methods
The genes hpaB and hpaC were amplified from E. coli W using the primers hpaB-F/hpaB-R and hpaC-F/hpaC-R, respectively. The hpaB fragment was cloned into the SacI/KpnI sites of pQE30 to obtain pQE30-hpaB. The hpaC fragment was cloned into the KpnI/SalI sites of pQE30-hpaB to obtain pQE30-hpaBC. The hpaBC genes were also amplified from pQE30-hpaBC using the primers hpaBC-F/hpaBC-R and then cloned into the SalI/HindIII sites of pQE30-hpaBC to obtain pQE30-2hpaBC. The hpaBC operon was amplied from E. coli W using the primers hpaB-F/hpaC-R and then cloned into the SacI/SalI to obtain pQE30-hpaBCN.
The knockouts of the csrA, tyrR and mutS genes were carried out according to the one-step inactivity method42 with the help of the pSIM6 plasmid43 expressing the lambda red recombination system. Gene knockouts were verified by colony PCR using appropriate primers (Supplementary Table 2).
The knockouts of other genes were carried out by a two-step recombination method using lambda red recombination and I-SceI cleavage as described as in Supplementary Fig. 1. The method was first reported by Yu et al.44. They used sacB as the counter-selectable marker. However, the efficiency of the first recombination is very low (24%) because sacB generally results in a certain number of false-positive colonies in the screening process due to mutation of sacB45. Thus, we used the IPTG-inducible ccdB gene as the counter-selectable marker. The ccdB gene was amplified from pOSIP-CH46 using the primers ccdBF/ccdBR, then cloned into the HindIII/XbaI sites of pXMJ1947 to obtain pXMJ-ccdB. The plasmid pXMJ-ccdB was digested by HindIII, blunted and self-ligated to obtain pEC-ccdB*. The IPTG-inducible ccdB gene was amplified from pXMJ-ccdB* using the primers ccdB*F/ccdB*R, then cloned into pMD18 to obtain pMD-lacI-PtacccdB. A kan resistance gene (encoding aminoglycoside 3′-phosphotransferase) containing I-SceI recognition sites was amplified from pK-JL48 using the primers kanF/kanR and then cloned into the XhoI/SpeI sites of pMD-lacI-PtrcccdB to obtain pMD-ccdBKanS. The I-Scel endonuclease gene was synthesized by Suzhou GENEWIZ, Inc. (Suzhou, China) and ligated into pUC57 to obtain pUC57-I-SceI. The I-Scel was cut from pUC57-I-SceI by EcoRI/KpnI and cloned into pBAD3049 to obtain pBAD30-I-SceI. The arabinose-inducible I-Scel was amplified from pBAD30-I-SceI using the primers IsceIF/IsceIR and cloned into the NdeI site of pSIM650 to obtain pSIMIS. The efficiency of the first recombination of the method reached 80.3%, which was much higher than that based on the sacB (24%, Supplementary Table 3).
