Abstract
The transcription factor sterol regulatory element-binding protein-1c (SREBP-1c) is a key regulator of lipogenesis and insulin sensitivity and is associated with non-alcoholic fatty liver disease (NAFLD). Here, we assessed the impact of common single nucleotide polymorphisms (SNPs) in SREBF-1c on NAFLD susceptibility and associated metabolic phenotypes in a Han Chinese population. Four common SNPs (rs62064119, rs2297508, rs11868035 and rs13306741) in the SREBP-1c gene were selected and genotyped in 593 patients with NAFLD and 593 healthy controls. Unconditional logistic regression was performed to assess the risk of NAFLD by determining odds ratios and 95% confidence intervals (CIs). No significant differences in genotype and allele frequencies of these four SNPs were found between the NAFLD population and the controls (all P > 0.05). In addition, we did not find any association between the SREBF-1c SNPs and the clinical and biochemical parameters, such as body mass index, total cholesterol, high density lipoprotein-and low density lipoprotein-cholesterol or systolic and diastolic blood pressure, except that the rs2297508 C-allele or rs11868035 G-allele showed significant associations with lower triglyceride levels in control subjects (P < 0.01). Our findings suggested that the four polymorphisms in SREBF-1c gene are not associated with risk of NAFLD in the Chinese Han population.
Similar content being viewed by others
Introduction
Sterol regulatory element-binding proteins (SREBPs) are a family of transcription factors that activate the entire program of cholesterol and fatty acid synthesis in the liver1,2. To date, three SREBP isoforms, i.e., SREBP-1a, -1c and -2, have been identified and characterized. SREBP-1a and SREBP-1c are encoded by a single gene named sterol regulatory element–binding protein gene (SREBF)-1 (via specific promoters and alternative splicing)3,4 and SREBP-2 is encoded by a separate gene, SREBF-25.
SREBP-1c is the main SREBF1 isoform, which is highly expressed in many tissues in mice and humans, including the liver, adipose tissue and skeletal muscle6 and can enhance transcription of genes involved in fatty acid and triacylglycerol synthesis7,8. Overexpression of SREBP-1c induced insulin resistance, diabetes, non-alcoholic fatty liver disease (NAFLD) and accelerated atherosclerosis in mice9,10,11,12. Given the mediating role of SREBF-1c in the transcription of genes involved in de novo lipogenesis and triglyceride-rich lipoprotein metabolism, the SREBF-1c gene could be considered as a good candidate indicator to predict metabolic syndromes disorders.
NAFLD is a spectrum of disorders ranging from simple fatty liver (NAFL) to non-alcoholic steatohepatitis (NASH), with or without fibrosis/cirrhosis13. The mechanism leading to NAFLD remains unclear; however, it is now largely accepted that the risk of NAFLD increases exponentially with the presence of metabolic syndrome or its components, including central abdominal obesity, hypertension, dyslipidaemia and type 2 diabetes (T2D)14,15. Therefore, it is well recognized that NAFLD is the hepatic manifestation of metabolic syndrome16. Single nucleotide polymorphisms (SNPs) within the SREBF-1c gene have been connected to metabolic syndrome disorders in humans17,18. Specifically, the SNP rs11868035 A/G, located in the intron of the SREBF-1c gene modulates the risk of type 2 diabetes, insulin resistance, obesity and hypercholesterolemia in different ethnicities19,20,21. In addition, a nearby-located synonymous SNP in exon 18 (rs2297508), located 216 bp downstream of rs11868035 A/G, was also found to be associated with obesity, diabetes and male-specific hypertriacylglycerolaemia in a French population22. Although the effects of SREBF-1c gene variants on metabolic syndromes are well documented, little is known about their effects on NAFLD. To date, only one study has reported that rs11868035 A allele conferred increased risk of severe steatosis and non-alcoholic steatohepatitis in a North-Western Italy population19.
