Abstract
Transforming growth factor β (TGF-β) signaling facilitates tumor development during the advanced stages of tumorigenesis, but induces cell-cycle arrest for tumor suppression during the early stages. However, the mechanism of functional switching of TGF-β is still unknown, and it is unclear whether inhibition of TGF-β signaling results amelioration or exacerbation of cancers. Here we show that the tumor suppressor p53 cooperates with Smad proteins, which are TGF-β signal transducers, to selectively activate plasminogen activator inhibitor type-1 (PAI-1) transcription. p53 forms a complex with Smad2/3 in the PAI-1 promoter to recruit histone acetyltransferase CREB-binding protein (CBP) and enhance histone H3 acetylation, resulting in transcriptional activation of the PAI-1 gene. Importantly, p53 is required for TGF-β-induced cytostasis and PAI-1 is involved in the cytostatic activity of TGF-β in several cell lines. Our results suggest that p53 enhances TGF-β-induced cytostatic effects by activating PAI-1 transcription, and the functional switching of TGF-β is partially caused by p53 mutation or p53 inactivation during cancer progression. It is expected that these findings will contribute to optimization of TGF-β-targeting therapies for cancer.
Similar content being viewed by others
Introduction
p53 is the most important tumor suppressor and is inactivated by mutations or deletions in approximately 50% of all malignancies1. p53 is activated by various types of stress, and can cause multiple outcomes through different modes of transcriptional activation of its target genes (e.g. cell-cycle arrest, DNA repair, and apoptosis)2,3,4,5,6. For example, p53 induces cell cycle arrest and DNA repair when cells are exposed to low levels of DNA damage, whereas it induces cell death when cells are exposed to extensive DNA damage. Although some p53 effects may be independent of transcription7, transcriptional regulation by p53 is important for tumor suppression and loss of its function strongly promotes tumor development8.
Transforming growth factor-β (TGF-β) is a multifunctional cytokine that regulates various cellular responses such as cell growth, cell motility, differentiation, apoptosis, and immune-regulation9. In cancer, TGF-β acts as tumor suppressor to induce growth arrest, senescence, and apoptosis at the early stages of tumorigenesis, but acts as a tumor promoter to induce epithelial-mesenchymal transition (EMT) and to promote angiogenesis in addition to loss of growth inhibitory effects at the advanced stages of cancer10. The tumor-facilitative functions of TGF-β signaling are crucial for high grade of malignancies, and increased TGF-β expression by tumor cells correlates with the progression of colorectal and prostate cancers11,12. In addition, activation of TGF-β signaling correlates with the resistance to multiple cancer drugs13,14. Thus, TGF-β signaling switches its functions from tumor suppressive to facilitative during cancer progression10. TGF-β signaling is considered to be an attractive molecular target for cancer therapy, and inhibitors of TGF-β signaling, such as receptor kinase inhibitors, neutralizing antibodies, and antisense oligonucleotides, have been used in pre-clinical trials15. However, the mechanism of functional switching of TGF-β is still not clear, and identifying this mechanism is important for establishment effective TGF-β-targeted therapeutic strategies for cancer.
TGF-β signaling is transduced into the nucleus by Smad proteins16,17,18,19. TGF-β binds a complex of receptors (the TGF-β type I receptor (TβRI) and the TGF-β type II receptor (TβRII)) and activates receptor serine/threonine kinase. Activated TβRI selectively phosphorylates Smad2 and Smad3, resulting in complex formation with Smad4. This complex translocates into the nucleus, where it regulates the transcription of TGF-β target genes through the recruitment of transcriptional coactivators and/or corepressors20. Since the affinity of the activated Smad complex to the DNA is insufficient to support association with the promoters of TGF-β target genes, the complex usually requires other DNA-binding factors, so-called Smad cofactors, for eliciting specific transcriptional regulation21,22,23.
Crosstalk between p53 and TGF-β signaling has been reported24. Specifically, p53 is required for TGF-β-induced mesoderm differentiation during Xenopus embryonic development25,26 and TGF-β-induced growth arrest in mammalian cells through cooperation with Smads25. Cordenonsi et al. have shown that several TGF-β target genes were under the joint control of p53 and Smads, and that p53 adjusted TGF-β-induced transactivation by interacting with a cognate binding site on the Mix.2 promoter25. They also found that p53 is required for expression of other TGF-β-induced genes (e.g. p21, PAI-1, and MMP2) through cooperation with Smads, and the presence of a p53 binding site in their promoters25. Recently, p53-dependent regulation of PAI-1 gene expression by TGF-β has been analyzed by the Higgins laboratory27. Overstreet et al. have shown that TGF-β regulated p53 activity by stimulating p53 phosphorylation and acetylation, promoting interaction with Smads and subsequent binding of the p53/Smads complex to the PAI-1 promoter27. However, the detailed molecular mechanism underlying the crosstalk between p53 and TGF-β signaling has not yet been fully elucidated. Based on these findings, we suggest that p53 acted as a Smad cofactor to enhance the tumor suppressive functions of TGF-β. Here, we focused on the plasminogen activator inhibitor type-1 (PAI-1) gene, whose promoter contains both Smad binding element (SBE) and p53 responsive element (p53RE)27,28,29. PAI-1 is required for p53- or TGF-β-induced cellular senescence30,31. In this study, we revealed that TGF-β induced complex formation between p53 and Smads in the PAI-1 promoter, and that p53 was required for the recruitment of histone acetyltransferase CREB binding protein (CBP) and the acetylation of histone H3. Moreover, p53 is required for TGF-β-induced cytostatic activity, and PAI-1 is also involved in its effect in several cell lines. These findings suggest that p53 plays an important role in TGF-β-induced cytostatic activity via full activation of PAI-1 transcription, and that p53 status is involved in the functional switching of TGF-β signaling.
Results
p53 enhances TGF-β-induced PAI-1 expression
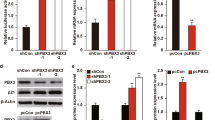
Firstly, we performed luciferase assay in HepG2 cells, human hepatoma cell lines expressing wild-type (WT) p53, to investigate the effects of p53 on PAI-1 transcription. PAI-1 promoter (−800~ + 71) (Fig. 1a) activity was enhanced by constitutively active TβRI (TβRI(T204D)) expression or p53 expression (Fig. 1b). Interestingly, p53 expression enhanced TGF-β-induced PAI-1 transcriptional activation. Conversely, transiently knockdown of p53 by siRNA almost completely suppressed TGF-β-induced PAI-1 transcription (Fig. 1c), indicating that TGF-β-induced PAI-1 transcription largely depended on p53. Of note, p53 affected the basal level of PAI-1 transcriptional activity. This is because p53 itself can activate PAI-1 transcription. To investigate the effect of p53 mutation on PAI-1 transcription, we used the mutant p53 R175H. Overexpression of p53 R175H resulted in diminished PAI-1 transactivation induced by TGF-β (Fig. 1d). Thus, both TGF-β and p53 are necessary for full activation of PAI-1 transcription in HepG2 cells.
p53 enhances TGF-β-induced PAI-1 expression.
