Abstract
Leaf senescence represents the final phase of leaf development and is characterized by a highly organized degenerative process involving the active translocation of nutrients from senescing leaves to growing tissues or storage organs. To date, a large number of senescence-associated transcription factors (sen-TFs) have been identified that regulate the initiation and progression of leaf senescence. Many of these TFs, including NAC (NAM/ATAF1/2/CUC2), WRKY, and MYB TFs, have been implicated in modulating the expression of downstream senescence-associated genes (SAGs) and chlorophyll degradation genes (CDGs) under the control of phytohormones. However, the involvement of basic helix-loop-helix (bHLH) TFs in leaf senescence has been less investigated. Here, we show that OsbHLH079 delays both natural senescence and dark-induced senescence: Overexpression of OsbHLH079 led to a stay-green phenotype, whereas osbhlh079 knockout mutation displayed accelerated leaf senescence. Similar to other sen-TFs, OsbHLH079 showed a gradual escalation in expression as leaves underwent senescence. During this process, the mRNA levels of SAGs and CDGs remained relatively low in OsbHLH079 overexpressors, but increased sharply in osbhlh079 mutants, suggesting that OsbHLH079 negatively regulates the transcription of SAGs and CDGs under senescence conditions. Additionally, we found that OsbHLH079 delays ABA-induced senescence. Subsequent RT-qPCR and dual-luciferase reporter assays revealed that OsbHLH079 downregulates the expression of ABA signaling genes, such as OsABF2, OsABF4, OsABI5, and OsNAP. Taken together, these results demonstrate that OsbHLH079 functions in delaying leaf yellowing by attenuating the ABA responses.
Similar content being viewed by others
Background
Leaves function as the primary organ for photosynthesis, converting light energy into chemical energy in the form of carbohydrate molecules (Woo et al. 2019). Leaf senescence, known as the final stage of leaf development, is a highly orchestrated degenerative process that entails the gradual breakdown of cellular components and macromolecules, including proteins, nucleic acids, and lipids (Pennell and Lam 1997; Gregersen et al. 2013; Schippers et al. 2015). The relocation of nutrients from senescing leaves towards developing tissues or storage organs is crucial for nutrient management and reproductive success (Lim et al. 2007; Masclaux-Daubresse et al. 2008). Crop yield and quality are greatly influenced by the timing of leaf senescence (Buchanan-Wollaston et al. 2005; Breeze et al. 2011; Su et al. 2017). Accelerated leaf senescence adversely impacts crop quality and yield by reducing the accumulation of photosynthetic assimilates in storage organs (Yang et al. 2016a; Mao et al. 2017). Conversely, delaying leaf senescence enables crops to prolong their photosynthetic lifespan, resulting in increased grain yield. However, it also reduces the protein level in grains due to delayed nitrogen remobilization from leaves (Zhao et al. 2015; Carmo-Silva et al. 2017; Havé et al. 2017). Hence, investigating the regulatory mechanisms of leaf senescence will offer important insights for crafting crop breeding strategies with a goal of enhancing agronomic traits.
