Abstract
The cytochrome b6 f complex occupies an electrochemically central position in the electron-transport chain bridging the photosynthetic reaction center of PS I and PS II. In plants, the subunits of these thylakoid membrane protein complexes are both chloroplast and nuclear encoded. How the chloroplast-encoded subunits of multi-spanning cytochrome b6 are targeted and inserted into the thylakoid membrane is not fully understood. Experimental approaches to evaluate the cytochrome b6 import mechanism in vivo have been limited to bacterial membranes and were not a part of the chloroplast environment. To evaluate the mechanism governing cytochrome b6 integration in vivo, we performed a comparative analysis of both native and synthetic cytochrome b6 insertion into purified thylakoids. Using biophysical and biochemical methods, we show that cytochrome b6 insertion into the thylakoid membrane is a non-spontaneous co-translational process that involves ALB3 insertase. Furthermore, we provided evidence that CSP41 (chloroplast stem–loop-binding protein of 41 kDa) interacts with RNC-cytochrome b6 complexes and may be involved in cytochrome b6 (petB) transcript stabilization or processing.
Similar content being viewed by others
Introduction
Photosynthetic electron transport is accomplished by three hetero-oligomeric integral oxygenic photosynthetic membrane protein complexes: Photosystems I (PS I) and II (PS II) and the cytochrome b6 f complex. The 220-kDa cytochrome b6 f complex dimer occupies an electrochemically central position in the electron-transport chain1. Hence, cytochrome b6 f provides an electronic connection between the two photosynthetic reaction centers of PS I and PS II, allowing linear electron transfer from the H2O electron donor to the NADP (nicotinamide-adenine dinucleotide phosphate) acceptor2. Furthermore, cytochrome b6 f and PS I complex participate in cyclic electron transfer, which generates an electrochemical proton gradient across the thylakoid membrane without net production of reducing equivalents3. In plants, these electron-transfer chain protein complexes are located in chloroplast thylakoid membranes, while their subunits are encoded by both nuclear and chloroplast genomes4. The proper thylakoid membrane assembly of PS I, PS II and cytochrome b6 f requires numerous regulatory factors for coordinated transport, insertion and assembly of these complexes subunits from both chloroplast and nuclear origin5. Although the electron-transfer chain function and structure have been extensively studied, the mechanism governing the assembly of these complexes in the thylakoid membrane is less understood. Specifically, little is known how their chloroplast-encoded subunits are targeted and inserted into the thylakoid membrane.
However, for the import into the thylakoid membrane of proteins from both nuclear and chloroplast origin, four independent precursor-specific transport pathways had been proposed (classified on the basis of their energy and stromal factor requirements)6. These four pathways have been categorized as “spontaneous”, signal recognition particle (SRP), secretory (Sec) and twin-arginine translocase-dependent (ΔpH/Tat)7. Integration of proteins into thylakoid membranes relies not only on the membrane translocation machinery, but also on the chloroplast stromal fraction. The Sec pathway requires the translocation ATPase and SecA proteins8. The cpSRP pathway uses GTP, cpSRP54 and cpSRP43 to target proteins to the thylakoid membrane, but the Tat pathway uses a trans-thylakoid pH gradient as its sole energy source7. All of these are found in the soluble stromal fraction. However, the “spontaneous” pathway does not seem to require any soluble factors or energy source for protein insertion into membrane9. The majority of proteins incorporated into the thylakoid membrane utilize the spontaneous or SRP pathway, while protein transport through the thylakoid membrane is mediated by the Sec or ΔpH/Tat pathway6.
Commonly, nuclear-encoded multi-spanning proteins are targeted to the thylakoid membrane by hydrophobic amino acid sequences from either their transmembrane segments or from a cleavable signal sequence (ss)10. Whereas, the spontaneous pathway seems to be the mainly utilized for the import of single-span proteins7. Importantly, the spontaneous mechanism was also shown to be active for some of nuclear-encoded multi-spanning thylakoid membrane proteins including the PsaK and PsaG subunits of PS I11,12.
The chloroplast-encoded cytochrome b6 binds a one covalently c-type haem as well as two non-covalently b-type haems and consists of four transmembrane helices, while the signal for this integral protein integration with thylakoid membrane remains unknown13,14. In the current model for assembly of cytochrome b6 f complex, the first step involves the transcriptional activation of the chloroplast petBD operon (encoding cytochrome b6 and subunit IV)13. Following the transcription, the petB and petD mRNAs are translated into the polypeptides that undergo insertion into the membrane and form the polytopic monomeric core of the cytochrome b6 f complex. In the next step the monomers form a dimer (CS) which is stabilized by lipids and simultaneously a Rieske ISP-cytochrome f sub-complex (RF) is formed. This sub-complex then interacts with the CS to form a cytochrome b6-subunit IV-ISP-cytochrome f sub-complex (CSRF). Regardless of the formation of the CSRF complex, small subunits (Pet G, L, M and N) form an additional sub-complex which may interact with the RF15. At last fully functional cytochrome b6 f complex is formed.
Hence, cytochrome b6 f complex assembly process requires a complex coordination between transcription, translation, chloroplast membrane transport, membrane insertion and sub-complexes assembly.
To date, experimental approaches to evaluate the cytochrome b6 import mechanism in vivo were limited to bacterial membrane and therefore did not involve the chloroplast environment16,17,18,19,20.
Hence, the objective of the present study was to examine the mechanism governing cytochrome b6 integration into the thylakoid membrane. Our comparative analysis of both native and synthetic cytochrome b6 revealed that an unfolded cytochrome b6 can be anchored into the thylakoid membrane by hydrophobic interactions that can be removed by chaotropic action. Hence, we excluded the spontaneous pathway for insertion of cytochrome b6 into the thylakoid membrane in vivo. Furthermore, our results indicate that the proper integration of cytochrome b6 is co-translationally mediated by other proteins. Indeed, we determined ALB3 insertase is a crucial protein for cytochrome b6 insertion into thylakoid membrane. Furthermore, we identified cpFtsY and CPS41 (chloroplast stem–loop-binding protein of 41 kDa) as other proteins involved in this process. These data are not limited to cytochrome b6, but also provide a new insight into the mechanisms involved in the insertion of integral membrane proteins integration into the thylakoid membrane.
Results
The primary aim of this work was to analyse the mechanism by which cytochrome b6 inserts into the thylakoid membrane. For these experiments, we used import assays for the insertion of different variants of cytochrome b6 proteins into purified Pea thylakoids. Our main criterion for correct insertion was that the cytochrome b6 has to be integrated with the membrane and thus cannot be extracted along with extrinsic, non-inserted proteins21. Furthermore, the cytochrome b6 has to be properly oriented in the membrane, with the C- terminus and N-terminus at the stromal side of the thylakoid membrane.
Cytochrome b6 integration with thylakoid membrane is not a spontaneous process
To follow the native cytochrome b6 integration with the Pea thylakoid membrane, the protein was purified from Synechocystis sp. PCC 6803 as described in ref. 22 and solubilised in the presence of n-dodecyl-β-D-maltoside (DDM). As shown in Supplemental Fig. 1, an amino acid consensus between cytochrome b6 protein sequences from Pea and Synechocystis sp. PCC 6803 was above 87% and positions with identical amino acid residues were above 79%. The circular dichroism (CD) spectra of isolated native cytochrome b6 indicated a high proportion of the α-helical structure characterised by negative maxima at 208 and 222 nm (Fig. 1A and Table 1). Following the import assay of native cytochrome b6, the protein integration with thylakoid membrane was tested with use of urea as chaotropic agent. Urea has been proven to be effective in removing extrinsic, non-inserted proteins from the thylakoid membranes21. Chaotropic agents are co-solutes that disrupt the van der Waals forces and hydrogen-bonding network between water molecules and reduce the stability of the native state of proteins by weakening the hydrophobic effect.
In order to distinguish the inserted protein from the original cytochrome b6 located in the isolated thylakoid membrane, prior to insertion assays, the exogenous cytochrome b6 was biotin labelled. Notably, thylakoid-associated native cytochrome b6 (isolated from Synechocystis sp. PCC 6803) was almost entirely chaotropic-extractable by a mild concentration of urea (4 M) and not detectable in the membrane fraction (Fig. 2A lane 3).
In vitro import of cytochrome b6 into thylakoid membrane.