(a) A schematic representation of the human PAI-1 promoter construct. (b,c) Effects of p53 overexpression (b) or p53 knockdown by siRNA (c) on the transactivation of PAI-1 promoter induced by TβRI(T204D) in HepG2 cells. Error bars represent s.d. (d) Effects of mutant p53 (R175H) on the transactivation of the PAI-1 promoter induced by treatment of TGF-β in HepG2 cells. Error bars represent s.d. (e) HepG2 cells and A549 cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 100 pM of TGF-β for the indicated periods. Expression of each gene was determined by semi-quantitative PCR. (f) HepG2 cells and A549 cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 100 pM of TGF-β for 6 h. The cell lysates were immunoblotted with the indicated antibodies. Uncropped images of gels/blots are shown in Supplementary Information, Figure S1.
Next, we examined the effects of p53 knockdown by siRNA on PAI-1 mRNA and protein expression in HepG2 cells. In addition to the results of the luciferase assay, p53 knockdown resulted in suppression of TGF-β-induced PAI-1 mRNA (Fig. 1e, left) and protein expression (Fig. 1f, left). TGF-β stimulation did not significantly affect p53 expression levels. The same result was also obtained using A549 cells, human lung cancer cell lines expressing WT p53 (Fig. 1e,f). Consistent with previous studies25,27, these results indicate that TGF-β and p53 synergistically regulate PAI-1 expression.
p53 selectively affects TGF-β target promoters containing both SBE and p53RE
As it has been reported that TGF-β signaling plays multiple roles in tumorigenesis, suppressively or facilitatively, we examined whether p53 affected the transactivation of other TGF-β target genes in HepG2 cells. Interestingly, p53 did not significantly affect TGF-β-induced Smad7 promoter (−557~ + 112) activation, which is a target promoter region of TGF-β and contains SBE, but not p53RE (Fig. 2a)32. This finding suggests that the regulation of TGF-β target genes by p53 depends on the existence of p53RE in these promoters. To demonstrate this more clearly, we constructed a PAI-1 promoter reporter in which p53RE was deleted (Fig. 2b). In contrast to the WT PAI-1 promoter, both overexpression and knockdown of p53 hardly affected the mutant PAI-1 promoter (Δp53RE) activity even after TGF-β stimulation (Fig. 2c,d). Moreover, p53RE-Luc, which contains only p53RE, was unresponsive to TGF-β (Fig. 2e).
p53 selectively affects TGF-β target promoters containing both SBE and p53RE.
(a) p53 did not significantly affect TGF-β-induced Smad7 promoter activation. HepG2 cells were transfected with Smad7-Luc in the presence or absence of p53 expression plasmid. After 24 h, cells were treated with 100 pM of TGF-β. After 18 h, luciferase activity was measured. The experiments were performed in triplicate, and the data are represented as the mean-fold activation ± s.d. (b) A schematic representation of the human PAI-1 promoter constructs. (c) HepG2 cells were transfected with the indicated constructs. After 24 h, luciferase activity was measured as in (a). (d) HepG2 cells were transfected with the indicated constructs and siRNAs. After 24 h, luciferase activity was measured as in (a). (e) TGF-β could not transactivate a p53-responsive reporter. HepG2 cells were transfected with p53RE-Luc. After 24 h, cells were treated with 100 pM of TGF-β for 18 h. The luciferase activity was measured as in (a). (f) HaCaT cells were transiently transfected with the indicated siRNAs. After 48 h, expression of each gene was determined by semi-quantitative PCR. (g) HaCaT cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 100 pM of TGF-β for 6 h. The cell lysates were immunoblotted with the indicated antibodies. Uncropped images of gels/blots are shown in Supplementary Information, Figure S1.
Tristetraprolin (TTP) promoter also contains responsive elements for both p53 and Smad33,34. Therefore, we examined the effects of p53 knockdown by siRNA on TTP mRNA expression in HepG2 and A549 cells. As expected, p53 knockdown also resulted in suppression of TGF-β-induced TTP mRNA (Fig. 1e). Taken together, these results suggested that the synergism of TGF-β signaling and p53 might occur in promoters containing both SBE and p53RE.
p53 inactivating mutations are found in approximately 50% of human cancers1. On the other hand, elevated expression of PAI-1 in tumors has been reported35. In fact, PAI-1 is up-regulated by TGF-β, even in cell types that carry mutations inactivating p53. This may be explained by other p53 family members, p63 and/or p73, which can compensate for p53 mutation in some cases. Cordenonsi et al. have previously shown that p63 knockdown blunted the induction of p21 by TGF-β in the human keratinocyte cell line, HaCaT (containing mutant H179Y/R828W in p53)25. Similarly, p63 knockdown resulted in suppression of TGF-β-induced PAI-1 expression in HaCaT cells (Fig. 2f,g). Thus, the regulatory function of p53 to express PAI-1 could be made redundant by the expression of other p53 family members. Alternatively, other Smad cofactor(s) may functionally compensate for mutations causing loss of function of the p53 gene by cooperating with Smads.
p53 does not significantly affect TGF-β signal transduction
To identify the detailed molecular mechanism of crosstalk between TGF-β signaling and p53 in PAI-1 transcription, we investigated the effects of p53 knockdown on Smad2 phosphorylation in HepG2 cells. Smad2 phosphorylation levels were detected by immunoblotting. One hour after TGF-β stimulation, Smad2 phosphorylation reached its peak, and then gradually decreased. TGF-β induced similar levels of Smad2 phosphorylation in both control cells and p53 knockdown cells (Fig. 3a). We also performed ChIP assay to examine the recruitment of Smad2/3 to the PAI-1 promoter in HepG2 cells. Consistent with the results in Fig. 3a, p53 knockdown did not significantly affect Smad2/3 recruitment to the PAI-1 promoter by TGF-β stimulation (Fig. 3b). Thus, p53 knockdown does not significantly affect TGF-β signal transduction in HepG2 cells. These findings suggest that p53 might selectively up-regulate PAI-1 gene transactivation after Smad2/3 binding to the SBE in response to TGF-β.
p53 affects neither Smad2 phosphorylation nor DNA-binding activity of Smad2/3.
