Abstract
Patients with multiple primary cancers (MPCs) are suspected to have a hereditary cancer syndrome. However, only a small proportion may be explained by mutations in high-penetrance genes. We investigate two unrelated MPC patients that met Hereditary Breast and Ovaria Cancer criteria, both presenting triple negative breast tumors and no mutations in BRCA1, BRCA2 and TP53 genes. Germline rearrangements on chromosome 7q, involving over 40 Mb of the same region, were found in both patients: one with mosaic loss (80% of cells) and the other with cnLOH (copy-neutral loss of heterozygosity) secondary to maternal allele duplication. Five children tested had no alterations on 7q. The patients shared 330 genes in common on 7q22.1-q34, including several tumor suppressor genes (TSGs) previously related to breast cancer risk and imprinted genes. The analysis of the triple negative BC from one patient revealed a mosaic gain of 7q translated for over-expressed cancer-related genes. The involvement of TSGs and imprinted genes, mapped on 7q, has the potential of being associated to MPC risk, as well as cancer progression. To our knowledge, this is the first description of patients with MPCs that harbor constitutive large alterations on 7q.
Similar content being viewed by others
Introduction
The incidence of cancer is continuously increasing, as is the number of cancer survivors1,2. Cancer patients have a higher risk of developing new malignancies when compared to the general population3. Data from the Surveillance, Epidemiology and End Results program estimated that subsequent primary cancers represent approximately 18% of all cancers in the USA4. The development of multiple primary cancers (MPCs) has been reported as being associated to the treatment received for the first cancer (chemotherapy and radiotherapy), personal lifestyle and genetic predisposition5.
Individuals who developed cancer at younger age, presented multiple primary tumors or reported several relatives with neoplasms are suspected of having a hereditary cancer predisposition syndrome6. Breast cancer (BC) falls within the tumor spectrum of several hereditary diseases, including Hereditary Breast and Ovarian Cancer syndrome (HBOC) and Li-Fraumeni syndrome (LFS)6. However, only a small proportion of familial BC cases can be explained by mutations in high-penetrance genes, such as BRCA1, BRCA2 and TP537.
Part of the missing heritability in BC may be explained by copy number variations (CNVs), which are chromosomal regions altered by gains or losses, or by copy-neutral loss of heterozygosity (cnLOH), defined as homozygous regions appearing as a result of inheritance of a pair of alleles from a single parent8,9. The contribution of rare germline CNVs to BC predisposition has been demonstrated in BRCA1/BRCA2 mutation-negative patients10,11,12. Moreover, an increased frequency of cnLOH in cases where no mutations are present in the mismatch repair genes suggests the involvement of unknown germline alterations in familial colorectal cancer risk13.
Deletions and cnLOH mapped on 7q have been widely described in both hematological malignancies; specifically myelodysplastic syndrome, acute myeloid leukemia (AML) and splenic marginal zone lymphoma14,15,16; and BC17,18. Furthermore, genomic deletions on chromosome 7q have also been associated with congenital defects, including developmental delay, learning difficulties, craniofacial dysmorphism and hypogenitalism19,20,21,22.
Herein, we report the molecular and clinical characterization of two unrelated MPC patients, both presenting triple negative BC, a positive family history of cancer, and without germline pathogenic mutations in BRCA1, BRCA2 and TP53 genes, showing large genomic rearrangements mapped on 7q.
Results
Patient 1 and relatives
The whole genomic analysis performed in the lymphocytic DNA from Patient 1 revealed a 43 Mb germline mosaic loss (80% of cells) of chromosome 7q22.1-q34 (Fig. 1) and a rare loss of 9q22.31 (Supplementary Table S1). Two children were evaluated for genomic alterations to assess the presence of 7q rearrangements. Her son inherited the rare deletion of 9q, while her daughter had only common CNVs. None of them presented any alteration of chromosome 7q (data not shown).
All alterations were confirmed by non-polymorphic probes (Log2 Ratio and smooth signal) and SNP probes (allele peaks). In the breast cancer tissue of Patient 2, an additional gain at a different region of chromosome 7q was detected. Moreover, almost all of the cnLOH region presented a mosaic gain, particularly at the 7q32-q34 region.
