Abstract
Candida albicans is one of the most dangerous pathogenic fungi in the world, according to the classification of the World Health Organization, due to the continued development of its resistance to currently available anticandidal agents. To overcome this problem, the current work provided a simple, one-step, cost-effective, and safe technique for the biosynthesis of new functionalized anticandidal selenium nanoparticles (Se NPs) against C. albicans ATCC10231 using the cell-free supernatant of Limosilactobacillus fermentum (OR553490) strain. The bacterial strain was isolated from yogurt samples available in supermarkets, in Damietta, Egypt. The mixing ratio of 1:9 v/v% between cell-free bacterial metabolites and sodium selenite (5 mM) for 72 h at 37 °C were the optimum conditions for Se NPs biosynthesis. Ultraviolet–visible spectroscopy (UV–Vis), Fourier transform infrared spectroscopy (FT-IR), transmission electron microscopy (TEM), X-ray diffraction (XRD), Zeta analyses, and elemental analysis system (EDS) were used to evaluate the optimized Se NPs. The Se NPs absorption peak appeared at 254 nm. Physicochemical analysis of Se NPs revealed the crystalline-shaped and well-dispersed formation of NPs with an average particle size of 17–30 nm. Se NPs have − 11.8 mV, as seen by the zeta potential graph. FT-IR spectrum displayed bands of symmetric and asymmetric amines at 3279.36 cm−1 and 2928.38 cm−1, aromatic and aliphatic (C–N) at 1393.32 cm−1 and 1237.11.37 cm−1 confirming the presence of proteins as stabilizing and capping agents. Se NPs acted as a superior inhibitor of C. albicans with an inhibition zone of 26 ± 0.03 mm and MIC value of 15 µg/mL compared to one of the traditional anticandidal agent, miconazole, which revealed 18 ± 0.14 mm and 75 µg/mL. The cytotoxicity test shows that Se NPs have a low toxic effect on the normal keratinocyte (IC50 ≈ 41.5 μg/mL). The results indicate that this green synthesis of Se NPs may have a promising potential to provide a new strategy for drug therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Background
The genus Candida comprises over 200 species of which 15 have been isolated from infections in humans and animals. The most prevalent pathogens are Candida albicans, C. glabrata, C. parapsilosis, C. tropicalis, and C. krusei [1]. C. albicans as an opportunistic fungal pathogen is a typical component of the microbiota of the human digestive tract. Even though there are only 200 species in the genus Candida, it can cause up to 75% of candidal infections. On the other hand, bacterial normal flora acquired host defense mechanisms to allow Candida organisms to proliferate and endure as commensals. Nonetheless, even a small alteration to the host’s biological milieu or defensive mechanism can let C. albicans develop into a pathogen that can cause potentially fatal infections [2]. The microbiota in healthy hosts often consists of a balanced mix of energy and metabolites. Potentially harmful germs are kept from growing out of control by the homeostatic balance. Dysbiosis, or the imbalance of the microbial community, is frequently linked to illnesses in humans [3]. While some oral bacteria promote C. albicans biofilm production, others hinder it. The expression of C. albicans virulence genes, as well as the growth of C. albicans hyphae and biofilm, were found to be up regulated by Streptococcus and Actinomyces species in an in vitro measurement of mixed species biofilm generated on the salivary pellicle. Additionally, S. oralis promoted the growth of C. albicans hyphae and biofilms [4]. C. albicans is the most dangerous Candida species according to the World Health Organization (WHO) fungal priority pathogens list [5]. Even though C. albicans is a typical microbiome, it can foster opportunistic infections that might endanger both human and animal lives. C. albicans typically affects the whole part of the body from the skin to the lower respiratory tract, oropharynx, gastrointestinal tract, and genitourinary system [6]. The term “candidiasis” refers to a group of illnesses that vary from severe infections to less significant, while invasive candidiasis (IC) includes serious conditions such as candidemia, endocarditis, disseminated infections, infections of the central nervous system, endophthalmitis, and osteomyelitis. Also, candidiasis risk factors include the use of corticosteroids, invasive procedures, and harsh chemotherapy [7]. Around 70% of fungal infections worldwide are brought on by C. albicans. Throughout the past few decades, it has consistently been the main contributor to invasive infections that can be fatal [8]. C. albicans can be treated through chemical anticandidal drugs like the Azole group which are two subgroups: imidazole and triazole subgroup, but these drugs have highly severe side effects such as hypertriglyceridemia, elevated liver enzymes, rash, pedal edema, hepatotoxicity [9]. The Candida sp. virulence increases due to its ability to resist the available antifungal drugs such as the azole group that occur due to modifications of the target enzyme or due to reduced access of the drug to the target [1, 5, 9].
Nanotechnology was a recent science field which studied matters at the nanoscale dimension and changes their properties. NPs are 1–100 nm in size and have advantages of chemical stability, potential antifungal effects, low toxicity, and low pathogen resistance [10, 11]. NPs such as silver, copper, gold, iron oxide, and selenium have critical roles in agriculture, food, the environment, and the nanomedicine field [12, 13]. Potential NPs could disrupt the cell membrane of the microbial cells through a mechanism of inhibiting the activity of the enzyme Lanosterol 14-α-demethylase, which is involved in the cholesterol analogue (ergosterol) biosynthesis and the largest sterol element of the fungal cell membrane. Also, NPs might induce reactive oxygen species (ROS), inhibition of spore germination, and protein regulation [14,15,16]. Among different NPs, selenium NPs (Se NPs) were preferred due to several advantages such as their essential role in improving human health, via seleno-proteins, antioxidant defense, cell signaling, immunological modulation, and other metabolic activities [17]. In addition, Se NPs display several benefits, such as low toxicity, high bioavailability, and degradability so, it’s safe for clinical administration and outstanding in nanomedicine as anticancer, antiviral activity, and antimicrobial [18, 19]. Also, Se is used in several pathophysiological diseases like cardiovascular diseases, cancer, diabetes, neurodegenerative diseases, and so on, due to its activity as an antioxidant and anti-inflammatory [20]. Through the previous studies, the biological methods for Se NPs synthesis were limitless. Thus, using microorganisms might gain great attention as new bio-nano factories that convert the ion metals into metal NPs through biotransformation [21].
These NPs can be synthesized by chemical, physical, and biological methods, but the chemical and physical methods are not preferred due to high thermal conditions, hazardous chemicals, acidic pH, and highly toxic and unsafe methods than the biological method [22]. The biological synthesis of NPs is preferable due to its low cost, simplicity, safety, increased biocompatibility and stability, and non-toxic, high-productivity method for the production of NPs [23]. The biological method depends on the green chemistry that can be performed by living organisms or their natural secondary metabolites such as proteins including plants [24, 25], fungi [26], yeast [27], algae [28], non-living viruses [29] and bacteria [23, 30, 31]. Among the significant categories of microorganisms involved in biosynthesis of NPs, the probiotic lactic acid bacteria (LAB) are recommended as a safe, rapid, easy to culture, available in dairy products and cheap bio-nano-factory for NPs production [32, 33]. They can be facultative anaerobes, even though they are sometimes classified as aerotolerant anaerobes. LABS, which are extensively dispersed in nature, is the main microflora of milk and its derivatives. LAB produce different antimicrobial substances such as formic acid, hydrogen peroxide, ethanol, acetone, and bacteriocins that might show synergistic action with NPs [34]. These compounds endow them with the ability to preserve food, acting as a natural means of competition to outcompete other microorganisms occupying the same niche [35]. Moreover, LAB was reported to produce a high yield of secondary metabolites and nourish with benefit proteins [36]. Different LAB strains were reported to biosynthsize NPs with varying sizes, ranging from 50 to 100 nm (S. thermofilus), 100–200 nm (Lactobacillus sp.), and 400–500 nm (Bifidobacter sp.) [32, 35]. They pay close attention to nutrition. These bacteria get their energy from substrate-level phosphorylation. The current study aimed to use Limosilactobacillus fermentum OR553490 to bio-reduce Se salt into selenium NPs (Se NPs) extracellularly, characterize Se NPs, and in vitro test their potential as an anticandidal agent. In addition, application of the biosynthesized Se NPs in different medical products such as topical cream and shampoo as newly combinations with chitosan (CS), alginate (Alg) and panthenol to enhance their antimicrobial action.