The onset of leaf senescence depends primarily on developmental age, but is also influenced by several endogenous factors and external cues. Endogenous factors such as phytohormone signals and nutrient status play a pivotal role, while the external cues include wounding, shading, high salinity, and pathogen infection (Smart 1994; Jing et al. 2002; Moore et al. 2003; Gepstein and Glick 2013; Zakari et al. 2020). As leaves mature, a significant change in gene expression occurs (Lin and Wu 2004; van der Graaff et al. 2006). Using comprehensive global gene expression profiling methods, scientists have identified several genes that are induced during leaf senescence. These genes are referred to as senescence-associated genes (SAGs) which control the progression of leaf senescence (Buchanan-Wollaston 1997; Guo and Gan 2005; Li et al. 2012). Over the past few decades, the physiological roles of SAGs in plants have been elucidated: they are involved in diverse biological processes, including nutrient redistribution, biomolecule degradation, and phytohormone signaling (Kong et al. 2006; Li et al. 2020). For instance, OsSAG12-2, encodes a functional proteolytic enzyme that is induced under senescence conditions and regulates stress-induced cell death (Singh et al. 2016). RLS1 (Rapid Leaf Senescence 1), which encodes an NB (nucleotide-binding site) containing protein with an ARM (armadillo) domain, is upregulated during dark-induced leaf senescence. It regulates the autophagy-like process leading to chloroplast degradation in senescent leaves (Jiao et al. 2012). Chlorophyll degradation genes (CDGs), which constitute a subset of extensively characterized SAGs, play a pivotal role in the gradual decline of chlorophyll levels in leaves during senescence. A total of seven CDGs have been identified in rice, namely OsNYC1, OsNOL, OsHCAR, OsSGR, OsNYC3, OsPAO, and OsRCCR1. These genes encode enzymes responsible for catalyzing sequential reactions within the chlorophyll degradation pathway and are induced when leaves enter senescence (Piao et al. 2017; Lee and Masclaux-Daubresse 2021). Loss-of-function mutations in any of these CDGs result in prominent stay-green phenotypes, primarily due to impaired chlorophyll catabolism (Kusaba et al. 2007; Park et al. 2007; Morita et al. 2009; Sato et al. 2009; Tang et al. 2011; Piao et al. 2017).
Phytohormones, including abscisic acid (ABA), ethylene (ET), jasmonic acid (JA), and salicylic acid (SA), serve as the primary regulators of leaf senescence (Lim et al. 2007; Woo et al. 2019). Among these phytohormones, ABA, classified as a sesquiterpenoid hormone, stands out as one of the extensively studied senescence-promoting phytohormones: the exogenous application of ABA noticeably accelerates leaf senescence by inducing several SAGs (Quiles et al. 1995). In recent decades, a number of ABA-related mutants have been isolated, and their senescence phenotypes have been thoroughly investigated. These studies comprehensively revealed that mutants with elevated ABA levels or enhanced ABA signaling display premature a leaf senescence phenotype, while mutants with decreased ABA levels or attenuated ABA signaling display a stay-green phenotype during senescence process (Gao et al. 2016; Huang et al. 2018; Kim et al. 2019; Piao et al. 2019). For example, a loss-of-function mutation in OsNCED3, an ABA biosynthetic gene in rice, results in delayed leaf senescence due to ABA deficiency, whereas OsNCED3-overexpressing lines with elevated ABA levels exhibit accelerated leaf senescence (Huang et al. 2018). Another example is OsWRKY5, a member of the WRKY transcription factor family in rice, which promotes leaf senescence by upregulating ABA biosynthesis genes such as OsNCED3 and OsNCED5 (Kim et al. 2019). In Arabidopsis, three ABA-responsive element binding factors (ABFs), including ABF2, ABF3, and ABF4, enhance ABA signaling and activate CDGs, thereby facilitating chlorophyll breakdown under ABA-induced senescence conditions (Gao et al. 2016). In contrast, OsMYB102, a rice MYB transcription factor, retards both natural leaf senescence and dark-induced senescence by attenuating ABA signaling (Piao et al. 2019).
To date, a large number of transcription factors (TFs) have been isolated in plants whose expression is activated during senescence (Guo et al. 2004). Many of these TFs, termed senescence-associated TFs (sen-TFs), operate downstream of phytohormone signaling networks and are involved in the regulation of SAGs, thereby influencing the onset and progression of leaf senescence (Bengoa Luoni et al. 2019; Woo et al. 2019). Among these TFs, NAC (NAM/ATAF1/2/CUC2), WRKY, and MYB TFs have been extensively studied, revealing their pivotal functions in orchestrating the significant reprogramming of gene expression during senescence (Miao et al. 2008; Balazadeh et al. 2011; Zhang et al. 2012; Park et al. 2018; Kim et al. 2019; Sakuraba et al. 2020). Despite these advances, the involvement of basic helix-loop-helix (bHLH) TFs in leaf senescence remains poorly understood. In this study, we shed light on this aspect by showing that OsbHLH079 retards both natural leaf senescence and dark-induced senescence. Similar to other sen-TFs, the expression of OsbHLH079 gradually increased during leaf senescence. During leaf senescence, the SAGs and CDGs were downregulated in OsbHLH079 overexpressors and upregulated in osbhlh079 knockout mutants. Moreover, OsbHLH079 was shown to delay ABA-induced senescence by attenuating ABA signaling. Collectively, our results suggest that OsbHLH079 delays leaf senescence through its role as a negative regulator in the ABA signaling pathway.