Patient 2 and relatives
A large cnLOH (49 Mb) of 7q22.1-q36.1 was detected in the lymphocytic DNA of Patient 2 (Fig. 1). The region covered by the large mosaic loss of Patient 1 was entirely contained within the region encompassed by the cnLOH of Patient 2, both sharing 330 genes. An additional 76 genes were also mapped exclusively in the cnLOH region (Supplementary Table S1). Moreover, three other rare alterations were identified in Patient 2: loss of 8q11.21, cnLOH of 19p13.11-p13.2 and loss of Xq25 (Supplementary Table S1). Of them, losses of 8q11.21 and Xq25 were inherited from her mother. Among the three children tested for genomic alterations, the son A inherited the rare loss of 8q11.21 from Patient 2 (Supplementary Table S1). No alteration mapped in chromosome 7q was identified in all relatives tested.
The genotype analysis using the SNPs contained in the 7q cnLOH region of Patient 2 revealed its maternal origin. Virtually all homozygous nucleotides present in Patient 2, mapped to 7q22.1-q36.1, were identified in her mother, either homozygous or heterozygous (Supplementary Table S2).
The triple negative BC tissue of the Patient 2 presented a large number of alterations (100) in almost all chromosomes, including large CNVs and cnLOH regions in mosaicism (Supplementary Fig. S1 and Table S3). The germline 7q22.1-q36.1 cnLOH was maintained in the tumor tissue. However, a large percentage of this region also exhibited a gain in mosaicism in approximately 50% of cells (Fig. 1). Specifically, a region close to 15 Mb (7q32.1-q34) presented two copy gains in mosaicism.
The gene expression analysis performed in the BC tissue showed 96 over- and 52 down-regulated genes among the 406 genes contained in the 7q cnLOH region (Supplementary Table S4). At least 62 of the 148 differentially expressed genes were cancer-related according to the Candidate Cancer Gene Database (CCGD, http://ccgd-starrlab.oit.umn.edu/about.php)23, and the Network of Cancer Genes 5.0 (NCG, http://ncg.kcl.ac.uk/query.php)24. Among the 312 genes mapped in this region with mosaic gain, 81 were up-regulated and from these, 34 genes were cancer-related (Supplementary Table S4).
The kinship between the Patients 1 and 2 was verified using the Mendelian Error Check tool (ChAS software v3.1), which considers the genotypable SNPs (749,157 probes) found in the CytoScan HD platform. The analysis showed 5.8% of errors (of all SNPs) and role validity equals zero, thus confirming their unrelatedness.
Discussion
The occurrence of MPCs in one individual may be due to environmental factors, aging, germline mutations or simply by chance5,6. We reported two unrelated patients with MPCs and family history of malignant neoplasms, being the triple negative BC the only tumor type common to both individuals. Of note, the emergence of three hematological malignancies (two lymphomas and leukemia) in Patient 1 occurred following treatment with radiotherapy and chemotherapy. The role of radio- and chemotherapy in subsequent malignant neoplasms is well recognized5. Myeloid neoplasms are among the most common cancers related to chemotherapy5.
Previous studies have reported a higher risk of lymphoma development following BC, as highlighted by Patient 125, or the development of BC after melanoma, as observed in Patient 226. However, the presence of multiple neoplasms, cancer at an early age (Patient 2) and a history of malignancy in several relatives strongly implies a hereditary factor as a cause of the cancer6. Indeed, both patients met the clinical criteria for HBOC27, having been tested for BRCA1, BRCA2 and TP53 mutations with negative results. A recent study evaluated 212 individuals with MPCs and identified a pathogenic variant in known cancer predisposition genes in less than 25% of the patients28.
In an effort to uncover the genetic basis of cancer in our patients, we performed molecular cytogenetic analyses. Large genomic rearrangements (over 40 Mb) on the long arm of chromosome 7 in both patients, starting at cytoband 7q22.1: a mosaic loss (80% of cells) in Patient 1 and a slightly larger DNA region with cnLOH in Patient 2. In a cohort of 100 health Brazilian individuals and 68 HBOC patients negative for mutations in BRCA1 and BRCA2, Krepischi et al.29 found only a limited number of common CNVs (<500 Kb) mapped in 7q. To our knowledge, there are no previous reports in literature of patients with cancer and constitutive alterations as large as those described here.
