Abstract
Psychotic disorders affect ~3% of the general population and are among the most severe forms of mental diseases. In early stages of psychosis, clinical aspects may be difficult to distinguish from one another. Undifferentiated psychopathology at the first-episode of psychosis (FEP) highlights the need for biomarkers that can improve and refine differential diagnosis. We investigated gene expression differences between patients with FEP–schizophrenia spectrum (SCZ; N=53) or FEP–Mania (BD; N=16) and healthy controls (N=73). We also verified whether gene expression was correlated to severity of psychotic, manic, depressive symptoms and/or functional impairment. All participants were antipsychotic-naive. After the psychiatric interview, blood samples were collected and the expression of 12 psychotic-disorder-related genes was evaluated by quantitative PCR. AKT1 and DICER1 expression levels were higher in BD patients compared with that in SCZ patients and healthy controls, suggesting that expression of these genes is associated more specifically to manic features. Furthermore, MBP and NDEL1 expression levels were higher in SCZ and BD patients than in healthy controls, indicating that these genes are psychosis related (independent of diagnosis). No correlation was found between gene expression and severity of symptoms or functional impairment. Our findings suggest that genes related to neurodevelopment are altered in psychotic disorders, and some might support the differential diagnosis between schizophrenia and bipolar disorder, with a potential impact on the treatment of these disorders.
Similar content being viewed by others
Introduction
Psychotic disorders, including schizophrenia, bipolar disorder and schizoaffective disorder, affect ~3% of the general population1, 2 and represent some of the most severe mental diseases. Characteristic symptoms include hallucinations, delusional beliefs, severe mood variations and cognitive impairment. However, during the early stages of psychosis, the clinical aspects may be difficult to distinguish from one another. The first-episode psychosis (FEP) is a critical period given that brain abnormalities and cognitive deficits are already present and progress faster, and more aggressively in the first years of the disorder,3 whereas the patients are not affected yet by factors related to disease progression, that is, duration of illness and exposure to antipsychotics.4, 5
More than a century has passed since Kraepelin first proposed the distinction between dementia praecox (schizophrenia) and manic-depressive insanity (bipolar disorder),6 but as both disorders may share the same psychotic symptoms, differentiating schizophrenia spectrum disorders from bipolar disorder is still a challenge. Therefore, different lines of research aim to identify biomarkers capable of distinguishing these disorders, including studies based on gene expression in peripheral tissues.7 On the other hand, some genes, including microRNAs, show a concordant expression and association for both schizophrenia and bipolar disorder in blood8 and also in brain tissues,9, 10 showing a possible common pathophysiological mechanism between these disorders, beyond the diagnostic boundaries. Moreover, previous studies revealed an effect of antipsychotics on gene expression.11, 12, 13 Therefore, assessing gene expression in early stages, such as FEP, is crucial, particularly before the administration of antipsychotics, but this is only feasible in peripheral tissues.
The majority of studies have focused on schizophrenia-spectrum psychosis, suggesting alterations in genes related to myelination, neurodevelopment and AKT pathway,14 although affective psychoses studies are under-represented in the literature, particularly early-stage affective psychoses. Very few studies on gene expression of antipsychotic-naive bipolar disorder patients have been conducted,15, 16, 17, 18 reporting alterations in inflammatory genes, such as TNF,15 and in genes of AKT1/mTOR pathway.18
Our objectives are to investigate differences in the messenger RNA (mRNA) levels of 12 genes among individuals with FEP of schizophrenia-spectrum disorder (SCZ), FEP with mania (BD) and healthy controls. We also want to verify whether gene expression is correlated to clinical features, including functional impairment and severity of psychotic, manic, and depressive symptoms. Particularly, we compared SCZ with BD to identify diagnostic specificity (genes related to manic symptoms), and FEP (both SCZ and BD) and healthy controls to find genes related to psychosis itself as a broad syndrome.
To our knowledge, this is the first study that compares gene expression between antipsychotic-naive FEP of schizophrenia spectrum disorder and FEP with mania. The study aims are to differentiate BD and SCZ, improving early diagnosis and adequate intervention.
We selected the genes based on their biological role in psychotic disorders. As a second criterion, these genes should be expressed in whole blood (according to information available in http://www.genecards.org). We selected genes related to dopamine neurotransmission (COMT), inflammation and the immune system (TNF), neurodevelopment (DISC1, PAFAH1B1 and NDEL1), myelination (MBP), cell signaling (AKT1), the microRNA machinery (DGCR8, DICER1 and DROSHA), protein degradation (UFD1L) and adhesion (DGCR2). Some genes were selected mainly because of their location in the 22q11.2 region (COMT, DGCR2, DGCR8 and UFD1L), as its deletion is one of the strongest known genetic risk factors for psychotic disorders.19 We previously analyzed the same genes in other studies comparing FEP (excluding individuals with bipolar disorder diagnosis) with ultra-high risk individuals and controls, with positive findings.11, 12, 20, 21, 22, 23 Here a comparison between FEP of schizophrenia spectrum disorder (SCZ) and FEP with mania (BD) is presented, as well as between BD and healthy controls.
Materials and methods
Study population
The Research Ethics Committee of UNIFESP approved the research protocol, and all participants and family members provided written informed consent prior to enrollment in the study (CEP 0603/10).
