Abstract
The Phosphatase and tensin homolog (PTEN) protein is a negative regulator of the Akt pathway, leading to suppression of apoptois and increased cell survival. Its role as a tumor-suppressor gene has been adequately substantiated, and PTEN hypermethylation has been demonstrated in familial and sporadic cancers. However, the association and clinical significance between PTEN hypermethylation and breast cancer remains unclear. In this study, we systematically reviewed studies of PTEN hypermethylation and breast cancer and quantify the association between PTEN hypermethylation and breast cancer using meta-analysis methods. The pooled OR, 22.30, 95% confidential intervals, CI = 1.98–251.51, P = 0.01, which demonstrates that loss of PTEN expression by hypermethylation plays a critical role in the early tumorigenesis of ductal carcinoma in situ (DCIS). In addition, PTEN hypermethylation also is detected in invasive ductal carcinomas (IDCs) and is significantly higher than in normal controls, OR = 23.32, 95% CI = 10.43–52.13, P < 0.00001. Further analysis did not show significant correlation between PTEN hypermethylation and the progression of breast cancer, estrogen receptor (ER), progesterone receptor (PgR), as well as HER2 status. These results indicate the PTEN hypermethylation is significantly associated with both DCIS and IDCs. The detection of PTEN hypermethylation could be an early tumorigenesis marker for breast cancer patients.
Similar content being viewed by others
Introduction
Breast cancer is the most common cancer among women worldwide. Breast cancer develops as a stepwise accumulation of genetic changes (point mutations, deletions, and gene amplifications) leading to oncogene activation or tumor suppressor gene inactivation1. In addition to genetic alterations, breast cancer progression is also controlled by epigenetic modifications such as DNA methylation, histone acetylation and nucleosomal remodeling2. Phosphatase and tensin homolog (PTEN) is a protein encoded by the tumor suppressor gene PTEN. It plays a critical role in cell growth, proliferation and differentiation3 and is involved in DNA repair and corresponding signaling in breast cancer4. First, PTEN is important in breast cancer initiating cells (CICs) survival. Knockdown of PTEN expression causes increases in normal and malignant mammary stem/progenitor cells, which was regulated by increased AKT/GSK-3beta/Wnt/beta-catenin pathway5,6. The resistance of breast cancers to herceptin (trastuzumab) and other chemotherapies are therefore related to loss of PTEN expression6,7,8,9,10. Second, multiple mechanisms of PTEN inactivation applies in breast cancer. Reduced expression of PTEN in breast cancer may result from PTEN gene as mutation, deletion and methylation9. The role of PTEN protein in cancers has been widely studied. For instance, a couple of meta-analyses demonstrated that PTEN was associated with poor survival in gastric, colorectal, as well as endometrial and prostate cancers11,12. A prognostic value of PTEN in breast cancer has also recently been investigated using meta-analysis. They solidified that PTEN inactivation is associated with unfavorable overall survival and disease-free survival13. However the systemic review on evaluation of PTEN methylation in breast cancer patients has not been documented. In the present study we performed a meta-analysis and systemic review on the relationship of PTEN hypermethylation and clinical parameters of human breast cancer to fill the gap of breast cancer research.
Material and Methods
Search Method
We systematically searched Embase, Pubmed and ISI web of knowledge to select studies from June 1, 1996 through April 1, 2016. Keywords used to search relevant publications included: “breast cancer”, “breast tumor”, and “breast carcinoma”, “methylation”, “hypermethylation”, “epigenetic changes”, “ phosphatase and tensin honolog”, and “PTEN”. We also scrutinized the reference lists of the relevant reviews and eligible articles for additional eligible articles. No restrictions were placed on language and publication status.
Selection criteria
All searched data were retrieved. The titles and abstracts of all searched records were scrutinized to judge their relevance and eligibility. After excluding of duplicates and publications with non-extractable data, the remaining papers were screened in the full text version for in- and exclusion criteria. The most complete investigations was included if the same patient populations were published in different resources. Cohort studies that fulfills the following criteria were considered eligible: (1) PTEN methylation examined in breast cancer patients, (2) PTEN methylation determined by polymerase chain reaction (PCR), (3) researches revealed the relationship between PTEN methylation status and breast cancer clinicopathological parameters and the outcomes, (4) studies which provided sufficient data to calculate Odds ratio (OR) and/or hazard ratio (HR) about overall survival (OS) and 95% confidence interval (CI). The exclusion criteria: (1) all the descriptive studies including reviews, letters, case studies, editorials, and conference abstracts, (2) all in vitro and in vivo laboratory works.