(A) The integration of the cytochrome b6 into the thylakoid membrane in the presence or absence of stromal fraction (SF) was analysed with Western blot. Urea was used as chaotropic agents (CH). Lane 1, purified native cytochrome b6 as a control; lanes 2 and 3, supernatant (S) and membrane pellet (P) after insertion of native cytochrome b6; lanes 4–7, supernatant and membrane pellet after insertion of denatured cytochrome b6. Cytochrome b6 was isolated from Synechocystis sp. PCC 6803, biotin labelled and anti-biotin antibodies was used for detection. (B) Lane 1, molecular weight standard; Lanes 2–3, membrane fraction after ss-cytochrome b6 insertion with and without chaotropic extraction, respectively. Antibodies against N-terminal residues of cytochrome b6 were used (10 μg of total protein per each lane was applied). (C) Lanes 1 and 2, supernatant (S) and membrane pellet (P) after insertion of ss-cytochrome b6. Urea was used as a chaotropic agents (CH) and anti-biotin antibodies were used for protein detection. All the experiments were repeated twice and 10 μg of total protein per each lane was applied.
Furthermore, in a control experiment, after insertion of cytochrome b6, we followed the native cytochrome folding during chaotropic extraction (Supplemental Fig. S2) and this protein was resistant to the treatment as reported23. As shown in Fig. 2B,C, both of the imported proteins, ss-cytochrome b6 and the native cytochrome b6, showed resistance to 4 M urea extraction. Hence, the native cytochrome b6 could not incorporate with thylakoid membranes spontaneously nor posttranslationaly through stromal proteins of Sec pathway.
A chemical denaturation of isolated cytochrome b6 was followed by UV-Vis spectroscopy and a circular dichroism analysis at 222 nm. The secondary structure of native cytochrome b6 was lost upon unfolding in the presence of GuHCl (guanidine hydrochloride). The GuHCl treatment led to a relatively flat spectrum indicating a substantial loss of secondary structure (Fig. 1A and Table 1). Furthermore, in the visible CD spectrum of the denatured cytochrome b6, a loss of heme and no Cotton effects in the Soret-band region were observed (Fig. 1B), as reported previously24.
Similarly, following the import assay of unfolded cytochrome b6, the protein integration with thylakoid membranes was tested with use of a chaotropic agent. As shown on Fig. 2A (lanes 4–7), independent of stromal fraction presence, no unfolded cytochrome b6 band was observed in thylakoid membranes after urea treatment. Hence, the denaturized cytochrome b6 neither integrated with thylakoid membranes utilizing spontaneous nor posttranslational Sec pathways. However under denaturing conditions, noncovalently bound haems dissociate from proteins25, although this did not affect the shape of the electrophoretic band for cytochrome b6 (Fig. 2A lane 4).
Spectroscopic analysis was conducted for integration into the thylakoid membranes of E. coli expressed spinach apocytochrome b618. Refolding of apocytochrome b6 was monitored by far-UV CD spectroscopy at 222 nm (Supplemental Fig. S3). As shown on Fig. 3A, in contrast to denaturized Synechocystis sp. PCC 6803 cytochrome b6, the denaturized apocytochrome b6 was only partially imported to the thylakoids (lane 3). Furthermore, both denaturized and refolded spinach apocytochrome b6 was sensitive to 4 M urea extraction (Fig. 3 lanes 4, 6 and 7) as well as by others chaotropes (Supplemental Fig. S4A). Hence spinach apocytochrome b6 did not integrate into the thylakoid membranes spontaneously, as well as the cyanobacteria native protein. On the other hand, ss-apocytochrome b6 was imported into the thylakoid membrane and properly oriented in the membrane with C-terminus and N-terminus at stromal side of thylakoid membrane (Fig. 3B lanes 2 and 3) and antibodies against cpSecY prevent cpSecA-dependent protein translocation into membrane by the Sec pathway26,27,28 (Supplemental Fig. S5).
Thylakoid membrane fractions after insertion of spinach apocytochrome b6 and treatment with chaotropic agents (CH).
(A) The integration of the cytochrome b6 into the thylakoid membrane in the presence or absence of stromal fraction (SF) was analysed with Western blot. Lane 1, thylakoid membrane; lane 2, purified apocytochrome b6; lane 3 pelleted membrane fraction with inserted denatured (unfolded) protein; lane 4, pelleted membrane fraction with inserted denatured protein after chaotropic treatment with urea; lane 5, control, membrane fraction with inserted into membrane ss-apocytochrome b6; lanes 6 and 7, supernatant and membrane pellet, respectively after centrifugation of refolded apocytochrome b6 and inserted into membrane. The experiments were repeated twice and 10 μg of total protein per lane was applied. (B) Thylakoid membrane fractions after insertion of spinach ss-apocytochrome b6 and treatment with carboxypeptidase B. Lane 1, purified ss-apocytochrome b6; lane 2, thylakoid membrane with inserted ss-apocytochrome b6; lane 3, membrane treated with carboxypeptidase B (depicted with (C) after protein insertion; lanes 4 and 5, membrane and supernatant fraction with inserted denatured protein after carboxypeptidase B treatment; lane 6, supernatant fraction similar to lane 5, but membrane with inserted denatured protein was treated with urea and carboxypeptidase B and an antibody against N-terminus of cytochrome b6 was used. Cytochrome b6 was biotin labelled and anti-biotin antibodies were used for detection with the exception of (B) line 6.
In the case of denaturized cytochrome b6, no biotin signal was detected after carboxypeptidase B treatment in both the membrane pellet and supernatant (Fig. 3B lanes 4 and 5), although the N-terminal signal of denaturized cytochrome b6 was observed in the supernatant.
PsbW integrates spontaneously with the isolated thylakoid membranes
In order to validate our cytochrome b6 insertion assays, we followed the thylakoid membrane integration of the cytosolic single span subunit W of PS II (PsbW), since this protein inserts into the thylakoid membrane by an apparently spontaneous pathway29. PsbW was used as an independent control for our experimental model. Since our experiments test whether the insertion of cytochrome b6 into the thylakoid membrane occurs spontaneously, the spontaneous insertion of mature PsbW into the thylakoid membrane observed in the same experimental model validates our methodological approach.
In order to probe the structure of the soluble form of the synthetic PsbW protein, biophysical analyses by CD and MS (mass spectroscopy) were performed. The CD spectra of denatured and DDM refolded PsbW differed significantly in secondary structure, indicating that the PsbW protein forms a transmembrane α-helix in hydrophobic environments of DDM (Fig. 4). The obtained spectra were overall in good agreement with previous studies29. The determined masses of the denatured and refolded protein protein’s (base on mass spectra), allowed us to establish the oligomeric stoichiometry of PsbW complex before insertion into the membrane. The MS analysis observed for biotynylated PsbW (6394.65 Da) agreed with the theoretical molecular weight of the monomeric species (6055.49 Da). Furthermore, MS and CD analysis showed that these proteins exist mainly as monomers (~89% for monomers and ~11% for oligomers).
As shown on Fig. 5, our in vitro experiments verified that synthetic PsbW is indeed spontaneously inserted into the isolated thylakoid membrane. The thylakoid import assays showed that the PsbW inserted into the thylakoid membranes and sorted efficiently also in an absence of a stromal fraction (quantified by densitometry analysis) and in the presence of apyrase (Supplemental Fig. S6). Apyrase is an ATP-diphosphohydrolase that catalyses the sequential hydrolysis of ATP to ADP and ADP to AMP and releases inorganic phosphate and prevents SecA de-insertion and further translocation across the thylakoid membrane by the ATP-dependent Sec pathway.
Thylakoid membrane fractions after insertion of PsbW.
The integration of the PsbW into the thylakoid membrane the presence or absence of stromal fraction was analysed by Western blot. Lane 1, thylakoid membrane before insertion; lanes 2–4 and 6–8, thylakoid membrane after insertion of PsbW; and lane 5, molecular weight standard. Antibodies against biotin were used for immunodetection. C - membrane treated with carboxypeptidase B after protein insertion, PK - membrane treated before protein insertion with proteinase K. On each lane, 10 μg of protein was applied. Identification of psbW protein in Western blot was also confirmed using MS.
Following the incubation of DDM vesicles of PsbW with carboxypeptidase B that catalyzes the hydrolysis of the basic amino acids from the C-terminal position of polypeptides (Fig. 5, lanes 4 and 7), the biotin labelled C-terminus of PsbW was completely sensitive to digestion and no biotin signal was detected after carboxypeptidase B treatment of PsbW. Hence incorporation of PsbW into the membrane was direct, with the N-terminus and the C-terminus on the opposite sides of the membrane. Furthermore, the thylakoid membranes proteinase K pretreatment did not inhibit insertion of the PsbW protein (Fig. 5, lane 6). Finally, the membrane integrated PsbW was completely insensitive to removal (Supplemental Fig. S7). These results confirmed previous reports of spontaneous insertion of PsbW into the thylakoid membrane29,30,31.