(a) HepG2 cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 100 pM of TGF-β for the indicated periods. The cell lysates were immunoblotted with the indicated antibodies. (b) HepG2 cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 100 pM of TGF-β for 2 h. The cell lysates were subjected to chromatin immunoprecipitation (ChIP) analysis with the indicated antibodies. Extracted DNA fragments were analyzed by semi-quantitative PCR. Uncropped images of gels/blots are shown in Supplementary Information, Figure S1.
The C-terminal domain of p53 interacts with the MH2 domain of Smad3
It has been reported that p53 interacts with Smad2/3 in the presence or absence of the TGF-β family25. As shown in Fig. 4a, p53 was co-precipitated with Smad2/3 in HepG2 cells. We next examined p53-Smad3 binding using the deletion mutants of p53 (Fig. 4b). The C-terminal domain of p53 (p53 C), but not other mutants, co-immnoprecipitated with FLAG-Smad3 (Fig. 4c). We also performed immunoprecipitation analysis using various deletion mutants of Smad3 (Fig. 4d). p53 co-immnoprecipitated with the MH2 domain of Smad3 (Smad3 C) (Fig. 4e). Taken together, these findings indicate that the C-terminal domain of p53 interacts with the MH2 domain of Smad3 (Fig. 4f).
The C-terminal domain of p53 interacts with the MH2 domain of Smad3.
(a) HepG2 cells were treated with or without 100 pM of TGF-β for 1.5 h. Cell lysates were immunoprecipitated (IP) with an anti-Smad2/3 antibody and then immunoblotted with the indicated antibodies. (b,d) A schematic representation of full-length and deletion mutants of p53 (b) and Smad3 (d). (c,e) H1299 cells were transiently transfected with the indicated constructs. After 24 h, the cell lysates were immunoprecipitated (IP) with an anti-FLAG antibody and then immunoblotted with the indicated antibodies. (f) A schematic representation of protein-protein interactions between p53 and Smad3. The interacting domains are connected with a blue line. Uncropped images of blots are shown in Supplementary Information, Figure S1.
TGF-β induces the complex formation between p53 and Smad3 in the PAI-1 promoter
We next performed ChIP assay to examine the recruitment of p53 and Smad2/3 in the PAI-1 promoter (Fig. 5a) by TGF-β stimulation in HepG2 cells. TGF-β stimulation resulted in the recruitment of Smad2/3 to SBE in the PAI-1 promoter (Fig. 5b, left panel). Interestingly, p53 was also recruited to SBE in response to TGF-β. Similarly, the recruitment of p53 and Smad2/3 to p53RE in the PAI-1 promoter was induced by TGF-β stimulation (Fig. 5b, middle panel). The recruitment of p53 to SBE suggests that p53 interacts with SBE through Smad2/3, because p53 cannot directly interact with SBE36. Similarly it is suggested that Smad2/3 interacts with p53RE through p53. Of note, similar amounts of p53 and Smad2/3 were recruited to both cis-elements by TGF-β stimulation. Consistent with previous studies27, it is suggested that TGF-β induces complex formation between p53 and Smad2/3 in the PAI-1 promoter, and this complex is necessary for TGF-β-induced PAI-1 transcription. Thus, p53 acts as a partner to Smad for PAI-1 gene transactivation induced by TGF-β.
TGF-β promotes p53/Smads complex formation in the PAI-1 promoter.
(a) A schematic diagram of a human PAI-1 gene promoter. (b) HepG2 cells were treated with 100 pM of TGF-β for 2 h. The cell lysates were subjected to ChIP analysis with the indicated antibodies. Extracted DNA fragments were analyzed by real-time PCR. Error bars represent s.d. (c,d) HepG2 cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 100 pM of TGF-β for 2 h. The cell lysates were subjected to ChIP analysis with the indicated antibodies. Extracted DNA fragments were analyzed by real-time PCR as in (b). (e) The mechanism of PAI-1 transcription by the p53/Smads complex. TGF-β induces Smad2/3 to translocate into the nucleus, and the Smad complex interacts with p53 in the PAI-1 promoter. In other words, p53 plays a role as a DNA-binding partner of Smad. The p53/Smads complex efficiently recruits CBP to the PAI-1 promoter. The CBP recruitment induces histone H3 acetylation and relaxation of the chromatin structure to activate the PAI-1 transcription.
The p53/Smads complex efficiently recruits the transcriptional coactivator CBP to the PAI-1 promoter
It is still unclear why complex formation between p53 and Smad2/3 is necessary for PAI-1 transcription. We hypothesized that the p53/Smads complex might be required for the recruitment of transcriptional coactivator(s) in the PAI-1 promoter. We next aimed to examine the coactivator(s) cooperating with the p53/Smads complex.
CBP is a transcriptional coactivator that has histone acetyltransferase activity and cooperates with various transcriptional factors including p53 and Smad37,38,39. Histone acetylation leads to a relaxation of the chromatin structure, and activates transcription. Therefore, we investigated the recruitment of CBP to the PAI-1 promoter by ChIP analysis in HepG2 cells. TGF-β stimulation resulted in the recruitment of CBP to the PAI-1 promoter in control siRNA cells. On the other hand, the CBP recruitment in response to TGF-β was diminished in p53 knockdown cells (Fig. 5c, left panel). In addition, we examined histone H3 acetylation levels using anti-acetylated histone H3 (AcH3) antibody. Similar to CBP recruitment, histone H3 acetylation levels were enhanced by TGF-β stimulation in control siRNA cells, but were suppressed in p53 knockdown cells (Fig. 5d, left panel). These findings suggest that p53 is necessary for TGF-β-induced CBP recruitment to the PAI-1 promoter leading to histone H3 acetylation in HepG2 cells. The p53/Smads complex is necessary for the recruitment of CBP and histone acetylation, leading to PAI-1 transcriptional activation induced by TGF-β (Fig. 5e).
p53 is required for TGF-β-induced cytostasis in several cell lines
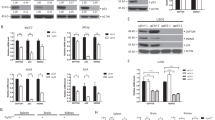
It has been shown that TGF-β has a cytostatic effect on human melanoma A375 cells40, which express the WT p53. p53 was also required for the induction of PAI-1 by TGF-β, similar to that seen in HepG2 and A549 cells (Fig. 6a,b). Therefore, we investigated whether p53 or PAI-1 was essential for TGF-β-induced cytostasis in A375 cells. As shown in Fig. 6c,e, p53 knockdown resulted in escape from growth arrest induced by TGF-β. Importantly, PAI-1 knockdown also resulted in bypass of TGF-β-mediated growth inhibition (Fig. 6d,e). On the other hand, p21 knockdown did not significantly alter the cytostatic effects of TGF-β.
p53 is required for TGF-β-induced cytostasis in several cell lines.