2 Materials and methods
2.1 Materials
Sodium selenite (Na2SeO3) and miconazole were purchased from Sigma Aldrich, St. Louis, MO, USA. Candida albicans ATCC10231 strain was used in the anticandidal activity investigations. Culture media was purchased from Difco Laboratories, Detroit, Mich. Chemicals were purchased from Oxoid Ltd., England.
2.2 Sample collection
Zabady, as previously mentioned by Altalhi et al. [37], is an Arabian yogurt produced from a 1:1 ratio of buffalo to cow milk. Samples of various Egyptian zabady (a total of 37 samples) were gathered in October and September from several supermarkets in the northern Egyptian cities of Damietta Governorate, which includes New Damietta, Kafr Saad, and Ras Elbar. Every sample was collected in sterile cups, brought to the Microbiology Laboratory, Faculty of Science, Damietta University, in an ice box, and kept in a refrigerator.
2.3 Isolation and purification of LAB bacteria
DE Man, Rogosa, Sharpe (MRS) agar medium was prepared and sterilized using an autoclave (121 °C, 1.5 atm). After sterilization, leave the media to cool down to (42–45 °C). Serial dilutions from different zabady samples were prepared (10–1 to 10–6). About 0.1 mL of each dilution was added into a sterile Petri dish and the autoclaved culture media were poured under aseptic conditions. The inoculated agar plates were mixed well and incubated at 37 °C for 24 h under anaerobic conditions. After incubation time, a single colony from each different bacterial colony was transferred into new agar plates [38].
2.4 Phenotypic characterization
After the purification steps, each single colony was stained using Gram stain followed by microscopic examination, morphological characterization, and testing of the catalase and oxidase activity. Only Gram-positive and catalase-negative bacterial isolates were selected, purified, and preserved on agar slants for further use [39].
2.5 Screening for Se NPs production
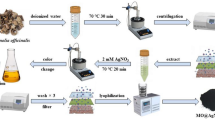
A 0.5 McFarland standard (1 × 106 cell/mL) from each different bacterial isolate was prepared in 100 mL MRS broth flasks for 24 h at 37 °C. The inoculated flasks were incubated overnight at 150 rpm and 37 °C. After the incubation period, bacterial cells were collected using centrifugation at 5000 rpm for 15 min while bacterial metabolites were filtered using a 0.22 μm syringe filter (Millex GV, Millipore) and transferred into another clean flask. 1 mM from Na2SeO3 solution was prepared and added into the flasks of bacterial metabolite by the ratio 1:1 (v/v), then incubated in a shaking incubator at 150 rpm at 37 °C until the color changed from colorless to red color. The produced Se NPs using different bacterial isolates were collected using centrifugation at 30,000 rpm for 20 min and lyophilized after being rinsed four times with distilled water [38]. All produced Se NPs reaction mixtures were measured spectrophotometrically to detect the Se NPs biosynthesis using double beam spectrum UV–Vis spectrophotometer V-760 (JASCO, UK).
2.6 Evaluation of anticandidal action of the biosynthesized Se NPs using LAB isolates
Aliquot 150 μg/mL of different produced Se NPs from each LAB isolate was prepared and tested as an anticandidal agent in comparison to the standard anticandidal; miconazole according to the guidelines of the Clinical and Laboratory Standards Institute (CLSI) [40]. A 1 × 106 cell/mL of C. albicans ATCC10231 Sabouraud dextrose agar (SDA) plates were prepared and used during the agar well diffusion method. After incubation at 28 °C for 24 h, zones of inhibition (ZOI) were measured in mm to determine the highest anticandidal Se NPs.
2.7 Molecular diagnosis of the selected LAB isolates
The selected lactobacillus isolate genotypic identification was done at the Animal Health Research Institute, Giza (Egypt). Bacterial cells of the isolate were collected from overnight bacterial growth on MRS broth medium using centrifugation at 5000 rpm for 15 min, washed 3 times TE buffer (10 mM tris chloride, 1 mM EDTA, pH8.0), and resuspended in 350 μL TE buffer and 20 mg/mL of lysozyme (Sigma, USA). Tubes were vortexed, 350 μl of 10% SDS, and 100 µg/mL proteinase-K (Vivantis Technologies, Malaysia) were added, along with 100 g per mL, and then incubated at 37 °C for 1 h. After centrifugation, 200 μl ethanol (96%) was added, vortexed, centrifugated, and then applied to a QIAamp mini spin column (QIAamp DNA Mini Kit, catalog no.51304). After centrifugation at 8000 rpm for 1 min, the QIAamp mini spin column was transferred into a new collection tube, and the filtrate was discarded. The QIAamp mini spin column was washed several times using a buffer. 100 μL 1/10 TE buffer was added and stored at – 20 °C until used for DNA sequencing [41].
The genomic DNA was detected using electrophoresis on 1% agarose gel electrophoresis prepared in TAE buffer (0.04 M Tris-acetate and 0.001 M EDTA, pH 8.0) and ethidium bromide (10 mg/mL). Then, the agarose gel was examined using a UV transilluminator. A Gel Pilot 100 bp plus ladder (cat. no. 239045, QIAGEN, USA) was used as a molecular mass marker.
Amplification was done using oligonucleotide primers sequences (Metabion, Germany) including F27 (5' AGAGTTTGATCMTGGCTCAG 3') and R1492 (5' TACGGYTACCTTGTTACGACTT 3') to identify the selected isolate. The initial denaturation was done at 94°C for 15 min followed by 35 cycles of denaturation at 94 °C for 30 s, annealing at 56°C for 1 min, and extension at 72°C for 1 min [42].
The PCR product was sequenced in forward and reverse directions using Applied Biosystems 3130 automated DNA Sequencer (ABI, 3130, USA), a ready reaction Bigdye Terminator V3.1 cycle sequencing kit (Perkin-Elmer/Applied Biosystems, No. 4336817, Foster City, CA).
A BLAST® analysis (Basic Local Alignment Search Tool) was performed to match the best similarities with other related sequences and establish sequence identity to GenBank accessions [43]. Phylogenetic analysis was performed using the CLUSTAL W multiple sequence alignment program, version 12.1 of MegAlign module of Lasergene DNAStar software Pairwise (Madison, Wisconsin, USA) [44].
The neighbor-joining was performed using the maximum composite likelihood methods and the phylogenetic tree analyses were viewed and analyzed using MEGA6. The bootstrap analysis was performed based on 100 replicates [45, 46]
2.8 Optimization of Se NPs production
Different concentrations from Na2SeO3 (1–7 mM), different ratios between cell-free bacterial metabolites and Na2SeO3 (1:1–1:11 v/v%), and different incubation times (24–96 h) were tested to determine the best conditions for the biosynthesis of Se NPs. The rate and concentration of Se NPs production were measured spectrophotometrically to elucidate the optimized parameters for the bio-formation of Se NPs.
2.9 Characterization of the optimized biosynthesized Se NPs
Optimized Se NPs were studied and characterized using FT/IR-4000 Series Fourier transform infrared spectrometer (FT-IR, JASCO, UK), LabX XRD-6000 X-ray diffractometer (XRD, Shimadzu, Japan), JEM-2100 transmission electron microscope (TEM, JEOL, Japan), Nano-ZS90 Zetasizer (Malvern, UK), and scanning electron microscope (SEM, JEOL JSM-6510, Japan) outfitted with an EDX Genesis energy dispersive x-ray elemental analysis system (EDS).
2.10 The anticandidal activity of the optimized Se NPs using the agar well diffusion method
The anticandidal activity of biosynthesized Se NPs was tested against C. albicans ATCC10231 in comparison to the standard anticandidal; miconazole using agar well diffusion method according to CLSI [40]. A 0.5 McFarland from the tested yeast was prepared and inoculated into sterile cooled molted SDA and then poured into a sterile Petri dish. Different concentrations (50 and 150 µg/mL) of optimized Se NPs and miconazole were prepared and added into 5 mm wells that were punched in the inoculated agar plates. Plates were incubated at 28 °C for 24 h and then ZOI was recorded in mm.
2.11 Minimal inhibition concentration (MIC) and minimal fungicidal concentration (MFC)
The MIC values of the optimized Se NPs and miconazole were determined using the broth dilution method [47]. Different concentrations (0–75 µg/mL) of Se NPs and miconazole were prepared and added into sterilized Sabouraud dextrose broth (SDB) media conical flasks inoculated by 0.5 McFarland of C. albicans ATCC10231. Flasks were incubated at 28 °C for 24 h. The growth rate of tested yeast was measured using a Beckman DU-40 UV–Vis spectrophotometer (USA) at wavelength 600 nm against an uninoculated broth medium as a blank.