Chromosomal integration was carried out by direct transformation as described by Chen et al.51 and Huang et al.52. The galP and glk genes were amplified from E. coli using the corresponding primers and cloned into pZSBP37 to obtain pZSBP-galP and pZSBP-glk, respectively. The glk gene under the control of the P37 promoter was digested with MluI/SalI from pZSBP-glk, then ligated into MluI/SalI-digested pHKKT5b to yield pHKKT5b-P37-glk. The galP gene under the control of the P37 promoter was digested with BglII/SalI from pZSBP-galP, then ligated into BamHI/SalI-digested pHKKT5b-P37-glk to yield pHKKT5b-P37-glk-P37-galP for chromosomal integration of P37-galP-P37-glk. The P37 promoter was amplified from pZSPB using the primers P37F/P37R and assembled into pZSPB by the BglBrick standard approach to produce pZSnP37 (n = 2, 3, 4, 5, 6 or 7), which has a tandem and stronger promoter. The tyrA and tyrB genes were amplified from E. coli using the corresponding primers and cloned into pMD-19T (simple) to obtain pMD-19T-tyrA and pMD-19T-tyrB, respectively. Site-directed mutagenesis was used to remove the BamHI/BglII sites and feedback inhibition of the tyrA to obtain pMD-19T-tyrAfbr. The hpaBC gene was amplified from pQE30-hpaBC using the primers hpaBCF1/hpaBCR2 and cloned into pMD-19T (simple) to obtain pMD-19T-hpaBC. The plasmid pMD-19T-tyrAfbr-tyrB-hpaBC containing the tyrAfbr-tyrB-hpaBC fusion protein chimera was assembled by the BglBrick standard approach. The fusion chimera fragment was cut from pMD-19T-tyrAfbr-tyrB-hpaBC by SphI/ApaI, then ligated into SphI/ApaI-digested pZS7P37 to yield pZS7P37-tyrAfbr-tyrB-hpaBC. The tyrAfbr-tyrB-hpaBC fragment under the control of the 7P37 promoter was cut from pZS.7P37-tyrAfbr-tyrB-hpaBC by MluI/BamHI, then cloned into the integration expression vector pP21KT5b to yield pP21KT5b-7P37-tyrAfbr-tyrB-hpaBC for chromosomal integration of 7P37-tyrAfbr-tyrB-hpaBC.
The replacement of 5′-UTR of the hpaC in E. coli DOPA-30 was carried out by the CRIPR-Cas method as described by Jang et al.53. The sgRNA fragment was amplified from pTargetF using the primers hpaCN20F/hpaCN20R and then cloned into the SpeI/XhoI sites of pTargetF to obtain the sgRNA plasmid pTargetF-hpaC. The target fragment was amplied from pQE30-hpaBCN using the primers hpaB/hpaBC.
MAGE and Screening of MAGE strains
Oligos were mixed in equimolar amounts to reach a final total oligo concentration of 1 μM. MAGE cycling was performed as previously described19,20,21. In brief, E. coli DOPA-3 harboring pSIM6 was grown in a 20-mL conical tube containing 5 mL of LB medium supplemented with 100 μg/mL ampicillin at 30 °C with 200 rpm agitation until the OD600 reached 0.5 to 0.7. Then, the cultures were heat-shocked in a shaking water bath at 42 °C for 15 min to induce the expression of λ Red recombination genes (gam, bet and exo). The cells were then chilled to 4 °C and centrifuged at 11,000 rpm for 30 s at 4 °C. The cultures were washed three times with ice-cold sterile 10% glycerol to remove salts. The cells were resuspended in 50 μL oligo mixture. Electroporation was carried out at 1.8 kV in 1-mm gap cuvettes on a Bio-Rad MicroPulser, BTX ECM-830. Cells were incubated in fresh LB low salt medium at 30 °C until their OD600 reached 0.4 to 0.6. The processes were repeated 30 times (30 MAGE cycles). After 5, 10, 15, 20, 25 and 30 cycles, the cells were grown overnight in 50 mL LB low salt medium and stored at −80 °C in a 15% (v/v) glycerol solution.
Cells from the 5th, 10th, 15th, 20th, 25th and 30th cycles were diluted, plated onto LB-agar plates with ampicillin and cultured overnight. Individual colonies were inoculated in individual wells of a 48-well deep-well microplate (4.6 mL) containing 600 mL of the fermentation medium without ascorbic acid and incubated at 30 °C with 200 rpm agitation for 48 h on a Microtron shaker (Infors). Because L-DOPA can be easily oxidized to dopachrome and then polymerized nonenzymatically to form melanin40, darker cultures were selected for HPLC analysis to determine L-DOPA concentration. Cultures with higher L-DOPA concentrations in the deep-well microplate analysis were selected for shake flask analysis. In the screening process, the culture temperature was set to 30 °C because the cells harbored pSIM6.