Results
Expression of OsbHLH079 is Increased during Leaf Senescence
OsbHLH079 (MSU locus ID, LOC_Os02g47660; RAP locus ID, Os02g0705500) is a 361 amino acid protein that belongs to the basic helix-loop-helix (bHLH) transcription factor family in rice (Li et al. 2006). The open reading frame (ORF) of OsbHLH079 spans 1,086 bp and consists of six exons (Seo et al. 2020). To investigate the effect of senescence on the expression of OsbHLH079, we initially measured the total chlorophyll contents and the mRNA levels of senescence-associated genes (SAGs) in the flag leaves of the japonica rice cultivar ‘Dongjin’ (hereafter referred to as wild type; WT) under natural senescence or dark-induced senescence conditions. As senescence progressed, the total chlorophyll contents decreased, and the mRNA levels of SAGs increased (Additional file 1: Figures 1, 2 and 3). Next, we monitored changes in OsbHLH079 mRNA levels in the flag leaves of WT during natural senescence. Reverse transcription and quantitative real-time PCR (RT-qPCR) analysis revealed a gradual increase in OsbHLH079 transcript levels during natural senescence (Fig. 1A). In addition, OsbHLH079 expression was upregulated in detached flag leaves during dark-induced senescence (DIS) (Fig. 1B). Furthermore, in naturally senescing flag leaves, OsbHLH079 transcripts accumulated to higher levels in the yellow sector compared to the green sector (Fig. 1C). These results suggest that OsbHLH079 is involved in the progression of leaf senescence in rice.
Expression profiles of OsbHLH079 during leaf senescence. A Temporal expression patterns of OsbHLH079 during natural senescence. Flag leaves were harvested at 10-day intervals from WT plants grown under natural long-day conditions in the field. The expression levels of OsbHLH079 were determined by RT-qPCR analysis and normalized to those of GAPDH. Averages and standard deviations were obtained from four biological samples, each consisting of approximately five flag leaves. Representative phenotypes of flag leaves at each time point are shown as images. DAH, days after heading. B Relative transcript levels of OsbHLH079 during dark-induced senescence. Leaf segments were collected from the flag leaves of WT plants at the heading stage and then floated, abaxial side up, on a 3 mM MES buffer (pH 5.8) at 30 °C in complete darkness. Leaf discs were sampled every 24 h at the indicated DDI. The expression levels of OsbHLH079 determined by RT-qPCR analysis were normalized to those of GAPDH. Data represent the mean ± SD of four biological replicates (approximately 4 leaf discs per sample). Representative phenotypes of leaf discs are shown for each time point. DDI, day(s) of dark incubation. C Spatial expression patterns of OsbHLH079 in senescing leaves. Each sector of naturally senescing leaves was sampled from flag leaves of WT at 50 DAH grown in the field. The mRNA levels of OsbHLH079 were quantified by RT-qPCR analysis with GAPDH as an internal control. Data are presented as mean ± SD (n = 4). A-C Significantly different values are indicated by distinct letters, as determined by one-way ANOVA and Duncan’s least significant range test (P < 0.05). These experiments were performed twice with independent biological replicates, and similar results were obtained