Interestingly, large germline deletions on 7q ranging from 10 Mb to 30 Mb, most of them entirely contained within the deletion found in mosaicism in Patient 1, have been reported in individuals with a significant spectrum of clinical features19,20,21,22. Although Patient 1 presented haploinsufficiency of various genes related to congenital anomalies (e.g. learning difficulties, speech delay, craniofacial alterations, cardiac defects and growth retardation), it seems that the percentage of normal cells (20%) on the 7q region was sufficient to avoid the manifestation of any malformation. Moreover, tissues derived from embryonic germ layers other than the mesoderm (blood lymphocytes) may have different levels of mosaicism, which may help to explain the absence of malformations in this patient. Recently, we reported a case with multiple tumors and deletion of one X chromosome in mosaicism, which did not present the clinical phenotype of Turner syndrome30.
Gains and losses involving genomic regions containing oncogenes or tumor suppressor genes (TSGs) may lead to cancer development8. Rare copy number alterations have been implicated in BC susceptibility in HBOC patients negative for mutations in BRCA1 and BRCA2 genes10,11,12. In addition, cnLOH may also contribute to cancer risk since they not occur randomly across the genome, but are associated with mutations in key cancer-related genes9,31. Recently, an increased level of cnLOHs involving 7q was reported in familial colorectal patients negative for mismatch repair genes compared to sporadic cases13.
The triple negative BC sample from Patient 2 was evaluated by molecular cytogenetic and transcriptomic analysis to compare with the germline alterations and to verify if the presence of 7q alterations could modify the dosage of genes mapped in this region. An extremely large number of genomic alterations, mainly gains and losses in mosaicism were found, thereby highlighting intra-tumor heterogeneity, and clonal evolution. In addition to the cnLOH of 7q found in lymphocytic DNA, two new cnLOHs and 10 mosaic cnLOHs were observed in the tumor, therefore suggesting the importance of cnLOHs in BC progression, as described in colorectal cancer32. Interestingly, cnLOH regions were detected more frequently in triple negative BC samples than in estrogen/progesterone/HER2 positive cases33. Moreover, there are various common fragile sites in 7q, which are regions prone to genomic rearrangement and have been previously associated with cancer development34,35.
The BC sample transcriptome analysis revealed 148 genes differentially expressed mapped in the same region of 7q cnLOH, most of them over-expressed (96 genes) and cancer-related (62 genes). These results indicated that large genomic rearrangements on 7q alter the gene dosage and give additional support for the functional relevance of this alteration found in both patients of our study.
The detected 7q deletion and cnLOH shared a large number of genes (330), including putative TSGs previously associated with BC risk (e.g. CAV1, MET and TES)18,36. Down-regulation of CAV1 gene was observed in the BC tissue of Patient 2. The presence of TSGs associated with hematological cancers, such as DOCK4, LUC7L2 and CUX1 were also observed16. Interestingly, Patient 1 developed two primary lymphomas and AML within 9 years. Recently, mutations in CUX1 have been identified in 20 different carcinoma types, making this gene an important candidate for tumorigenesis37. In addition, the oncogene BRAF; previously reported as involved in melanoma, papillary thyroid carcinoma and colorectal cancer38; was included in the altered genomic region mapped to 7q and may be associated with the emergence of melanoma three years before the BC in Patient 2.
Besides its involvement in unmasking homozygous cancer-related mutations, cnLOHs may also disrupt genomic imprinting, an epigenetic process that leads to monoalleic expression depending on parental origin, resulting in two active or repressed alleles9,39. The deregulation of imprinted genes has been described in malignant neoplasms40. According to the Geneimprint database (http://www.geneimprint.com/site/genes-byspecies), five imprinted genes are mapped to 7q32: COPG2IT1, MEST, CPA4, MESTIT1 and KLF14, of which, a loss of imprinting (biallelic expression) of MEST has been associated with BC development40,41,42. Interestingly, MEST was apparently silenced in the blood sample of Patient 2 (maternal allele dissomy and maternal imprinting) and the BC of Patient 2 presented MEST over-expression, suggesting the loss of imprinting. Similar mechanism has been described for IGF2 (maternal imprinting) in colon cancer cells43.