Given the strong association between NAFLD and multiple metabolic diseases, the aim of this study was to identify whether polymorphisms in the SREBF-1c gene are associated with increased risk of NAFLD in a population from China. Subsequently, we also examined the association of each SNP with the metabolic phenotypes.
Results
Descriptive characteristics of the study population
The descriptive characteristics of the study participants according to NAFLD status (NAFLD patients vs. controls) are shown in Table 1. Briefly, there were no statistically significant differences between the NAFLD patients and control subjects in terms of age, sex, marital status, education, smoking or income status (all P > 0.05, see), indicating that the frequency matching was adequate. However, the NAFLD patients had a lower percentage of tea drinkers, as well as higher occurrence of most of the risk factors for the metabolic syndrome, including elevated blood pressure, body mass index (BMI), fasting plasma glucose (FPG), total cholesterol (TC), trigylcerides (TGs), low density lipoprotein (LDL)-C and decreased high density lipoprotein (HDL)-C (each P < 0.05), whereas the opposite was observed for controls. Additionally, levels of alanine transaminase (ALT) and aspartate transaminase (AST) were significantly higher in the patients with NAFLD than in the controls (both P < 0.05).
Association between individual genotypes and risk of NAFLD
SNPs information for the SREBF-1c gene is shown in Table 2. These SNPs were genotyped in all 1,186 individuals using iPlex technology based on a MassARRAY platform and were commonly distributed in the study samples. The corresponding frequencies of the rs62064119 G, rs2297508 C, rs11868035 G and rs13306741 A alleles among the participants were 0.560, 0.130, 0.133 and 0.047 in NAFLD patients and 0.556, 0.120, 0.121 and 0.044 in controls, respectively. In the univariate analysis, after multiple comparison correction by permutation tests, there were no significant differences in the allele frequency of these four SNPs between the control group and NAFLD patients (empirical P-values = 0.39–0.87).
Distributions of the genotypes of these four polymorphisms in NAFLD patients and control subjects and their associations with NAFLD risk are summarized in Table 3. All of the observed genotype frequencies conformed to the Hardy–Weinberg equilibrium (HWE), both in NAFLD patients and the controls (PNAFLD = 0.130–0.054; Pcontrol = 0.350–0.882, respectively). There were no significant differences in the genotype distribution between the two groups (P > 0.05). Variables such as age, sex, income, marital status, education, smoking, tea drinking, BMI and other clinical features might affect the development of NAFLD; therefore, unconditional logistic regression analysis was further used to estimate associations between the SREBF-1c genotypes and the risk of NAFLD. After controlling for the effect of these confounding factors, we also did not find any significant effects of SREBF-1c SNPs on the risk of NAFLD.
We further investigated the effects of SREBF1c variants on liver function tests. As shown in Table 4, both the A-allele of rs11868035 and G-allele of rs2297508 were associated with significantly increased ALT levels (P < 0.05) in all studied subjects (i.e. n = 1186, without separating them into controls and cases). However, no statistical significance for the effect of SREBF-1c gene variants on liver function was found in both NAFLD and control subjects (each P > 0.05).
The linkage disequilibrium (LD) pattern among SREBF-1c polymorphisms was further evaluated in our study population. The disequilibrium statistics between rs62064119, rs2297508, rs11868035 and rs13306741 showed a very strong LD (D′ = 0.90–0.99) with each other and a haplotype block including rs2297508 and rs11868035 was detected. As shown in Table 5, we did not detect any significant difference in the overall distribution of haplotypes between cases and controls (Psim for global score test = 0.29).