(a,b) A375 cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 100 pM of TGF-β for 6 h. Expression of each gene was determined by semi-quantitative PCR (a). The cell lysates were immunoblotted with the indicated antibodies (b). (c) A375 cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 100 pM of TGF-β for 96 h and then stained with crystal violet. (d) A375 cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 100 pM of TGF-β for 6 h. The cell lysates were immunoblotted with the indicated antibodies. (e) A375 cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 100 pM of TGF-β for 72 h and then the cell viability was examined using the CellTiter-Glo assay. (f) Cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 100 pM of TGF-β for 6 h. The cell lysates were immunoblotted with the indicated antibodies. (g,i) Cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 300 pM of TGF-β for 72 h and then the cell viability was examined using the CellTiter-Glo assay. (h) NMuMG cells were transiently transfected with the indicated siRNAs. After 48 h, cells were treated with 100 pM of TGF-β for 6 h. Expression of each gene was determined by semi-quantitative PCR. Uncropped images of gels/blots are shown in Supplementary Information, Figure S1.
To strengthen the significance of PAI-1 in TGF-β-mediated cytostasis, the same experiments were performed in other cell lines. HepG2 cells and non-tumorigenic human breast MCF10A cells possess WT p53 and exhibit a strong growth inhibitory response to TGF-β41,42. As shown in Fig. 6g, the growth of both cell lines was potently inhibited by TGF-β. Similar to A375 cells, PAI-1 knockdown also decreased the antiproliferative effect of TGF-β (Fig. 6f,g). Consistent with the findings of Kortlever et al.31, we also found that TGF-β-induced cytostasis was impaired in HaCaT cells (mutant p53 H179Y/R828W) when PAI-1 was knocked down (Fig. 6f,g). Furthermore, we investigated whether PAI-1 is involved in the cytostatic response to TGF-β in a non-transformed mouse mammary epithelial cell line, NMuMG. NMuMG cells biallelically express a WT and a missense mutant (R277C) form of p5343. In agreement with a previous report44, treatment with TGF-β resulted in decreased proliferation of NMuMG cells (Fig. 6i). p53 knockdown partially, but significantly, recovered the growth arrest induced by TGF-β (Fig. 6h,i). Likewise, PAI-1 knockdown in NMuMG cells also partially rescued the decreased proliferation in response to TGF-β (Fig. 6h,i). Of note, p21 knockdown in NMuMG cells partially reversed the cytostatic effect of TGF-β, in contrast to A375 cells. Collectively, these findings suggest that p53 plays an important role in TGF-β-induced cytostasis via full activation of PAI-1 transcription, and that loss of p53 function confers resistance to the growth inhibitory activity of TGF-β.
Discussion
In this study, we reveal the molecular mechanism of TGF-β-induced PAI-1 transcriptional activation. Specifically, TGF-β induces p53/Smads complex formation in the PAI-1 promoter, and this complex efficiently recruits CBP to the PAI-1 promoter, consequently leading to histone acetylation to relax the chromatin structure and activate PAI-1 transcription (Fig. 5e). Furthermore, these findings suggest that p53 plays a significant role in TGF-β-induced cellular senescence via full activation of PAI-1 expression.
Not only CBP but also other coactivators possibly enhance TGF-β-induced PAI-1 transcription
Although we clearly show that TGF-β stimulation induced CBP recruitment to the PAI-1 promoter, it seems that the quantity of CBP recruitment might be insufficient. We hypothesize that p300 or PCAF, which are other transcriptional coactivators of TGF-β signaling38,39,45,46, would be recruited to the PAI-1 promoter in response to TGF-β. Overstreet et al. have recently shown transcriptional complex formation involving p53/Smad3/p300 in response to TGF-β27. Therefore, it is plausible that TGF-β also causes p53/Smads/p300 complex formation, subsequently resulting in histone acetylation leading to PAI-1 transcriptional activation. Similarly, it is possible that other transcriptional coactivators and chromatin modifications regulate TGF-β-induced PAI-1 transcription.
The mechanism of p53 recruitment by TGF-β signaling
Various cellular stresses activate p53-induced transcription by stabilizing and recruiting p53 to target promoters2,3,4,5,6. In this study, we reveal that TGF-β stimulation induces p53 recruitment to the PAI-1 promoter without p53 stabilization, but how TGF-β signaling activates p53 is unclear.
We hypothesize that Smad2/3 binding to SBE in the PAI-1 promoter enhances the affinity between p53 and p53RE. TGF-β signaling could not recruit p53 to the promoter without SBE. TGF-β also induced Smad2/3 recruitment to the PAI-1 promoter in both control cells and p53 knockdown cells. Thus, it can be suggested that Smad2/3 binding to the promoter enhances the affinity between p53 and p53RE. In addition, it is also suggested that promoter DNA is necessary for p53/Smads complex formation. Thus, the promoter DNA likely acts as the scaffold for p53 and Smad2/3 to stabilize the p53/Smads complex. This mechanism has been supported by the findings of Cordenonsi et al.25. p53 altered TGF-β-induced transactivation by interacting with a cognate binding site on the Mix.2 promoter25. Alternatively, it is possible that posttranslational modifications of p53 occur in response to TGF-β. In this regard, it has been reported that TGF-β regulates p53 activity by stimulating p53 phosphorylation and acetylation27,47.
p53 selectively enhances TGF-β-mediated tumor suppression
It has been revealed that the promoter of p21 contains both SBE and p53RE, as well as PAI-148,49. p21 is a cyclin-dependent kinase inhibitor and induces G1 arrest49. p21 is also an effector for TGF-β-mediated tumor suppression. The TTP promoter contains responsive elements for both p53 and Smad33,34, and p53 is required for the induction of TTP by TGF-β. TTP is also known to act as a potent tumor suppressor50. We hypothesize that these genes are regulated by the same molecular mechanisms as PAI-1 transcription. Although further studies are needed to clarify this prediction, we suggest that p53 likely enhances transcriptional activation of various TGF-β target genes related to tumor suppressive functions, such as cellular senescence, cell cycle arrest, and apoptosis.