Patient 1 and her son also presented a rare deletion of 9q22.31 encompassing the genes CENPP and OGN. The absence of OGN protein has been associated to colorectal carcinogenesis44, yet no alterations within these genes have been described as associated with cancer development risk. Three additional rare alterations were detected in Patient 2: two losses covering no genes on chromosomes 8q11.21, also present in her mother and one of her sons, and Xp25; as well as a cnLOH of chromosome 19p13.11-p13.2 encompassing 161 genes. Copy number aberrations on 19p13.2, including PKN1, SMARCA4 and LYL1 genes have been associated with triple negative BC and AM45,46.
Potential weakness of this study was the limited access of other relatives with cancer. In addition, neither histologically normal samples to evaluate the mosaicism nor other tumor samples from the probands were available. Nevertheless, the germline rearrangements detected in both probands, the cnLOH and deletion, are very large and encompass several TSGs previously associated with neoplasms, including BC, making this region an important candidate for MPCs risk.
In summary, we identified two large genomic rearrangements of chromosome 7q (mosaic loss and cnLOH) in two unrelated patients with MPCs, both presenting triple negative breast neoplasms and family history of cancer. The genomic analysis of the BC sample from Patient 2 indicated that the 7q region is prone to genomic instability and may include genes related to cancer growth and progression. The whole gene expression analysis of this BC revealed deregulated genes in 7q cnLOH region, including several cancer-related genes, giving additional support that genomic alterations of 7q have functional role in the tumor risk development.
Methods
Patients
The two probands (Patient 1 and Patient 2) included in this study were selected from a cohort of 27 MPC patients negative for pathogenic mutations in the TP53, BRCA1 and BRCA2 genes. Clinicopathological features of each individual were re-evaluated to exclude recurrence or metastases, confirming the diagnosis of multiple primary tumors. Sixteen of the 27 patients fulfilled criteria for hereditary cancer syndromes, including LFS (three cases), HBOC (five cases) and LFS/HBOC simultaneously (two cases)27,47. Female BC was the most common tumor type found in this set of MPC patients (N = 10) (Supplementary Table S5).
Additionally, five children (two of Patient 1 and three of Patient 2), the mother and the breast cancer tissue of Patient 2 were evaluated. This study was approved by the Human Research Ethics Committee of A.C. Camargo Cancer Center. Sao Paulo, Brazil (CEP No. 1726112). Written informed consent was obtained from all participants prior to sample collection. All experiments were carried out in accordance with the approved guidelines.
Patient 1 was first diagnosed with triple-negative ductal carcinoma in situ on the right breast at 58 years of age. She underwent a segmental mastectomy, axillary lymphadenectomy and radiotherapy. At age 61, she presented with lymphocytic lymphoma, for which, she received chemotherapy and underwent axillary dissection. Orbital non-Hodgkin lymphoma of the left eye was diagnosed at 63 years of age, which was enucleated and treated with chemotherapy. At age 65, a squamous cell carcinoma of the lower lip was diagnosed and completely resected. Furthermore, the patient also developed chronic myeloid leukemia, dying at the age of 70. The patient met the criteria for HBOC (Fig. 2A)27. Multiple relatives from both the paternal and maternal sides of the family had been diagnosed with different neoplasms. Both of her children (one son and one daughter with 36 and 44 years age, respectively) have no cancer diagnosed in the last follow-up (November 2016).
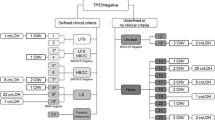
Pedigrees of Patient 1 (A) and Patient 2 (B). The patients fulfilled criteria for HBOC and were negative for germline mutations in the BRCA1, BRCA2 and TP53 genes. The black and grey arrows indicate the probands and the relatives submitted to molecular karyotyping analysis, respectively. The numbers above or below the symbols of individuals with cancer represent age at diagnosis. SCC: Squamous cell carcinoma.
Patient 2 developed in situ melanoma of the left foot at 42 years of age, and was treated by surgical resection. Three years later (aged 45) she was diagnosed with invasive ductal carcinoma of the left breast, characterized as pT2N1M0, triple-negative and with p53 positivity in 90% of the tumor cells. The patient was treated with modified radical mastectomy and adjuvant chemo-radiotherapy. The patient’s sister and paternal aunt both died from breast cancer at ages 42 and 43, respectively. The family also fulfilled the criteria for HBOC (Fig. 2B)27. To date, none of her three children (two sons and one daughter) have developed any neoplasms (aged 32, 33 and 37 years, respectively, November 2016).