Association of Clinical Parameters with different SREBF-1c gene variants in NAFLD Patients and controls
Given that SREBF-1c gene variants have been reported to be associated with the components of metabolic syndrome and the high association between NAFLD and multiple metabolic diseases, we further investigated whether SREBF-1c variants might be associated with alterations in metabolic parameters related to NAFLD in our population. We assessed the relationship between the four SREBF-1c variants and FPG, BMI and plasma lipids in the NAFLD and control subjects, respectively. As shown in Tables S1–4, no statistical significance for the associations between the SREBF-1c gene variants and all selected clinical parameters was found in both NAFLD and control subjects (each P > 0.05), except that the rs2297508 C-allele or rs11868035 G-allele showed significant associations with lower TG level in control subjects (rs2297508 GG vs. GC + CC: 1.27 ± 0.69 vs. 1.08 ± 0.44, p = 0.003, rs11868035 AA vs. AG + GG: 1.26 ± 0.68 vs. 1.08 ± 0.45, p = 0.004).
Discussion
In this study, to evaluate whether SNPs in the whole SREBF-1c gene are associated with NAFLD or metabolic parameters related to this disease, we enrolled a total of 1186 unrelated Chinese adults in a case-control study to investigate the relationship between four SREBF-1c polymorphisms (rs62064119, rs2297508, rs11868035 and rs13306741) and the risk of NAFLD. Our results failed to reveal an association between SNPs or common haplotypes of SREBF-1c and NAFLD in the Chinese Han population. In addition, we did not find any association between the SREBF-1c SNPs and BMI, TC, HDL-and LDL-cholesterol or systolic and diastolic blood pressure, both in NAFLD and control subjects, except that the rs2297508 C-allele or rs11868035 G-allele showed significant associations with a lower TG level in control subjects. Thus, our results suggested that these four SNPs in SREBF-1c gene are not associated with the risk of NAFLD in the Chinese Han population.
SREBP-1c is a well-known regulator of hepatic lipogenesis and insulin sensitivity. Animal studies have shown that increased hepatic SREBP-1c is associated with insulin resistance, diabetes and hepatic steatosis, partially by the enhanced synthesis of lipotoxic saturated fatty acids and increased hepatic gluconeogenesis23,24. Two SREBP-1c SNPs (rs11868035 A/G and rs2297508 G/C) were evaluated previously in relation to metabolic diseases risk and carriers of the rs11868035 A allele and rs2297508 G allele have been reported to significantly increase the risk of type 2 diabetes18,21. However, little is known about the effect of SREBF-1c gene variants on NAFLD, except that Musso et al. investigated the relationship between SREBF-1c rs11868035 A/G polymorphism and risk of NAFLD and found that rs11868035 A allele conferred increased risk of severe steatosis and non-alcoholic steatohepatitis in a North-Western Italy population19. This association, however, was not observed in our study: we did not observe any significant difference in rs11868035 genotypes between NAFLD cases and controls. Moreover, no association with the other three SNPs or any common haplotypes of the SREBPF-1c gene and NAFLD risk was observed. Since lack of association can always be due to lack of power, a small effect on NAFLD cannot be ruled out. However, based on our sample size and the minor allele frequency, the power of detecting an OR of 1.2 at a two-sided α = 0.05, were >80% respectively. These results suggest that these four SNPs in the SREBF-1c gene may not influence fatty liver disease in our population. The inconsistency may be explained by the following reasons: Firstly, the most prevalent alleles of rs11868035 and rs2297508 are different for various populations of different genetic background18,19,25. Genome diversity data extracted from http://www.ncbi.nlm.nih.gov/SNP/ shows that the frequencies of rs2297508 G allele and rs11868035 A allele in Chinese population are much higher than those in CEU population (rs2297508 G allele: 82.2% vs 25.8%; rs11868035 A: 83.7% vs 28.1%), which may partially explained the inconsistency. Secondly, it is now largely accepted that NAFLD is multifactorial and various genetic alterations and environmental factors influence the development and progression of NAFLD26,27. A genome-wide association study (GWAS) showed that the rs738409 G allele of Padiponutrin/patatin-like phospholipase domain-containing protein 3 (PNPLA3) was significantly associated with increased risk of NAFLD28, which was further replicated in several populations29,30. Moreover, Krawczyk et al. reported that carriers of both PNPLA3 rs738409 and SREBP1c rs2297508 risk genotypes significantly promoted hepatic fibrogenesis, as compared to patients carrying none of these variants31. Silico analysis also revealed the presence of a sterol response element (SRE) binding site on the PNPLA3 gene promoter and PNPLA3 expression is induced directly by SREBP1c32. In fact, we previously demonstrated that the rs738409 GG genotype of PNPLA3 significantly increased the risk of NAFLD in our cohort (adjusted OR = 2.25, 95% CI: 1.46–3.45 for GG genotype vs CC genotype; P = 0.001)30.However, in the present study, we did not find any joined effects between PNPLA3 rs738409 and SREBP1c polymorphisms on NAFLD (data not shown). Finally, the inconsistent results may be partially explained by the differences in the study design or statistical power. Thus, further studies in large population-based cohorts from different ethnicities are warranted to identify the role of SREBP1-c polymorphisms in NAFLD.