MIC flasks were inoculated into sterile cooled molted SDA medium and then poured into a sterile Petri dish. The inoculated agar plates were incubated at 28 °C for 24 h. After incubation, the growth of yeast colonies was examined, and the total was counted in colony-forming units per mL (CFU. mL−1) to determine MFC values [48].
2.12 Ultra-structure study of Se NPs-treated C. albicans ATCC10231
Exponential-phase cultures of C. albicans ATCC10231 were treated by MIC value of Se NPs for 2 h at 28°C. Yeast cells were collected using centrifugation at 5000 rpm for 15 min, washed with distilled water 3 times, and then fixed using a solution with 3% glutaraldehyde. Similarly, untreated C. albicans ATCC10231 was prepared and used as a control. Fixed samples were dehydrated using a series of pure ethanol dilutions (10–90%), filtered using acetone, embedded in an Epon‐Araldite (1:1) mixture for 1 h, and then polymerized at 65°C for 24 h. Using copper grids, the ultrathin sections of samples were collected, stained, and observed with a TEM [49].
2.13 Cytotoxicity effect of the biosynthesized Se NPs
The cytotoxic effect of the Se NPs was tested using normal mice keratinocyte cells. Keratinocyte cells were cultivated in a serum-free medium at the Regional Centre for Mycology and Biotechnology (Al-Azhar University, Cairo) according to the CELLnTEC protocol, Advanced Cell Systems AG (Bern, Switzerland). The experimental protocol was approved by the local ethical committee of AUHA (Al-Azhar University Housing Animals) and was conducted in compliance with the IACUC (Institutional Animal Care and Use Committee). A total of 15 × 103 mouse keratinocyte cells were taken during the exponential phase of growth, planted into 96-well tissue culture plates, and left to adhere for a full day. Se NPs were then added to the appropriate wells to obtain a final concentration of 0–200 μg/mL. After incubation for 24 h, each well was filled with 20 μl of DMEM medium containing 0.4% trypan blue solution and incubated for an additional 4 h. Then, the medium was examined using the hemocytometer to count live (unstained) and dead cells (stained) under a microscope to determine the half-maximal inhibitory concentration (IC50) of the Se [50].
2.14 In vitro application of the biosynthesized Se NPs as a based topical anticandidal drug
Hydrated sodium alginate (2%) was prepared in distilled water and stirred for 30 min at room temperature (25 °C). A MIC of Se NPs was added to the mixture and then stirred for 15 min to obtain alginate/Se NPs (Alg/Se). Also, the previous mixture was prepared in the presence of 1% CS to obtain Alg/CS/Se. Alg/Se and Alg/CS/Se were tested against C. albicans ATCC10231 using the agar well diffusion method and compared to solo Se NPs, CS, and Alg/CS.
In addition, Se NPs (MIC) was mixed with 2% panthenol cream and tested against C. albicans ATCC10231 compared to solo 2% panthenol cream. The increase in fold area for Se NPs mixed with panthenol or CS was determined using the equation (B2 − A2)/A2, where A and B were inhibition zones of the anticandidal agent alone (2% panthenol) and combined with Se NPs, respectively [51].
2.15 Statistical analysis
Statistical analysis was performed on the data using SPSS version 18 software. Using One-way Analysis of Variance (ANOVA), all experiment values were presented as the mean ± standard deviation (SD). A significant threshold of p < 0.05 was applied [52].
3 Results
3.1 Isolation, purification, and phenotypic characterization of bacteria isolates
Thirteen Gram-positive, catalase-negative, non-spore-forming bacteria were recorded from the total bacterial isolates from zabady. Citric acid was positive in only 4 of them. When examined under a microscope, different morphological characteristics of bacterial cells were noted such as coccus, coccobacillus, or bacillus which are found in clusters, variable-length chains. Based on the difference in cell morphology, six LAB isolates (ESA1, ESA3, ESA5, ESA8, ESA12, and ESA13) were selected and tested for the biosynthesis of Se NPs. ESA1 and ESA12 isolates could grow at various temperatures ranging from 15 to 45 °C. At 2–6% salt, all bacterial isolates grew, except for ESA5 and ESA12 which were suppressed at 6% salt (Table 1).
3.2 Extracellular biosynthesis of Se NPs using LAB isolates
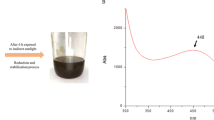
The first indication in the testing of Se NPs production by the LAB isolates is changing the medium color from pale yellow at the beginning of the experiment to red color at the end of the incubation period (Fig. 1). Among the selected isolates, ESA5 was able to produce Se NPs within 72 h at higher concentrations and faster rates than other isolates according to spectroscopy analysis results. The UV–Vis spectrum of ESA5-Se NPs showed an adsorption peak at 254 nm which matched with the characteristic properties of colloidal Se NPs [53].
3.3 Anticandidal action of LAB-Se NPs
Dried Se NPs produced using different LAB isolates were tested against C. albicans using the agar well diffusion method compared to the standard drug, miconazole. All LAB-Se NPs revealed inhibition zones higher than miconazole confirming their potentiality as a strong anticandidal agent (Fig. 2). However, Se NPs produced by ESA5 showed the highest inhibition zone compared to other LAB-Se NPs.
3.4 Molecular characterization of ESA5 strain
The ESA5 isolate was chosen as a potent bio-nano-factory for Se NPs production. To verify the identification of the bacterial isolate, a phylogenetic tree was constructed using the neighbor-joining method, and the 16S rDNA sequence was analyzed. After the full 16S rRNA gene of the ESA5 isolate was sequenced, BLAST analysis revealed a relationship between the strain and the Bacillota phyla based on a comparison with sequences found in the NCBI database. The Limosilactobacillus genus was associated with the ESA5 strain. Strain ESA5 indeed showed 100% similarity with the strain of the type Limosilactobacillus fermentum (Fig. 3). The partial 16S rDNA sequence gene of L. fermentum ESA5 was deposited to GenBank under accession number OR553490.
Phylogenetic tree based on 16S rRNA sequences of bacterial strain (Limosilactobacillus sp. ESA5). The number of branch nodes were bootstrap values (from 100 replicates) [54]. Bootstrap values above 50% are displayed. The genus Bacillus type strain was used as an out-group
3.5 Optimization of Se NPs produced by L. fermentum OR553490
Different parameters were tested for optimal biosynthesis of Se NPs including concentrations of Na2SeO3, the ratio between cell-free bacterial metabolites and Na2SeO3, and incubation periods (Fig. 4). The red color intensity of Se NPs production increases by increasing the concentration of Na2SeO3 from 1 to 5 mM which enhanced the biosynthesis formation of Se NPs (Fig. 4A). The mixing ratio between the cell-free bacterial metabolites and 5 mM Na2SeO3 in a 1:9 (v/v%) had the best and optimal production conditions for Se NPs formation compared to 1:11 (v/v%) (Fig. 4B). It was found that the production of Se NPs increased by increasing the incubation period (Fig. 4C).