SRP-related chloroplast proteins are responsible for the cytochrome b6 integration into thylakoid membrane
The results of the import assays questioned both spontaneous and Sec-dependent mechanisms for cytochrome b6 import into thylakoid membrane. Furthermore, since efficient import was observed for unfolded cytochrome b6 only, the involvement of posttranslational SRP mechanism seemed unlikely. Hence, to confirm directly that the cytochrome b6 is a chloroplast protein targeted in a GTP-dependent (guanosine triphosphates) process termed co-translational translocation, chloroplast import experiments were performed using cell free in vitro system.
Cell-free native spinach cytochrome b6 expression was performed in the linked system and transcription and translation reactions were separated. Translations were carried out in the presence of thylakoid membrane or thylakoid membrane with a stroma fraction. As shown in Fig. 6 (lanes 2 and 3), the translation product was detected in the thylakoid membrane. Pretreatment of thylakoids and stroma with proteinase K (Fig. 6, lane 4) prevented insertion of cytochrome b6 into membrane due to degradation of thylakoid and stromal translocation proteins. Furthermore, in Fig. 6, lane 5, a significant level of insertion was achieved in the presence of cpSecY antibody. Antibodies against cpSecY prevent cpSecA-dependent protein translocation into membrane by the Sec pathway26,27,28, suggesting that integration of the cytochrome b6 is Sec-independent. Following the import assay of native cytochrome b6, protein integration with thylakoid membrane was tested with the use of a chaotropic agent (Supplemental Fig. S4B). Furthermore, the membrane integrated cytochrome b6 was completely insensitive to removal by urea, KSCN and NaOH.
Autoradiograph of cytochrome b6 expressed in cell-free assay in the presence of thylakoid membrane and stroma.
(A) Lane 1, thylakoid membrane as a control; lanes 2 and 3, translation of cytochrome b6 in the presence of thylakoid membrane and stromal fraction, supernatant (S) and membrane pellet (P) after fractionation; (B) Lanes 1 and 2 translation of cytochrome b6 in the presence of thylakoid membrane and stromal fraction, supernatant (S) and membrane pellet (P) after fractionation; lane 3 and 4, translation of cytochrome b6 in the presence of thylakoid membrane, stroma and cpSecY antibody, membrane pellet (P) and supernatant after fractionation; (C) Lane 1, thylakoid membrane as a control; lane 2 and 3, translation of cytochrome b6 in the presence of thylakoid membrane and stromal fraction, supernatant (S) and membrane pellet (P) after fractionation, lane 4 and 5, translation of cytochrome b6 in the presence of thylakoid membrane, stroma and cpSecY antibody, membrane pellet (P) and supernatant (S) after fractionation; lane 6 and 7 same as in lane 2 and 3 but endogenous RNA in stroma was removed by enzymatic digestion before use in translation reaction (reaction were performed in the presence of RNasin ribonuclease inhibitor).
Next, to identify chloroplast proteins that could govern cytochrome b6 import a chemical cross-linking analysis combined with a mass spectrometry approach was applied. Cytochrome b6 cell-free translations were carried out in the presence of thylakoid membrane with a stroma fraction. Interacting proteins were identified with the cytochrome b6 targeted ribosome-nascent chain complexes (RNCs) after immunoprecipitation of cross-linked proteins with an antibody against cytochrome b6. These proteins were identified with MS using Mascot Distiller software as shown in Table 2 and Supplemental Table S1. A MS analysis of proteins co-immunoprecipitated with an antibody against cytochrome b6 without crosslinker was used as a control of the specificity of the cross-linking (Supplemental Table S2).
The identified proteins were previously recognized either as crucial for SRP-mediated co-translational transport into thylakoid membrane (cpSRP54, cpFtsY and ALB3) or for untranslated plastid mRNA stabilization (CSP41). The cytochrome b6 elongating nascent chain is known to interact with the cpSRP54 but not with cpSecY19. Our results indicate that targeting and insertion of cytochrome b6 protein into the thylakoid membrane occurs through co-translational SRP pathway in vivo.
The MS analysis of cross-linking products shown that cpSRP, cpFtsY and ALB3 form a complex with RNC-cytochrome b6 complexes at the membrane. Furthermore, we have demonstrated that GTP hydrolysis is required for cytochrome b6 insertion. As presented in Fig. 7B, (lanes 3 and 4) the GMP-PNP (5′ guanylylimidodiphosphate, non-hydrolysable GTP analogue) inhibits cytochrome b6 integration, suggesting that cpSRP–cpFtsY complex occupies functional ALB3 integration sites.
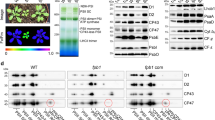
Autoradiograph of cytochrome b6 expressed in cell free assay in the presence of thylakoid membrane and stroma.
(A) Lane 1, translation of cytochrome b6 - control; lane 2 and 3, translation of cytochrome b6 in the presence of thylakoid membrane, stromal fraction and ALB3 antibody, supernatant (S) and membrane pellet (P) after fractionation; lane 4, translation of cytochrome b6 in the presence of thylakoids membrane and stroma; lane 5 and 6, translation of cytochrome b6 in the presence of thylakoid membrane and stromal fraction, supernatant (S) and membrane pellet (P) after fractionation. (B) Lane 1, translation of cytochrome b6, membrane pellet fraction after fractionation - control; lane 2, translation of cytochrome b6 in the presence of 0.5 mM non-hydrolysable ATP analogue (AMP-PNP, 5′ adenylylimidodiphosphate), membrane pellet; lane 3, translation of cytochrome b6 in the presence of 0.1 mM GMP-PNP, membrane pellet; lane 4, translation of cytochrome b6 in the presence of 0.5 mM GMP-PNP.
ALB3 is crucial for SRP-dependent cytochrome b6 import into the thylakoid membrane
The SRP-dependent pathway delivers RNC complexes to the Sec/ALB3 translocon32,33,34 or via independent ALB3 insertase in the cytoplasmic membrane33. Hence, the results of MS detection of ALB3 suggested that cytochrome b6 import into the thylakoid membrane was by the Sec/ALB3 insertion site or by ALB3 itself. To test this hypothesis, cell-free native spinach cytochrome b6 translations were carried out in the presence of thylakoid membrane with the stroma fraction in a presence of ALB3 antibody. We did not observe this effect in the presence of antibody against cytochrome f (negative control) or against C-terminus of cytochrome b6 (Supplemental Fig. S8). The anti-ALB3 serum was able to inhibit the association of a cpSRP–cpFtsY complex with ALB328,35,36. As shown in Fig. 7A, the translation product import into the thylakoid membrane was significantly reduced if ALB3 function was compromised (Fig. 7A, lane 3). In order to verify the specificity of the anti-ALB3 and to show that it does not block protein insertion into membrane by itself, we tested for PsbW protein spontaneous insertion into membrane in the presence of the anti-ALB3 (Supplemental Fig. S9). This study suggests that ALB3 is crucial for the cytochrome b6 import into the thylakoid membrane in vivo.
Discussion
The mechanisms responsible for targeting and insertion into the thylakoid membrane of both nuclear and chloroplast-encoded proteins are still not well understood. The presence of signalling sequence, protein structure and origin, are crucial for the choice of transport pathway involved in import process10. However, in case of some crucial thylakoid membrane proteins like cytochrome b6, the situation is less clear. The chloroplast-encoded cytochrome b6 is multi-spanning protein without a defined thylakoid membrane signal sequence37. Interestingly, it was reported that this protein is able to spontaneously integrate into the bacterial membrane16. Furthermore, cytochrome b6 fusion with an exogenic signalling sequence directed this protein import into a bacterial membrane via the Sec mechanism18. The main limitation of these experimental models was the substitution of thylakoid membrane with bacterial as well as the lack of stromal proteins that may facilitate the import process. Furthermore, in the case of the co-translational mechanism of membrane import, the presence of chloroplast-specific proteins may be crucial for cytochrome b6 folding and orientation within the thylakoid membrane19. Hence, to properly examine this protein import into the thylakoid membrane, the model should resemble the chloroplast environment in vivo. To address this problem and evaluate the mechanism governing cytochrome b6 integration in vivo, we performed a comparative analysis of both native and synthetic cytochrome b6 insertion into purified thylakoids. Our criteria for determining the correct mechanism of insertion was (i) that the protein should be integrated with the membrane and hence resistant to chaotropic extraction, (ii) and that it should be properly oriented within the membrane.