We further investigated the effects of SREBF1c variants on liver function tests (LFTs) and the results showed that both the A-allele of rs11868035 and G-allele of rs2297508 were associated with significantly increased ALT levels (P < 0.05) in the entire cohort (i.e. n = 1186, without separating them into controls and cases), but not in separate analysis for cases and controls. The explanation for this inconsistency was that, we conducted a case-control study of which the subjects were recruited based on their disease status, therefore the entire subjects (including cases and control) were not a random sampling from source population and the association between SREBF1c variants and increased LFTs in the entire subjects may be spurious due to selection bias. Up to date, there was no report about the association between SREBF-1c variants and LFTs, further studies in a large random sampling are warranted to identify the role of SREBP1-c polymorphisms in LFTs.
Previous reports regarding associations between SREBF-1c variants and quantitative metabolism have been inconclusive. Previous studies have indicated an association of the rs11868035 variant with increased fasting total and LDL, cholesterol levels in Caucasian and Danish populations, respectively21,22. In contrast, studies from Austrian and French populations did not show any association of the rs2297508 variant with fasting cholesterol level33,34 In the present study, we also evaluated the influence of SREBF-1c SNPs on metabolic syndrome component traits and the results showed that the rs2297508 GC + CC genotypes or rs11868035 TC + CC genotypes showed significant associations with lower TG levels compared with wild-type genotypes in the control group, which suggests that these variant have stronger effects on some metabolic traits than on NAFLD. Rs2297508 is a synonymous mutation and rs11868035 is intronic SNP. Although these two SNPs associated with serum TG do not directly change protein structure, they could influence various aspects of mRNA metabolism35. Further functional studies should provide additional insights into the role of these variants in lipid metabolism.
Our study had strengths and limitations. The strengths of our study include our ability to match cases of NAFLD with controls to enable us to find variants that might be associated with NAFLD. Secondly, the moderate sample size employed in this study could have reduced type II errors. Finally, extensive information on anthropometrics and lifestyle factors were collected in this study to adjust for confounding factors. However, the potential limitations of the present study should also be considered. Firstly, there was a recruitment bias resulting from the retrospective case–control study design that could be a possible confounding factor in our results; however, the results did not appear to be seriously affected by this bias, because the control group was in HWE. Secondly, the ultrasonographic definition of steatosis may have missed some cases of milder steatosis. However, recent standardized criteria have significantly improved the ultrasonographic diagnostic accuracy to detect even minor degrees of steatosis36. Finally, our study was performed on a specific population (i.e., a health examination population of Han Chinese) and thus might not be applicable to the entire population or other ethnicities. Further population-based association studies are warranted to verify our results, especially those including different ethnic groups.
In summary, our study confirmed that the polymorphisms of SREBF-1c gene are not related to the risk of NAFLD in the Chinese Han population. However, other genetic risk factors for this disease may exist. Further research is needed to screen novel biomarkers and explore the mechanism underlying the pathogenesis of NAFLD.