OsbHLH079 Delays Dark-Induced Leaf Senescence
To investigate the physiological role of OsbHLH079 in leaf senescence, we used two independent transgenic lines overexpressing OsbHLH079 (referred to as osbhlh079-D and OsbHLH079-OE) (See Additional file 1: Fig. S4 for details). In addition, we generated two independent knockout mutant lines (designated as osbhlh079-1 and osbhlh079-2) (See Additional file 1: Fig. S5 for details). The osbhlh079-D line carries an activation-tagged T-DNA located 815 nucleotides upstream of the initiation codon of OsbHLH079 (Additional file 1: Fig. S4A), and detailed genetic information on osbhlh079-D has been reported previously (Seo et al. 2020). The OsbHLH079-OE line contains 35S::OsbHLH079 constructs integrated into its genome (Additional file 1: Fig. S4B). RT-qPCR analysis revealed the significant accumulation of OsbHLH079 transcripts in both lines compared to the WT, with the OsbHLH079-OE line showing even higher expression levels than the osbhlh079-D line (Additional file 1: Fig. S4C). The two independent osbhlh079 mutants, osbhlh079-1 and osbhlh079-2, were generated by CRISPR/Cas9-mediated mutagenesis (Miao et al. 2013), followed by the isolation of null segregants (Additional file 1: Fig. S5A, B). Chromatograms from direct sequencing of OsbHLH079 revealed that osbhlh079-1 and osbhlh079-2 carry a 5-bp deletion and a 1-bp deletion, respectively, within the coding region of OsbHLH079, as shown in Additional file 1: Fig. S5C-F.
We then evaluated the phenotypes of osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 grown under natural long-day conditions in the field. At the heading stage, the top four leaves of osbhlh079-D and OsbHLH079-OE displayed wider leaf angles, while those of osbhlh079-1 and osbhlh079-2 showed narrower leaf angles compared to the WT (Additional file 1: Fig. S6), as previously reported (Seo et al. 2020). To evaluate the effects of OsbHLH079 on the progression of dark-induced senescence (DIS), we incubated the detached flag leaf discs from WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 plants on a 3 mM MES buffer (pH 5.8) at 30 °C in complete darkness. Compared to WT, the leaf discs of OsbHLH079 overexpressors retained their green color for a longer period of time, whereas those of osbhlh079 mutants exhibited accelerated leaf yellowing compared to WT (Fig. 2A). Consistent with the visible phenotypes, the total chlorophyll levels in the leaves of osbhlh079-D and OsbHLH079-OE remained higher, while those of osbhlh079-1 and osbhlh079-2 were lower than WT during DIS (Fig. 2B). Furthermore, the ion leakage rate, an indicator of membrane disintegration, was significantly lower in the leaf discs of OsbHLH079 overexpressors and higher in those of osbhlh079 mutants than in WT at 4 and 5 days of dark incubation (DDI) (Fig. 2C). In addition, the senescence-associated genes (SAGs) and the chlorophyll degradation genes (CDGs) were down-regulated in OsbHLH079-overexpressing lines and up-regulated in osbhlh079 knock-out mutant lines under DIS conditions (Additional file 1: Fig. S7). Taken together, these results suggest a negative role of OsbHLH079 in leaf yellowing during DIS.