Molecular cytogenetic analysis
Genomic DNA was extracted from the peripheral blood of the probands, the mother of Patient 2, and five children of both probands following the standard protocol of the Gentra Puregene Blood Kit (Qiagen, Valencia, Ca, USA). Genomic DNA of the breast tumor tissue of Patient 2 was isolated using the phenol-chloroform method. Molecular karyotyping was carried out for all samples using the CytoScan HD Array (Affymetrix, Santa Clara, CA, USA) following the manufacturer’s instructions. The detection of CNVs and cnLOHs was performed using the Chromosome Analysis Suite (ChAS) software v.3.1 (Affymetrix). The criteria used for analysis considered at least 50 markers for gains/mosaic gains, 25 for losses/mosaic losses and cnLOHs with a minimum of 5 Mb. The alterations were visually confirmed, with poor quality data excluded. The CNVs detected were compared to the Affymetrix Database of Variants (aDGV), consisting of 2,421 phenotypically healthy individuals evaluated by Cytoscan HD, as well as to the Database of Genomic Variants (DGV, http://dgv.tcag.ca/dgv/app/home, updated in May 2016). CNVs with a frequency <0.5% in the aDGV and <0.05% in the DGV were considered rare. The microarray data are accessible at NCBI’s Gene Expression Omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo/) database, accession number GSE 77138.
Transcriptome analysis
Total RNA was isolated from the BC tissue of Patient 2 using the RNeasy Mini Kit (Qiagen, Valencia, Ca, USA), according to the manufacturer’s instructions. The Human Universal Reference RNA (Stratagene, Santa Clara, USA) was used as a reference sample for comparison with the tumor tisue. All samples were analyzed in duplicate. Whole genome gene expression analysis was assessed using the Human Transcriptome Array (HTA) 2.0 platform (Affymetrix) following the manufacturer’s recommendations. Microarray data were normalized with the Expression Console software (Affymetrix). Differentially expressed genes were obtained with the Transcriptome Analysis Console (TAC) software v3.1 (Affymetrix), considering the ANOVA test (P < 0.05) and fold change (FC) >1.5 or <−1.5. We limited our analysis to genes mapped to cnLOH of 7q detected in Patient 2. The gene expression data is also available in the GEO database (GSE 77138).
Additional Information
How to cite this article: Villacis, R. A. R. et al. Germline large genomic alterations on 7q in patients with multiple primary cancers. Sci. Rep. 7, 41677; doi: 10.1038/srep41677 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Torre, L. A. et al. Global cancer statistics, 2012. CA Cancer J Clin 65, 87108 (2015).
DeSantis, C. E. et al. Cancer treatment and survivorship statistics, 2014. CA Cancer J Clin 64, 252–271 (2014).
Utada, M., Ohno, Y., Hori, M. & Soda, M. Incidence of multiple primary cancers and interval between first and second primary cancers. Cancer Sci 105, 890–896 (2014).
Altekruse, S. F. et al.–2007. National Cancer Institute. Available at: http://seer.cancer.gov/csr/1975_2007/ (Accessed in March 2016) (2010).
Travis, L. B., Demark Wahnefried, W., Allan, J. M., Wood, M. E. & Ng, A. K. Aetiology, genetics and prevention of secondary neoplasms in adult cancer survivors. Nat Rev Clin Oncol 10, 289–301 (2013).
Cybulski, C., Nazarali, S. & Narod, S. A. Multiple primary cancers as a guide to heritability. Int J Cancer 135, 1756–1763 (2014).
Rich, T. A., Woodson, A. H., Litton, J. & Arun, B. Hereditary breast cancer syndromes and genetic testing. J Surg Oncol 111, 66–80 (2015).
Kuiper, R. P., Ligtenberg, M. J., Hoogerbrugge, N. & Geurts van Kessel, A. Germline copy number variation and cancer risk. Curr Opin Genet Dev 20, 282–289 (2010).
Lapunzina, P. & Monk, D. The consequences of uniparental disomy and copy number neutral loss-of-heterozygosity during human development and cancer. Biol Cell 103, 303–317 (2011).
Pylkäs, K. et al. Rare copy number variants observed in hereditary breast cancer cases disrupt genes in estrogen signaling and TP53 tumor suppression network. PLoS Genet 8, e1002734 (2012).