Antipsychotic-naive FEP patients (N =69) were recruited from a psychiatric emergency unit in São Paulo, Brazil. The diagnosis of a psychotic disorder was established by trained psychiatrists based on the criteria of the Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition (DSM-IV), using the Structured Clinical Interview of the DSM-IV. Patients were assessed at baseline and followed up for at least 2 months. At the end of follow-up, patients with a schizophrenia or schizophreniform disorder diagnosis were classified as FEP of schizophrenia spectrum disorder (SCZ, N =53) and patients who met bipolar disorder (with psychotic symptoms) diagnostic criteria were classified as FEP with mania (BD, N =16).
Inclusion criteria were age between 16 and 40 years and no previous history of antipsychotic medication. Prior treatment with benzodiazepines was allowed. Patients with psychotic episodes due to a general medical condition, substance-induced psychotic disorder, intellectual disability or psychotic episodes that were associated with major depressive disorder were excluded.
The patients were assessed by: (a) PANSS (Positive and Negative Syndrome Scale);24 (b) CGI (Clinical Global Impression Scale);25 (c) GAF (Global Assessment of Functioning Scale), (d) CDSS (Calgary Depression Scale for Schizophrenia);26 and (e) YMRS (Young Mania Rating Scale). PANSS dimensions were derived from a previous study in a Brazilian population.27
The healthy control group (N =73) comprised age- and gender-matched volunteers with no current or previous psychiatric diagnoses or first-degree family history of psychotic disorders.
Peripheral blood samples were collected prior to the administration of antipsychotics (for patients) or after psychiatric interview (for controls).
Analysis of transcript levels of selected genes
A total of 5 ml of whole blood was collected in PAXgene RNA tubes (PreAnalytix, Hombrechtikon, Switzerland). RNA was isolated using a PAXgene Blood RNA kit (Qiagen, Germantown, MD, USA) according to the manufacturer’s instructions. RNA integrity was determined through electrophoresis on a 1.0% agarose gel, and the quality and quantity of the RNA samples were determined using a NanoDrop ND-1000 spectrophotometer (Nanodrop, Wilmington, DE, USA).
Approximately 400 ng of each RNA sample was reverse-transcribed using a High-Capacity cDNA Reverse Transcription Kit (Life Technologies, Foster City, CA, USA). Then, 20 to 100 ng of complementary DNA (cDNA) was diluted in H2O, mixed with TaqMan Universal PCR Master Mix (Life Technologies), and loaded on Taqman Low-Density Array (TLDA) microfluidic cards (Life Technologies). Probes and primers of 12 target genes, two housekeeping genes (ACTB and GAPDH), and one positive control for the reaction (18S) were preloaded in the 384 wells (in triplicates) of each TLDA card (Life Technologies). Assays and the exons and transcripts that they recognize are described in Supplementary Table 1. The experiments were performed in accordance with the manufacturer's instructions using the ViiA 7 Real-Time PCR System (Life Technologies).
Genes were selected based on their previously reported association with psychotic disorders and their expression in blood (http://www.genecards.org/).
Gene expression was quantified using the relative threshold method (Crt) with the geometric mean (GM) between ACTB and GAPDH as the endogenous control. Delta cycle relative threshold values (ΔCrt=Crttarget gene−CrtGM) were calculated for each sample and 2-ΔCrt values were included in the PASW Statistics (version 18.0, SPSS, Chicago, IL, USA) data set.
Statistical analysis
Sample size was chosen using G*Power 3.1.6 software (Heinrich-Heine-Universität Düsseldorf, Düsseldorf, Germany) and considering an analysis of variance test, with medium effect size f=0.30, α=0.05; power=0.80 and 3 groups. Two-sided tests were used for statistical analyses. For continuous variables, normality and homogeneity was checked with Shapiro–Wilk and Levene tests and log-transformed as needed. Gender and age differences between the SCZ, BD and control group were verified using the χ2-test and analysis of variance test, respectively. Differences in severity of symptoms and functional impairment (CDSS, CGI, GAF and YMRS total scores and PANSS scores) were compared using Student’s t-test.
2-ΔCrt values were compared among SCZ, BD and control groups using general linear model (GLM), with gender as a fixed factor if necessary and with Bonferroni correction for multiple comparisons (12, considering the number of genes assessed). Post hoc comparisons were carried out using the Bonferroni test. In addition, for genes that were differentially expressed, we verified the correlation between the 2-ΔCrt values and the clinical aspects using linear regression and inserting group (SCZ or BD) as an independent variable. In order to evaluate the predictive power of these genes, receiver-operating characteristic curves were obtained, and the respective statistical significance were assessed by χ2-test on the area under the curve (AUC). The significance level was set at 5%.
Results
The clinical and demographic characteristics of the participants are presented in Table 1.
SCZ patients exhibited higher scores on the negative dimension of the PANSS, and lower scores on the excitement compound of the PANSS and YMRS, than BD patients (Table 1). In addition, BD patients presented with poorer global function, as indicated by GAF (Table 1), compared with SCZ patients. No significant age or gender differences were observed among the different groups.