3.6 Characterization of the optimized biosynthesized Se NPs
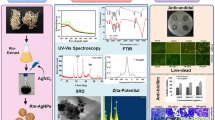
The FT-IR and XRD spectra were used to characterize the optimized Se NPs as well as TEM and Zeta analyses were used to study their shape, size, distribution, and potential (Fig. 5). FT-IR results of Se NPs displayed two main regions: functional group and fingerprint region (Fig. 5A). Stretching bands of symmetric (3392 cm−1 and 3292 cm−1) and asymmetric amines (2932 cm−1, 2899 cm−1 and 2299 cm−1) appeared at L. fermentum metabolites and Se NPs spectra, respectively. Aromatic and aliphatic C-N were assigned at 1643 cm−1, 1589 cm−1, 1535 cm−1, 1390 cm−1, and 1235 cm−1, respectively confirming the presence of proteins as stabilizing and capping agents during the biosynthesis process. Peaks at 2932 cm−1, 1643 cm−1, 1589 cm−1, 1535 cm−1, 1390 cm−1, and 1235 cm−1 in cell-free bacterial metabolites of L. fermentum were shifted to superior frequencies in the Se NPs pattern indicating the interaction of L. fermentum proteins with Se through the amine groups. The presence of NPs as a Se–O was confirmed by the metal–oxygen stretching vibrations that appeared at 1057 cm−1 [55]. The crystalline structure of Se NPs produced by L. fermentum was characterized by the XRD analysis (Fig. 5B). The results exhibited distinct characteristic peaks of Se NPs at angels 26.8°, 31.1°, 37.64°, 43.94°, 44.92°, 55.97°, 65.76°, 74.83° and 77.39° which corresponds to crystallographic planes of (100), (101), (110), (102), (111), (112), (210), (113), and (301), respectively. Lattice constants a = 4.357Å and c = 4.945Å of the obtained diffraction peaks assigned to the hexagonal structure of Se NPs which agree with the standard JCPDS data (JCPDS No. 06-0362). The absence of impurity diffraction peaks suggests that the final product was pure. Scherrer’s equation was used to evaluate the crystallite size of Se NPs: D = Kλ/(β cos θ); D: particle size, K: constant = 0.94, λ: x-ray wavelength, β: full width at half maximum and θ: the angle of diffraction. The crystallite size of Se NPs has been found to be ≈26 nm. TEM micrograph of Se NPs showed the successful biosynthesis of homogeneous rod-shaped NPs with average sizes ranging from 17 to 30 nm (Fig. 5C). The Zeta potential results showed the negative charge of the biosynthesized Se NPs (− 11.8 mV) as shown in Fig. 5D. The green synthetic Se NPs' energy dispersive X-ray spectrum confirms the presence of signals for nitrogen, elemental Se, sodium, oxygen, and carbon, as shown in Fig. 5E. The existence of strong signals proportional to elemental Se was exhibited by EDX spectrum. The signal of strong principal absorption, 1.37 keV was obtained for Se NPs. The peaks at 12.2, 11.2 keV, and 1.5 keV indicated the presence of elemental Se. Thus, elemental Se's existence verified the green reduction of selenium salt to elemental form. The remaining peaks were attributed to nitrogen (8.56%), carbon (33.66%), oxygen (19.81%), selenium (4.81%), and sodium (15.69%). The bioactive components, such as polysaccharides and proteins, that were attached to the surface of Se NPs were responsible for the extra peaks that followed those of nitrogen, oxygen, and carbon.
3.7 The anticandidal activity of the optimized Se NPs
The in vitro study showed a stronger anticandidal effect of the biosynthesized Se NPs than miconazole (Table 2 and Fig. 6). The MIC test was conducted for both Se NPs and Miconazole drug against C. albicans in the range of 5–40 µg/mL. The results showed that biosynthesized Se NPs had a MIC value of 25 µg/mL while the miconazole MIC value was 40 µg/mL (Fig. 6). The MFC results matched with MIC concentrations which confirmed the superior biocidal action of the biosynthesized Se NPs against C. albicans.
3.8 TEM study of C. albicans treated by Se NPs
The anticandidal effect of Se NPs against C. albicans was investigated using a TEM micrograph of Se NPs-treated C. albicans cells compared to untreated cells (Fig. 7). The normal cells of C. albicans appeared to have a normal cell wall, and cell membrane, intact cytoplasm with uniformly dense and homogeneous microstructure as well as small vacuole (Fig. 7A). In contrast, Se NPs exhibit several different morphological variations in the treated C. albicans cells. These changes included rupture of the cell wall, wrinkling appearance of the cell wall, noticeable separation between cell and cytoplasmic membrane, and formation of a big vacuole (Fig. 7B).
The biocidal action of Se NPs on the ultrastructure of C. albicans cells. A Untreated control cells (without NPs). B Treated cells with 25 µg/mL Se NPs (MIC). CW cell wall, CM cytoplasmic membrane, CY cytoplasm, L lipids formation, and V vacuoles. Note the visible separation between the cell and cytoplasmic membrane (white arrows) and the accumulation of NPs inside the treated cells. Bar scale = 500 nm
3.9 Cytotoxicity study of Se NPs
The biosynthesized Se NPs were applied and tested against normal keratinocyte skin cells to study their toxicity (Fig. 8). Se NPs revealed an IC50 ≈ 41.5 ± 0.9 μg/mL. This result confirmed the safety of using lower concentrations from the biosynthesized Se NPs including MIC concentration of Se NPs (25 μg/mL).
3.10 In vitro anticandidal potential of Se NPs as a based topical drug
The anticandidal action of Se NPs, Alg/Se, and Alg/CS/Se was studied against C. albicans using the agar well diffusion method (Table 3 and Fig. 9A). In addition, Se NPs mixed with panthenol (2%) had a stronger anticandidal action against C. albicans (43 mm) than solo panthenol cream (20 mm) as shown in Fig. 9B. The biosynthesized Se NPs displayed a significant effect when companies with CS and panthenol causing an increase in the fold area reached to 9.3 and 3.6, respectively.
4 Discussion
Candida albicans is one of the most serious fungal infections due to causing several diseases for the whole part of the body and the most dangerous problem is the resistance to antifungal agents. It can infect animals as well as humans and cause serious diseases such as candidiasis, which cause serious infection according to the infection site, cutaneous candidiasis, oral and gastrointestinal mucosal candidiasis, and vaginal canal [8]. Although there are several types of antifungal drugs such as clotrimazole, econazole, miconazole, terbinafine, fluconazole, ketoconazole, and amphotericin, were extreme irritants and might be lethal, many studies reported that azole antifungal resistance cases of C. albicans [56, 57]. This study aims to find an alternative green approach to overcome candidal infection problems especially C. albicans resistance to current commercial drugs.
Nanomaterials have piqued the interest of scientists due to their applications in chemistry, medicine, and other sciences. Among nanomaterials, nanometals of selenium, silver, copper, and gold display novel chemical, physical, and biological characteristics that were used as strong anticandidal agents [58,59,60,61]. Recently, biological methods for NPs production have made tremendous progress due to their low cost, simplicity, and quick procedure. Selenite reduction by microorganisms has received considerable attention among the many forms of selenium due to its toxicity [62]. In the current work, Se NPs were extracellularly biosynthesized using the cell-free supernatant of L. fermentum OR553490 as a safe, cheap, and simple one-step. Classical, biochemical, and molecular techniques have been used to confirm the identification of the L. fermentum strain at the species level. The formation of red color by the cell-free supernatant of L. fermentum implies that this bacterium's crude metabolites may bio-reduce poisonous and colorless selenite to nontoxic and red metallic Se NPs. The production of a distinctive red color was caused by the excitation of surface plasmon vibrations of Se [63]. Even so, the exact approach by which bacteria produce Se NPs has not yet been recognized. Previous investigations demonstrated that NADH and the NADH-dependent nitrate reductase enzyme play essential roles in the forming of metal nanoparticles. Dwivedi et al. [64] suggested that NADH and NADH-dependent reductases as redox agents are responsible for the reduction of metal ions of selenite to selenium nanospheres using the culture supernatant of Pseudomonas aeruginosa. Reductases can also act as capping agents, ensuring the production of thermodynamically stable nanostructures [65]. L. fermentum was reported to produce NADH and NADH-dependent reductase enzymes [66,67,68,69]. As a result, it is anticipated that a multi-component redox system in the cell-free supernatant of L. fermentum, comprising NADH, and most likely NADH-dependent reductases, would function to catalyze the biosynthesis of Se NPs.
The biosynthesis of Se NPs was optimized by testing different parameters such as concentrations of Na2SeO3, the ratio between cell-free bacterial metabolites and Na2SeO3, and incubation periods. It was recorded that mixing of mixing of cell-free bacterial metabolites with 5 mM Na2SeO3 in a 1:9 (v/v%) for 72 h were the best conditions for the biosynthesis of Se NPs whereas higher mixing ratio (1:11 v/v%) did not favorable for the Se NPs formation due to the excess amount of reducing agents [70, 71]. El-Dein et al. [72] recorded that the mixing ratio of 1:3 v/v% enhanced the production of NPs using bacterial metabolites. However, the red color intensity increased by the incubation time which refers to the high reduction rate, L. fermentum started to change the yellowish media color after the first 12 h and gave a high yield of Se NPs after 72 h. The small amount production of Se NPs at lower concentrations of Na2SeO3 solutions (1, 3 mM) might be due to the low amounts of enzymes' substrate, while higher concentration of Na2SeO3 solution (7 mM) did not enhance the Se NPs formation due to its toxicity [65, 73]. El-Saadony et al. [64] obtained different results in the optimization condition for the probiotic bacteria L. paracasei. The best concentration of Na2SeO3 was 4 mM and the reaction mixture was incubated for 32 h. The ratio between Na2SeO3 and the cell-free supernatant of L. fermentum was increasable relation, increasing the salt concentration with the cell-free supernatant of L. fermentum that is rich with reducing enzymes led to increasing the Se NPs production.