L-DOPA production in shake flasks
For L-DOPA production, a single colony was inoculated into 5 mL of LB medium in a 20-mL conical tube which was cultured overnight at 37 °C in a rotary shaker at 200 rpm. The overnight seed culture was then inoculated into 50 mL of fermentation medium with a starting OD600 of 0.1. The fermentation medium (pH 7.0) contains (g/L): peptone 10, yeast extract 5, NaCl 10, glucose 14, ascorbic acid 0.45 and 10 mL of trace element solution. The trace element solution contains (g/L): FeSO4·7H2O 10, ZnSO4·7H2O 2.2, MnSO4·4H2O 0.58, CuSO4·5H2O 1, (NH4)6Mo7O24·4H2O 0.1, Na2B4O7·10H2O 0.2 and HCl 10 mL. The main cultures were incubated at 37 °C for 48 h in a rotary shaking incubator at 150 rpm. IPTG was added as an inducer to a final concentration of 0.1 mM after 6 h when needed.
Fed-batch culture for L-DOPA production
The seed culture produced in 5 mL of LB medium was subcultured in 6 × 50 mL LB medium for 10–12 h with shaking at 200 rpm at 37 °C. The seed culture (~300 mL) was inoculated into a 5 L fermenter (Biostat B5, B. Braun, Germany) containing 3 L of fermentation medium with an initial OD600 of approximately 0.4. The fermentation medium (pH 7.0) contains (g/L): peptone 10, yeast extract 5, NaCl 10, glucose 25, (NH4)2SO4 15, KH2PO4 2, MgSO4·7 H2O 2, CaCl2 14.7 mg, thiamine 0.1 mg, ascorbic acid 1.8 and 1 mL of trace element solution. The trace element solution contains (mg/L): EDTA 8, CoCl2·6 H2O 2.5, MnCl2·4H2O 15, CuCl2·2H2O 1.5, H3BO3 3.0, Na2MoO4·2H2O 2.5, Zn(CH3COO)2·2H2O 13.0, Fe(III)citrate 100, thiamine·HCl 4.5. Fermentation was carried out at 37 °C with an airflow of 3 L/min and agitation rate of 600 rpm. IPTG was added as an inducer to a final concentration of 0.1 mM after 24 h. The pH was controlled at 7.0 by automatic addition of NH4OH. The feed solution (pH 7.0,) contains (g/L): glucose 500, tryptone 25, yeast extract 50, MgSO4·7H2O 17.2, (NH4)SO4 7.5, ascorbic acid 18. The feed was introduced continuously into the fermenter by using the pH-stat feeding strategy. Once the glucose is exhausted, the pH rises rapidly. When the pH was higher than 7.0 by 0.1 U, the feed was automatically added to the fermenter. A total of 680 mL feed solution was added. Samples were periodically withdrawn and the following parameters were measured: OD600, residual glucose concentration, tyrosine concentration and L-DOPA concentration. Fermentation experiments were carried out in triplicate.
Analytical methods
Growth was monitored by measuring the optical density at 600 nm. Tyrosine and L-DOPA in the supernatants were analyzed using a Shimadzu HPLC system (LC-20 A, Shimadzu, Japan) equipped with an Inertsil ODS-SP column (5 μm, 4.6 × 150 mm, GL Sciences Inc., Tokyo, Japan). The mobile phase was 0.2% TFA in 40% methanol, with a flow rate of 0.5 mL/min, at 30 °C. A photodiode array detector (SPD-M20A) operating at 280 nm was used and a standard curve was constructed from serial dilutions of a standard stock solution. Glucose concentration was determined by using glucose oxidase and a glucose assay kit (Shanghai Rongsheng Biotech Corporation, Shanghai, China).
Additional Information
How to cite this article: Wei, T. et al. Genome engineering Escherichia coli for L-DOPA overproduction from glucose. Sci. Rep. 6, 30080; doi: 10.1038/srep30080 (2016).
References
Min, K., Park, K., Park, D. H. & Yoo, Y. J. Overview on the biotechnological production of L-DOPA. Appl Microbiol Biot. 99, 575–584 (2015).
Krishnaveni, R., Rathod, V., Thakur, M. S. & Neelgund, Y. F. Transformation of L-Tyrosine to L-DOPA by a Novel Fungus, Acremonium rutilum, Under Submerged Fermentation. Curr Microbiol 58, 122–128 (2009).