Data extraction and methodological assessment
Three authors independently completed the data extraction from eligible studies by evaluating all of records that meet the inclusion and exclusion criteria. The following data were extracted from each study: (1) bibliographic data including the first author name, year of publication, and country; (2) data on clinicopathological characteristics including number of cases, sample source, age, state of disease, histology, ER status, PR status and HER2 status; (3) methylation detection methods, methylation rate, and/or expression, and follow up. Data for study characteristics were summarized in a table format. Investigation heterogeneity was evaluated to determine whether or not the data of the various studies could be analyzed for a meta-analysis.
Three investigators read through each publication independently for the methodological evaluation of the studies, and assessed and scored them according to REMARK guidelines and ELCWP quality scale14,15. Then they provided the quality scores, compared them, and then reached a consensus value for each item if disagreements existed between the investigators.
Statistical analysis
Analysis was performed using the STATA 12.0 (Stata Corporation, TX, USA) and Review Manager 5.2 (Cochrane Collaboration, Oxford, UK). The pooled rate of PTEN hypermethylation and 95% confidence intervals (95% CI) from different studies were combined to obtain a summary estimate. A summary Odds Ratio was estimated and compared for the inter-relationship between PTEN and each tumor factors/characteristics. Heterogeneity among studies was examined with Cochran’s Q test16 and the I2 statistic17,18. A fixed effect model was used for meta-analyses when heterogeneity was not an issue (I2 values <50% or P value ≤0.10), while a random-effects model was used to pool data and attempt to identify potential sources of heterogeneity based on subgroup analyses, if there was substantial heterogeneity (I2 values ≥50%). P values tailed less than 0.05 were considered statistically significant.
The analysis of meta-regression and publication bias was performed using STATA version 10.0. The possibility of publication bias was determined by using Begg’s funnel plot and Egger’s regression test19. For statistical heterogeneity, we determined the reasons using meta-regression, subgroup analysis and sensitivity analysis.
Patient survival analysis
An online database20 was used to assess relevance of PTEN expression to relapse free survival. The database was established using gene expression data and survival information of 3,557 patients downloaded from Gene Expression Omnibus (GEO) (Affymetrix HGU133A and HGU133+2 microarrays). Briefly, PTEN gene was entered into the database (http://kmplot.com/breast/) to obtain Kaplan-Meier survival plots where the number-at-risk is indicated below the main plot. Hazard ratio (and 95% confidence intervals) and logrank P were calculated and displayed on the webpage.
Another online tool we used is OncoLnc, a tool for interactively exploring survival correlations, and for downloading clinical data coupled to expression data for mRNAs, miRNAs, or long noncoding RNAs (lncRNAs)21. OncoLnc contains survival data for 8,647 patients from 21 cancer studies performed by The Cancer Genome Atlas (TCGA), along with RNA-SEQ expression for mRNAs and miRNAs from TCGA, and lncRNA expression from MiTranscriptome beta. This dataset contains 995 breast invasive cancer (BRCA) patients, OncoLnc is available at http://www.oncolnc.org. The multivariate cox regressions were performed followed by a Kaplan-Meier analysis for BRCA.
Results
Initially, 488 articles were identified by the literature search. Among these references, 9 studies were considered potentially eligible, and 8 studies published between 2004 to 2015 were finally included in this meta-analysis. The flow chart of study is shown in Fig. 1. The articles were excluded due to reviews, in vitro and in vivo laboratory studies.
Included study characteristics
The basic characteristics of included studies were summarized in Table 1. Eight studies were conducted in United States, Norway, Spain, China, India and Iran. The sample sizes ranged from 28 to 199, with a total of 927 breast cancer patients and 588 corresponding controls. PTEN status was consistently examined by methylation specific PCR (MSP) using breast cancer tissues across the included studies.