A previous report has indicated that TGF-β induces EMT in A549 cells, which express WT p5351. p53 suppresses the transcription of an EMT-inducing transcriptional factor Snail via the induction of micro RNA-34a/b/c genes52. Consistent with this, p53 knockdown enhanced TGF-β-induced Snail expression, but suppressed TGF-β-induced PAI-1 expression, in A549 cells (data not shown). Therefore, p53 mutation or loss might promote TGF-β-mediated cancer progression and metastasis. Moreover, Huang et al. have reported that the activation of TGF-β signaling confers drug resistance in cancer cells via MAPK activation13. They examined the effects of TGF-β stimulation on drug resistance in various cancer cell lines. However, the p53 status in those cell lines did not significantly affect TGF-β-induced drug resistance (e.g. WT p53 SKCO-1 cells versus mutant p53 PC-9 cells). Thus, p53 may not be involved in TGF-β-mediated drug resistance, another tumor promoter effect of TGF-β.
Finally, we demonstrated that PAI-1 is involved in the cytostatic response to TGF-β in several cell lines (Fig. 6). Kortlever et al. have shown that the induction of PAI-1 by TGF-β is critical for the induction of cellular senescence in HaCaT cells and primary mouse embryonic fibroblasts31. They also clarified that PAI-1 is not merely a marker of senescence, but is both necessary and sufficient for the induction of cellular senescence downstream of p5330. Mechanistically, PAI-1 expression leads to down-regulation of PI3K-Akt signaling and nuclear exclusion of cyclin D1. Loss of PAI-1 expression or uPA overexpression results in a bypass of cellular senescence30. Importantly, although PAI-1 is induced by TGF-β, the growth of most tumor cells is poorly inhibited by TGF-β. One possible explanation is that cells no longer respond to TGF-β-induced cytostatic effects when the PAI-1:uPA (urokinase plasminogen activator) balance shifts toward excess uPA31. Overexpression of uPA is often observed in several malignant tumors, and a higher level of uPA expression is associated with poor prognosis53. The cellular uPA/PAI-1 ratio would determine whether TGF-β suppresses the tumor growth or not31. Another possible explanation is that some cancer driver genes can lead to poor response to PAI-1-mediated cytostatic effects. Further studies are necessary to clarify these possibilities.
In summary, we found p53 to play a crucial role as a Smad partner in TGF-β-mediated tumor suppression, and the functional switching of TGF-β is partially caused by p53 loss or its mutation during tumor development. In future, further investigation into the regulation of TGF-β-mediated tumor-facilitative effects by p53, and identification of the effects of p53 loss or its mutation on TGF-β signaling are needed. In conclusion, we have identified a detailed molecular mechanism in which p53 acts in partnership with Smad to selectively enhance PAI-1 transcription. Furthermore, p53 might induce selective activation of TGF-β-mediated tumor suppression. Our study helps to clarify the mechanisms of TGF-β functional switching, and ultimately to establish effective TGF-β target therapies for high grade malignancies.
Methods
Cell Lines, Plasmids, and Transfections
HepG2, 293, A549, HaCaT, and NMuMG cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM) (Nacalai Tesque) supplemented with 4.5 g/L glucose, 10% fetal bovine serum (FBS) (SIGMA), 100 U/ml of penicillin G, and 100 μg/ml of streptomycin. For culture of NMuMG cells, media were also supplemented with 10 μg/ml of insulin (Wako). H1299 cells were cultured in RPMI1640 medium (Nacalai Tesque) containing 10% FBS and penicillin/streptomycin. MCF10A cells were cultivated in Mammary Epithelial Cell Growth Medium containing bovine pituitary extract, human EGF, human insulin, hydrocortisone (Promocell), penicillin/streptomycin, and choleratoxin (Wako). Cells were grown in a 5% CO2 atmosphere at 37 °C.
The original constructs encoding the human p53, Smad3, and TβRI were described previously54,55. Smad7-Luc was kindly provided by Dr. K. Miyazono (University of Tokyo)32. pCMV-β-galactosidase (β-gal) was purchased from Clontech. pGL4/p53RE (p53RE-Luc) construct was generated by subcloning a Asp718/HindIII fragment of pp53-TA-Luc (Clontech) into Asp718/HindIII of pGL4.10 (Promega). pGL4/PAI-1 (PAI-1-Luc) was generated by ligating the human PAI-1 promoter region (−800~ + 77)56 with pGL4.10. pGL4/PAI-1 (∆p53RE) (PAI-1-Luc (∆p53RE)) was also constructed by a polymerase chain reaction (PCR)-based approach using pGL4/PAI-1 as a template. All constructs were verified by sequencing.
For DNA transfection, plasmids were transiently transfected with Lipofectamine2000 regent (Invitrogen). For short interfering RNA (siRNA) transfection, siRNAs were transfected using Lipofectamine RNAiMAX reagent (Invitrogen) according to manufacturer’s protocol. siRNA oligo targeting human p53 has been previously described54. The siRNA duplexes were as follows: human PAI-1 siRNA sense strand, 5′-CCUGGGAAUGACCGACAUGTT-3′30; human p63 siRNA sense strand, 5′-CACACAUGGUAUCCAGAUGTT-3′57 mouse p53 siRNA sense strand, 5′-GUACAUGUGUAAUAGCUCCTT-3′58 mouse PAI-1 siRNA sense strand, 5′-GAACAAGAAUGAGAUCAGUTT-3′30 and mouse p21 siRNA sense strand, 5′-AGACCAGCCUGACAGAUUUTT-3′59(SIGMA). siRNA oligo targeting human p21 (VHS40202) and Stealth RNAi™ siRNA Negative Control Med GC Duplex #2 were obtained from Invitrogen.
Luciferase Assay
Cells were transfected with the luciferase reporter plasmid, expression plasmids, β-gal expression plasmid, and empty vector. The total amount of transfected DNA was the same in each experiment. Luciferase activity in cell lysates was measured. Luciferase activity was normalized against β-gal activity60.
RNA Extraction and Reverse-transcription
Total RNA extractions were performed as previously described61. First-strand cDNA was synthesized with PrimeScript first-strand cDNA Synthesis Kit (TaKaRa Bio Inc.) as previously described60.