Ali, S. & Ikram-ul-Haq Kinetic basis of celite (CM 2:1) addition on the biosynthesis of 3,4-dihydroxyphenyl-L-alanine (L-DOPA) by Aspergillus oryzae ME2 using L-tyrosine as a basal substrate. World J Microb Biot. 22, 347–353 (2006).
Ali, S., Ikram-ul-Haq, Qadeer, M. A. & Rajoka, M. I. Double mutant of Aspergillus oryzae for improved production of L-DOPA (3,4-dihydroxyphenyl-L-alanine) from L-tyrosine. Biotechnol Appl Bioc. 42, 143–149 (2005).
Ali, S., Shultz, J. L. & Ikram-ul-Haq High performance microbiological transformation of L-tyrosine to L-dopa by Yarrowia lipolytica NRRL-143. BMC Biotechnol. 7, 50 (2007).
Surwase, S. N. & Jadhav, J. P. Bioconversion of L-tyrosine to L-DOPA by a novel bacterium Bacillus sp. JPJ. Amino Acids 41, 495–506 (2011).
Surwase, S. N., Patil, S. A., Apine, O. A. & Jadhav, J. P. Efficient microbial conversion of L-tyrosine to L-DOPA by Brevundimonas sp. SGJ. Appl Biochem Biotech 167, 1015–1028 (2012).
Surwase, S. N., Patil, S. A., Jadhav, S. B. & Jadhav, J. P. Optimization of l-DOPA production by Brevundimonas sp SGJ using response surface methodology. Microb Biotechnol. 5, 731–737 (2012).
Koyanagi, T. et al. Effective production of 3,4-dihydroxyphenyl-L-alanine (L-DOPA) with Erwinia herbicola cells carrying a mutant transcriptional regulator TyrR. J Biotechnol. 115, 303–306 (2005).
Foor, F., Morin, N. & Bostian, K. A. Production of L-dihydroxyphenylalanine in Escherichia coli with the tyrosine phenol-lyase gene cloned from Erwinia herbicola. Appl Environ Microbiol. 59, 3070–3075 (1993).
Bielecki, S. & Bolek, R. Immobilization of recombinant E. coli cells with phenol-lyase activity. In Progress in Biotechnology vol. 11 (eds Wijffels, R. H. et al. ), 472–478 (Elsevier 1996).
Lee, S.-G., Ro, H.-S., Hong, S.-P., Kim, E.-H. & Sung, M.-H. Production of L-DOPA by thermostable tyrosine phenol-lyase of a Themophilic Symbiobacterium species overexpressed in recombinant Escherichia coli. J Microbiol Biotechnol. 6, 98–102 (1996).
Park, H.-S., Lee, J.-Y. & Kim, H.-S. Production of L-DOPA(3,4- dihydroxyphenyl-L-alanine) from benzene by using a hybrid pathway. Biotechnol Bioeng. 58, 339–343 (1998).
Lee, J.-Y. & Xun, L. Novel biological process for L-DOPA production from L-tyrosine by P-hydroxyphenylacetate 3-hydroxylase. Biotechnol Lett. 20, 479–482 (1998).
Nakagawa, A. et al. A bacterial platform for fermentative production of plant alkaloids. Nat Commun. 2, 326 (2011).
Muñoz, A. J. et al. Metabolic engineering of Escherichia coli for improving L-3,4-dihydroxyphenylalanine (L-DOPA) synthesis from glucose. J Ind Microbiol Biot. 38, 1845–1852 (2011).
Song, C. W., Lee, J. & Lee, S. Y. Genome engineering and gene expression control for bacterial strain development. Biotechnol J 10, 56–68 (2015).
Esvelt, K. M. & Wang, H. H. Genome-scale engineering for systems and synthetic biology. Mol Syst Biol. 9, 641 (2013).
Wang, H. H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–U133 (2009).