PTEN hypermethylation and clinicopathological characteristics
The PTEN hypermethylation in ductal carcinoma in situ (DCIS) and invasive ductal carcinomas (IDCs)
The patients with DCIS showed high level of PTEN hypermethylation compared to normal individuals. The pooled OR from 3 studies including 71 of DCIS and 124 of normal breast tissues are shown in Fig. 2A (odds ratios, OR = 22.30, 95% confidential intervals, CI = 1.98–251.51, P = 0.01), which demonstrates that loss of PTEN expression by hypermethylation plays a critical role in the early tumorigenesis of DCIS. In addition, PTEN hypermethylation also is detected in IDCs and is significantly higher than in normal controls (OR = 23.32, 95% CI = 10.43–52.13, P < 0.00001), as shown in Fig. 2B. There is no significant difference between PTEN hypermethylation between individuals with IDCs and with DCIS, OR = 0.79 95% CI = 0.29–2.18, P = 0.65 (Fig. 2C). The blue rectangle in each figure showed the individual OR where as the diamond showed the total OR.
The loss of PTEN expression through hypermethylation in ductal carcinoma in situ (DCIS) and invasive ductal carcinomas (IDCs).
(A) The patients with DCIS showed high level of PTEN hypermethylation compared to normal individuals. The pooled OR from 3 studies including 71 of DCIS and 124 of normal breast tissues are shown (odds ratios, OR = 22.30, 95% confidential intervals, CI = 1.98–251.51, P = 0.01), which demonstrates that loss of PTEN expression by hypermethylation plays a critical role in the early tumorigenesis of DCIS. (B) PTEN hypermethylation also is detected in IDC and is significantly higher than in normal controls (OR = 23.32, 95% CI = 10.43–52.13, P < 0.00001). (C) There is no significant difference between PTEN hypermethylation between individuals with IDC and with DCIS, OR = 0.79 95% CI = 0.29–2.18, P = 0.65.
PTEN hypermethylation in the progression of breast cancer
We then determined 540 breast cancer patients pooled in 4 studies to evaluate the role of inactivation of PTEN via hypermethylation in the progression of breast cancer. In Fig. 3, aberrant PTEN hypermethylation is not significantly higher in advanced stage of breast cancer (III) than that in early staged breast cancer (I & II), OR = 0.87, 95% CI = 0.20–3.72, P = 0.85. These results indicate that the inactivation of PTEN gene by hypermethylation may not play an important role in breast cancer progression from initial stage to advanced stage. The blue rectangle in each figure showed the individual OR where as the diamond showed the total OR.
PTEN hypermethylation in the progression of breast cancer.
540 breast cancer patients pooled in 4 studies to evaluate the role of inactivation of PTEN via hypermethylation in the progression of breast cancer. The aberrant PTEN hypermethylation is not significantly higher in advanced stage of breast cancer (III) than that in early staged breast cancer (I & II), OR = 0.87, 95% CI = 0.20–3.72, P = 0.85. These results indicate that the inactivation of PTEN gene by hypermethylation may not play an important role in breast cancer progression from initial stage to advanced stage.
Correlation with PTEN hypermethylation and hormone receptor status and HER2 status
No significant association with estrogen receptor (ER) status was observed for PTEN methylaltion, OR = 1.20 with 95%CI: 0.82–1.77 (P = 0.35, Fig. 4A). PgR and HER2 status were not associated with PTEN methylation either OR = 1.10 95% CI = 0.73–1.63, P = 0.65; OR = 1.33 95% CI = 0.88–2.02, P = 0.17 respectively (Fig. 4B,C). The blue rectangle in each figure showed the individual OR where as the diamond showed the total OR.
Correlation with PTEN hypermethylation and hormone receptor status and HER2 status.
(A) No significant association with estrogen receptor (ER) status was observed for PTEN methylaltion, OR = 1.20 with 95%CI: 0.82–1.77 (P = 0.35). (B) No correlation with PTEN hypermethylation and PR. PR receptor status were not associated with PTEN methylation either OR = 1.10 95% CI = 0.73–1.63, P = 0.65. (C) No correlation with PTEN hypermethylation and HER2. HER2 receptor status were not associated with PTEN methylation either OR = 1.33 95% CI = 0.88–2.02, P = 0.17 respectively.
The role of PTEN in breast cancer patient survival
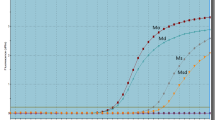
This assessment of clinical relevance was performed in a patient survival analysis using an online database containing the expression of 54,675 genes and 20-year survival information of 4142 patients20, including survival information of 3557 breast cancer patients (http://www.kmplot.com/analysis/). PTEN downregulation was not found to correlate with shorter relapse free survival (RFS) for all breast cancer patients followed for 20 years (Fig. 5, hazardous ratio 1.04, p = 0.65).