The biosynthesis of Se NPs was confirmed by the presence of an absorbance peak at ≈254 nm. Several studies reported different absorbance peaks for Se NPs for example, Hemalatha et al. [73] studied Se NPs which have an absorbance peak of 290 nm while El-Saadony et al. [74] reported other results at 263 nm and 300 nm, respectively. The absorbance peak changes towards longer wavelengths as particle size increases [53]. Only when the particle size is 100 nm or larger, Se NPs display a consistent absorption peak in the wavelength band above 300 nm. The absorption peak typically changes towards red, and the peak intensity diminishes as nanoparticle size increases [75].
There are common problems related to the stability of NPs. Agglomeration and aggregation of NPs are some of the biggest problems that decrease their use in different applications. Capping agents have a noticeable role in preventing NPs aggregation [76]. The FT-IR analysis confirmed the presence of proteins as capping agents that surrounded the NPs. Some studies confirm that the binding of protein capping agents increases the stability of NPs through cysteine and amine residues, which prevent the accumulation and aggregation of NPs [77]. Also, the amide III band of proteins was documented during the use of L. acidophilus in the biosynthesis of Se NPs [18]. This capping agent may have negative or positive charges that also have a relative role in the stability of NPs and important characteristics of the colloidal dispersion of NPs. Se NPs-L. fermentum was found to have a negative charge of -11.8mV which causes a repulsion force between NPs grains and increases their stability. Laslo et al. [34] reported that L. casei produced Se NPs with a maximum value of Zeta potential of – 23 mV. It is worth noting that the zeta potential surface charge of NPs also has a great role in antimicrobial activity through the electrostatic adhesion interaction between NPs and microbial cell membranes [78]. Different bacteria such as L. lactis, Lactobacillus sp., and Bifidobacter sp. were used in Se NPs production as a green nano-factory, however, one of their disadvantages was the large nano-sized particles which ranged from 100 nm-550 nm [33]. In contrast, L. fermentum produced Se NPs with a size range of 17–30 nm. The XRD results confirmed purity and crystalline nature of the produced Se NPs. The XRD patterns of the L. fermentum-Se NPs matched with L. paracasei-Se NPs which showed eight peaks at 2θ of 28.61°, 31.19°, 40.01°, 45.02°, 56.21°, 66.23°, 75.11°, and 84.74° which corresponds to crystallographic planes of 100, 101, 110, 102, 111, 201, 112 and 202 [64].
The biosynthesized Se NPs were reported by several studies as a strong antimicrobial agent [53, 77, 79, 80]. Similarly, the biosynthesized Se NPs using L. fermentum had stronger biocidal action against C. albicans with MIC and MFC values of 25 µg/mL compared to the standard drug miconazole (40 µg/mL). El-Saadony et al. [64] reported a MIC and MFC of 55 and 80 μg/mL, respectively of LAB-Se NPs against C. albicans. Shakibaie et al. [81] biosynthesized Se NPs by using Bacillus species Msh1 and had MIC of 70 μg/mL against C. albicans. Besides the low MIC value of the biosynthesized Se NPs, the cytotoxic effect was performed on the normal keratinocyte cells by the trypan blue method. A low toxic effect of Se NPs with IC50 value ≈ 41.5 μg/mL.
Nevertheless, the exact biocidal mode of action of NPs has not been determined, several studies recommended that selenium can increase the ROS formation. Small-sized reduced selenium ions can infuse easily through the cell wall of the fungus, attach, and affect negatively the respiratory sequence and ATP [82]. In addition, selenium has a denaturation action by increasing the superoxide radical levels and causes an increase the fungal cell death [83]. TEM micrographs of treated C. albicans by Se NPs showed the accumulation of Se NPs inside the treated cells in the cytoplasmic membrane which might cause the interactions between the microbial cellular components and NPs. This treatment led to numerous modifications in the treated cells compared to normal cells. These changes involved morphological abnormalities in the cell wall and cytoplasmic membrane, decreasing cytoplasmic and DNA content, large vacuoles, and lipids formation.
The current study also investigated the potent anticandidal potential of Se NPs in combination with CS and panthenol. Both CS and panthenol were recommended and used as anticandidal agents against different pathogenic microbes [16, 84]. However, many Candida species recently showed several resistant mechanisms against these antimicrobial agents include active efflux systems and decrease the antimicrobial agent accumulation inside their cells [85, 86]. Alg was reported to enhance the antifungal activity against candidal biofilms [87]. Alg was also used to reduce the toxicological effects of NPs and to expand its clinical uses in nanocarriers and drug delivery systems [88]. Kalińska et al. [89] in vitro study displayed that the combination of NPs such silver and copper NPs with cosmetics such as panthenol, vitamin C, sodium lactate, and marigold flower extract act as strong antimicrobial agents compared to their solo components. When CS and panthenol were combined with Se NPs, there was a strong anticandidal effect against C. albicans with an increase in fold area of 9.3 and 3.6, respectively. These results increase the possibilities of using the biosynthesized Se NPs in different medical and industrial applications as promising solo or combined anticandidal agents.
5 Conclusion
The current study highlighted a novel treatment for dangerous candidal infections using Se NPs. A cell-free supernatant of L. fermentum (OR553490) was used in the extracellular biosynthesis of Se NPs. 5 mM Na2SeO3 and 1:9 (v/v%) mixing ratio between Na2SeO3 and L. fermentum metabolite enhanced the bio-production of Se NPs for 72 h. The biosynthesized Se NPs using L. fermentum revealed stronger anticandidal action against C. albicans compared to Se NPs synthesized by other LABs. They mark their effectiveness as anticandidal at lower concentrations, moreover, having a low toxic effect on normal cells. The obtained results recommended the combination between L. fermentum Se NPs with CS and/or panthenol as developed potential anticandidal agents to compete the C. albicans resistance problems. Furthermore, the toxicity and anticandidal action of Se NPs needs to be studied in vivo with an animal model.
Data availability
The partial sequence of the 16S ribosomal RNA gene of Limosilactobacillus fermentum strain ESA5 obtained in the current study was deposited in the NCBI GenBank database under accession number: OR553490 and is available at the following URL: https://www.ncbi.nlm.nih.gov/nuccore/OR553490. The datasets generated during the current study are available from the corresponding author upon reasonable request.
Abbreviations
- Se NPs:
-
Selenium nanoparticles
- CS:
-
Chitosan
- Alg:
-
Alginate
- UV–Vis:
-
Ultraviolet–visible
- FT-IR:
-
Fourier transform-infrared spectroscopy
- XRD:
-
The X-ray diffraction
- TEM:
-
Transmission electron microscopy
- EDS:
-
Elemental analysis system
- IC50 :
-
Half maximal inhibitory concentration
- ANOVA:
-
One-way analysis of variance
References
Contaldo M, Di Stasio D, Romano A, Fiori F, Della Vella F, Rupe C, Lajolo C, Petruzzi M, Serpico R, Lucchese A. Oral candidiasis and novel therapeutic strategies: Antifungals, phytotherapy, probiotics, and photodynamic therapy. Curr Drug Deliv. 2023;20(5):441–56. https://doi.org/10.2174/1567201819666220418104042.
Jenkinson HF, Douglas LJ. Interactions between Candida species and bacteria in mixed infections. In: Brogden KA, Guthmiller JM (Eds). Polymicrobial diseases, 2002; 357–373. ASM Press, Wiley, Inc https://doi.org/10.1128/9781555817947.ch18
Li H, Miao MX, Jia CL, Cao YB, Yan TH, Jiang YY, Yang F. Interactions between Candida albicans and the resident microbiota. Front Microbiol. 2022;13:930495. https://doi.org/10.3389/fmicb.2022.930495.
Cavalcanti IM, Del Bel Cury AA, Jenkinson HF, Nobbs AH. Interactions between Streptococcus oralis, Actinomyces oris, and Candida albicans in the development of multispecies oral microbial biofilms on salivary pellicle. Mol Oral Microbiol. 2017;32(1):60–73. https://doi.org/10.1111/omi.12154.
Parums DV. The World Health Organization (WHO) fungal priority pathogens list in response to emerging fungal pathogens during the COVID-19 pandemic. Med Sci Monit. 2022;28:939088–91. https://doi.org/10.12659/MSM.939088.
Yapar N. Epidemiology and risk factors for invasive candidiasis. Ther Clin Risk Manag. 2014;10:95–105. https://doi.org/10.2147/TCRM.S40160.