As a control for our experimental model, we tested if subunit W (PsbW) of the photosystem II is spontaneously inserted properly into the thylakoid membrane. PsbW is a single-span thylakoid membrane protein that is synthesized with a cleavable hydrophobic signal peptide and integrated into the thylakoid membrane by an apparently spontaneous mechanism29. The relatively simple insertion mechanism used by the PsbW together with the notable insertion-competence of the in vitro translation products suggested that this protein may be inherently stable in aqueous phases and hence a good model system for studying membrane proteins in general29. CD spectroscopy in the ultraviolet wavelength region can be used to estimate this protein secondary structural content and thus the protein folding state. An essential part of the control experiment (insertion of PsbW into the isolated thylakoid membrane) was to show that the factors and methods are actually responsible for the effects observed. This study confirmed previous findings38 that for the PsbW proper insertion into the thylakoid membrane, the presence of additional membrane proteins is not required. Furthermore, we also confirmed that PsbW is indeed inserted into the thylakoid membrane by the spontaneous pathway and does not interfere with the ALB3 protein during insertion.
Recently, it was reported that cytochrome b6 can spontaneously insert into the bacterial cytoplasmic membrane16. White and Wimley39 proposed a model in which the protein could follow one of two basic spontaneous insertion pathways: a “water path” in which the protein folds (forms α-helices) in the aqueous phase prior to insertion as a folded entity, or an “interface path” where the unfolded protein binds to the membrane interface where α-helix formation is promoted and insertion ensues29. Studies of the “water path” are precluded in most cases since the tendency membrane proteins is to aggregate in aqueous buffers. Although, recent studies have shown that the cytochrome b6 protein is largely unstructured in aqueous solution, it acquires a highly α-helical structure upon interaction with certain types of lipid prior to full insertion40.
In our study, the folding of cytochrome b6 was induced by dialysis of solubilised cytochrome b6 to buffer with DDM. The disordered structure of the solubilised form of cytochrome b6 in the presence of DDM was converted into a folded structure20.
The formation of α-helices is a key event during spontaneous protein insertion into the membrane. Energetic considerations strongly suggest that these must form prior to full protein insertion39,41. In contrast, insertion of partially unfolded cytochrome b6 isolated from Synechocystis sp. PCC 6803 was observed nevertheless. Furthermore, completely denatured cytochrome b6 was only partially imported to the thylakoids and could also be easily removed by urea washes. The formation of α-helices is a key event during membrane protein insertion, but the precise timing of α-helix formation is unclear in most cases39. An in vitro insertion is extremely difficult to address usually because of the insoluble nature of membrane proteins in aqueous buffer. Herein we show that DDM stabilizes native cytochrome b6. While soluble cytochrome b6 showed no detectable α-helix content, upon addition of DDM, there was a rapid increase in secondary structure formation. This indicates that the hydrophobic regions of cytochrome b6 are able to form α-helical structures in a hydrophobic environment, leading to protein insertion into the membrane29. This observation provides strong evidence that apocytochrome b6 is not able to insert into the thylakoids by a spontaneous pathway.
The Sec-dependent pathway for the integration of cytochrome b6 into the thylakoid membrane has been also suggested17,24. However, a lack of the N-terminal presequence in cytochrome b6 and the opposite orientation in the cytoplasmic membrane after expression in E. coli suggests that other pathways are utilized. Indeed, previous studies in bacteria show that the fusion of cytochrome b6 to MBP (maltose binding protein) directs the cytochrome b6 onto the Sec-dependent pathway17. Hence, topogenic signals in the amino acid sequence of cytochrome b6 protein are recognised by the E. coli Sec translocon leading to integration of this protein into the bacterial inner membrane although in an opposite orientation as compared to that in the thylakoid18.
Cytochrome b6 is not a typical passenger for unassisted integration since it lacks the cleavable N-terminal hydrophobic domain, which is missing in the prokaryotic or plastid-encoded counterparts. Consequently, a “helical hairpin”-type loop insertion mechanism is not feasible for cytochrome b6. Two variants of an SRP-dependent pathway exist, the posttranslational and the co-translational variant42. The SRP-dependent pathway delivers RNC complexes to the Sec translocon or Sec/ALB3 insertion site or to an independent ALB3 in the cytoplasmic membrane. The translocon is an essential component in cotranslational translocation because it binds the RNC and allows the translocation of the nascent chain and coordinates the insertion of proteins into the membrane.
In order to confirm directly that the cytochrome b6 is a chloroplast protein targeted in a GTP-dependent process termed co-translational translocation, we performed in vitro chloroplast import experiments. Although the cytochrome b6 protein was not synthesized in the wheat germ extracts but in the human in vitro protein system, the protein was efficiently transported into the thylakoid membranes. Mass spectrometry analysis of the cytochrome b6 crosslinked proteins allowed identification of cpSRP54, cpFtsY (both GTPase), ALB3 and CSP41 as potentially involved in targeting and insertion of cytochrome b6 protein into the thylakoid membrane. As reported previously, cytochrome b6 elongating nascent chain interacts with the cpSRP54 but not with cpSecY19. Hence, the result of crosslinking experiments suggests that in thylakoid, the RNCs contact the SRP in the stroma and then contact the ALB3 protein. Hence, cytochrome b6 insertion could be co-translational and mediated by cpFtsY and TMH (trans membrane helices) in order to enter the ALB3. cpFtsY would interact with the thylakoid membrane to support cpSRP-dependent targeting. Furthermore, a conserved amphipathic helix located at the N-terminus of cpFtsY that is both necessary and sufficient for interaction with the thylakoid membrane43.
We also show that impairment of ALB3 limits cytochrome b6 import into the thylakoid membrane. These results are consistent with earlier studies showing that nuclease pretreatment did not remove the cytochrome b6-RNC complexes from the membrane44,45. Moore, et al.35 indicated that cpSRP54/cpFtsY copurifies with ALB3 and proposed that cpFtsY and ALB3 form a complex together to insert thylakoid membrane proteins independently of cpSecY. Hence, the ALB3 protein appears to act in two functionally separate pools: one associated with cpSecYEG may serve in cotranslational integration activities and second functionally independent of cpSecYEG, it mediates integration posttranslationaly35. In the chloroplast, ALB3 can function as an insertase independent of other components (cpSecY). ALB3 is exposed rather to the stroma and therefore displays a higher accessibility for stromal components, also beyond the chloroplast SRP (cpSRP) pathway46. Nuclear-encoded light-harvesting chlorophyll proteins (LHCP) integrate into the thylakoid membrane using chloroplast SRP proteins and ALB3, but independently of cpSecY34. After import into the chloroplast stroma, the LHCP forms with the chloroplast cpSRP the LHCP-cpSRP54-cpSRP43 transit complex, then being directed to the SRP receptor (cpFtsY) at the thylakoid membrane. Binding both cpSRP54 and cpFtsY is a GTP dependent process35. At the membrane, the cpSRP43 protein within the transit complex is recognized by ALB3, by utilizing its C-terminal and membrane-embedded domain47. In addition to acting independently, ALB3 may insert proteins co-translationally into the thylakoid membrane using cpSec translocase. Specific ALB3′s substrates, that interact with ALB3 using the split-ubiquitin system, are D1, D2, CP43, PSI-A and the ATPase subunit CF0III48.
However, so far, it has not been shown that these proteins or any other membrane protein strictly requires both ALB3 and cpSecY for insertion into the thylakoid membrane of plants. Moreover, it was also shown that the D1 nascent peptide may also have the ability to localize to the membrane independently of cpFtsY and cpSecY via ALB3. However, the subsequent integration events may require the use of cpSecY for lateral movement or assembly of the D1 protein from the ALB3 translocon into the lipid bilayer49.
Herein, we show that thylakoid membrane treatment with anti-cpSecY antibody was not able to prevent the cytochrome b6 integration. However, antibodies against cpSecY prevent cpSecA-dependent protein translocation26,27,28. This suggest that cpSecY is likely not a part of the functional complex and cpSRP54 and cpFtsY form the GTP dependent targeting/translocation complex with ALB3.
Neither this nor previous studies have shown interaction of cytochrome b6 or RNC-cytochrome b6 complexes with cpSecY19. Hence, we propose that ALB3 insertase independently of cpSecY imports cytochrome b6 into the thylakoid membrane as depicted in Fig. 8. Although we cannot exclude the involvement of SecYEG translocon and do not rule out a situation that ALB3 protein may only be involved in an early stage of insertion as assembly factor.