PTEN mRNA does not correlate with disease-free survival in breast cancer patients.
This assessment of clinical relevance was performed in a patient survival analysis using an online database containing the expression of 54,675 genes and 20-year survival information of 4142 patients20, including survival information of 3557 breast cancer patients (http://www.kmplot.com/analysis/). PTEN downregulation was not found to correlate with shorter relapse free survival (RFS) for all breast cancer patients followed for 20 years (Fig. 5, hazardous ratio 1.04, p = 0.65).
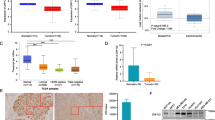
By using the tool of OncoLnc to link TCGA survival data of breast invasive cancer (BRCA) patients to mRNAs, patients were split into non-overlapping upper and lower slices, namely upper 25 percent and lower 25 percent (Fig. 6). Similarly low expression of PTEN was not correlated with disease free survival (logrank p = 0.593).
PTEN mRNA does not correlate with disease-free survival in breast invasive cancer patients (BRCA).
By using the tool of OncoLnc to link TCGA survival data of breast invasive cancer (BRCA) patients to mRNAs, patients were split into non-overlapping upper and lower slices, namely upper 25 percent and lower 25 percent (Fig. 6). Similarly low expression of PTEN was not correlated with disease free survival (logrank p = 0.593).
Publication bias and sensitivity analyses
In the current meta-analysis of PTEN hypermethylation and clinicopathological features, the publication biases were evaluated by the funnel plots. The publication biases were ruled out by the symmetric funnel plot (Fig. 7). The sensitivity analyses were conducted by removing one study at a time to determine the stability. The pooled ORs and HRs are not significantly changed, suggesting the stability of this meta-analyses.
Publication bias and sensitivity analyses.
In the current meta-analysis of PTEN hypermethylation and clinicopathological features, the publication biases were evaluated by the funnel plots. The publication biases were ruled out by the symmetric funnel plot. The funnel plots were largely symmetric suggesting there were no publication biases in the meta-analysis of PTEN hypermethylation and breast cancer clinicopathological features. The funnel plot from 3 studies comparing 91 ductal carcinoma in situ (DCIS) and 124 normal breast tissue (A). The funnel plot from 6 studies comparing invasive ductal carcinomas (IDCs, n = 981) and normal breast tissue (n = 282) (B). The funnel plot from 4 studies comparing IDCs (n = 210) and DCIS (n = 458) (C). The funnel plot from 4 studies comparing different staged [III (n = 152) VS. I/II (n = 388)] (D). The funnel plot from 4 studies in determining ER-positive (n = 442) and ER-negative (n = 268) and PTEN hypermethylation in breast cancer patients (E). The funnel plot from 3 studies in determining PR-positive (n = 400) and PR-negative (n = 251) and PTEN hypermethylation in breast cancer patients (F). The funnel plot from 4 studies in determining HER2-positive (n = 263) and HER2-negative (n = 321) and PTEN hypermethylation in breast cancer patients (G).
Discussion
Phosphatase and tensin homologue (PTEN) is one of the tumor suppressor which atagonizes the phosphatidylinositol 3-kinase-AKT pathway via the lipid phosphatase activity22. Genetic changes of PTEN, such as mutations and deletion, and/or epigenetic alterations are often observed in a variety of cancers23. In addition, PTEN functions are well known to be regulated by various posttranslational modifications including phosphorylation, unbiquitylation, acetylation, and oxidation24. Recent findings from Hamamoto’s group showed that SMYD2-mediated PTEN methylation at lysine 313 increased phosphorylation at serine 380 of PTEN, consequently inactivated PTEN and activated AKT activity and promoted cell growth and proliferation25. They confirmed and extended the previous notions that cross-talk of different types of posttranslational modification on histone proteins and non-histone proteins such as methylation is commonly existed1,26,27,28,29,30. Our meta-analysis demonstrated that 1). The loss of PTEN expression through hypermethylation plays an important role in the formation of both DCIS and IDCs, 2) PTEN hypermethylation does not contribute to the progression of breast cancer, 3) No correlation with PTEN hypermethylation was found for hormone receptor status and HER2 status.