Semi-quantitative PCR and Quantitative Real-time PCR
Semi-quantitative PCR was performed as previously described55. PCR was performed using AmpliTaq Gold 360 Mater Mix (Applied Biosystems) and a 2720 Thermal Cycler 2700 (Applied Biosystems). The following primer sequences were used: human p53, 5′-CTCACCATCATCACACTGGAAGAC-3′ (forward) and 5′-AGAGGAGCTGGTGTTGTTGGGCAG-3′ (reverse); human GAPDH, 5′-TGAAGGTCGGAGTCAACGGATTTGGT-3′ (forward) and 5′-CATGTGGGCCATGAGGTCCACCAC-3′ (reverse)61; human PAI-1, 5′-CATGGGGCCATGGAACAAGG-3′ (forward) and 5′-CTTCCTGAGGTCGACTTCAG-3′ (reverse); human TTP, 5′-TCATCCACAACCCTAGCGAA-3′ (forward) and 5′-GATGCGATTGAAGATGGGGA-3′ (reverse)61; human p63, 5′-CCAGACTCAATTTAGTGAGC-3′ (forward) and 5′-ACTTGCCAGATCATCCATGG-3′ (reverse)57; mouse p53, 5′-GATGACTGCCATGGAGGAGT-3′ (forward) and 5′-CTCGGGTGGCTCATAAGGTA-3′ (reverse)62; mouse GAPDH, 5′-GGCATTGTGGAAGGGCTCA-3′ (forward) and 5′-TCCACCACCCTGTTGCTGT-3′ (reverse)63; mouse PAI-1, 5′-GGGAAAAGGGGCTGTGTGAC-3′ (forward) and 5′-GTACACGGTGTGTGGCTGTC-3′ (reverse)64; and mouse p21, 5′-TGTCTTGCACTCTGGTGTCTGAGC-3′ (forward) and 5′-TCTTGCAGAAGACCAATCTGCG-3′ (reverse)65. PCR amplification was performed in the linear range and PCR products were separated by 1.5–2% agarose gel electrophoresis55.
Antibodies
The following commercially available antibodies used were: anti-PAI-1 (clone 41/PAI-1; BD Biosciences), anti-p53 (DO-1; Calbiochem), anti-β-actin (AC-15; Sigma), anti-phospho-Smad2 (Ser465/467) (138D4; Cell Signaling Technology), anti-Smad2/3 (clone 18/Smad2/3; BD Bioscience), anti-CBP (A-22; Santa Cruz Biotechnology), anti-acetyl-Histone H3 (catalog no. 06–599; EMD Millipore), anti-Myc (4A6; EMD Millipore), anti-FLAG (M2; Sigma), anti-HA (Y-11; Santa Cruz Biotechnology), and anti-GFP (B-2; Santa Cruz Biotechnology). Mouse immunoglobulin G1 (IgG1) (MB002; R & D Systems) and rabbit IgG (Southern Biotech) were used as controls.
Immunoprecipitation, and Immunoblotting
Immunoprecipitation and immunoblotting were performed as previously described60,66. The immunoprecipitated FLAG-protein complexes were eluted using 3xFLAG peptide (Sigma) for 30 min on ice.
Chromatin immunoprecipitation (ChIP) assay
ChIP assay was performed as previously described54,60. The purified DNA was analyzed by quantitative real-time PCR or semi-quantitative PCR. Quantitative real-time PCR was performed using GeneAce SYBR qPCR Mix α (NIPPON GENE) and a 7300 Real-Time PCR System (Applied Biosystems). For real-time PCR amplification, the following primer sequences were used: human PAI-1 promoter (SBE), 5′-GCAGGACATCCGGGAGAGA-3′ (forward) and 5′-CCAATAGCCTTGGCCTGAGA-3′ (reverse)67; human PAI-1 promoter (p53RE/TSS), 5′-CCAAGAGCGCTGTCAAGAAGA-3′ and 5′-AGGAATTCAGCTGCTGGAGG-3′ (reverse)68; and human HPRT1 first intron, 5′-TGTTTGGGCTATTTACTAGTTG-3′ (forward) and 5′-ATAAAATGACTTAAGCCCAGAG-3′ (reverse)54. For semi-quantitative PCR amplification, human PAI-1 promoter (SBE), 5′-CCTCCAACCTCAGCCAGACAAG-3′ (forward) and 5′-CCCAGCCCAACAGCCACAG-3′ (reverse)69 primers were used.
Cell Viability assay
Cell viability was measured using the CellTiter-Glo luminescent cell viability assay (Promega) according to manufacturer’s protocol54.
Additional Information
How to cite this article: Kawarada, Y. et al. TGF-β induces p53/Smads complex formation in the PAI-1 promoter to activate transcription. Sci. Rep. 6, 35483; doi: 10.1038/srep35483 (2016).
References
Hainaut, P. & Hollstein, M. p53 and human cancer: the first ten thousand mutations. Adv. Cancer Res. 77, 81–137 (2000).
Beckerman, R. & Prives, C. Transcriptional regulation by p53. Cold Spring Harb. Perspect. Biol. 2, a000935 (2010).
Kruse, J. P. & Gu, W. Modes of p53 regulation. Cell 137, 609–622 (2009).
Vousden, K. H. & Lane, D. P. p53 in health and disease. Nat. Rev. Mol. Cell Biol. 8, 275–283 (2007).
Riley, T., Sontag, E., Chen, P. & Levine, A. Transcriptional control of human p53-regulated genes. Nat. Rev. Mol. Cell Biol. 9, 402–412 (2008).
Oren, M. Decision making by p53: life, death and cancer. Cell Death Differ. 10, 431–442 (2003).
Green, D. R. & Kroemer, G. Cytoplasmic functions of the tumor suppressor p53. Nature 458, 1127–1130 (2009).
Hollstein, M., Sidransky, D., Vogelstein, B. & Harris, C. C. p53 mutations in human cancers. Science 253, 49–53 (1991).
Siegel, P. M. & Massagué, J. Cytostatic and apoptotic actions of TGF-beta in homeostasis and cancer. Nat. Rev. Cancer 3, 807–821 (2003).
Massagué, J. TGFβ signalling in context. Nat. Rev. Mol. Cell Biol. 13, 616–630 (2012).
Tsushima, H. et al. High levels of transforming growth factor beta 1 in patients with colorectal cancer: association with disease progression. Gastroenterology 110, 375–382 (1996).
Wikström, P., Stattin, P., Franck-Lissbrant, I., Damber, J. E. & Bergh, A. Vascular density is predictor of cancer-specific survival in prostatic carcinoma. Prostate 37, 19–29 (1998).
Huang, S. et al. MED12 controls the response to multiple cancer drugs through regulation of TGF-β receptor signaling. Cell 151, 937–950 (2012).
Sun, C. et al. Reversible and adaptive resistance to BRAF(V600E) inhibition in melanoma. Nature 508, 118–122 (2014).