Analysis of transcript levels of selected genes among antipsychotic-naive SCZ, BD patients and healthy controls
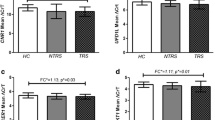
Gene expression results comparing the three groups (SCZ, BD and healthy controls) are reported in Table 2. Four genes were found to be differentially expressed among the groups after Bonferroni correction for 12 comparisons. Of them, two genes (MBP and NDEL1) exhibited higher expression levels in antipsychotic-naive patients (both SCZ and BD) than in healthy controls (Table 2). Moreover, the expression of two other genes, AKT1 and DICER, was higher in BD than in both SCZ patients and controls (Table 2).
The expression of these four genes was not correlated with prior benzodiazepine administration (P>0.05) or age (P>0.05).
We analyzed the receiver-operating characteristic curve and we found the same results: AKT1 and DICER1 2-▵Crt values differentiated BD from SCZ patients (AKT1 AUC=0.768, P=0.001; DICER1 AUC=0.812, P=1.706 × 10-4; Supplementary Figure 1) and healthy controls (AKT1 AUC=0.798, P=2.129 × 10-4; DICER1 AUC=0.865, P=5.525 × 10-6; Supplementary Figure 2), but not SCZ from healthy controls (AKT1 AUC=0.505, P=0.927; DICER1 AUC=0.561, P=0.246; Supplementary Figure 3). On the other hand, MBP and NDEL1 ▵Crt values did not distinguish SCZ from BD (MBP AUC=0.597, P=0.267; NDEL1 AUC=0.553, P=0.525; Supplementary Figure 1), but both could differentiate SCZ (MBP AUC=0.677, P=0.006; NDEL1 AUC=0.660, P=0.002; Supplementary Figure 3) and BD patients (MBP AUC=0.805, P=2.856 × 10-4; NDEL1 AUC=0.728, P=0.004; Supplementary Figure 2) from healthy controls.
Correlation between gene expression and clinical characteristics
We tested the correlation of differential gene expression (AKT1, DICER, MBP and NDEL1) and the severity of symptoms and functional impairment (CDSS, CGI, GAF and YMRS total scores, and PANSS scores) using linear regression and inserting group as an independent variable. However, no significant correlation was detected after Bonferroni correction for multiple comparisons.
Discussion
In the present study, we identified gene expression alterations that might be linked to manic and psychotic features. We primarily detected that AKT1 and DICER1 expression levels were higher in BD patients compared with SCZ patients and controls, suggesting that the expression of these genes is associated more specifically to manic features. In addition, MBP and NDEL1 expression levels were higher in both SCZ and BD patients than in healthy controls, indicating that these genes may be related to psychosis per se (independently of diagnosis). We described similar results comparing FEP (excluding BD) and controls in a previous study.22 Here we focused on verifying if these changes in gene expression are also observed in BD, and moreover, if they can discriminate BD from SCZ patients. Although all these four genes have a role in central nervous system, this study aimed to find potential biomarkers, even if they might not be related to the pathophysiology of the disease. A brief description of each comparison (SCZ × BD and BD × control) and the relationship of each gene with psychotic disorders are described below.
SCZ and BD
Two genes were differentially expressed in BD, when compared with both SCZ and healthy controls, namely AKT1 (V-Akt Murine Thymoma Viral Oncogene Homolog 1) and DICER1 (Dicer 1, ribonuclease type III).
AKT1, which encodes a serine–threonine protein kinase, is involved in a variety of central nervous system functions such as neurodevelopment, synaptic plasticity and protein synthesis.28 Moreover, AKT1 was shown to facilitate dopamine signaling,29, 30 and to regulate a wide array of cellular processes, such as metabolism, growth, proliferation and apoptosis.31 AKT1 participates in the PI3K/Akt/mTOR pathway, which is important for many immunological mechanisms.32
In addition, lithium, antidepressants, antipsychotics and other mood stabilizers seem to increase phosphorylation of AKT.33, 34, 35 A decrease in AKT1 protein and mRNA levels was found in postmortem brain tissue and lymphocyte-derived cells of individuals with schizophrenia and bipolar disorder, compared with healthy controls.35, 36, 37 Although these studies primarily suggest an AKT1 deficiency, a recent study14 found increased AKT1 expression levels in peripheral blood of antipsychotic-naive schizophrenia patients. Kumarasinghe et al. (2013) also observed that antipsychotic pharmacotherapy could partially compensate for this upregulation, providing further evidence for a link between AKT1 and dopaminergic transmission.
Although we did not find differences in AKT1 mRNA levels between SCZ and controls, we found that this gene was upregulated specifically in BD patients. A previous study in a Brazilian sample of unmedicated, depressed individuals with bipolar disorder showed decreased AKT1 expression in the blood.18 Our finding of increased AKT1 expression in patients on the opposite pole of the spectrum (in mania) may suggest an association between AKT1 expression and mood pole.