OsbHLH079 delays leaf yellowing during dark-induced senescence. A-C Changes in leaf color (A), total chlorophyll content (B), and ion leakage rate (C) in complete darkness. Rice plants were grown under natural day-night conditions in the paddy field, and leaf discs were collected from the flag leaves of rice plants at the heading stage. The leaf discs were then incubated on a 3 mM MES buffer (pH 5.8), abaxial side up, at 30 °C in complete darkness until the indicated DDI. The data in (B, C) were obtained from four independent samples, with approximately 10 mg of leaf discs per sample in (B) and around five leaf discs per sample in (C), and are presented as mean ± SD. Statistical analysis was performed using the two-tailed Student’s t-test (*P < 0.05, **P < 0.01, and ***P < 0.001). These experiments were repeated twice with similar results. Chl, chlorophyll; DDI, day(s) of dark incubation; FW, fresh weight; NS, not significant
OsbHLH079 Retards Natural Leaf Senescence
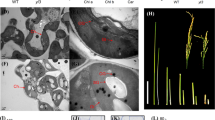
To elucidate the physiological role of OsbHLH079 in natural leaf senescence, we monitored the progression of age-dependent leaf senescence in WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 grown in the natural field. At the heading stage, there were no noticeable differences in leaf color among these plants (Fig. 3A, B). However, osbhlh079-D and OsbHLH079-OE displayed a delayed leaf yellowing phenotype, whereas osbhlh079-1 and osbhlh079-2 showed a premature leaf yellowing phenotype during grain filling (Fig. 3A, B). Consistent with these observations, the flag leaves of OsbHLH079 overexpressors maintained a higher concentration of total chlorophyll, whereas those of osbhlh079 mutants showed a significant decrease in total chlorophyll content compared to the WT flag leaves during natural senescence (Fig. 3C). In addition, we measured the Fv/Fm ratio, which serves as an indicator of photosystem II efficiency, to compare the photosynthetic performance of the leaves under natural senescence conditions. The flag leaves of OsbHLH079 overexpressors sustained higher Fv/Fm ratios than those of WT during grain filling (Fig. 3D). In contrast, the Fv/Fm ratios in the flag leaves of osbhlh079 null mutants decreased drastically after the heading stage compared to those of WT (Fig. 3D). Next, we performed a comparative analysis of the chloroplast structure in senescing flag leaves among WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 using transmission electron microscopy (TEM). At the heading stage, the chloroplasts of all plants appeared intact, with highly stacked grana thylakoids (Fig. 4). However, at 30 days after heading (DAH), osmiophilic globuli, which are plastid-localized lipoprotein particles associated with senescent chloroplasts (Besagni and Kessler 2013), began to accumulate in the chloroplasts of osbhlh079-1 and osbhlh079-2, whereas the chloroplasts of WT, osbhlh079-D, and OsbHLH079-OE did not show any accumulation of osmiophilic globuli (Fig. 4). At 50 DAH, the chloroplasts of OsbHLH079 overexpressors remained intact and free of osmiophilic globuli, whereas those of WT plants accumulated osmiophilic globuli (Fig. 4). Meanwhile, the thylakoid membranes of chloroplasts in osbhlh079-1 and osbhlh079-2 were almost completely disintegrated, and the number and size of osmiophilic globuli were significantly increased (Fig. 4). Collectively, these results suggest that OsbHLH079 functions as a negative regulator of natural leaf senescence.
OsbHLH079 negatively regulates natural leaf senescence in rice. A Phenotypes of WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 at 0, 30, and 50 DAH grown in the field. Scale bars = 20 cm. B Comparison of leaf phenotypes among WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 plants during natural senescence. Flag leaves from the plants in (A) were photographed, and the images shown are representative of three independent flag leaves. Scale bars = 2 cm. C, D Changes in total chlorophyll content (C) and Fv/Fm ratio (D) of flag leaves under natural senescence conditions. Means and standard deviations were obtained from four independent plants. Asterisks denote statistically significant differences compared to WT, as determined by the two-tailed Student’s t-test (**P < 0.01 and ***P < 0.001). These experiments were performed twice with similar results. Chl, chlorophyll; DAH, days after heading; FW, fresh weight; NS, not significant
Transmission electron micrograph (TEM) of chloroplasts during natural senescence. Leaf segments, approximately 2 cm away from the leaf tip, were sampled from the flag leaves of WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 grown under natural field conditions and subjected to TEM analysis. Scale bars = 500 nm. The images are representative of four independent chloroplasts. DAH, days after heading; G, grana thylakoid; O, osmiophilic globule; S, starch granule
OsbHLH079 Modulates the Expression of Senescence-Associated Genes during Natural Senescence
Senescence-associated genes (SAGs) are upregulated under senescing conditions and play a crucial role in controlling the progression of leaf senescence (Lee et al. 2001; Lee and Masclaux-Daubresse 2021). To investigate the effect of OsbHLH079 on the expression of SAGs during leaf senescence, we conducted RT-qPCR analysis to compare the mRNA levels of representative SAGs in senescing flag leaves of WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 grown in the natural field. Representative SAGs included Osl2 (aminotransferase), Osl20 (E1-α subunit of branched-chain α-keto dehydrogenase), Osl55 (biotinylated subunit of β-methylcrotonyl-CoA carboxylase), Osl57 (3-ketoacyl-CoA thiolase), Osl85 (isocitrate lyase), Osh69 (seed imbibition protein), and OsSAG12-2 (proteolytic enzyme) (Lee et al. 2001; Singh et al. 2016) (see Additional file 2: Table S1). At the heading stage, there were no detectable differences in the expression levels of any of the genes among WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 (Fig. 5). However, during grain filling, the SAGs were downregulated in OsbHLH079-overexpressing lines and upregulated in osbhlh079 loss-of-function mutants (Fig. 5). These results indicate that OsbHLH079 retards the progression of leaf senescence by negatively regulating the transcription of SAGs under senescent conditions.