Ikushima, H. & Miyazono, K. TGFbeta signalling: a complex web in cancer progression. Nat. Rev. Cancer 10, 415–424 (2010).
Wrana, J. L. Signaling by the TGFβ superfamily. Cold Spring Harb. Perspect. Biol. 5, a011197 (2013).
Derynck, R. & Zhang, Y. E. Smad-dependent and Smad-independent pathways in TGF-beta family signalling. Nature 425, 577–584 (2003).
Massagué, J., Seoane, J. & Wotton, D. Smad transcription factors. Genes Dev. 19, 2783–2810 (2005).
Heldin, C. H., Miyazono, K. & ten Dijke, P. TGF-beta signalling from cell membrane to nucleus through SMAD proteins. Nature 390, 465–471 (1997).
Miyazawa, K., Shinozaki, K., Hara T., Furuya, T. & Miyazono, K. Two major Smad pathways in TGF-beta superfamily signalling Genes Cells 7, 1191–1204 (2002).
Ikushima, H. & Miyazono, K. Cellular context-dependent “colors” of transforming growth factor-beta signaling. Cancer Sci. 101, 306–312 (2010).
Massagué, J. & Gomis, R. R. The logic of TGFbeta signaling. FEBS Lett. 580, 2811–2820 (2006).
Seoane, J. Escaping from the TGF-beta anti-proliferative control. Carcinogenesis 27, 2148–2156 (2006).
Dupont, S. et al. Convergence of p53 and TGF-beta signaling networks. Cancer Lett. 213, 129–138 (2004).
Cordenonsi, M. et al. Links between tumor suppressors: p53 is required for TGF-beta gene responses by cooperating with Smads. Cell 113, 301–314 (2003).
Takebayashi-Suzuki, K. et al. Interplay between the tumor suppressor p53 and TGF beta signaling shapes embryonic body axes in Xenopus. Development 130, 3929–3939 (2003).
Overstreet, J. M., Samarakoon, R., Meldrum, K. K. & Higgins, P. J. Redox control of p53 in the transcriptional regulation of TGF-β1 target genes through SMAD cooperativity. Cell Signal. 26, 1427–1436 (2014).
Dennler, S. et al. Direct binding of Smad3 and Smad4 to critical TGF beta-inducible elements in the promoter of human plasminogen activator inhibitor-type 1 gene. EMBO J. 17, 3091–3100 (1998).
Kunz, C., Pebler, S., Otte, J. & von der Ahe, D. Differential regulation of plasminogen activator and inhibitor gene transcription by the tumor suppressor p53. Nucleic Acid Res. 23, 3710–3717 (1995).
Kortlever, R. M., Higgins, P. J. & Bernards, R. Plasminogen activator inhibitor-1 is a critical downstream target of p53 in the induction of replicative senescence. Nat. Cell Biol. 8, 888–884 (2006).
Kortlever, R. M., Nijwening, J. H. & Bernards, R. Transforming growth factor-beta requires its target plasminogen activator inhibitor-1 for cytostatic activity. J. Biol. Chem. 283, 24308–24313 (2008).
Nagata, M. et al. Nuclear and cytoplasmic c-Ski differently modulate cellular functions. Genes Cells 11, 1267–1280 (2006).
Ogawa, K., Chen, F., Kim, Y. J. & Chen, Y. Transcriptional regulation of tristetraprolin by transforming growth factor-beta in human T cells. J. Biol. Chem. 278, 30373–30381 (2003).
Lee, J. Y. et al. Tumor suppressor p53 plays a key role in induction of both tristetraprolin and let-7 in human cancer cells. Nucleic Acids Res. 41, 5614–5625 (2013).
Czekay, R. P. et al. PAI-1: An Integrator of Cell Signaling and Migration. Int. J. Cell Biol. 2011, 562481 (2011).
Menendez, D., Inga, A. & Resnick, M. A. The expanding universe of p53 targets. Nat. Rev. Cancer 9, 724–737 (2009).
Gu, W., Shi, X. L. & Roeder, R. G. Synergistic activation of transcription by CBP and p53. Nature 387, 819–823 (1997).
Janknecht, R., Wells, N. J. & Hunter, T. TGF-beta-stimulated cooperation of smad proteins with the coactivators CBP/p300. Genes Dev. 12, 2114–2119 (1998).
Feng, X. H., Zhang, Y., Wu, R. Y. & Derynck, R. The tumor suppressor Smad4/DPC4 and transcriptional adaptor CBP/p300 are coactivators for smad3 in TGF-beta-induced transcriptional activation. Genes Dev. 12, 2153–2163 (1998).
Larisch-Bloch, S., Yarom, R. & Sulitzeanu, D. Degenerative changes in the A375 melanoma line induced by transforming growth factor beta 1. Immunol Lett. 33, 151–156 (1992).
Tsukada, Y., Tanaka, T., Miyazawa, K. & Kitamura, N. Involvement of down-regulation of Cdk2 activity in hepatocyte growth factor-induced cell cycle arrest at G1 in the human hepatocellular carcinoma cell line HepG2. J. Biochem. 136, 701–709 (2004).
Chen, C. R., Kang, Y. & Massagué, J. Defective repression of c-myc in breast cancer cells: A loss at the core of the transforming growth factor beta growth arrest program. Proc. Natl. Acad Sci. USA. 98, 992–999 (2001).
Termén, S., Tan, E. J., Heldin, C. H. & Moustakas, A. p53 regulates epithelial-mesenchymal transition induced by transforming growth factor β. J. Cell Physiol. 228, 801–813 (2013).
Miettinen, P. J., Ebner, R., Lopez, A. R. & Derynck, R. TGF-beta induced transdifferentiation of mammary epithelial cells to mesenchymal cells: involvement of type I receptors. J. Cell Biol. 127, 2021–2036 (1994).
Nishihara, A. et al. Role of p300, a transcriptional coactivator, in signalling of TGF-beta. Genes Cells 7, 613–623 (1998).
Itoh, S., Ericsson, J., Nishikawa, J., Heldin, C. H. & ten Dijke, P. The transcriptional co-activator P/CAF potentiates TGF-beta/Smad signaling. Nucleic Acids Res. 28, 4291–4298 (2000).
Cordenonsi, M. et al. Integration of TGF-beta and Ras/MAPK signaling through p53 phosphorylation. Science 315, 840–843 (2007).
Seoane, J., Le, H. V., Shen, L., Anderson, S. A. & Massagué, J. Integration of Smad and forkhead pathways in the control of neuroepithelial and glioblastoma cell proliferation. Cell 117, 211–223 (2004).
el-Deiry, W. S. et al. WAF1, a potential mediator of p53 tumor suppression. Cell 75, 817–825 (1993).