The progression from DCIS to IDC still remains poorly understood. However it is well accepted that cancer development is arising from the accumulation of genomic changes leading to oncogene overexpression and/or loss of expression of tumor suppressor genes. On one hand, some copy number changes present in DCIS tend to increase in IDCs31, thus the distinct stage-specific markers have been identified in each stage32,33,34. On the other hand, the frequency of some copy numbers is similar to what is observed in DCIS and IDC suggesting the same cellular origin of the two components35,36,37. The case of PTEN methylation in breast cancer fits into the latter scenario, which showed no difference in PTEN methylation frequency in DCIS and IDC.
The results from Klajic et al. demonstrated that PTEN methylation levels were low and increased significantly in late stage III and IV38, however the results from the pooled study1,39,40,41 did not show the significant difference between different stage of breast cancer. Our current meta-analysis on PTEN methylation in breast cancer implied that PTEN methylation is an early tumorigenic marker for breast cancer and stays positive and stable through the whole process of malignancy from very early to advanced stage.
Only one study investigated the prognostic significance of PTEN hypermethylation in breast cancer. By univariate analysis they found that the high expression of PTEN which is equivalent to low hypermethylation status of PTEN was significantly associated with a longer relapse-free survival in patients with ER-negative an PTEN-positive status1. We were unable to perform quantitative analysis due to only one publication. Instead, we used 1) gene expression data and survival information of 3,557 breast cancer patients downloaded from Gene Expression Omnibus (GEO) (Affymetrix HGU133A and HGU133+2 microarrays), and 2) survival data for 995 breast invasive cancer (BRCA) patients from The Cancer Genome Atlas (TCGA), along with RNA-SEQ expression for mRNAs and miRNAs from TCGA, and lncRNA expression from MiTranscriptome beta. The online tools http://kmplot.com/breast/ and http://www.oncolnc.org were used to obtain Kaplan-Meier survival plots correspondingly. No correlation was found between low expression of PTEN and disease-free survival. However we need carefully interpret these survival data since all these data are base on mRNA expression, they may not accurately reflect the protein expression which are more crucial to gene function.
In contrast to the data from mRNA expression, a group in China recently conducted a meta-analysis to assess the association of PTEN negativity by immunohistochemistry method with overall survival and disease-free survival. They found that PTEN negativity was significantly associated with unfavourable prognosis in terms of overall survival in breast cancer although the association of PTEN negativity and disease-free survival was not validated by their study13.
In conclusion, the meta-analysis demonstrated that PTEN could be an early tumorigenesis marker for breast cancer, whether PTEN methylation with negative protein expression (not only mRNA expression) was probably associated with poor survival status needs more clinical cohort studies to confirm.
Additional Information
How to cite this article: Lu, Y.-M. et al. The association between phosphatase and tensin homolog hypermethylation and patients with breast cancer, a meta-analysis and literature review. Sci. Rep. 6, 32723; doi: 10.1038/srep32723 (2016).
References
Zhang, H. Y., Liang, F., Jia, Z. L., Song, S. T. & Jiang, Z. F. mutation, methylation and expression in breast cancer patients. Oncol Lett 6, 161–168 (2013).
Jovanovic, J., Ronneberg, J. A., Tost, J. & Kristensen, V. The epigenetics of breast cancer. Mol Oncol 4, 242–254 (2010).
Vazquez, F. & Sellers, W. R. The PTEN tumor suppressor protein: an antagonist of phosphoinositide 3-kinase signaling. Biochim Biophys Acta 1470, M21–35 (2000).
Davis, N. M. et al. Deregulation of the EGFR/PI3K/PTEN/Akt/mTORC1 pathway in breast cancer: possibilities for therapeutic intervention. Oncotarget 5, 4603–4650 (2014).
Korkaya, H. et al. Regulation of mammary stem/progenitor cells by PTEN/Akt/beta-catenin signaling. PLoS Biol 7, e1000121 (2009).
Razis, E. et al. Evaluation of the association of PIK3CA mutations and PTEN loss with efficacy of trastuzumab therapy in metastatic breast cancer. Breast Cancer Res Treat 128, 447–456, doi: 10.1007/s10549-011-1572-5 (2011).
Rexer, B. N., Shyr, Y. & Arteaga, C. L. Phosphatase and tensin homolog deficiency and resistance to trastuzumab and chemotherapy. J Clin Oncol 31, 2073–2075 (2013).