Materials and Methods
Study population
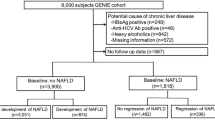
A 1:1 frequency-matched case–control Study of men and women aged 20–78 years was performed in a health examination centre of the Union Hospital of Fujian Medical University, from June of 2008 to May of 2012. Participants with anyone status of the following were excluded: 1) Alcohol abuse in the past year (weekly ethanol consumption ≥140 grams for men; ≥70 grams for women); 2) hepatitis B or hepatitis C virus infection; and 3) had received lipid-lowering treatment or any other drug-modifying lipid measures. The study design has been previously described in detail elsewhere37. Briefly, the case–control study comprised 593 NAFLD cases and 593 controls, each matched in terms of age (within 5 years), sex, ethnicity, occupation, geographical location and recruitment date. A case of NAFLD was defined by an abdominal ultrasonography made by two experienced radiologists blinded to the participants’ data. An ultrasonographic diagnosis of steatosis was based on recently proposed standardized criteria, which have substantially improved the diagnostic accuracy of ultrasonography36. Patients with secondary causes of steatosis, including total parenteral nutrition and the use of drugs known to precipitate steatosis were excluded. In addition, patients with any of the following diseases were excluded from participation in this study: autoimmune hepatitis, drug-induced liver disease, primary biliary cirrhosis and primary sclerosing cholangitis. Healthy individuals without steatosis by abdomen ultrasonography (n = 593) were recruited randomly from the same population during the same study period: these subjects underwent a routine health check and were free of any known major diseases. All enrolled subjects were of Chinese Han ethnicity. Informed, written consent was obtained from all participants. The studies were conducted in accordance with the Declaration of Helsinki II and were approved by the local ethics committees of the Union Hospital of Fujian Medical University and Fujian Medical University.
Demographic and clinical data
All subjects underwent a complete physical examination in the morning after a 12-hour overnight fast and 48-hour alcohol abstinence. Health examination included anthropometric measurements, abdominal ultrasonography, a questionnaire on health-related behaviour (smoking and tea drinking) and biochemical determinations. Smokers were defined as subjects who had smoked at least 100 cigarettes during their lifetime and were classified as smokers vs, non-smokers. Tea drinkers were defined as individuals who drank at least one cup of green tea per day for more than half a year and classified as drinkers vs, non-drinkers. Weight and standing height were measured in a standardized fashion by a trained examiner. The BMI was calculated as the ratio of weight (kilograms) to the square of height (meters) (weight (kg) divided by height squared (m2)). Overweight and/or obesity were defined as having a BMI ≥25 kg/m2. Three resting blood pressure readings were obtained at 1-min intervals; the second and third systolic and diastolic pressure readings were averaged and used for analysis. After an overnight fast, venous blood samples were collected from an antecubital vein into vacutainer tubes containing EDTA. Plasma was separated by centrifugation (2,500 × g for 10 min at 4 °C) and samples were aliquoted and frozen for subsequent analyses. The levels of TC, HDL-C and LDL-C, TGs, FPG) and uric acid (UA), in addition to liver function tests, were measured by standard clinical laboratory techniques, which have been described previously37.