Pappas PG, Lionakis MS, Arendrup MC, Ostrosky-Zeichner L, Kullberg BJ. Invasive candidiasis. Nat Rev Dis Primers. 2018;4(1):1–20. https://doi.org/10.1038/nrdp.2018.26.
Talapko J, Juzbašić M, Matijević T, Pustijanac E, Bekić S, Kotris I, Škrlec I. Candida albicans-the virulence factors and clinical manifestations of infection. J Fungi. 2021;7(2):1–19. https://doi.org/10.3390/jof7020079.
Moudgal V, Sobel J. Antifungals to treat Candida albicans. Expert Opin Pharmacother. 2010;11(12):2037–48. https://doi.org/10.1517/14656566.2010.493875.
Du W, Gao Y, Liu L, Sai S, Ding C. Striking back against fungal infections: the utilization of nanosystems for antifungal strategies. Int J Mol Sci. 2021. https://doi.org/10.3390/ijms221810104.
Manimaran M, Muthuvel A, Said NM. Microwave-assisted green synthesis of SnO2 nanoparticles and their photocatalytic degradation and antibacterial activities. Nanotechnol Environ Eng. 2023;8:413–23. https://doi.org/10.1007/s41204-022-00297-3.
Robles-García MA, Rodríguez-Félix F, Marquez-Rios E, Aguilar JA, Barrera-Rodríguez A, Aguilar J, Ruiz-Cruz S, Del-Toro-Sánchez CL. Applications of nanotechnology in the agriculture, food, and pharmaceuticals. JNN. 2016;16(8):8188–207. https://doi.org/10.1166/jnn.2016.12925.
Saqib S, Nazeer A, Ali M, Zaman W, Younas M, Shahzad A, Sunera NM. Catalytic potential of endophytes facilitates synthesis of biometallic zinc oxide nanoparticles for agricultural application. Biometals. 2022;35:967–85. https://doi.org/10.1007/s10534-022-00417-1.
Saqib S, Zaman W, Ayaz A, Habib S, Bahadur S, Hussain S, Muhammad S, Ullah F. Postharvest disease inhibition in fruit by synthesis and characterization of chitosan iron oxide nanoparticles. ISBAB. 2020;28: 101729. https://doi.org/10.1016/j.bcab.2020.101729.
Muthuvel A, Adavallan K, Balamurugan K, Krishnakumar N. Biosynthesis of gold nanoparticles using Solanum nigrum leaf extract and screening their free radical scavenging and antibacterial properties. Biomed Prev Nutr. 2014;4(2):325–32. https://doi.org/10.1016/j.bionut.2014.03.004.
Slavin YN, Bach H. Mechanisms of antifungal properties of metal nanoparticles. Nanomaterials. 2022;12:4470. https://doi.org/10.3390/nano12244470.
Lin W, Zhang J, Xu J-F, Pi J. The advancing of selenium nanoparticles against infectious diseases. Front Pharmacol. 2021. https://doi.org/10.3389/fphar.2021.682284.
Alam H, Khatoon N, Khan MA, Husain SA, Saravanan M, Sardar M. Synthesis of selenium nanoparticles using probiotic bacteria Lactobacillus acidophilus and their enhanced antimicrobial activity against resistant bacteria. J Clust Sci. 2020;31(5):1003–11. https://doi.org/10.1007/s10876-019-01705-6.
Salem SS. A mini review on green nanotechnology and its development in biological effects. Arch Microbiol. 2023;205(4):128. https://doi.org/10.1007/s00203-023-03467-2.
Ferreira RLU, Sena-Evangelista KCM, de Azevedo EP, Pinheiro FI, Cobucci RN, Pedrosa LFC. Selenium in human health and gut microflora: bioavailability of selenocompounds and relationship with diseases. Front Nutr. 2021;8: 685317. https://doi.org/10.3389/fnut.2021.685317.
Balakrishnaraja R. Comparison studies on the synthesis of selenium nanoparticles by various micro-organisms. Int J Pure Appl Biosci. 2014;2:112–7.
Salem SS, Fouda A. Green synthesis of metallic nanoparticles and their prospective biotechnological applications: an overview. Biol Trace Elem Res. 2021;199:344–70. https://doi.org/10.1007/s12011-020-02138-3.
Fayed R, Elnemr A-M, El-Zahed MM. Synthesis, characterization, antimicrobial, and electrochemical studies of biosynthesized zinc oxide nanoparticles using the probiotic Bacillus coagulans (ATCC 7050). J Microbiol Biotechnol Food Sci. 2023;13(3):9962. https://doi.org/10.55251/jmbfs.9962.
Ahmad N, Ali S, Abbas M, Fazal H, Saqib S, Ali A, Ullah Z, Zaman S, Sawati L, Zada A, Sohail. Antimicrobial efficacy of Mentha piperata-derived biogenic zinc oxide nanoparticles against UTI-resistant pathogens. Sci Rep. 2023;13(1):14972. https://doi.org/10.1038/s41598-023-41502-w.
Al-Zaqri N, Muthuvel A, Jothibas M, Alsalme A, Alharthi FA, Mohana V. Biosynthesis of zirconium oxide nanoparticles using Wrightia tinctoria leaf extract: characterization, photocatalytic degradation and antibacterial activities. Inorg Chem Commun. 2021;127: 108507. https://doi.org/10.1016/j.inoche.2021.108507.
Saqib S, Faryad S, Afridi MI, Arshad B, Younas M, Naeem M, Zaman W, Ullah F, Nisar M, Ali S, Elgorban AM. Bimetallic assembled silver nanoparticles impregnated in Aspergillus fumigatus extract damage the bacterial membrane surface and release cellular contents. Coatings. 2022;12(10):1505. https://doi.org/10.3390/coatings12101505.
Moghaddam AB, Moniri M, Azizi S, Rahim RA, Bin AA, Saad WZ, Namvar F, Navaderi M, Mohamad R. Biosynthesis of ZnO nanoparticles by a new Pichia kudriavzevii yeast strain and evaluation of their antimicrobial and antioxidant activities. Molecules. 2017. https://doi.org/10.3390/molecules22060872.
Sharif MS, Hameed H, Waheed A, Tariq M, Afreen A, Kamal A, Mahmoud EA, Elansary HO, Saqib S, Zaman W. Biofabrication of Fe3O4 nanoparticles from Spirogyra hyalina and Ajuga bracteosa and their antibacterial applications. Molecules. 2023;28(8):3403. https://doi.org/10.3390/molecules28083403.
Krishnan S, Bhardwaj SK, Liu S, Xing R, Chavali M. Virus-assisted biological methods for greener synthesis of nanomaterials. Handbook of Greener synthesis of nanomaterials and compounds. Fundam Prin Methods. 2021;1:785–806. https://doi.org/10.1016/B978-0-12-821938-6.00025-6.
Bisht N, Phalswal P, Khanna PK. Selenium nanoparticles: a review on synthesis and biomedical applications. Mater Adv. 2022;3(3):1415–31. https://doi.org/10.1039/d1ma00639h.
El-Zahed MM, Baka ZA, Abou-Dobara MI, El-Sayed AK. Biogenic synthesis of chitosan/silver nanocomposite by Escherichia coli D8 (MF062579) and its antibacterial activity. Jordan J Biol Sci. 2023;16(2):279–88. https://doi.org/10.54319/jjbs/160212.
Heidari Z, Faezi Ghasemi M, Modiri L. The synergistic antibacterial effect of bacteriocin produced by Lactobacillus casei ATCC 39392 and iron oxide nanoparticles (IONPs) on selected foodborne pathogens. IJMCM. 2020;10(1):1301–11.
Stabnikova O, Khonkiv M, Kovshar I, Stabnikov V. Biosynthesis of selenium nanoparticles by lactic acid bacteria and areas of their possible applications. World J Microbiol Biotechnol. 2023;39(9):230. https://doi.org/10.1007/s11274-023-03673-6.
Laslo V, Pinzaru SC, Zaguła G, Kluz M, Vicas SI, Cavalu S. Synergic effect of selenium nanoparticles and lactic acid bacteria in reduction cadmium toxicity. J Mol Struct. 2022;1247: 131325. https://doi.org/10.1016/j.molstruc.2021.131325.
Qiao L, Dou X, Song X, Chang J, Zeng X, Zhu L, Xu C. Selenite Bioremediation by food-grade probiotic Lactobacillus casei ATCC 393: insights from proteomics analysis. Microbiol Spectr. 2023;11: e0065923. https://doi.org/10.1128/spectrum.00659-23.