Fujita, T. et al. PTEN activity could be a predictive marker of trastuzumab efficacy in the treatment of ErbB2-overexpressing breast cancer. Br J Cancer 94, 247–252, doi: 6602926 (2006).
Nagata, Y. et al. PTEN activation contributes to tumor inhibition by trastuzumab, and loss of PTEN predicts trastuzumab resistance in patients. Cancer Cell 6, 117–127, doi: 10.1016/j.ccr.2004.06.022 (2004).
Esteva, F. J. et al. PTEN, PIK3CA, p-AKT, and p-p70S6K status: association with trastuzumab response and survival in patients with HER2-positive metastatic breast cancer. Am J Pathol 177, 1647–1656, doi: S0002-9440(10)60218-0 (2010).
Chen, J. et al. Clinical and prognostic significance of HIF-1alpha, PTEN, CD44v6, and survivin for gastric cancer: a meta-analysis. PLoS One 9, e91842 (2014).
Ocana, A. et al. Activation of the PI3K/mTOR/AKT pathway and survival in solid tumors: systematic review and meta-analysis. PLoS One 9, e95219 (2014).
Yang, Z. Y. et al. The prognostic value of phosphatase and tensin homolog negativity in breast cancer: A systematic review and meta-analysis of 32 studies with 4393 patients. Crit Rev Oncol Hematol 18, 30014–30012 (2016).
McShane, L. M. et al. Reporting recommendations for tumor marker prognostic studies (REMARK). Journal of the National Cancer Institute 97, 1180–1184, doi: 10.1093/jnci/dji237 (2005).
Steels, E. et al. Role of p53 as a prognostic factor for survival in lung cancer: a systematic review of the literature with a meta-analysis. Eur Respir J 18, 705–719 (2001).
DerSimonian, R. & Laird, N. Meta-analysis in clinical trials. Control Clin Trials 7, 177–188, doi: 0197-2456(86)90046-2 (1986).
Higgins, J. P., Thompson, S. G., Deeks, J. J. & Altman, D. G. Measuring inconsistency in meta-analyses. BMJ (Clinical research ed.) 327, 557–560, doi: 10.1136/bmj.327.7414.557 (2003).
DerSimonian, R. Meta-analysis in the design and monitoring of clinical trials. Statistics in medicine 15, 1237-1248; discussion 1249–1252, doi: 10.1002/(sici)1097-0258(19960630)15:12<1237::aid-sim301>3.0.co;2-n (1996).
Egger, M., Davey Smith, G., Schneider, M. & Minder, C. Bias in meta-analysis detected by a simple, graphical test. BMJ 315, 629–634 (1997).
Gyorffy, B. et al. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat 123, 725–731 (2010).
Tomczak, K., Czerwinska, P. & Wiznerowicz, M. The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol 19, A68–77, doi: 10.5114/wo.2014.47136 (2015).
Lee, J. O. et al. Crystal structure of the PTEN tumor suppressor: implications for its phosphoinositide phosphatase activity and membrane association. Cell 99, 323–334 (1999).
Hollander, M. C., Blumenthal, G. M. & Dennis, P. A. PTEN loss in the continuum of common cancers, rare syndromes and mouse models. Nature reviews. Cancer 11, 289–301, doi: 10.1038/nrc3037 (2011).
Okumura, K. et al. PCAF modulates PTEN activity. The Journal of biological chemistry 281, 26562–26568, doi: 10.1074/jbc.M605391200 (2006).
Nakakido, M. et al. Dysregulation of AKT Pathway by SMYD2-Mediated Lysine Methylation on PTEN. Neoplasia (New York, N.Y.) 17, 367–373, doi: 10.1016/j.neo.2015.03.002 (2015).
Cho, H. S. et al. RB1 methylation by SMYD2 enhances cell cycle progression through an increase of RB1 phosphorylation. Neoplasia (New York, N.Y.) 14, 476–486 (2012).
Huang, J. et al. Repression of p53 activity by Smyd2-mediated methylation. Nature 444, 629–632, doi: 10.1038/nature05287 (2006).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science (New York, N.Y.) 293, 1074–1080, doi: 10.1126/science.1063127 (2001).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705, doi: 10.1016/j.cell.2007.02.005 (2007).
Yang, X. D. et al. Negative regulation of NF-kappaB action by Set9-mediated lysine methylation of the RelA subunit. The EMBO journal 28, 1055–1066, doi: 10.1038/emboj.2009.55 (2009).