SNP selection and genotyping
The genomic region harbouring SREBF-1c was examined to select SNPs based on the LD patterns of Chinese Han population in Beijing (CHB) from the International HapMap Project SNP database [HapMap Data Rel. 28 Phase II + III, August 10, on National Center for Biotechnology Information (NCBI) B36 assembly, dbSNP b126, accessed April 2013]. We then used the pairwise tagging method of the Haploview v4.2 software (Broad institute, Cambridge, MA, USA) to capture SNPs with a minimum minor allele frequency (MAF) of > 0.02 and a minimum r2 of > 0.8. Additionally, the SNPs were prioritized according to the following criteria i): known SNPs associated with metabolic traits from the literature; ii) SNPs within coding regions (exon); iii) SNPs within the promoter region (2,500 bp before the start codon); iv) SNPs within 3′ untranslated region (UTR) (500 bp after the stop codon); and v) SNPs within 100 bp before an exon-intron splicing boundaries. As result, four SNPs (rs11868035, rs2297508, rs13306741 and rs62064119) were selected from unrelated Han Chinese individuals in Beijing. DNA was isolated from EDTA-treated blood samples following standard procedures. These polymorphisms were genotyped by iPlex technology based on a MassARRAY platform in all subjects. Primers for the amplification and extension reactions were designed using the Mass Array Assay Design Version 3.1 software (Sequenom, San Diego, CA, USA) and SNP genotypes were determined according to the iPLEX protocol provided by the manufacturer. Genotyping assays were performed by laboratory personnel who were blinded to the NAFLD status of each subject. The genotyping quality was examined using a detailed QC procedure consisting of a > 95% successful call rate, duplicate calling of genotypes, internal positive control samples and HWE testing.
Statistical analysis
Unless otherwise indicated, phenotypic quantitative data were expressed as mean ± standard deviation. To evaluate differences in clinical characteristics between cases and controls, Student’s t-test, Pearson’s Chi-squared test or the nonparametric Mann–Whitney U tests were used as appropriate. HWE was checked both in cases and controls by using the chi-squared test. In the case–control studies, each SNP was tested for association with NAFLD in a logistic regression analysis, adjusted for age, sex, marriage, education or income, smoking, tea drinking and clinical characteristics. When the genotype frequency for homozygotes of the minor allele was less than 5%, carriers (heterozygotes and homozygotes individuals) of the minor allele were grouped. P-values were adjusted for multiple tests.
LD between SNPs was estimated in the case–control studies using Haploview4.2 (http://www.broad.mit.edu/mpg/haploview). The Haplo.stats software package was developed using the R language and was used to estimate adjusted ORs and 95% CIs for each haplotype38. To assess statistical significance, we performed permutation procedures to correct the P-value of single-locus association results for multiple testing. Simulations were run 1,000 times for empirical P-values. All statistical analyses were performed using R statistical software.
Additional Information
How to cite this article: Peng, X.-E. et al. Lack of association between SREBF-1c gene polymorphisms and risk of non-alcoholic fatty liver disease in a Chinese Han population. Sci. Rep. 6, 32110; doi: 10.1038/srep32110 (2016).
References
Yokoyama, C. et al. SREBP-1, a basic-helix-loop-helix-leucine zipper protein that controls transcription of the low density lipoprotein receptor gene. Cell 75, 187–197 (1993).
Brown, M. S. & Goldstein, J. L. The SREBP pathway: regulation of cholesterol metabolism by proteolysis of a membrane-bound transcription factor. Cell 89, 331–340 (1997).
Ferre, P. & Foufelle, F. SREBP-1c transcription factor and lipid homeostasis: clinical perspective. Horm Res 68, 72–82 (2007).
Raghow, R., Yellaturu, C., Deng, X., Park, E. A. & Elam, M. B. SREBPs: the crossroads of physiological and pathological lipid homeostasis. Trends Endocrinol Metab 19, 65–73 (2008).
Miserez, A. R., Cao, G., Probst, L. C. & Hobbs, H. H. Structure of the human gene encoding sterol regulatory element binding protein 2 (SREBF2). Genomics 40, 31–40 (1997).
Shimomura, I., Shimano, H., Horton, J. D., Goldstein, J. L. & Brown, M. S. Differential expression of exons 1a and 1c in mRNAs for sterol regulatory element binding protein-1 in human and mouse organs and cultured cells. J Clin Invest 99, 838–845 (1997).
Horton, J. D., Goldstein, J. L. & Brown, M. S. SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J Clin Invest 109, 1125–1131 (2002).