Our mass spectrometry analysis of cytochrome b6 in vitro translation mixtures provided a first evidence that CSP41 interacts with RNC-cytochrome b6 complexes and maybe be involved in petB transcript stabilization or in a process of monocistronic petB mRNA synthesis. CSP41 proteins are highly abundant chloroplast proteins50 and ribosome 70S association of CSP41 a and b was reported. The first report on CSP41 suggested that it binds in vitro to the 3′ end of the petD mRNA51. To date multiple functions have been proposed for CSP41proteins: RNase activity, ribosomal biogenesis and plastid transcriptions52. Recently, it was demonstrated by RIP-chip analysis that CSP41 can bind to various chloroplast RNAs and the CSP41 proteins might serve to stabilize RNAs and to protect them against degradation53. This includes transcripts for the large Rubisco subunit (rbcL), PSI (psaA, psaB) and PSII (psbA, psbC, psbD) core proteins and 16S and 23S rRNAs53. The lack of CSP41 proteins decreases transcripts for photosynthetic proteins and of some ribosomal RNAs. This included petD mRNA, encoding subunit IV from the cytochrome b6 f complex and is transcribed as a part of the polycistronic cluster psbB-psbH-petB-petD, which is subsequently processed to monocistronic mRNAs54. In CSP41 compromised mutant petD mRNA of the cytochrome b6 f complex was 20% reduced comparing to WT53. However, the role of CSP41 in cytochrome b6 insertion into thylakoid membranes requires further studies.
Herein, we provided evidence for the role of SRP54, FtsY and ALB3 in targeting and insertion of cytochrome b6 protein into the thylakoid membrane. Our data demonstrated that the formation of a complex between cpSRP–cpFtsY and ALB3 is a necessary step in the integration of cytochrome b6 into thylakoid membranes. However, future studies that combine more exact membrane fractionation approaches with RNC profiling are required to understand the interplay between cytochrome b6 and ALB3 insertase and/or ALB3/SecYEG translocon as well as the role of CSP41.
Methods
Isolation Thylakoid Membranes and Stroma from Intact Chloroplasts
Seeds of Pea (Pisum Sativum, cv Calvedon) were germinated, planted in plastic trays and grown hydroponically as described in ref. 19. Fully developed leaves were harvested 2 h after the lights were turned on. Intact chloroplast from Pea leaves were isolated as described in ref. 19 and total chlorophyll (CHL) content was measured according to55.
Washed thylakoids and stromal extract were prepared from isolated chloroplasts as described in ref. 9 except that purified thylakoid membranes (at 1 mg mL−1 CHL) were incubated (5 min on ice) with micrococcal nuclease (8 units μL−1) in the presence of 10 mM CaCl2, protease inhibitor cocktail (P9599, Sigma-Aldrich) and 1 mM DTT (dithiothreitol). The digestion was stopped with 20 mM EGTA. Finally, the isolated thylakoid membranes were washed twice and resuspended in HMS buffer (50 mM HEPES-KOH, pH 8.0 with 10 mM MgCl2, 100 mM sorbitol, protease inhibitors cocktail and 1 mM DTT buffer at 4 mg mL−1 of CHL. Reconstituted lysates were prepared fresh prior the experiments by suspending the thylakoid membranes in the stroma.
Isolation of cytochrome b6
The isolation of spinach (GenBank: NC_002202.1) apocytochrome b6 from E. coli cells and the protein refolding/reconstitution assays were previously in refs 20 and 24, respectively. Native cytochrome b6 was isolated from Synechocystis sp. PCC 6803 membranes according to22. Cytochrome b6, both native and overexpressed, were biotinylated using the Thermo Scientific EZ-Link PFP-biotin assay.
Synthetic proteins
The PsbW protein (GenBank: CAA59409.1) was obtained from GenScript at purity above 85%. Cytochrome b6 as a fusion protein (ss-cytochrome b6) to the C-proximal part of the signal pre-sequence of the Oxygen Evolving Complex (SLQSDFKELAHKCEASKIAGFALATSALVASGASA, OE33, GenBank: BAA02554.1).
The Ala-X-Ala consensus sequence that is recognised by a thylakoid processing peptidase (TPP) and that cleaves off this transit sequence was changed to protect fusion protein from being cleaved (SLQSDFKELAHKCEASKIAGFALVTSALVASGRSA). An additional Lys was incorporated into C-terminus in order to allow for biotin labelling. The molecular weight of synthetic protein was assessed with the MALDI and ESI-MS techniques (electrospray ionization mass spectroscopy, QTOF Premier mass spectrometer, Waters Corp), using the Mascot database search engine (version 2.1, Matrix Science)56.
Mass spectroscopy
For identification, the proteins were analysed in electrophoresis, stained with Coomassie solution and blotted. Furthermore, protein bands were cut out and subjected to in-gel digestion. After reduction of cysteine residues with DTT and alkylation with iodoacetamide, proteins were digested with trypsin and the resulting peptide mixtures analyzed by liquid chromatography/MS. After preprocessing of the raw data using Mascot Distiller software, output lists of precursor and product ions were compared for identification to the NCBI database using the Mascot database search engine (version 2.1, Matrix Science)56. Protein scores are derived from ions scores as a non-probabilistic basis for ranking protein hits. Ions score is −10*Log(P), where P is the probability that the observed match is a random event56. We chose only proteins with unique queries. The number of matches MS/MS spectra that uniquely match to the accession and are not shared with other accessions were identified. These matched spectra pass the minimal criteria for ion score and have a false positive rate of less 1%. Furthermore, the analysis resulted in the identification of several hundreds of peptides that were impossible to assign to a specific protein. Therefore, the final analysis score cut off was set at 20 to eliminate low-score peptides and 40 to eliminate low-score proteins. Individual ions score > 41 indicate identity or extensive homology (p < 0.05)56. To calculate total score, the individual ions score with expect to a value lower than 0.05 was chosen for identified peptide (http://www.matrixscience.com/help/scoring_help.html)56.
Protein insertion into thylakoid membrane
In order to remove external endogenous proteins domains, thylakoid membranes were pre-incubated with proteinase K (40 μg mL−1 final) for 2 h on ice. The digestion was terminated with 4 mM PMSF (phenylmethylsulfonyl fluoride) and the thylakoid membranes collected (10 min, 11,000 × g). The proteins (0.3 mM final concentration) were incubated with the thylakoid membranes for approximately 1 h. Import incubation mixtures contained the excess of thylakoid membrane (50 μg CHL), stromal extract (30 μg protein) and 10 mM ATP (adenosine 5′-triphosphates). All experiments were carried out at 25 °C under a green safe light and a microstirrer/heater thermocouple with the reaction medium was used to ensure that the proteins were in a homogeneous distribution and there was a proper temperature within the subphase below the thylakoid membrane. If present, the stromal extract was equivalent to 1.3 times the CHL concentration. The import assays were terminated and unincorporated proteins removed by washing the thylakoid membrane three times in 2 mL of ice-cold 50 mM HEPES-KOH buffer, pH 8.0 with 330 mM sorbitol (3 min, 5,000 × g).
Proteolytic assessment of protein orientation in thylakoid membranes
The Michl, et al.57 procedure was used with the following modifications: thylakoid membranes after protein import assays were resuspended in HM buffer (20 mM HEPES-KOH, pH 8.5 mM MgCl2, 10 mM KCl and 10 mM DTT) and exposed to light (~300 μmol photons m−2 s−1) at 4 °C, in order to restore the ΔpH. Subsequently the thylakoid membranes were incubated with trypsin (60 μg mL−1, 1,000–2,000 BAEE (Na-benzoyl-arginine ethyl ester) units mg−1) for 10 min on ice. The digestion was stopped with trypsin inhibitor (120 μg mL−1, Sigma-Aldrich) and membranes were washed twice in HM enriched with 60 μg mL−1 trypsin inhibitor (20,000 × g for 5 min at 4 °C). The thylakoids were finally collected by centrifugation at 20,000 × g for 10 min at 4 °C and resuspended in stromal extract or in 15 μL of HM buffer containing 5 μg of trypsin inhibitor.