DICER1 synthesizes DICER, a member of the ribonuclease III protein family that is involved in the generation of microRNAs (miRNAs), which regulate gene expression at the posttranscriptional level.38 MiRNAs are 22-nt-long RNAs generated from longer precursor RNAs. In general, they repress translation, but they can also acquire other functions after binding to their target RNA. Notably, many studies implicated miRNAs in the development of psychotic disorders.39, 40
DICER has an important role in the development and function of the immune41 and central nervous systems.42 DICER1 is upregulated in the dorsolateral prefrontal cortex42, 43 and lymphoblastoid cell lines44 of schizophrenia cases. In addition, DICER1 single-nucleotide polymorphisms45 and copy-number variations46 are associated to schizophrenia. Notably, valproic acid, a mood stabilizer used to treat bipolar disorder, induces DICER degradation.47
BD and healthy controls
Four genes were upregulated in BD patients compared with healthy controls (AKT1, DICER1, MBP and NDEL1). AKT1 and DICER1 expression levels were different between BD and SCZ patients, whereas the expression levels of MBP (myelin basic protein) and NDEL1 (nuclear distribution of protein nudE-like 1) were similar. Notably, in our previous findings in a larger cohort of FEP without bipolar disorder patients we found that MBP and NDEL1 were upregulated in antipsychotic-naive patients compared with controls.22 Here we show that these genes are also dysregulated in FEP with mania.
Myelin-related pathways are involved in the aetiologies of both schizophrenia and bipolar disorder.48 The MBP gene produces two families of structurally related proteins from different promoters: the MBP and the Golli (gene in the oligodendrocyte lineage) isoforms. In our study, we used an assay that can detect both types of isoforms (one Golli—NM_001025101, and two classic MBP isoforms—NM_001025090 and NM_001025092; Supplementary Table 1). Owing to the higher expression of Golli isoforms in the immune system,49 we may assume that they represent the expression detected in our experiments. The biological function of Golli isoforms involves myelin formation and maintenance,50 oligodendrocyte proliferation and migration,51 and calcium homeostasis.52 This specific calcium pathway is altered in schizophrenia and bipolar disorder53 and is also affected by antipsychotic medications.54
MBP expression studies in postmortem tissues from different brain regions revealed an association with psychotic disorders, suggesting that MBP mRNA and protein levels (of the classic isoforms) are decreased in patients with schizophrenia 55, 56, 57 or bipolar disorder.58 However, no differences were found in the Golli-MBP mRNA levels in postmortem dorsolateral prefrontal cortex samples of schizophrenia patients compared with controls.59 Notably, MBP expression is affected by antipsychotic treatment,60, 61 and hence, analyzing antipsychotic-naive patients is essential to attenuate the effects of these medications on gene expression.
In our study, both antipsychotic-naive SCZ and BD patients exhibited increased MBP expression levels (most likely Golli isoforms) compared with healthy controls, suggesting that this gene might be associated to psychosis per se, as both patient groups exhibited psychotic symptoms. Indeed, when we analyzed FEP patients in a larger sample, excluding those with manic features, MBP was also upregulated compared with healthy controls22 and to ultra-high risk individuals,23 supporting that MBP might have a role in psychosis. Moreover, in an independent report by Kumarasinghe et al. (2013), antipsychotic-naive schizophrenia patients exhibited higher MBP mRNA levels, compared with controls. After 6 to 8 weeks of risperidone or haloperidol treatment, the MBP mRNA levels were similar to that of controls.14 However, another study did not reveal significant changes in MBP in the peripheral blood of first-episode schizophrenia and bipolar disorder patients.17
NDEL1 is encoded by a gene located at chromosome 17p13.1, and is robustly expressed in developing and mature neurons in the brain. It has been suggested to have a role in neuronal migration during embryogenesis.62 Other well-known functions of NDEL1 include cytoskeleton organization, cell proliferation and survival regulation, oligopeptidase activity (potential neuropeptidase activity), neuritogenesis, neuronal migration and cell signaling.63, 64, 65 This protein was first discovered due to its enzyme activity on neuropeptides,66 and second because of its ability to form complexes with other proteins such as LIS1, which is encoded by the PAFAH1B1 (platelet-activating factor acetylhydrolase 1b, regulatory Subunit 1 (45 kDa)) gene that is mutated in lissencephaly, a rare brain formation disorder.67, 68 In addition, NDEL1 is able to bind DISC1,65 a psychosis-associated protein that is the product of a well-known schizophrenia risk factor gene,69 and is associated with bipolar disorder.70
Gene expression studies conducted in both brain and peripheral tissues revealed reduced NDEL1 expression levels in schizophrenia patients.71, 72 However, none of these studies have investigated antipsychotic-naive subjects or patients at the first stages of psychotic disorders. In a similar analysis, we have previously reported an upregulation of NDEL1 gene expression in FEP patients compared with healthy controls.22As we have included BD patients in the present study, we suggest that NDEL1 expression is also altered in psychotic disorders with manic features, in the same way as observed in SCZ patients. Moreover, a subgroup of FEP patients with depression showed lower levels of NDEL1 expression,20 opposite to our finding in BD, which is also at the opposite pole of depression. Thus, higher NDEL1 expression might be characteristic of BD and FEP without depression, while lower NDEL1 expression levels could be associated to FEP with depression.