Gallagher, R. R., Li, Z., Lewis, A. O. & Isaacs, F. J. Rapid editing and evolution of bacterial genomes using libraries of synthetic DNA. Nat Protoc. 9, 2301–2316 (2014).
Carr, P. A. et al. Enhanced multiplex genome engineering through co-operative oligonucleotide co-selection. Nucleic Acids Res. 40, e132 (2012).
Wang, H. H. & Church, G. M. Multiplexed Genome Engineering and Genotyping Methods: Applications for Synthetic Biology and Metabolic Engineering. Method Enzymol. 498, 409–426 (2011).
Pittard, J., Camakaris, H. & Yang, J. The TyrR regulon. Mol Microbiol. 55, 16–26 (2005).
Suzuki, K., Babitzke, P., Kushner, S. R. & Romeo, T. Identification of a novel regulatory protein (CsrD) that targets the global regulatory RNAs CsrB and CsrC for degradation by RNase E. Gene Dev 20, 2605–2617 (2006).
Na, D. et al. Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs. Nat Biotechnol. 31, 170–174 (2013).
Yakandawala, N., Romeo, T., Friesen, A. D. & Madhyastha, S. Metabolic engineering of Escherichia coli to enhance phenylalanine production. Appl Microbiol Biot. 78, 283–291 (2008).
Jiang, M. & Zhang, H. Engineering the shikimate pathway for biosynthesis of molecules with pharmaceutical activities in E. coli. Curr Opin Biotechnol. 42, 1–6 (2016).
Rodriguez, A. et al. Engineering Escherichia coli to overproduce aromatic amino acids and derived compounds. Microb Cell Fact. 13, 126 (2014).
Carmona, S. B., Moreno, F., Bolivar, F., Gosset, G. & Escalante, A. Inactivation of the PTS as a strategy to engineer the production of aromatic metabolites in Escherichia coli. J Mol Microb Biotech. 25, 195–208 (2015).
Yi, J., Draths, K. M., Li, K. & Frost, J. W. Altered glucose transport and shikimate pathway product yields in E. coli. Biotechnol Progr. 19, 1450–1459 (2003).
Balderas-Hernandez, V. E. et al. Metabolic engineering for improving anthranilate synthesis from glucose in Escherichia coli. Microb Cell Fact 8, 19 (2009).
Hernandez-Montalvo, V. et al. Expression of galP and glk in a Escherichia coli PTS mutant restores glucose transport and increases glycolytic flux to fermentation products. Biotechnol Bioeng. 83, 687–694 (2003).
Zhao, J., Baba, T., Mori, H. & Shimizu, K. Effect of zwf gene knockout on the metabolism of Escherichia coli grown on glucose or acetate. Metab Eng. 6, 164–174 (2004).
Patnaik, R. & Liao, J. C. Engineering of Escherichia coli central metabolism for aromatic metabolite production with near theoretical yield. Appl Environ Microbiol. 60, 3903–3908 (1994).
Santos, C. N., Xiao, W. & Stephanopoulos, G. Rational, combinatorial and genomic approaches for engineering L-tyrosine production in Escherichia coli. Proc Natl Acad Sci USA 109, 13538–13543 (2012).
Olson, M. M. et al. Production of tyrosine from sucrose or glucose achieved by rapid genetic changes to phenylalanine-producing Escherichia coli strains. Appl Microbiol Biotechnol. 74, 1031–1040 (2007).
Zhang, Y. H. Substrate channeling and enzyme complexes for biotechnological applications. Biotechnol Adv. 29, 715–725 (2011).
Li, X. R., Tian, G. Q., Shen, H. J. & Liu, J. Z. Metabolic engineering of Escherichia coli to produce zeaxanthin. J Ind Microbiol Biotechnol. 42, 627–636 (2015).
Santos, C. N. S., Xiao, W. H. & Stephanopoulos, G. Rational, combinatorial and genomic approaches for engineering L-tyrosine production in Escherichia coli. P Natl Acad Sci USA 109, 13538–13543 (2012).