Thylakoid membranes prior to carboxypeptidase B (C9584, Sigma-Aldrich) treatment were supplemented with procaine and Triton X-100 at final concentrations of 3 mM and 0.01%, respectfully. Then, carboxypeptidase B (5 mg/mL, ≥125 units mg−1 protein) in 0.1 M Na+ citrate buffer, pH 6.5) was added for 1 h at 30 °C. Subsequently, after this treatment an equal aliquot of enzyme solution was added and the incubation continued for another 30 min. Controls received 0.1 M Na+ citrate buffer, pH 6.5, instead of the enzyme solution. The final pH of the samples was kept at 6.5–7.0. The proteolytic reaction was stopped by addition of the carboxypeptidase inhibitor (C0279, Sigma-Aldrich).
Assessment of integration of the targeted proteins into the thylakoid membrane
The proteins associated with but not integrated into the thylakoid membranes are not resistant to chaotropic agents23. To monitor integration of the targeted proteins, the thylakoid membranes (at 0.1 mg mL−1 CHL) were incubated for 15 to 30 min on ice in either 0.1 M NaHCO3-NaOH, pH 12.5 or 4 M urea in 50 mM HEPES-KOH pH 7.7, 50 mM potassium acetate, 100 mM mannitol, 5 mM magnesium acetate, 2 mM DTT and a protease inhibitor cocktail. After these incubations, the membranes were washed and collected in HMS buffer (16,000 × g for 3 min).
Cell free transcription-translation assay
Coupled transcription/translation reactions of spinach cytochrome b6 construct were performed with the Human in vitro Protein Expression Kit for DNA Templates (Thermo Scientific). For in vitro transcription of the full length cytochrome b6 plasmid pT7CFE1-b6 containing the 5′ UTR consisting of EMCV internal ribosome entry site (IRES) was used. pT7CFE1-b6 was isolated using a standard mini prep protocol (GeneJET Plasmid Miniprep Kit, K0502, ThermoFisher Scientific). To avoid compromising protein expression yield, RNase A was used during the purification. Transcription with T7 polymerase was performed according to Human in vitro Protein Expression Kit for DNA. The reaction was performed at 32 °C for 90 min in a 20 μL reaction mixture and a total of 1 μg of DNA template was used. The mRNA (2 μg) generated from the transcription was added to each translation reaction mix containing all of the machinery for protein expression (cell lysate for protein expression, accessory proteins, amino acid minus methionine, energy mix). Translation reactions were performed essentially according to manufacturer’s recommendations for a 50 μL translation reaction. However, some modifications were made for insertion of cytochrome b6 into thylakoid membrane. Radioactive [35S]methionine (1,000 Ci/mmol) and additional RNase and protease inhibitors (RNasin, antipain, pepstatin and leupeptin, 0.2 mM each) were used during the translation assays. Mannitol was added (100 mM) to stabilize thylakoid membrane. Cytochrome b6 was synthesized in vitro in the presence of thylakoid membrane (250 μg mL−1 of CHL) with or without the fresh prepared stromal fraction (equivalent to thylakoid membrane containing 250 μg mL−1 of CHL) for 2 h at 30 °C. To increase protein yield, 2 μL more of the transcription mixture (1.0 μg μL−1) and 1 μL of energy mix was added for each reaction being performed after 2 h of translation and each reaction mixture was incubated for another 2 hours. The total reaction volume was 100 μL. Each translational sample (25 μL) was analysed by SDS-PAGE and the autoradiography.
In order to remove external domains of endogenous proteins, thylakoid membranes or stroma were pre-incubated with proteinase K (40 μg mL−1 final) for 2 h on ice. The digestion was terminated with 4 mM PMSF and the thylakoid membranes collected (10 min, 11,000 × g). In order to remove endogenous RNA during translation reactions thylakoid membranes or stroma were pre-incubated with RNase A (5 ng RNase A) for 15 min at 37 °C. Prior to translation reaction ribonuclease inhibitor (RNasin, Promega) was added (2 μl RNasin ribonuclease inhibitor (40 units/μL).
Co-immunoprecipitation
To isolate targeted RNCs, translation reactions were diluted with one volume of ice-cold 50 mM HEPES-KOH, pH 7.8, 1 M potassium acetate, 10 mM magnesium acetate buffer (HMS) and centrifuged for 1 min at 12,000 × g. The thylakoid membranes pellets were re-suspended to the original volume (100 μl) in HMS buffer and treated (for 30 min at 0 °C) with the membrane-permeable crosslinkers: BSOCOES (homobifunctional N-hydroxysuccinade ester bis[2-(succinimidyloxycarbonyloxy) ethyl] sulfone, Pierce) or DMA (homobifunctional imidoester dimethyladipimidate, Pierce). Reactions were then quenched with a large molar excess of DTT in an equal volume of HMS buffer. The thylakoid membranes were than incubated with 1% DDM (30 min at 0 °C) and insoluble materials were removed by ultracentrifugation at 50,000 rpm (Beckman SW60 Ti rotor) at 4 °C for 1 h. The resultant supernatant was further incubated (30 min) with an excess (5 μg mL−1) of antibody against cytochrome b6 N-terminus19. Subsequently, protein A- Sepharose CL-4B (GE Healthcare) was added and incubation was continued for an additional 4 h with rotation at 4 °C. The resin was collected (500 × g, 5 min, 4 °C) and washed three times with two volumes of HMS buffer. After the final wash, the bound protein fraction was eluted (in 0.25 M Tris-HCl, pH 6.8 with 100 μL 5% SDS, 8 M urea, for 1 h at 37 °C), analysed in SDS-PAGE and subjected to autoradiography58. Furthermore, the detected bands were excised from a polyacrylamide gel and analysed by mass spectroscopy with fingerprints.
Spectra measurements and circular dichroism analyses
PsbW and cytochrome b6 were solubilised in 1% DDM. To remove the free detergent, samples were extensively dialyzed against HM buffer and then centrifuged at 100,000 × g to remove any insoluble aggregates. Far UV and VIS CD spectra were performed as described in ref. 24. To gain information on the secondary structure, the CD spectra were analysed59.
Autoradiography and Western blot
The protein concentration was determined with the BCA (bicinchoninic acid) reagent according to the manufacturer’s instructions (Pierce). Following the normalization of protein concentrations, samples were mixed with an equal volume of 2X Laemmli sample buffer and incubated for 5 min at 95 °C prior to separation by Tricine/Tris SDS-PAGE as described in ref. 60. Prior to autoradiography, gels were stained with Coomassie brilliant blue, to confirm equal protein loading and dried under vacuum. The X-ray film was placed over the dry gel and exposed for 24 h. Importantly, we always used the same exposure time so that the image intensity was used as a consistent guide to the exposure required for subsequent autoradiography. Following SDS-PAGE, the proteins were transferred to nitrocellulose membranes (180 mA for 30 min at 4 °C). The membranes were then blocked with bovine serum albumin (BSA, Sigma-Aldrich) in PBS/Tween-20 (3% BSA, 0.5% Tween-20) for 1–2 hours, followed by immunoblotting with the primary antibody specified for each experiment: antibodies against N-terminal residues of cytochrome b6 as well as against the interhelical part of cytochrome b6 (between helix 1 and 2 and between helix 3 and 4)19 and against ALB336 were prepared in rabbits injected with the synthetic peptide (GenScript); biotin antibody was purchased from Abcam® (ab53468). After washing steps (3 washes with 20 mM sodium bicarbonate buffer pH 7.4, 5 min each at room temperature, followed by 3 washes with 100 mM glycine pH 2.4, 10 min each, at room temperature) the membranes were incubated with goat anti-rabbit IgG (H + L) secondary antibodies (BioRad) and detected using ECL (enhanced chemiluminescence, Amresco). Densitometry was performed using Image Lab software v. 4.1 (BioRad). We use the same exposure time for all autoradiography.
Additional Information
How to cite this article: Króliczewski, J. et al. ALB3 Insertase Mediates Cytochrome b6 Co-translational Import into the Thylakoid Membrane. Sci. Rep. 6, 34557; doi: 10.1038/srep34557 (2016).
References
Cramer, W. A., Yamashita, E. & Hasan, S. S. In Encyclopedia of Biological Chemistry Vol. 4 (eds Lane, M. D. & Lennarz, W. J. ) 167–171 (Academic Press, Waltham MA, 2013).
Tikhonov, A. N. The cytochrome b6f complex at the crossroad of photosynthetic electron transport pathways. Plant Physiol. Biochem. 81, 163–183, doi: 10.1016/j.plaphy.2013.12.011 (2014).
Joliot, P. & Johnson, G. N. Regulation of cyclic and linear electron flow in higher plants. Proc. Natl. Acad. Sci. USA 108, 13317–13322, doi: 10.1073/pnas.1110189108 (2011).