Kuusisto, K. M. et al. Copy number variation analysis in familial BRCA1/2-negative Finnish breast and ovarian cancer. PLoS One 8, e71802 (2013).
Masson, A. L. et al. Expanding the genetic basis of copy number variation in familial breast cancer. Hered Cancer Clin Pract 12, 15 (2014).
Middeldorp, A. et al. Increased frequency of 20q gain and copy-neutral loss of heterozygosity in mismatch repair proficient familial colorectal carcinomas. Int J Cancer 130, 837–846 (2012).
O’Keefe, C., McDevitt, M. A. & Maciejewski, J. P. Copy neutral loss of heterozygosity: a novel chromosomal lesion in myeloid malignancies. Blood 115, 2731–2739 (2010).
Watkins, A. J. et al. Splenic marginal zone lymphoma: characterization of 7q deletion and its value in diagnosis. J Pathol 220, 461–474 (2010).
Honda, H., Nagamachi, A. & Inaba, T. -7/7q- syndrome in myeloid-lineage hematopoietic malignancies: attempts to understand this complex disease entity. Oncogene 34, 2413–2425 (2015).
Zeng, W. R. et al. Refined mapping of the region of loss of heterozygosity on the long arm of chromosome 7 in human breast cancer defines the location of a second tumor suppressor gene at 7q22 in the region of the CUTL1 gene. Oncogene 18, 2015–2021 (1999).
Miller, B. J., Wang, D., Krahe, R. & Wright, F. A. Pooled analysis of loss of heterozygosity in breast cancer: a genome scan provides comparative evidence for multiple tumor suppressors and identifies novel candidate regions. Am J Hum Genet 73, 748–767 (2003).
Feuk, L. et al. Absence of a paternally inherited FOXP2 gene in developmental verbal dyspraxia. Am J Hum Genet 79, 965–972 (2006).
Chen, C. P. et al. Prenatal diagnosis and molecular cytogenetic characterization of a de novo interstitial deletion of 7q (7q22.1 → q31.1). Gene 521, 311–315 (2013).
Martínez-Jacobo, L. et al. Delineation of a de novo 7q21.3q31.1 Deletion by CGH-SNP arrays in a girl with multiple congenital anomalies including severe glaucoma. Mol Syndromol 4, 285–291 (2013).
Del Refugio Rivera-Veja, M. et al. A novel 23.1 Mb interstitial deletion involving 7q22.3q32.1 in a girl with short stature, motor delay, and craniofacial dysmorphism. Cytogenet Genome Res 145, 1–5 (2015).
Abbott, K., Nyre, E., Abrahante, J., Ho, Y., Vogel, R. & Starr, T. The Candidate Cancer Gene Database: a database of cancer driver genes from forward genetic screens in mice. Nucleic Acids Res 43, D844–D848 (2015).
An, O., Dall’Olio, G. M., Mourikis, T. P. & Ciccarelli, F. D. NCG 5.0: updates of a manually curated repository of cancer genes and associated properties from cancer mutational screenings. Nucleic Acids Res 44, D992–D999 (2016).
Ricceri, F. et al. Risk of second primary malignancies in women with breast cancer: Results from the European prospective investigation into cancer and nutrition (EPIC). Int J Cancer 137, 940–948 (2015).
Robsahm, T. E., Karagas, M. R., Rees, J. R. & Syse, A. New malignancies after squamous cell carcinoma and melanomas: a population-based study from Norway. BMC Cancer 14, 210 (2014).
NCCN - National Comprehensive Cancer Network. Clinical Practice Guidelines in Oncology. Genetic/Familial High-Risk Assessment: BRCA-Related Breast and/or Ovarian Cancer Syndrome (BRCA-1) Version 1. Available at: http://www.nccn.org/professionals/physician_gls/pdf/genetics_screening.pdf (Accessed in November 2016) (2017).
Whitworth, J. et al. A clinical and genetic analysis of multiple primary cancer referrals to genetics services. Eur J Hum Genet 23, 581–587 (2015).
Krepischi, A. C. et al. Germline DNA copy nymber variation in familial and early-onset breast cancer. Breast Cancer Res 14, R24 (2012).