He L, Chen Y, Zhang H, Wang H, Chen S, Liu S, Liu A, Li Q, Ao X, Liu Y. Isolation and identification of Lactobacillus and yeast species and their effect on the quality of fermented rice cakes. Innov Food Sci Emerg Technol. 2022;77: 102984. https://doi.org/10.1016/J.IFSET.2022.102984.
Altalhi AA. Antilisterial activity of plantaricin UG1 during manufacture of Zabady and Kareesh cheese: two Arabian dairy products. Int J Biomed Sci. 2008;4(4):319–22.
Rajasree SR, Sathyamoorthy G. Extracellular biosynthesis of selenium nanoparticles using some species of Lactobacillus. Indian J Mar Sci. 2015;44:766–75.
Gordana Z, Zeljka R, Valentina V, Jelena B, Topisirovic L, Ivana S. Characterization and antimicrobial activity of vaginal Lactobacillus isolate. Arch Biol Sci Belgrade. 2011;63(1):29–35. https://doi.org/10.2298/ABS1101029Z.
Clinical and Laboratory Standards Institute (CLSI). Method for antifungal disk diffusion susceptibility testing of yeasts; approved guideline, 2nd ed., M44-A2. Clinical and Laboratory Standards Institute, 2009; Wayne, PA, USA.
Abdulamir AS, Yoke TS, Nordin N, Bakar FA. Detection and quantification of probiotic bacteria using optimized DNA extraction, traditional and real-time PCR methods in complex microbial communities. Afr J Biotechnol. 2010;9(10):1481–92. https://doi.org/10.5897/AJB09.1322.
Lagacé L, Pitre M, Jacques M, Roy D. Identification of the bacterial community of maple sap by using amplified ribosomal DNA (rDNA) restriction analysis and rDNA sequencing. Appl Environ Microbiol. 2004;70(4):2052–60. https://doi.org/10.1128/AEM.70.4.2052-2060.2004.
Altschul SF, Gish W, Miller W, Myers EW, Lipmanl DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–10. https://doi.org/10.1016/S0022-2836(05)80360-2.
Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22(22):4673–80. https://doi.org/10.1093/nar/22.22.4673.
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol. 2013;30:2725–9. https://doi.org/10.1093/molbev/mst197.
Urshev Z, Doynova D, Prasev I, Denkova-Kostova R, Koleva A, Denkova Z, Goranov B, Kostov G. identification of lactic acid bacteria strains isolated from Sourdoughs prepared with different flour types. Appl Sci. 2024;14(5):2093. https://doi.org/10.3390/app14052093.
CLSI. Reference method for broth dilution antifungal susceptibility testing of yeasts; fourth informational supplement, M27-S4. 2012; Wayne, PA, USA
Cantón E, Pemán J, Viudes A, Quindós G, Gobernado M, Espinel-Ingroff A. Minimum fungicidal concentrations of amphotericin B for bloodstream Candida species. Diagn Microbiol Infect Dis. 2003;45(3):203–6. https://doi.org/10.1016/S0732-8893(02)00525-4.
Staniszewska M, Bondaryk M, Siennicka K, Kurzatkowski W. Ultrastructure of Candida albicans pleomorphic forms: phase-contrast microscopy, scanning and transmission electron microscopy. Pol J Microbiol. 2012;61(2):129.
Strober W. Trypan blue exclusion test of cell viability. Curr Protoc Immunol. 1997;21(1):A3B. https://doi.org/10.1002/0471142735.ima03bs21.
Birla SS, Tiwari VV, Gade AK, Ingle AP, Yadav AP, Rai M. Fabrication of silver nanoparticles by Phoma glomerata and its combined effect against Escherichia coli, Pseudomonas aeruginosa and Staphylococcus aureus. Lett Appl Microbiol. 2009;48:173–9. https://doi.org/10.1111/j.1472-765X.2008.02510.x.
OConnor BP. SPSS and SAS programs for determining the number of components using parallel analysis and Velicer’s MAP test. Behav Res Methods Instrum Comput. 2000;32(3):396–402. https://doi.org/10.3758/BF03200807.
Shah CP, Dwivedi C, Singh KK, Kumar M, Bajaj PN. Riley oxidation: a forgotten name reaction for synthesis of selenium nanoparticles. Mater Res Bull. 2010;45(9):1213–7. https://doi.org/10.1016/j.materresbull.2010.05.013.
Felsenstein J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution. 1985;39(4):783–91. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x.
Gunti L, Dass RS, Kalagatur NK. Phytofabrication of selenium nanoparticles from Emblica officinalis fruit extract and exploring its biopotential applications: antioxidant, antimicrobial, and biocompatibility. Front Microbiol. 2019;10:931. https://doi.org/10.3389/fmicb.2019.00931.
Vanden-Bossche H, Engelen M, Rochette F. Antifungal agents of use in animal health—chemical, biochemical and pharmacological aspects. J Vet Pharmacol Ther. 2003;26(1):5–29. https://doi.org/10.1046/j.1365-2885.2003.00456.x.
Whaley SG, Berkow EL, Rybak JM, Nishimoto AT, Barker KS, Rogers PD. Azole antifungal resistance in Candida albicans and emerging non-albicans Candida Species. Front Microbiol. 2017;7(1):2173. https://doi.org/10.3389/fmicb.2016.02173.
Carmo PHFD, Garcia MT, Figueiredo-Godoi LMA, Lage ACP, Silva NSD, Junqueira JC. Metal nanoparticles to combat Candida albicans infections: an update. Microorganisms. 2023;11(1):138. https://doi.org/10.3390/microorganisms11010138.
Kokila K, Elavarasan N, Sujatha V. Diospyros montana leaf extract-mediated synthesis of selenium nanoparticles and their biological applications. New J Chem. 2017;41(15):7481–90. https://doi.org/10.1039/C7NJ01124E.
Lara HH, Romero-Urbina DG, Pierce C, Lopez-Ribot JL, Arellano-Jiménez MJ, Jose-Yacaman M. Effect of silver nanoparticles on Candida albicans biofilms: an ultrastructural study. J nanobiotech. 2015;13(1):1–12. https://doi.org/10.1186/s12951-015-0147-8.
Tabassum N, Jeong GJ, Jo DM, Khan F, Kim YM. Treatment of Staphylococcus aureus and Candida albicans polymicrobial biofilms by phloroglucinol-gold nanoparticles. Microb Pathog. 2023. https://doi.org/10.1016/j.micpath.2023.106416.
Zhang J, Wang H, Yan X, Zhang L. Comparison of short-term toxicity between Nano-Se and selenite in mice. Life Sci. 2005;76(10):1099–109. https://doi.org/10.1016/j.lfs.2004.08.015.
Lin ZH, Wang CC. Evidence on the size-dependent absorption spectral evolution of selenium nanoparticles. Mater Chem Phys. 2005;92(2–3):591–4. https://doi.org/10.1016/j.matchemphys.2005.02.023.
El-Saadony MT, Saad AM, Taha TF, Najjar AA, Zabermawi NM, Nader MM, AbuQamar SF, El-Tarabily KA, Salama A. Selenium nanoparticles from Lactobacillus paracasei HM1 capable of antagonizing animal pathogenic fungi as a new source from human breast milk. Saudi J Biol Sci. 2021;28(12):6782–94. https://doi.org/10.1016/J.SJBS.2021.07.059.
Dwivedi S, AlKhedhairy AA, Ahamed M, Musarrat J. Biomimetic synthesis of selenium nanospheres by bacterial strain JS-11 and its role as a biosensor for nanotoxicity assessment: a novel Se-bioassay. PLoS ONE. 2013;8(3): e57404. https://doi.org/10.1371/journal.pone.0057404.
He S, Guo Z, Zhang Y, Zhang S, Wang J, Gu N. Biosynthesis of gold nanoparticles using the bacteria Rhodopseudomonas capsulata. Mater Lett. 2007;61(18):3984–7. https://doi.org/10.1016/j.matlet.2007.01.018.
Aarnikunnas J, Von Weymarn N, Rönnholm K, Leisola M, Palva A. Metabolic engineering of Lactobacillus fermentum for production of mannitol and pure l-lactic acid or pyruvate. Biotechnol Bioeng. 2003;82(6):653–63. https://doi.org/10.1002/bit.10615.