Fey, V., Wagner, R., Bräutigam, K. & Pfannschmidt, T. Photosynthetic redox control of nuclear gene expression. J. Exp. Bot. 56, 1491–1498, doi: 10.1093/jxb/eri180 (2005).
Chi, W., Ma, J. & Zhang, L. Regulatory factors for the assembly of thylakoid membrane protein complexes. Philos. Trans. R. Soc. Lond. B Biol. Sci. 367, 3420–3429, doi: 10.1098/rstb.2012.0065 (2012).
Aldridge, C., Cain, P. & Robinson, C. Protein transport in organelles: Protein transport into and across the thylakoid membrane. FEBS J. 276, 1177–1186, doi: 10.1111/j.1742-4658.2009.06875.x (2009).
Schuenemann, D. Mechanism of protein import into thylakoids of chloroplast. Biol. Chem. 388, 907–915, doi: 10.1515/BC.2007.111 (2007).
Yuan, J. & Cline, K. Plastocyanin and the 33-kDa subunit of the oxygen-evolving complex are transported into thylakoids with similar requirements as predicted from pathway specificity. J. Biol. Chem. 269, 18463–18467 (1994).
Cline, K., Henry, R., Li, C. J. & Yuan, J. G. Multiple Pathways for Protein Transport into or Across the Thylakoid Membrane. EMBO J. 12, 4105–4114 (1993).
Robinson, C., Woolhead, C. & Edwards, W. Transport of proteins into and across the thylakoid membrane. J. Exp. Bot. 51, 369–374, doi: 10.1093/jexbot/51.suppl_1.369 (2000).
Mant, A., Woolhead, C. A., Moore, M., Henry, R. & Robinson, C. Insertion of PsaK into the thylakoid membrane in a “Horseshoe” conformation occurs in the absence of signal recognition particle, nucleoside triphosphates, or functional albino3. J. Biol. Chem. 276, 36200–36206, doi: 10.1074/jbc.M102914200 (2001).
Zygadlo, A., Robinson, C., Scheller, H. V., Mant, A. & Jensen, P. E. The properties of the positively charged loop region in PSI-G are essential for its “spontaneous” insertion into thylakoids and rapid assembly into the photosystem I complex. J. Biol. Chem. 281, 10548–10554, doi: 10.1074/jbc.M512687200 (2006).
Zak, E., Sokolenko, A., Unterholzner, G., Altschmied, L. & Herrmann, R. G. On the mode of integration of plastid-encoded components of the cytochrome bf complex into thylakoid membranes. Planta 201, 334–341, doi: 10.1007/s004250050075 (1997).
Zhang, L., Paakkarinen, V., Suorsa, M. & Aro, E. M. A SecY homologue is involved in chloroplast-encoded D1 protein biogenesis. J. Biol. Chem. 276, 37809–37814, doi: 10.1074/jbc.M105522200 (2001).
Saif Hasan, S., Yamashita, E. & Cramer, W. A. Transmembrane signaling and assembly of the cytochrome b6f-lipidic charge transfer complex. Biochim Biophys Acta 1827, 1295–1308, doi: 10.1016/j.bbabio.2013.03.002 (2013).
Dreher, C., Prodohl, A., Weber, M. & Schneider, D. Heme binding properties of heterologously expressed spinach cytochrome b(6): implications for transmembrane b-type cytochrome formation. FEBS Lett. 581, 2647–2651, doi: 10.1016/j.febslet.2007.05.007 (2007).
Kroliczewski, J., Gubernator, B., Rogner, M. & Szczepaniak, A. On the mode of integration of the thylakoid membrane protein cytochrome b(6) into cytoplasmic membrane of Escherichia coli. Acta Biochim. Pol. 58, 335–343 (2011).
Kroliczewski, J., Hombek-Urban, K. & Szczepaniak, A. Integration of the thylakoid membrane protein cytochrome b6 in the cytoplasmic membrane of Escherichia coli. Biochemistry 44, 7570–7576, doi: 10.1021/bi047422w (2005).
Piskozub, M., Kroliczewska, B. & Kroliczewski, J. Ribosome nascent chain complexes of the chloroplast-encoded cytochrome b6 thylakoid membrane protein interact with cpSRP54 but not with cpSecY. J. Bioenerg. Biomembr. 47, 265–278, doi: 10.1007/s10863-014-9598-0 (2015).
Surma, M. A., Szczepaniak, A. & Króliczewski, J. Comparative Studies on Detergent-Assisted Apocytochrome b6 Reconstitution into Liposomal Bilayers Monitored by Zetasizer Instruments. PLoS ONE 9, e111341, doi: 10.1371/journal.pone.0111341 (2014).
Kim, S. J., Jansson, S., Hoffman, N. E., Robinson, C. & Mant, A. Distinct “assisted” and “spontaneous” mechanisms for the insertion of polytopic chlorophyll-binding proteins into the thylakoid membrane. J. Biol. Chem. 274, 4715–4721, doi: 10.1074/jbc.274.8.4715 (1999).
Boronowsky, U., Wenk, S., Schneider, D., Jager, C. & Rogner, M. Isolation of membrane protein subunits in their native state: evidence for selective binding of chlorophyll and carotenoid to the b(6) subunit of the cytochrome b(6)f complex. Biochim. Biophys. Acta 1506, 55–66, doi: 10.1016/S0005-2728(01)00184-0 (2001).
Breyton, C., de Vitry, C. & Popot, J. L. Membrane association of cytochrome b6f subunits. The Rieske iron-sulfur protein from Chlamydomonas reinhardtii is an extrinsic protein. J. Biol. Chem. 269, 7597–7602 (1994).
Kroliczewski, J. & Szczepaniak, A. In vitro reconstitution of the spinach chloroplast cytochrome b6 protein from a fusion protein expressed in Escherichia coli. Biochim. Biophys. Acta 1598, 177–184, doi: 10.1016/S0167-4838(02)00369-2 (2002).
Kuras, R. et al. Molecular genetic identification of a pathway for heme binding to cytochrome b(6). J. Biol. Chem. 272, 32427–32435, doi: 10.1074/jbc.272.51.32427 (1997).
Mori, H., Summer, E. J., Ma, X. & Cline, K. Component specificity for the thylakoidal Sec and Delta pH-dependent protein transport pathways. J. Cell Biol. 146, 45–56, doi: 10.1083/jcb.146.1.45 (1999).
Schuenemann, D., Amin, P., Hartmann, E. & Hoffman, N. E. Chloroplast SecY Is Complexed to SecE and Involved in the Translocation of the 33-kDa but Not the 23-kDa Subunit of the Oxygen-evolving Complex. J. Biol. Chem. 274, 12177–12182 (1999).
Jiang, F. et al. Chloroplast YidC homolog Albino3 can functionally complement the bacterial YidC depletion strain and promote membrane insertion of both bacterial and chloroplast thylakoid proteins. J Biol Chem 277, 19281–19288, doi: 10.1074/jbc.M110857200 (2002).
Woolhead, C. A., Mant, A., Kim, S. J., Robinson, C. & Rodger, A. Conformation of a purified “spontaneously” inserting thylakoid membrane protein precursor in aqueous solvent and detergent micelles. J. Biol. Chem. 276, 14607–14613, doi: 10.1074/jbc.M009600200 (2001).
Lorkovic, Z. J. et al. Molecular characterization of PsbW, a nuclear-encoded component of the photosystem II reaction center complex in spinach. Proc. Natl. Acad. Sci. USA 92, 8930–8934 (1995).
Thidholm, E. et al. Novel approach reveals localisation and assembly pathway of the PsbS and PsbW proteins into the photosystem II dimer. FEBS lett 513, 217–222, doi: 10.1016/s0014-5793(02)02314-1 (2002).
Sundberg, E. et al. ALBINO3, an Arabidopsis nuclear gene essential for chloroplast differentiation, encodes a chloroplast protein that shows homology to proteins present in bacterial membranes and yeast mitochondria. Plant Cell 9, 717–730, doi: 10.1105/tpc.9.5.717 (1997).
Hennon, S. W., Soman, R., Zhu, L. & Dalbey, R. E. YidC/Alb3/Oxa1 Family of Insertases. J. Biol. Chem. 290, 14866–14874, doi: 10.1074/jbc.R115.638171 (2015).
Klostermann, E., Droste Gen Helling, I., Carde, J. P. & Schunemann, D. The thylakoid membrane protein ALB3 associates with the cpSecY-translocase in Arabidopsis thaliana. Biochem. J. 368, 777–781, doi: 10.1042/bj20021291 (2002).