Ross, C. R., Brennan-Laun, S. E. & Wilson, G. M. Tristetraprolin: roles in cancer and senescence. Ageing Res Rev. 11, 473–484 (2012).
Kawata, M. et al. TGF-β-induced epithelial-mesenchymal transition of A549 lung adenocarcinoma cells is enhanced by pro-inflammatory cytokines derived from RAW264.7 macrophage cells. J. Biochem. 151, 205–216 (2012).
Kim, N. H. et al. A p53/miRNA-34 axis regulates Snail1-dependent cancer cell epithelial-mesenchymal transition. J. Cell. Biol. 195, 417–433 (2011).
Choi, Y. K. et al. Oberexpression of urokinase-type plasminogen activator in human gastric cancer cell line (AGS) induces tumorigenicity in severe combined immunodeficient mice. Jpn. J. Cancer Res. 93, 151–156 (2002).
Inoue, Y., Iemura, S., Natsume, T., Miyazawa, K. & Imamura, T. Suppression of p53 activity through the cooperative action of Ski and histone deacetylase SIRT1. J. Biol. Chem. 286, 6311–6320 (2011).
Inoue, Y. et al. Smad3 is acetylated by p300/CBP to regulate its transactivation activity. Oncogene 26, 500–508 (2007).
Keeton, M. R., Curriden, S. A., van Zonneveld, A. J. & Loskutoff, D. J. Identification of regulatory sequences in the type 1 plasminogen activator inhibitor gene responsive to transforming growth factor beta. J. Biol. Chem. 266, 23048–23052 (1991).
Gu, X., Coates, P. J., Laurell, G. & Nylander, K. p63 contributes to cell invasion and migration in squamous cell carcinoma of head and neck. Cancer Lett. 263, 26–34 (2008).
Dirac, A. M. & Bernards, R. Reversal of senescence in mouse fibroblasts through lentiviral suppression of p53. J. Biol. Chem. 278, 11731–11734 (2003).
Fujiwara, K. et al. Pivotal role of the cyclin-dependent kinase inhibitor p21WAF1/CIP1 in apoptosis and autophagy. J. Biol. Chem. 283, 388–397 (2008).
Miyajima, C., Inoue, Y. & Hayashi, H. Pseudokinase tribbles 1 (TRB1) negatively regulates tumor-suppressor activity of p53 through p53 deacetylation. Biol. Pharm. Bull. 38, 618–624 (2015).
Inoue, Y., Abe, K., Onozaki, K. & Hayashi, H. TGF-β decreases the stability of IL-18-induced IFN-γ mRNA through the expression of TGF-β-induced tristetraprolin in KG-1 cells. Biol. Pharm. Bull. 38, 536–544 (2015).
Kumar, D., Singla, S. K., Puri, V. & Puri, S. The restrained expression of NF-kB in renal tissue ameliorates folic acid induced acute kidney injury in mice. PLoS One 10, e115947 (2015).
Ezzi, S. A., Urushitani, M. & Julien, J. P. Wild-type superoxide dismutase acquires binding and toxic properties of ALS-linked mutant forms through oxidation. J. Neurochem. 102, 170–178 (2007).
Thuault, S. et al. Transforming growth factor-beta employs HMGA2 to elicit epithelial-mesenchymal transition. J. Cell Biol. 174, 175–183 (2006).
Matsumoto, A., Takeishi, S. & Nakayama, K. I. p57 regulates T-cell development and prevents lymphomagenesis by balancing p53 activity and pre-TCR signaling. Blood 123, 3429–3439 (2014).
Inoue, Y., Kitagawa, M. & Taya, Y. Phosphorylation of pRB at Ser612 by Chk1/2 leads to a complex between pRB and E2F-1 after DNA damage. EMBO J. 26, 2083–2093 (2007).
Koinuma, D. et al. Chromatin immunoprecipitation on microarray analysis of Smad2/3 binding sites reveals roles of ETS1 and TFAP2A in transforming growth factor beta signaling. Mol. Cell. Biol. 29, 172–186 (2009).
López-Díaz, F. J. et al. Coordinate transcriptional and translational repression of p53 by TGF-β1 impairs the stress response. Mol. Cell 50, 552–564 (2013).
Kurisaki, K. et al. Nuclear factor YY1 inhibits transforming growth factor beta- and bone morphogenetic protein-induced cell differentiation. Mol. Cell. Biol. 23, 4494–4510 (2003).
Acknowledgements
The authors are grateful to Drs Kohei Miyazono and Keiji Miyazawa for reagents. The authors thank members of the Hayashi laboratory for helpful discussions. This work was supported by a Grant-in-Aid for Scientific Research (C) (No. 24590085, 15K07936, 15K07937) from the Japan Society for the Promotion of Science (JSPS), a Grant-in-Aid for Young Scientists (B) (No. 24700983) from JSPS, and a Grant-in-Aid for Research in Nagoya City University. Y.I. was supported by the Mochida Memorial Foundation for Medical and Pharmaceutical Research, the Takeda Science Foundation, the Nakatomi Foundation, the Hori Sciences & Arts Foundation, the YOKOYAMA Foundation for Clinical Pharmacology (No. YRY1408), and the INAMORI Foundation.
Author information
Authors and Affiliations
Contributions
Y.K., Y.I. and H.H. designed the experiments. Y.K., Y.I., F.K., K.F., K.S., T.T. and Y.I. performed the experiments. Y.I. and H.H. supervised the study. Y.I. directed the study. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kawarada, Y., Inoue, Y., Kawasaki, F. et al. TGF-β induces p53/Smads complex formation in the PAI-1 promoter to activate transcription. Sci Rep 6, 35483 (2016). https://doi.org/10.1038/srep35483
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep35483
- Springer Nature Limited
This article is cited by
-
SET8 is a novel negative regulator of TGF-β signaling in a methylation-independent manner
Scientific Reports (2023)
-
Oncogenic RAS sensitizes cells to drug-induced replication stress via transcriptional silencing of P53
Oncogene (2022)
-
Emerging role of tumor suppressor p53 in acute and chronic kidney diseases
Cellular and Molecular Life Sciences (2022)
-
TRPM7 restrains plasmin activity and promotes transforming growth factor-β1 signaling in primary human lung fibroblasts
Archives of Toxicology (2022)
-
Periplocin and cardiac glycosides suppress the unfolded protein response
Scientific Reports (2021)