The results of this study need to be replicated in additional studies and should be interpreted in light of some limitations. First, the sample size of the groups and particularly the BD patient group was small, and, hence, it may lack power to identify some gene expression differences. Second, it was a cross-sectional study, and the follow-up of these patients would confirm the diagnosis and may help to find markers for response to treatment. Third, we did not have a group of patients with mania without psychotic symptoms. Such a group would help to define which genes are specifically related to mania. Fourth, we cannot assure if our findings in whole blood translate to what occurs in the brain. Considering that whole blood presents a mixture of various leukocyte subtypes, our gene expression findings may also be partially confounded by various proportions of leukocyte subtypes.73 However, an important strength is our focus on unmedicated patients in the first stages of the disorder. Consequently, we needed to use a biological material that can be easily collected, though we acknowledge that peripheral markers may not necessarily reflect brain pathophysiology.
Conclusions
To our knowledge, this is the first study that compares gene expression in antipsychotic-naive FEP of SCZ and FEP with mania (BD), suggesting potential diagnostic specificities. On the basis of an integrated model, we propose that MBP and NDEL1 are upregulated in SCZ and BD patients, who all exhibit psychotic symptoms. Moreover, two other genes, AKT1 and DICER1, were upregulated in BD patients only, indicating that these genes could be related to mania, independently of psychotic symptoms. Although further validation in a large sample is still needed, our findings suggest that genes related to neuronal development are altered in psychotic disorders, and some of them might support the differential diagnosis between schizophrenia and bipolar disorder in the near future, which in turn could have an impact on the treatment of these disorders.
References
Perala J, Suvisaari J, Saarni SI, Kuoppasalmi K, Isometsa E, Pirkola S et al. Lifetime prevalence of psychotic and bipolar I disorders in a general population. Arch Gen Psychiatry 2007; 64: 19–28.
Bogren M, Mattisson C, Isberg PE, Nettelbladt P . How common are psychotic and bipolar disorders? A 50-year follow-up of the Lundby population. Nord J Psychiatry 2009; 63: 336–346.
Birchwood M, Todd P, Jackson C . Early intervention in psychosis. The critical period hypothesis. Br J Psychiatry Suppl 1998; 172: 53–59.
Demjaha A, MacCabe JH, Murray RM . How genes and environmental factors determine the different neurodevelopmental trajectories of schizophrenia and bipolar disorder. Schizophr Bull 2012; 38: 209–214.
Fuste M, Pinacho R, Melendez-Perez I, Villalmanzo N, Villalta-Gil V, Haro JM et al. Reduced expression of SP1 and SP4 transcription factors in peripheral blood mononuclear cells in first-episode psychosis. J Psychiatr Res 2013; 47: 1608–1614.
Kraepelin E . Manic-Depressive Insanity and Paranoia. E. & S. Livingstone: Edinburgh, UK, 1919.
van de Leemput J, Glatt SJ, Tsuang MT . The potential of genetic and gene expression analysis in the diagnosis of neuropsychiatric disorders. Expert Rev Mol Diagn 2016; 16: 677–695.
Wirgenes KV, Tesli M, Inderhaug E, Athanasiu L, Agartz I, Melle I et al. ANK3 gene expression in bipolar disorder and schizophrenia. Br J Psychiatry 2014; 205: 244–245.
Zhao Z, Xu J, Chen J, Kim S, Reimers M, Bacanu SA et al. Transcriptome sequencing and genome-wide association analyses reveal lysosomal function and actin cytoskeleton remodeling in schizophrenia and bipolar disorder. Mol Psychiatry 2015; 20: 563–572.
Smalheiser NR, Lugli G, Zhang H, Rizavi H, Cook EH, Dwivedi Y . Expression of microRNAs and other small RNAs in prefrontal cortex in schizophrenia, bipolar disorder and depressed subjects. PLoS ONE 2014; 9: e86469.
Ota VK, Noto C, Gadelha A, Santoro ML, Ortiz BB, Andrade EH et al. Evaluation of neurotransmitter receptor gene expression identifies GABA receptor changes: a follow-up study in antipsychotic-naive patients with first-episode psychosis. J Psychiatr Res 2014; 56: 130–136.
Ota VK, Noto C, Gadelha A, Santoro ML, Silva PN, Melaragno MI et al. Neurotransmitter receptor and regulatory gene expression in peripheral blood of Brazilian drug-naive first-episode psychosis patients before and after antipsychotic treatment. Psychiatry Res 2013; 210: 1290–1292.
Santoro ML, Ota VK, Stilhano RS, Silva PN, Santos CM, Diana MC et al. Effect of antipsychotic drugs on gene expression in the prefrontal cortex and nucleus accumbens in the spontaneously hypertensive rat (SHR). Schizophr Res 2014; 157: 163–168.
Kumarasinghe N, Beveridge NJ, Gardiner E, Scott RJ, Yasawardene S, Perera A et al. Gene expression profiling in treatment-naive schizophrenia patients identifies abnormalities in biological pathways involving AKT1 that are corrected by antipsychotic medication. Int J Neuropsychopharmacol 2013; 16: 1483–1503.
Padmos RC, Hillegers MH, Knijff EM, Vonk R, Bouvy A, Staal FJ et al. A discriminating messenger RNA signature for bipolar disorder formed by an aberrant expression of inflammatory genes in monocytes. Arch Gen Psychiatry 2008; 65: 395–407.
Clelland CL, Read LL, Panek LJ, Nadrich RH, Bancroft C, Clelland JD . Utilization of never-medicated bipolar disorder patients towards development and validation of a peripheral biomarker profile. PLoS ONE 2013; 8: e69082.