Expression profiles of senescence-associated genes in naturally senescing flag leaves. A-G Relative transcript levels of Osl2 (A), Osl20 (B), Osl55 (C), Osl57 (D), Osl85 (E), Osh69 (F), and OsSAG12-2 (G) in WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2. The middle parts of the flag leaves were harvested from the rice plants grown under natural long-day conditions in the paddy field at the indicated DAH and were used for RT-qPCR analysis. The expression levels of each gene were normalized to that of GAPDH, an internal control. The values presented are the means of four biological replicates, with each replicate consisting of five independent flag leaves, and the error bars indicate the standard deviations. Significance of differences between means was analyzed by the two-tailed Student’s t-test (*P < 0.05, **P < 0.01, and ***P < 0.001). These analyses were performed twice independently with similar results. DAH, days after heading; NS, not significant
OsbHLH079 Downregulates Chlorophyll Degradation Genes during Leaf Senescence
Chlorophyll degradation is a crucial process in leaf senescence, leading to the gradual loss of green pigments (Pružinská et al. 2005; Gan and Hörtensteiner 2013). Chlorophyll degradation genes (CDGs) encode enzymes that facilitate sequential reactions in the chlorophyll catabolic pathway, and CDGs are known to be induced under senescence conditions (Piao et al. 2017; Lee and Masclaux-Daubresse 2021). To investigate the role of OsbHLH079 in chlorophyll degradation, we compared the transcript levels of CDGs, including OsNYC1 (chlorophyll b reductase; Kusaba et al. 2007), OsNOL (chlorophyll b reductase; Sato et al. 2009), OsHCAR (7-hydroxy methyl chlorophyll a reductase; Piao et al. 2017), OsSGR (Mg2+ dechelatase; Park et al. 2007), OsNYC3 (pheophytinase; Morita et al. 2009), OsPAO (pheophorbide a oxygenase; Tang et al. 2011), and OsRCCR1 (red chlorophyll catabolite reductase; Tang et al. 2011), in flag leaves of WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 grown under natural long-day conditions. Consistent with previous reports (Piao et al. 2017; Lee and Masclaux-Daubresse 2021), their transcript abundances gradually increased in WT as senescence progressed. At the heading stage, the expression levels of the CDGs in osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 were similar to those in WT (Fig. 6). However, the mRNA levels of the CDGs in senescing flag leaves were lower in OsbHLH079 overexpressors and higher in osbhlh079 knockout mutants compared to those in WT (Fig. 6). These results suggest that OsbHLH079 functions as a repressor of CDG expression during natural senescence, thereby inhibiting the progression of leaf yellowing.