Park YC, Oh EJ, Jo JH, Jin YS, Seo JH. Recent advances in biological production of sugar alcohols. Curr Opin Biotechnol. 2016;37:105–13. https://doi.org/10.1016/j.copbio.2015.11.006.
Balakumaran MD, Ramachandran R, Balashanmugam P, Mukeshkumar DJ, Kalaichelvan PT. Mycosynthesis of silver and gold nanoparticles: optimization, characterization and antimicrobial activity against human pathogens. Microbiol Res. 2016;182:8–20. https://doi.org/10.1016/j.micres.2015.09.009.
Xu SY, Zhou L, Xu Y, Hong HY, Dai C, Wang YJ, Zheng YG. Recent advances in structure-based enzyme engineering for functional reconstruction. Biotechnol Bioeng. 2023;120(12):3427–45. https://doi.org/10.1002/bit.28540.
Wen S, Hui Y, Chuang W. Biosynthesis and antioxidation of nano-selenium using lemon juice as a reducing agent. Green Process Synth. 2021;10(1):178–88. https://doi.org/10.1515/gps-2021-0018.
El-Dein MM, Baka ZAM, Abou-Dobara MI, El-Sayed AK, El-Zahed MM. Extracellular biosynthesis, optimization, characterization and antimicrobial potential of Escherichia coli D8 silver nanoparticles. J Microbiol Biotechnol Food Sci. 2021;10(4):648–56. https://doi.org/10.15414/jmbfs.2021.10.4.648-656.
Hemalatha T, Krithiga G, Santhosh Kumar B, Sastry TP. Preparation and characterization of hydroxyapatite-coated selenium nanoparticles and their interaction with osteosarcoma (SaOS-2) cells. Acta Metall Sin. 2014;27(6):1152–8. https://doi.org/10.1007/s40195-014-0153-0.
El-Saadony MT, Saad AM, Najjar AA, Alzahrani SO, Alkhatib FM, Shafi ME, Selem E, Desoky E-SM, Fouda SEE, El-Tahan AM, Hassan MAA. The use of biological selenium nanoparticles to suppress Triticum aestivum L. crown and root rot diseases induced by Fusarium species and improve yield under drought and heat stress. Saudi J Biol Sci. 2021;28(8):4461–71. https://doi.org/10.1016/j.sjbs.2021.04.043.
Kaur G, Bakshi MS. Nonideal mixing of Se–Te in aqueous micellar phase for nanoalloys over the whole mole mixing range with morphology control from nanoparticles to nanoribbons. J Phys Chem C. 2010;114(1):143–54. https://doi.org/10.1021/jp9086249.
El-Zahed MM, Baka ZA, Abou-Dobara MI, El-Sayed AK, Aboser MM, Hyder A. In vivo toxicity and antitumor activity of newly green synthesized reduced graphene oxide/silver nanocomposites. Bioresour Bioprocess. 2021;8(1):44. https://doi.org/10.1186/s40643-021-00400-7.
Escobar-Ramírez MC, Castañeda-Ovando A, Pérez-Escalante E, Rodríguez-Serrano GM, Ramírez-Moreno E, Quintero-Lira A, Contreras-López E, Añorve-Morga J, Jaimez-Ordaz J, González-Olivares LG. Antimicrobial activity of se-nanoparticles from bacterial biotransformation. Fermentation. 2021. https://doi.org/10.3390/fermentation7030130.
Hnain A, Brooks J, Lefebvre DD. The synthesis of elemental selenium particles by Synechococcus leopoliensis. Appl Microbiol Biotechnol. 2013;97(24):10511–9. https://doi.org/10.1007/s00253-013-5304-0.
Pescuma M, Aparicio F, Zysler RD, Lima E, Zapata C, Marfetán JA, Vélez ML, Ordoñez OF. Biogenic selenium nanoparticles with antifungal activity against the wood-rotting fungus Oligoporus pelliculosus. Biotechnol Rep. 2023;37:00787. https://doi.org/10.1016/j.btre.2023.e00787.
Yang J, Wang J, Yang K, Liu M, Qi Y, Zhang T, Fan M, Wei X. Antibacterial activity of selenium-enriched lactic acid bacteria against common food-borne pathogens in vitro. J Dairy Sci. 2018;101:1930–42. https://doi.org/10.3168/jds.2017-13430.
Shakibaie M, Mohazab NS, Ayatollahi Mousavi SA. Antifungal activity of selenium nanoparticles synthesized by Bacillus species Msh-1 against Aspergillus fumigatus and Candida albicans. Jundishapur J Microbiol. 2015;8(9): e26381. https://doi.org/10.5812/jjm.26381.
Ogunsona EO, Muthuraj R, Ojogbo E, Valerio O, Mekonnen TH. Engineered nanomaterials for antimicrobial applications: a review. Appl Mater Today. 2020;18: 100473. https://doi.org/10.1016/j.apmt.2019.100473.
Vera-González N, Shukla A. Advances in biomaterials for the prevention and disruption of Candida biofilms. Front Microbiol. 2020;11: 538602. https://doi.org/10.3389/fmicb.2020.538602.
Mencucci R, Favuzza E, Bottino P, Mazzantini C, Zanotto E, Pellegrini-Giampietro DE, Landucci E. A new ophthalmic formulation containing antiseptics and dexpanthenol: in vitro antimicrobial activity and effects on corneal and conjunctival epithelial cells. Exp Eye Res. 2020;201: 108269. https://doi.org/10.1016/j.exer.2020.108269.
Garcia LG, de Melo Guedes GM, da Silva ML, Castelo-Branco DS, Sidrim JJ, de Aguiar CR, Rocha MF, Vieira RS, Brilhante RS. Effect of the molecular weight of chitosan on its antifungal activity against Candida spp. in planktonic cells and biofilm. Carbohydr Polym. 2018;195:662–9. https://doi.org/10.1016/j.carbpol.2018.04.091.
Kołaczkowska A, Kołaczkowski M. Drug resistance mechanisms and their regulation in non-albicans Candida species. J Antimicrob Chemother. 2016;71(6):1438–50. https://doi.org/10.1093/jac/dkv445.
Powell LC, Adams JY, Quoraishi S, Py C, Oger A, Gazze SA, Francis LW, von Ruhland C, Owens D, Rye PD, Hill KE. Alginate oligosaccharides enhance the antifungal activity of nystatin against candidal biofilms. Front Cell Infect Microbiol. 2023;13:1122340. https://doi.org/10.3389/fcimb.2023.1122340.
Spadari CD, de Bastiani FW, Lopes LB, Ishida K. Alginate nanoparticles as non-toxic delivery system for miltefosine in the treatment of candidiasis and cryptococcosis. Int J Nanomedicine. 2019;12:5187–99. https://doi.org/10.2147/IJN.S205350.
Kalińska A, Jaworski S, Wierzbicki M, Kot M, Radzikowski D, Smulski S, Gołębiewski M. Silver and copper nanoparticles as the new biocidal agents used in pre-and post-milking disinfectants with the addition of cosmetic substrates in dairy cows. Int J Mol Sci. 2023;24(2):1658. https://doi.org/10.3390/ijms24021658.
Acknowledgements
The authors are grateful to the Microbiology Lab, Botany and Microbiology Department, Damietta University for providing the facilities for conducting this research.
Funding
Supported by the Academy of Scientific Research and Technologies (ASRT).
Author information
Authors and Affiliations
Contributions
Conceptualization and Methodology: Mohamed El‑Zahed, Esraa Mohamed; Data Curation and Formal Analysis: Mohamed El‑Zahed, Esraa Mohamed; Investigation and Resources: Mohamed El‑Zahed, Esraa Mohamed; Original draft preparation: Mohamed El‑Zahed, Esraa Mohamed; Writing, Reviewing and Editing: Mohamed El‑Zahed, Esraa Mohamed; Supervision and Project Administration: Mohamed El‑Zahed. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The cytotoxicity experiments on the mice keratinocyte cells were conducted in compliance with the IACUC (The Institutional Animal Care and Use Committee) statement for using animals in research and teaching by the local ethical committee of AUHA (Al-Azhar University Housing Animals).
Consent for publication
All authors give consent for the publication of the manuscript in the Journal of BMC Microbiology.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mohamed, E.A., El‑Zahed, M.M. Anticandidal applications of selenium nanoparticles biosynthesized with Limosilactobacillus fermentum (OR553490). Discover Nano 19, 115 (2024). https://doi.org/10.1186/s11671-024-04055-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s11671-024-04055-z