Gutierrez-Fernandez A, Gonzalez-Pinto A, Vega P, Barbeito S, Matute C . Expression of oligodendrocyte and myelin genes is not altered in peripheral blood cells of patients with first-episode schizophrenia and bipolar disorder. Bipol Disord 2010; 12: 107–109.
Machado-Vieira R, Zanetti MV, Teixeira AL, Uno M, Valiengo LL, Soeiro-de-Souza MG et al. Decreased AKT1/mTOR pathway mRNA expression in short-term bipolar disorder. Eur Neuropsychopharmacol 2015; 25: 468–473.
Christofolini DM, Bellucco FT, Ota VK, Belangero SI, Cernach MC, Gadelha A et al. Assessment of 22q11.2 copy number variations in a sample of Brazilian schizophrenia patients. Schizophr Res 2011; 132: 99–100.
Noto C, Ota VK, Santoro ML, Gouvea ES, Silva PN, Spindola LM et al. Depression, cytokine, and cytokine by treatment interactions modulate gene expression in antipsychotic naive first episode psychosis. Mol Neurobiol 2015; e-pub ahead of print 22 October 2015; doi:10.1007/s12035-015-9489-3.
Ota VK, Noto C, Gadelha A, Santoro ML, Spindola LM, Gouvea ES et al. Changes in gene expression and methylation in the blood of patients with first-episode psychosis. Schizophr Res 2014; 159: 358–364.
Ota VK, Noto C, Santoro ML, Spindola LM, Gouvea ES, Carvalho CM et al. Increased expression of NDEL1 and MBP genes in the peripheral blood of antipsychotic-naive patients with first-episode psychosis. Eur Neuropsychopharmacol 2015; 25: 2416–2425.
Santoro ML, Gadelha A, Ota VK, Cunha GR, Asevedo E, Noto CS et al. Gene expression analysis in blood of ultra-high risk subjects compared to first-episode of psychosis patients and controls. World J Biol Psychiatry 2015 16 441–446.
Vessoni AL . Adaptação e estudo de confiabilidade da escala de avaliação das síndromes positiva e negativa para a esquizofrenia no Brasil. São Paulo Escola Paulista de Medicina 1993.
Lima MS, Soares BG, Paoliello G, Machado Vieira R, Martins CM, Mota Neto JI et al. The Portuguese version of the Clinical Global Impression-Schizophrenia Scale: validation study. Rev Bras Psiquiatr 2007; 29: 246–249.
Bressan RA, Chaves AC, Shirakawa I, de Mari J . Validity study of the Brazilian version of the Calgary Depression Scale for Schizophrenia. Schizophr Res 1998; 32: 41–49.
Higuchi CH, Ortiz B, Berberian AA, Noto C, Cordeiro Q, Belangero SI et al. Factor structure of the Positive and Negative Syndrome Scale (PANSS) in Brazil: convergent validation of the Brazilian version. Rev Bras Psiquiatr 2014; 36: 336–339.
Zheng W, Wang H, Zeng Z, Lin J, Little PJ, Srivastava LK et al. The possible role of the Akt signaling pathway in schizophrenia. Brain Res 2012; 1470: 145–158.
Alessi DR, Andjelkovic M, Caudwell B, Cron P, Morrice N, Cohen P et al. Mechanism of activation of protein kinase B by insulin and IGF-1. EMBO J 1996; 15: 6541–6551.
Beaulieu JM, Gainetdinov RR, Caron MG . Akt/GSK3 signaling in the action of psychotropic drugs. Ann Rev Pharmacol Toxicol 2009; 49: 327–347.
Brazil DP, Yang ZZ, Hemmings BA . Advances in protein kinase B signalling: AKTion on multiple fronts. Trends Biochem Sci 2004; 29: 233–242.
Weichhart T, Saemann MD . The PI3K/Akt/mTOR pathway in innate immune cells: emerging therapeutic applications. Annals of the rheumatic diseases. Ann Rheum Dis 2008; 67 (Suppl 3): iii70–iii74.
Chalecka-Franaszek E, Chuang DM . Lithium activates the serine/threonine kinase Akt-1 and suppresses glutamate-induced inhibition of Akt-1 activity in neurons. Proc Natl Acad Sci USA 1999; 96: 8745–8750.
De Sarno P, Li X, Jope RS . Regulation of Akt and glycogen synthase kinase-3 beta phosphorylation by sodium valproate and lithium. Neuropharmacology 2002; 43: 1158–1164.
Emamian ES, Hall D, Birnbaum MJ, Karayiorgou M, Gogos JA . Convergent evidence for impaired AKT1-GSK3beta signaling in schizophrenia. Nat Genet 2004; 36: 131–137.
Thiselton DL, Vladimirov VI, Kuo PH, McClay J, Wormley B, Fanous A et al. AKT1 is associated with schizophrenia across multiple symptom dimensions in the Irish study of high density schizophrenia families. Biol Psychiatry 2008; 63: 449–457.
van Beveren NJ, Buitendijk GH, Swagemakers S, Krab LC, Roder C, de Haan L et al. Marked reduction of AKT1 expression and deregulation of AKT1-associated pathways in peripheral blood mononuclear cells of schizophrenia patients. PLoS ONE 2012; 7: e32618.