Horton, J. D. et al. Activation of cholesterol synthesis in preference to fatty acid synthesis in liver and adipose tissue of transgenic mice overproducing sterol regulatory element-binding protein-2. J Clin Invest 101, 2331–2339 (1998).
Vitto, M. F. et al. Reversion of steatosis by SREBP-1c antisense oligonucleotide did not improve hepatic insulin action in diet-induced obesity mice. Horm Metab Res 44, 885–890 (2012).
Muraoka, T. et al. Ezetimibe decreases SREBP-1c expression in liver and reverses hepatic insulin resistance in mice fed a high-fat diet. Metabolism 60, 617–628 (2011).
Shimomura, I., Bashmakov, Y. & Horton, J. D. Increased levels of nuclear SREBP-1c associated with fatty livers in two mouse models of diabetes mellitus. J Biol Chem 274, 30028–30032 (1999).
Karasawa, T. et al. Sterol regulatory element-binding protein-1 determines plasma remnant lipoproteins and accelerates atherosclerosis in low-density lipoprotein receptor-deficient mice. Arterioscler Thromb Vasc Biol 31, 1788–1795 (2011).
Farrell, G. C. & Larter, C. Z. Nonalcoholic fatty liver disease: from steatosis to cirrhosis. Hepatology 43, S99–S112 (2006).
Marchesini, G. et al. Nonalcoholic fatty liver, steatohepatitis and the metabolic syndrome. Hepatology 37, 917–923 (2003).
Gao, X. & Fan, J. G. Diagnosis and management of non-alcoholic fatty liver disease and related metabolic disorders: consensus statement from the Study Group of Liver and Metabolism, Chinese Society of Endocrinology. J Diabetes 5, 406–415 (2013).
Marchesini, G. et al. Nonalcoholic fatty liver disease: a feature of the metabolic syndrome. Diabetes 50, 1844–1850 (2001).
Zhang, Z. et al. Associations of the SREBP-1c gene polymorphism with gender-specific changes in serum lipids induced by a high-carbohydrate diet in healthy Chinese youth. Appl Physiol Nutr Metab 36, 226–232 (2011).
Liu, J. X. et al. Association of sterol regulatory element-binding protein-1c gene polymorphism with type 2 diabetes mellitus, insulin resistance and blood lipid levels in Chinese population. Diabetes Res Clin Pract 82, 42–47 (2008).
Musso, G., Bo, S., Cassader, M., De Michieli, F. & Gambino, R. Impact of sterol regulatory element-binding factor-1c polymorphism on incidence of nonalcoholic fatty liver disease and on the severity of liver disease and of glucose and lipid dysmetabolism. Am J Clin Nutr 98, 895–906 (2013).
Liu, J. X., Liu, J. & Guo, Q. Association of sterol regulatory element binding protein-1c genetic polymorphisms rs2297508 and rs11868035 with type 2 diabetes mellitus in Gansu Han and Dongxiang population. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 29, 328–333 (2012).
Grarup, N. et al. Association of variants in the sterol regulatory element-binding factor 1 (SREBF1) gene with type 2 diabetes, glycemia and insulin resistance: a study of 15,734 Danish subjects. Diabetes 57, 1136–1142 (2008).
Laudes, M. et al. Genetic variants in human sterol regulatory element binding protein-1c in syndromes of severe insulin resistance and type 2 diabetes. Diabetes 53, 842–846 (2004).
Knebel, B. et al. Liver-specific expression of transcriptionally active SREBP-1c is associated with fatty liver and increased visceral fat mass. Plos One 7, e31812 (2012).
Chu, X. et al. Sterol regulatory element-binding protein-1c mediates increase of postprandial stearic acid, a potential target for improving insulin resistance, in hyperlipidemia. Diabetes 62, 561–571 (2013).