Claus, H. & Decker, H. Bacterial tyrosinases. Syst Appl Microbiol. 29, 3–14 (2006).
Ronda, C., Pedersen, L. E., Sommer, M. O. A. & Nielsen, A. T. CRMAGE: CRISPR Optimized MAGE Recombineering. Sci Rep-Uk 6, 19452 (2016).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. P Natl Acad Sci USA 97, 6640–6645 (2000).
Sharan, S. K., Thomason, L. C., Kuznetsov, S. G. & Court, D. L. Recombineering: a homologous recombination-based method of genetic engineering. Nat Protoc. 4, 206–223 (2009).
Yu, B. J. et al. Rapid and efficient construction of markerless deletions in the Escherichia coli genome. Nucleic Acids Res. 36, e84 (2008).
Mizoguchi, H., Tanaka-Masuda, K. & Mori, H. A simple method for multiple modification of the Escherichia coli K-12 chromosome. Biosci Biotech Bioch. 71, 2905–2911 (2007).
St-Pierre, F. et al. One-step cloning and chromosomal integration of DNA. ACS Synth Biol. 2, 537–541 (2013).
Jakoby, M., Ngouoto-Nkili, C. E. & Burkovski, A. Construction and application of new Corynebacterium glutamicum vectors. Biotechnol Tech. 13, 437–441 (1999).
Jiang, L. Y., Chen, S. G., Zhang, Y. Y. & Liu, J. Z. Metabolic evolution of Corynebacterium glutamicum for increased production of L-ornithine. Bmc Biotechnol. 13, 47 (2013).
Guzman, L. M., Belin, D., Carson, M. J. & Beckwith, J. Tight regulation, modulation and high-level expression by vectors containing the arabinose PBAD promoter. J Bacteriol. 177, 4121–4130 (1995).
Sharan, S. K., Thomason, L. C., Kuznetsov, S. G. & Court, D. L. Recombineering: a homologous recombination-based method of genetic engineering. Nat Protoc. 4, 206–223 (2009).
Chen, Y. Y. et al. Chromosomal evolution of Escherichia coli for the efficient production of lycopene. BMC Biotechnol. 13, 6 (2013).
Huang, M. T., Chen, Y. Y. & Liu, J. Z. Chromosomal engineering of Escherichia coli for efficient production of Coenzyme Q(10). Chinese J Chem Eng. 22, 559–569 (2014).
Jiang, Y. et al. Multigene editing in the Escherichia coli genome via the CRISPR-Cas9 system. Appl Environ Microb. 81, 2506–2514 (2015).
Acknowledgements
We are grateful to the National Natural Science Foundation of China (Grant Nos 30970089, 21276289, J1310025), the Natural Science Foundation of Guangdong Province (No. 2015A030311036), the Project of the Scientific and Technical Program of Guangdong Province (No. 2015A010107004) and the Project of the Scientific and Technical Program of Guangzhou (No. 201607010028) for their financial support.
Author information
Authors and Affiliations
Contributions
T.W. performed the experiments. B.-Y.C. developed the markerless deletion approach and performed gene deletions. J.-Z.L. directed the project and wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wei, T., Cheng, BY. & Liu, JZ. Genome engineering Escherichia coli for L-DOPA overproduction from glucose. Sci Rep 6, 30080 (2016). https://doi.org/10.1038/srep30080
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep30080
- Springer Nature Limited
This article is cited by
-
Biosynthesis of fragrance 2-phenylethanol from sugars by Pseudomonas putida
Biotechnology for Biofuels and Bioproducts (2024)
-
Overlapping promoter library designed for rational heterogenous expression in Cordyceps militaris
Microbial Cell Factories (2022)
-
Advances in engineering microbial biosynthesis of aromatic compounds and related compounds
Bioresources and Bioprocessing (2021)
-
Tyrosinase-based production of l-DOPA by Corynebacterium glutamicum
Applied Microbiology and Biotechnology (2021)
-
Application of CRISPR/Cas System in the Metabolic Engineering of Small Molecules
Molecular Biotechnology (2021)