Carthew RW . Gene regulation by microRNAs. Curr Opin Genet Dev 2006; 16: 203–208.
Forstner AJ, Hofmann A, Maaser A, Sumer S, Khudayberdiev S, Muhleisen TW et al. Genome-wide analysis implicates microRNAs and their target genes in the development of bipolar disorder. Transl Psychiatry 2015; 5: e678.
Caputo V, Ciolfi A, Macri S, Pizzuti A . The emerging role of MicroRNA in schizophrenia. CNS Neurol Disord Drug Targets 2015; 14: 208–221.
Devasthanam AS, Tomasi TB . Dicer in immune cell development and function. Immunol Invest 2014; 43: 182–195.
Santarelli DM, Beveridge NJ, Tooney PA, Cairns MJ . Upregulation of dicer and microRNA expression in the dorsolateral prefrontal cortex Brodmann area 46 in schizophrenia. Biol Psychiatry 2011; 69: 180–187.
Beveridge NJ, Gardiner E, Carroll AP, Tooney PA, Cairns MJ . Schizophrenia is associated with an increase in cortical microRNA biogenesis. Mol Psychiatry 2010; 15: 1176–1189.
Sanders AR, Goring HH, Duan J, Drigalenko EI, Moy W, Freda J et al. Transcriptome study of differential expression in schizophrenia. Hum Mol Genet 2013; 22: 5001–5014.
Zhou Y, Wang J, Lu X, Song X, Ye Y, Zhou J et al. Evaluation of six SNPs of MicroRNA machinery genes and risk of schizophrenia. J Mol Neurosci 2013; 49: 594–599.
Xu B, Roos JL, Levy S, van Rensburg EJ, Gogos JA, Karayiorgou M . Strong association of de novo copy number mutations with sporadic schizophrenia. Nat Genet 2008; 40: 880–885.
Zhang Z, Convertini P, Shen M, Xu X, Lemoine F, de la Grange P et al. Valproic acid causes proteasomal degradation of DICER and influences miRNA expression. PLoS ONE 2013; 8: e82895.
Yu H, Bi W, Liu C, Zhao Y, Zhang D, Yue W . A hypothesis-driven pathway analysis reveals myelin-related pathways that contribute to the risk of schizophrenia and bipolar disorder. Progress Neuropsychopharmacol Biol Psychiatry 2014; 51: 140–145.
Feng JM, Hu YK, Xie LH, Colwell CS, Shao XM, Sun XP et al. Golli protein negatively regulates store depletion-induced calcium influx in T cells. Immunity 2006; 24: 717–727.
Jacobs EC, Pribyl TM, Feng JM, Kampf K, Spreur V, Campagnoni C et al. Region-specific myelin pathology in mice lacking the golli products of the myelin basic protein gene. J Neurosci 2005; 25: 7004–7013.
Paez PM, Fulton D, Spreuer V, Handley V, Campagnoni AT . Modulation of canonical transient receptor potential channel 1 in the proliferation of oligodendrocyte precursor cells by the golli products of the myelin basic protein gene. J Neurosc 2011; 31: 3625–3637.
Feng JM, Fernandes AO, Campagnoni CW, Hu YH, Campagnoni AT . The golli-myelin basic protein negatively regulates signal transduction in T lymphocytes. J Neuroimmunol 2004; 152: 57–66.
Heyes S, Pratt WS, Rees E, Dahimene S, Ferron L, Owen MJ et al. Genetic disruption of voltage-gated calcium channels in psychiatric and neurological disorders. Progress Neurobiol 2015; 134: 36–54.
Lidow MS . Calcium signaling dysfunction in schizophrenia: a unifying approach. Brain Res Brain Res Rev 2003; 43: 70–84.
Martins-de-Souza D, Gattaz WF, Schmitt A, Maccarrone G, Hunyadi-Gulyas E, Eberlin MN et al. Proteomic analysis of dorsolateral prefrontal cortex indicates the involvement of cytoskeleton, oligodendrocyte, energy metabolism and new potential markers in schizophrenia. J Psychiatr Res 2009; 43: 978–986.
Matthews PR, Eastwood SL, Harrison PJ . Reduced myelin basic protein and actin-related gene expression in visual cortex in schizophrenia. PLoS ONE 2012; 7: e38211.
Tkachev D, Mimmack ML, Ryan MM, Wayland M, Freeman T, Jones PB et al. Oligodendrocyte dysfunction in schizophrenia and bipolar disorder. Lancet 2003; 362: 798–805.
Wesseling H, Gottschalk MG, Bahn S . Targeted multiplexed selected reaction monitoring analysis evaluates protein expression changes of molecular risk factors for major psychiatric disorders. Int J Neuropsychopharmacol 2015; 18: 1.
Baruch K, Silberberg G, Aviv A, Shamir E, Bening-Abu-Shach U, Baruch Y et al. Association between golli-MBP and schizophrenia in the Jewish Ashkenazi population: are regulatory regions involved? Int J Neuropsychopharmacol 2009; 12: 885–894.