Expression analysis of chlorophyll degradation genes during natural senescence of flag leaves. A-G Relative mRNA levels of OsNYC1 (A), OsNOL (B), OsHCAR (C), OsSGR (D), OsNYC3 (E), OsPAO (F), and OsRCCR1 (G) were compared among WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2. Total RNA samples from Fig. 5 were subjected to RT-qPCR analysis, using GAPDH as a reference for normalization. Data are presented as mean ± SD of four independent biological samples, each with approximately five flag leaves per sample. Asterisks on the bars indicate statistically significant differences compared to WT transcript levels (Student’s t-test; *P < 0.05, **P < 0.01, and ***P < 0.001). These experiments were performed twice with similar results. DAH, days after heading; NS, not significant
OsbHLH079 Regulates ABA-induced Leaf Senescence in Rice
Various phytohormones, such as abscisic acid (ABA), ethylene (ET), jasmonic acid (JA), and salicylic acid (SA), play important roles in the progression of leaf senescence (Jan et al. 2019; Guo et al. 2021; Huang et al. 2022). To investigate the effects of these phytohormones on OsbHLH079 expression, we measured the mRNA levels of OsbHLH079 in the leaves of 10-day-old WT seedlings treated with ABA, 1-aminocyclopropane-1-carboxylic acid (ACC, the immediate precursor of ET; Ververidis and John 1991), methyl jasmonate (MeJA, the methylated derivative of JA; Peng and Zhang 2017), or SA. RT-qPCR analysis revealed that OsbHLH079 transcription was exclusively induced in response to ABA: OsbHLH079 expression increased to approximately 2.5-fold and 3.9-fold after 3 and 6 h of ABA treatment, respectively (Fig. 7A). Next, we monitored the progression of leaf yellowing in detached flag leaves of WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 during treatment with ABA, ACC, MeJA, or SA. Before the application of phytohormones, leaf colors were nearly uniform in each plant line (Fig. 7B and Additional file 1: Fig. S8). After 3 days of treatments, we observed that OsbHLH079 overexpressors and osbhlh079 knockout mutants showed hyposensitivity and hypersensitivity, respectively, to ABA-induced leaf senescence: the leaves of osbhlh079-D and OsbHLH079-OE retained their green colors for a longer period of time compared to those of WT, while the leaf discs of osbhlh079-1 and osbhlh079-2 exhibited an early leaf yellowing phenotype (Fig. 7B). In support of these observations, the osbhlh079-D and OsbHLH079-OE maintained high levels of total chlorophyll, whereas the total chlorophyll contents in osbhlh079-1 and osbhlh079-2 sharply decreased during ABA-induced leaf senescence (Fig. 7C). Moreover, the ion leakage rates of leaf discs from OsbHLH079 overexpressors remained at a low level, while those of osbhlh079 knockout mutants dramatically increased under ABA-mediated senescence conditions (Fig. 7D). In addition, the transcript levels of SAGs and CDGs were lower in OsbHLH079-overexpressing lines and higher in osbhlh079 knock-out mutant lines compared to those in WT at 3 days of ABA treatment (Additional file 1: Fig. S9). However, during ACC-, MeJA-, or SA-mediated senescence, leaf colors, total chlorophyll levels, and ion leakage rates showed almost no discernible differences among WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 (Additional file 1: Figs. S8 and S10). Taken together, these results indicate that OsbHLH079 plays a crucial role in delaying ABA-induced leaf senescence in rice.