Yao, J. et al. Combined cDNA array comparative genomic hybridization and serial analysis of gene expression analysis of breast tumor progression. Cancer research 66, 4065–4078, doi: 10.1158/0008-5472.can-05-4083 (2006).
Urbich, C. et al. Fluid shear stress-induced transcriptional activation of the vascular endothelial growth factor receptor-2 gene requires Sp1-dependent DNA binding. FEBS Lett 535, 87–93, doi: S0014579302038796 (2003).
Porter, D. et al. Molecular markers in ductal carcinoma in situ of the breast. Molecular cancer research: MCR 1, 362–375 (2003).
Schuetz, C. S. et al. Progression-specific genes identified by expression profiling of matched ductal carcinomas in situ and invasive breast tumors, combining laser capture microdissection and oligonucleotide microarray analysis. Cancer research 66, 5278–5286, doi: 10.1158/0008-5472.can-05-4610 (2006).
Done, S. J., Eskandarian, S., Bull, S., Redston, M. & Andrulis, I. L. p53 missense mutations in microdissected high-grade ductal carcinoma in situ of the breast. Journal of the National Cancer Institute 93, 700–704 (2001).
Lukas, J., Niu, N. & Press, M. F. p53 mutations and expression in breast carcinoma in situ. The American journal of pathology 156, 183–191, doi: 10.1016/s0002-9440(10)64718-9 (2000).
Zhou, W. et al. Full sequencing of TP53 identifies identical mutations within in situ and invasive components in breast cancer suggesting clonal evolution. Mol Oncol 3, 214–219, doi: 10.1016/j.molonc.2009.03.001 (2009).
Klajic, J. et al. Quantitative DNA methylation analyses reveal stage dependent DNA methylation and association to clinico-pathological factors in breast tumors. BMC Cancer 13, 456 (2013).
Garcia, J. M. et al. Promoter methylation of the PTEN gene is a common molecular change in breast cancer. Genes Chromosomes Cancer 41, 117–124 (2004).
Siddiqui, S., Akhter, N., Deo, S. V., Shukla, N. K. & Husain, S. A. A study on promoter methylation of PTEN in sporadic breast cancer patients from North India. Breast Cancer 11, 11 (2016).
Yari, K., Payandeh, M. & Rahimi, Z. Association of the hypermethylation status of PTEN tumor suppressor gene with the risk of breast cancer among Kurdish population from Western Iran. Tumour Biol 29, 29 (2015).
Khan, S. et al. PTEN promoter is methylated in a proportion of invasive breast cancers. Int J Cancer 112, 407–410 (2004).
Muggerud, A. A. et al. Frequent aberrant DNA methylation of ABCB1, FOXC1, PPP2R2B and PTEN in ductal carcinoma in situ and early invasive breast cancer. Breast Cancer Res 12, R3 (2010).
Zhao Yingfang, S. S. & Jiang Jianjying . A study on methylation of gene PTEN 5′ CpG island in breast carcinomas and tissues adjacent to breast cancer. Capital Medical Network 3, 1–3 (2010).
Author information
Authors and Affiliations
Contributions
Y.-M.L. and F.C. contributed equally; Y.-M.L. and L.-S.T. participated in the design of the study; Y.-M.L. and F.C. performed experiments; Y.-M.L. and F.C. analyzed data; L.-S.T. wrote the manuscript; All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Lu, YM., Cheng, F. & Teng, LS. The association between phosphatase and tensin homolog hypermethylation and patients with breast cancer, a meta-analysis and literature review. Sci Rep 6, 32723 (2016). https://doi.org/10.1038/srep32723
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep32723
- Springer Nature Limited
This article is cited by
-
Alterations of PTEN and SMAD4 methylation in diagnosis of breast cancer: implications of methyl II PCR assay
Journal of Genetic Engineering and Biotechnology (2021)
-
Reversal of increased mammary tumorigenesis by valproic acid and hydralazine in offspring of dams fed high fat diet during pregnancy
Scientific Reports (2019)
-
The LXR-623-induced long non-coding RNA LINC01125 suppresses the proliferation of breast cancer cells via PTEN/AKT/p53 signaling pathway
Cell Death & Disease (2019)
-
Isoliquiritigenin modulates miR-374a/PTEN/Akt axis to suppress breast cancer tumorigenesis and metastasis
Scientific Reports (2017)