Narayan S, Kass KE, Thomas EA . Chronic haloperidol treatment results in a decrease in the expression of myelin/oligodendrocyte-related genes in the mouse brain. J Neurosci Res 2007; 85: 757–765.
Sugai T, Kawamura M, Iritani S, Araki K, Makifuchi T, Imai C et al. Prefrontal abnormality of schizophrenia revealed by DNA microarray: impact on glial and neurotrophic gene expression. Ann N Y Acad Sci 2004; 1025: 84–91.
Chansard M, Hong JH, Park YU, Park SK, Nguyen MD. Ndel1 Nudel . (Noodle): flexible in the cell? Cytoskeleton 2011; 68: 540–554.
Sasaki S, Mori D, Toyo-oka K, Chen A, Garrett-Beal L, Muramatsu M et al. Complete loss of Ndel1 results in neuronal migration defects and early embryonic lethality. Mol Cell Biol 2005; 25: 7812–7827.
Chansard M, Wang J, Tran HC, Neumayer G, Shim SY, Park YU et al. The cytoskeletal protein Ndel1 regulates dynamin 2 GTPase activity. PLoS ONE 2011; 6: e14583.
Hayashi MA, Portaro FC, Bastos MF, Guerreiro JR, Oliveira V, Gorrao SS et al. Inhibition of NUDEL (nuclear distribution element-like)-oligopeptidase activity by disrupted-in-schizophrenia 1. Proc Natl Acad Sci USA. 2005; 102: 3828–3833.
Hayashi MA, Portaro FC, Tambourgi DV, Sucupira M, Yamane T, Fernandes BL et al. Molecular and immunochemical evidences demonstrate that endooligopeptidase A is the predominant cytosolic oligopeptidase of rabbit brain. Biochem Biophys Res Commun 2000; 269: 7–13.
Reiner O . LIS1. let's interact sometimes... (part 1). Neuron 2000; 28: 633–636.
Reiner O, Cahana A, Escamez T, Martinez S . LIS1-no more no less. Mol Psychiatry 2002; 7: 12–16.
Tomppo L, Hennah W, Lahermo P, Loukola A, Tuulio-Henriksson A, Suvisaari J et al. Association between genes of Disrupted in schizophrenia 1 (DISC1) interactors and schizophrenia supports the role of the DISC1 pathway in the etiology of major mental illnesses. Biol Psychiatry 2009; 65: 1055–1062.
Palo OM, Antila M, Silander K, Hennah W, Kilpinen H, Soronen P et al. Association of distinct allelic haplotypes of DISC1 with psychotic and bipolar spectrum disorders and with underlying cognitive impairments. Hum Mol Genet 2007; 16: 2517–2528.
Lipska BK, Mitkus SN, Mathew SV, Fatula R, Hyde TM, Weinberger DR et al. Functional genomics in postmortem human brain: abnormalities in a DISC1 molecular pathway in schizophrenia. Dialogues Clin Neurosci 2006; 8: 353–357.
Rampino A, Walker RM, Torrance HS, Anderson SM, Fazio L, Di Giorgio A et al. Expression of DISC1-interactome members correlates with cognitive phenotypes related to schizophrenia. PLoS ONE 2014; 9: e99892.
Whitney AR, Diehn M, Popper SJ, Alizadeh AA, Boldrick JC, Relman DA et al. Individuality and variation in gene expression patterns in human blood. Proc Natl Acad Sci USA. 2003; 100: 1896–1901.
Acknowledgements
We thank the patients, their families, and the psychiatrists and nurses for their participation in this study. This study was supported by Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP 2010/08968-6; 2011/50740-5; 2014/50830-2, 2014/07280-1, 2013/10498-6, 2012/12686-1), Brazil; and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Brazil.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
CN has received a scholarship from Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and has served as a consultant or advisory board member for Janssen. AG was on the speakers’ bureau and/or has acted as a consultant for Janssen-Cilag in the last 12 months and has also received research support from Brazilian government institutions (CNPq). EB has been supported by CNPq, CAPES and FAPESP. RAB has received research funding from FAPESP, CNPq, CAPES, Fundação Safra, Fundação ABADS, Janssen, Eli Lilly, Lundbeck, Novartis and Roche, has served as a speaker for Astra Zeneca, Bristol, Janssen, Lundbeck and Revista Brasileira de Psiquiatria and is a shareholder of Radiopharmacus Ltda and Biomolecular Technology. The remaining authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Translational Psychiatry website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Gouvea, E., Ota, V., Noto, C. et al. Gene expression alterations related to mania and psychosis in peripheral blood of patients with a first episode of psychosis. Transl Psychiatry 6, e908 (2016). https://doi.org/10.1038/tp.2016.159
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tp.2016.159
- Springer Nature Limited
This article is cited by
-
NKCC1 to KCC2 mRNA Ratio in Schizophrenia and Its Psychopathology: a Case–Control Study
Journal of Molecular Neuroscience (2022)
-
Widespread transcriptional disruption of the microRNA biogenesis machinery in brain and peripheral tissues of individuals with schizophrenia
Translational Psychiatry (2020)
-
Gene expression over the course of schizophrenia: from clinical high-risk for psychosis to chronic stages
npj Schizophrenia (2019)
-
Accessing Gene Expression in Treatment-Resistant Schizophrenia
Molecular Neurobiology (2018)