Harding, A. H. et al. Polymorphisms in the gene encoding sterol regulatory element-binding factor-1c are associated with type 2 diabetes. Diabetologia 49, 2642–2648 (2006).
Suzuki, A. et al. Effect of changes on body weight and lifestyle in nonalcoholic fatty liver disease. J Hepatol 43, 1060–1066 (2005).
Harrison, S. A. & Day, C. P. Benefits of lifestyle modification in NAFLD. Gut 56, 1760–1769 (2007).
Rotman, Y., Koh, C., Zmuda, J. M., Kleiner, D. E. & Liang, T. J. The association of genetic variability in patatin-like phospholipase domain-containing protein 3 (PNPLA3) with histological severity of nonalcoholic fatty liver disease. Hepatology, Md 52, 894–903 (2010).
Santoro, N. et al. A common variant in the patatin-like phospholipase 3 gene (PNPLA3) is associated with fatty liver disease in obese children and adolescents. Hepatology, Md 52, 1281–1290 (2010).
Peng, X. E. et al. Genetic variants in PNPLA3 and risk of non-alcoholic fatty liver disease in a Han Chinese population. Plos one 7, e50256 (2012).
Krawczyk, M., Grunhage, F. & Lammert, F. Identification of combined genetic determinants of liver stiffness within the SREBP1c-PNPLA3 pathway. International journal of molecular sciences 14, 21153–21166 (2013).
Dubuquoy, C. et al. Distinct regulation of adiponutrin/PNPLA3 gene expression by the transcription factors ChREBP and SREBP1c in mouse and human hepatocytes. Journal of hepatology 55, 145–153 (2013).
Eberle, D. et al. SREBF-1 gene polymorphisms are associated with obesity and type 2 diabetes in French obese and diabetic cohorts. Diabetes 53, 2153–2157 (2004).
Hernaez, R. et al. Diagnostic accuracy and reliability of ultrasonography for the detection of fatty liver: a meta-analysis. Hepatology 54, 1082–1090 (2011).
Cazzola, M. & Skoda, R. C. Translational pathophysiology: a novel molecular mechanism of human disease. Blood 95, 3280–3288 (2000).
Hernaez, R. et al. Diagnostic accuracy and reliability of ultrasonography for the detection of fatty liver: a meta-analysis. Hepatology 54, 1082–1090 (2011).
Peng, X. E., Wu, Y. L., Lu, Q. Q., Hu, Z. J. & Lin, X. Two genetic variants in FABP1 and susceptibility to non-alcohol fatty liver disease in a Chinese population. Gene 500, 54–58 (2012).
Lake, S. L. et al. Estimation and tests of haplotype-environment interaction when linkage phase is ambiguous. Hum Hered 55, 56–65 (2003).
Acknowledgements
We thank all study participants for their cooperation and also acknowledge the efforts of our recruiting and technical staff. This work was supported by grants from the National Natural Science Foundation of China (No. 81473047 and No. 81271822), the Natural Science Foundation of Fujian province (No. 2015J01303), the Training Program Foundation for Middle-aged and Young Talents from Sanitation System of Fujian province (2014-ZQN-ZD-23) and the New Century Excellent Talents in Fujian Province University (No. JA14127).
Author information
Authors and Affiliations
Contributions
X.-E.P. and X.L. conceived and supervised the project. F.-L.C. and W.L. performed the experiments. X.-E.P. and Z.H. conducted data analysis. X.-E.P. and X.L. contributed to experiments. All authors discussed the results and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Peng, XE., Chen, FL., Liu, W. et al. Lack of association between SREBF-1c gene polymorphisms and risk of non-alcoholic fatty liver disease in a Chinese Han population. Sci Rep 6, 32110 (2016). https://doi.org/10.1038/srep32110
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep32110
- Springer Nature Limited
This article is cited by
-
Genetic and metabolic aspects of non-alcoholic fatty liver disease (NAFLD) pathogenicity
Egyptian Journal of Medical Human Genetics (2023)