Basso, T. R. et al. Genomic profile of a Li-Fraumeni-like syndrome patient with a 45,X/46,XX karyotype, presenting neither mutations in TP53 nor clinical stigmata of Turner syndrome. Cancer Genet 208, 341–344 (2015).
Torabi, K. et al. Patterns of somatic uniparental disomy identify novel tumor suppressor genes in colorectal cancer. Carcinogenesis 36, 1103–1110 (2015).
Melcher, R. et al. LOH and copy neutral LOH (cnLOH) act as alternative mechanism in sporadic colorectal cancers with chromosomal and microsatellite instability. Carcinogenesis 32, 636–642 (2011).
Tuna, M., Smid, M., Zhu, D., Martens, J. W. & Amos, C. I. Association between acquired uniparental disomy and homozygous mutations and HER2/ER/PR status in breast cancer. PLoS One 5, e15094 (2010).
Lukusa, T. & Fryns, J. P. Human chromosome fragility. Biochim Biophys Acta 1779, 3–16 (2008).
Dillon, L. W., Burrow, A. A. & Wang, Y. H. DNA instability at chromosomal fragile sites in cancer. Curr Genomics 11, 326–337 (2010).
Zhu, J. et al. Testin is a tumor suppressor and prognostic marker in breast cancer. Cancer Sci 103, 2092–2101 (2012).
Wong, C. C. et al. Inactivating CUX1 mutations promote tumorigenesis. Nat Genet 46, 33–38 (2014).
Pakneshan, S., Salajegheh, A., Smith, R. A. & Lam, A. K. Clinicopathological relevance of BRAF mutations in human cancer. Pathology 45, 346–356 (2013).
Uribe-Lewis, S., Woodfine, K., Stojic, L. & Murrell, A. Molecular mechanisms of genomic imprinting and clinical implications for cancer. Expert Rev Mol Med 13, e2 (2011).
Kim, J., Bretz, C. L. & Lee, S. Epigenetic instability of imprinted genes in human cancers. Nucleic Acids Res 43, 10689–10699 (2015).
Pedersen, I. S. et al. Frequent loss of imprinting of PEG/MEST in invasive breast cancer. Cancer Res 59, 5449–5451 (1999).
Pedersen, I. S. et al. Promoter switch: a novel mechanism causing biallelic PEG1/MEST expression in invasive breast cancer. Hum Mol Genet 11, 1449–1453 (2002).
Wang, H. et al. Restoration of IGF2 imprinting by polycomb repressive complex 2 docking factor SUZ12 in colon cancer cells. Exp Cell Res 338, 214–221 (2015).
Wang, Y. et al. Differential expression of mimecan and thioredoxin domain-containing protein 5 in colorectal adenoma and cancer: a proteomic study. Exp Biol Med (Maywood) 232, 1152–1159 (2007).
Turner, N. et al. Integrative molecular profiling of triple negative breast cancers identifies amplicon drivers and potential therapeutic targets. Oncogene 29, 2013–2023 (2010).
Dirse, V. et al. A population-based single nucleotide polymorphism array analysis of genomic aberrations in younger adult acute lymphoblastic leukemia patients. Genes Chromosomes Cancer 54, 326–333 (2015).
Bougeard, G. et al. Revisiting Li-Fraumeni Syndrome From TP53 Mutation Carriers. J Clin Oncol 33, 2345–2352 (2015).
Acknowledgements
We thank the patients who participated in this study and the Nucleic Acid Bank of A.C. Camargo Cancer Center (São Paulo, Brazil). This study was supported by grants from the National Institute of Science and Technology in Oncogenomics (INCITO - São Paulo Research Foundation, FAPESP 2008/57887-9, and National Council for Scientific and Technological Development, CNPq 573589/08-9).
Author information
Authors and Affiliations
Contributions
S.R.R. and M.I.A. conceived and designed the experiments. R.A.R.V., T.R.B. and L.M.C. performed the experiments and analyzed the data. S.R.R., A.F.N. and M.I.A. contributed with reagents/materials/analysis tools; S.R.R., R.A.R.V. and L.M.C. wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Villacis, R., Basso, T., Canto, L. et al. Germline large genomic alterations on 7q in patients with multiple primary cancers. Sci Rep 7, 41677 (2017). https://doi.org/10.1038/srep41677
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep41677
- Springer Nature Limited