Moore, M., Goforth, R. L., Mori, H. & Henry, R. Functional interaction of chloroplast SRP/FtsY with the ALB3 translocase in thylakoids: substrate not required. J. Cell Biol. 162, 1245–1254, doi: 10.1083/jcb.200307067 (2003).
Moore, M., Harrison, M. S., Peterson, E. C. & Henry, R. Chloroplast Oxa1p Homolog Albino3 Is Required for Post-translational Integration of the Light Harvesting Chlorophyll-binding Protein into Thylakoid Membranes. J. Biol. Chem. 275, 1529–1532, doi: 10.1074/jbc.275.3.1529 (2000).
Schnell, D. J. Protein targeting to the thylakoid membrane. Annu Rev Plant Physiol Plant Mol Biol. 49, 97–126, doi: doi: 10.1146/annurev.arplant.49.1.97 (1998).
Woolhead, C. A. et al. Distinct Albino3-dependent and -independent Pathways for Thylakoid Membrane Protein Insertion. J. Biol. Chem. 276, 40841–40846, doi: 10.1074/jbc.M106523200 (2001).
White, S. H. & Wimley, W. C. Membrane protein folding and stability: physical principles. Annu. Rev. Biophys. Biomol. Struct. 28, 319–365, doi: 10.1146/annurev.biophys.28.1.319 (1999).
Bryson, E. A., Rankin, S. E., Carey, M., Watts, A. & Pinheiro, T. J. Folding of apocytochrome c in lipid micelles: formation of alpha-helix precedes membrane insertion. Biochemistry 38, 9758–9767, doi: 10.1021/bi990119o (1999).
Ladokhin, A. S. & White, S. H. Folding of amphipathic alpha-helices on membranes: energetics of helix formation by melittin. J. Mol. Biol. 285, 1363–1369, doi: 10.1006/jmbi.1998.2346 (1999).
Eichacker, L. A. & Henry, R. Function of a chloroplast SRP in thylakoid protein export. Biochim. Biophys. Acta 1541, 120–134, doi: 10.1016/s0167-4889(01)00151-3 (2001).
Marty, N. J. et al. The Membrane-binding Motif of the Chloroplast Signal Recognition Particle Receptor (cpFtsY) Regulates GTPase Activity. J. Biol. Chem. 284, 14891–14903, doi: 10.1074/jbc.M900775200 (2009).
Zoschke, R. & Barkan, A. Genome-wide analysis of thylakoid-bound ribosomes in maize reveals principles of cotranslational targeting to the thylakoid membrane. Proc. Natl. Acad. Sci. USA 112, E1678–E1687, doi: 10.1073/pnas.1424655112 (2015).
Barkan, A. Proteins encoded by a complex chloroplast transcription unit are each translated from both monocistronic and polycistronic messenger-RNAs. EMBO J. 7, 2637–2644 (1988).
Asakura, Y., Kikuchi, S. & Nakai, M. Non-identical contributions of two membrane-bound cpSRP components, cpFtsY and Alb3, to thylakoid biogenesis. Plant J 56, 1007–1017, doi: 10.1111/j.1365-313X.2008.03659.x (2008).
Falk, S., Ravaud, S., Koch, J. & Sinning, I. The C terminus of the Alb3 membrane insertase recruits cpSRP43 to the thylakoid membrane. J Biol Chem 285, 5954–5962, doi: 10.1074/jbc.M109.084996 (2010).
Pasch, J. C., Nickelsen, J. & Schünemann, D. The yeast split-ubiquitin system to study chloroplast membrane protein interactions. Appl Microbiol Biotechnol 69, 440–447, doi: 10.1007/s00253-005-0029-3 (2005).
Walter, B., Hristou, A., Nowaczyk, M. M. & Schunemann, D. In vitro reconstitution of co-translational D1 insertion reveals a role of the cpSec-Alb3 translocase and Vipp1 in photosystem II biogenesis. Biochem. J. 468, 315–324, doi: 10.1042/BJ20141425 (2015).
Zybailov, B. et al. Sorting signals, N-terminal modifications and abundance of the chloroplast proteome. PLoS One 3, e1994, doi: 10.1371/journal.pone.0001994 (2008).
Yang, J. & Stern, D. B. The Spinach Chloroplast Endoribonuclease CSP41 Cleaves the 3′-Untranslated Region of petD mRNA Primarily within Its Terminal Stem-Loop Structure. J. Biol. Chem. 272, 12874–12880, doi: 10.1074/jbc.272.19.12874 (1997).
Leister, D. Complex(iti)es of the ubiquitous RNA-binding CSP41 proteins. Front. Plant Sci. 5, doi: 10.3389/fpls.2014.00255 (2014).
Qi, Y. et al. Arabidopsis CSP41 proteins form multimeric complexes that bind and stabilize distinct plastid transcripts. J. Exp. Bot. 63, 1251–1270, doi: 10.1093/jxb/err347 (2012).
Stoppel, R. & Meurer, J. Complex RNA metabolism in the chloroplast: an update on the psbB operon. Planta 237, 441–449, doi: 10.1007/s00425-012-1782-z (2012).
Arnon, D. I. Copper enzymes in isolated chloroplasts. Polyphenoloxidase in Beta vulgaris. Plant Physiol. 24, 1–15, doi: 10.1104/pp.24.1.1 (1949).
Perkins, D. N., Pappin, D. J., Creasy, D. M. & Cottrell, J. S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567, doi: 10.1002/(SICI)1522-2683(19991201)20:18<3551::AID-ELPS3551>3.0.CO;2-2 (1999).
Michl, D., Robinson, C., Shackleton, J. B., Herrmann, R. G. & Klosgen, R. B. Targeting of proteins to the thylakoids by bipartite presequences: CFoII is imported by a novel, third pathway. EMBO J. 13, 1310–1317 (1994).
Bonifacino, J. S., Dell’Angelica, E. C. & Springer, T. A. Immunoprecipitation. Curr. Protoc. Protein Sci., doi: 10.1002/0471140864.ps0908s18 (2001).
Gubernator, B., Kroliczewski, J., Kallas, T. & Szczepaniak, A. Iron-sulfur cluster reconstitution of spinach chloroplast Rieske protein requires a partially prefolded apoprotein. Biochim. Biophys. Acta 1764, 735–742, doi: 10.1016/j.bbapap.2005.12.013 (2006).
Schagger, H. & von Jagow, G. Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal. Biochem. 166, 368–379 (1987).
Wang, P. & Dalbey, R. E. Inserting membrane proteins: The YidC/Oxa1/Alb3 machinery in bacteria, mitochondria and chloroplasts. Biochim Biophys Acta 1808, 866–875, doi: 10.1016/j.bbamem.2010.08.014 (2011).
Acknowledgements
This work was supported by grants NN301464434 from the Ministry of Science and Higher Education to J.K. We would like to thank Prof. Matthias Roegner and his group members for helpful in cytochrome b6 preparation from Synechocystis sp. PCC 6803 and Prof. James Collawn for critical reading of the manuscript. We would also like to acknowledge Prof. Andrzej Szczepaniak for his kind help and laboratory support. This publication was supported by Wroclaw Centre of Biotechnology Programme. The Leading National Research Centre (KNOW) for years 2014–2018.
Author information
Authors and Affiliations
Contributions
J.K. envisioned the project and supervised the research. Conceived and designed the experiments: J.K. Performed the experiments: J.K. and M.P. Analysed the data: R.B., J.K. and M.P. Contributed reagents/materials/analysis tools: R.B., B.K. and J.K. Wrote the paper: R.B., B.K. and J.K. All authors reviewed the manuscript.
Ethics declarations
Competing interests
Małgorzata Piskozub has paid employment at Amplicon Sp. z o. o. The following authors have no competing interests: Rafał Bartoszewski, Bożena Króliczewska, Jarosław Króliczewski.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Króliczewski, J., Piskozub, M., Bartoszewski, R. et al. ALB3 Insertase Mediates Cytochrome b6 Co-translational Import into the Thylakoid Membrane. Sci Rep 6, 34557 (2016). https://doi.org/10.1038/srep34557
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep34557
- Springer Nature Limited
This article is cited by
-
De-etiolation-induced protein 1 (DEIP1) mediates assembly of the cytochrome b6f complex in Arabidopsis
Nature Communications (2022)
-
Chloroplast PetD protein: evidence for SRP/Alb3-dependent insertion into the thylakoid membrane
BMC Plant Biology (2017)
-
Structural disorder in plant proteins: where plasticity meets sessility
Cellular and Molecular Life Sciences (2017)