OsbHLH079 delays ABA-induced leaf senescence in rice. A Expression of OsbHLH079 in response to senescence-promoting phytohormones. WT seedlings were grown on half-strength Murashige and Skoog (MS) solid medium (pH 5.7) under constant light conditions at 30 °C for 10 days. Seedlings were then transferred to half-strength MS liquid medium supplemented with 100 µM ABA, 10 mM ACC, 100 µM MeJA, or 100 µM SA. Leaves were harvested at 3 and 6 h after treatments, and their total RNAs were used for RT-qPCR analysis, with expression levels of OsbHLH079 normalized to those of GAPDH. Means and standard deviations were obtained from four biological replicates consisting of approximately three leaves per sample. Statistically significant differences compared to the mock-treated samples are marked by asterisks, as determined by the two-tailed Student’s t-test (***P < 0.001). ABA, abscisic acid; ACC, 1-aminocyclopropane-1-carboxylic acid; MeJA, methyl jasmonate; NS, not significant; SA, salicylic acid. B-D Leaf color (B), total chlorophyll content (C), and ion leakage rate (D) of WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 under ABA-induced senescence conditions. Flag leaf discs were collected at the heading stage from the rice plants grown under natural field conditions. The leaf discs were then treated with a 3 mM MES buffer (pH 5.8) containing 50 µM ABA for the indicated time periods under continuous light conditions at 30 °C. Data presented in (C, D) represent the average ± SD of four biological replicates [around 10 mg of leaf discs per replicate in (C) and approximately five leaf discs per replicate in (D)]. Asterisks indicate significantly different values according to Student’s t-test (**P < 0.01 and ***P < 0.001). ABA, abscisic acid; Chl, chlorophyll; DT, day(s) of treatment; FW, fresh weight; NS, not significant. These experiments were performed twice with similar results
OsbHLH079 Suppresses the Expression of ABA Signaling Genes under Senescing Conditions
Since OsbHLH079 delays ABA-induced leaf senescence (Fig. 7B-D), we hypothesized that OsbHLH079 might be related to ABA metabolism and/or signaling. To investigate the role of OsbHLH079 in ABA metabolism, we compared the endogenous ABA levels in flag leaves among WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2. However, no significant differences in endogenous ABA levels were found among the plant lines (Additional file 1: Fig. S11). We then focused on the relationship between OsbHLH079 and ABA signaling genes. Specifically, we analyzed the expression patterns of ABA signaling genes, including OsABF2, OsABF4, OsABI5, OsbZIP23, OsEEL, and OsNAP (Xiang et al. 2008; Lu et al. 2009; Hossain et al. 2010a, b; Liang et al. 2014; Yang et al. 2019), in senescing flag leaves of WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2. At the heading stage, the mRNA levels of OsABF2, OsABF4, OsABI5, and OsNAP were downregulated in OsbHLH079 overexpressors and upregulated in osbhlh079 knock-out mutants (Fig. 8). During grain filling, the transcript abundances of all the investigated genes remained at a relatively low level in flag leaves of OsbHLH079 overexpressors and sharply increased in the flag leaves of osbhlh079-1 and osbhlh079-2 knockout mutants compared to WT (Fig. 8): their expression showed an increase in WT as senescence progressed, consistent with previous studies (Hossain et al. 2010a; Liang et al. 2014; Kang et al. 2019; Piao et al. 2019; Sakuraba et al. 2020) (Fig. 8). To further confirm whether OsbHLH079 inhibits the transcription of ABA signaling genes, we conducted a dual-luciferase reporter assay. For the reporter constructs, the promoter regions of OsABF2 (-1,553 to -1), OsABF4 (-1,709 to + 7), OsABI5 (-1,971 to + 58), or OsNAP (-1,542 to + 84) were fused with the LUC reporter gene (Fig. 9A). The LUC activities of the protoplasts transformed with proOsABF2::LUC, proOsABF4::LUC, proOsABI5::LUC, and proOsNAP::LUC constructs were significantly reduced when each of them was co-transfected with the pUbi::OsbHLH079-MYC effector plasmid (Fig. 9B). Taken together, these results suggest that OsbHLH079 desensitizes ABA signaling by down-regulating the expression of ABA signaling genes, thereby delaying leaf senescence in rice.
Transcriptional analysis of genes involved in ABA signaling during natural leaf senescence in rice. A-F Relative transcript abundances of OsABF2 (A), OsABF4 (B), OsABI5 (C), OsbZIP23 (D), OsEEL (E), and OsNAP (F) were measured in flag leaves of WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2. Total RNAs isolated from the samples shown in Fig. 5 were subjected to RT-qPCR analysis. The mRNA levels of GAPDH, a reference gene, were used to normalize. The values shown in the graphs are averages of four independent samples (around 5 flag leaves per sample), and the error bars represent standard deviations. Differences between the means were statistically analyzed using two-tailed Student’s t-test (*P < 0.05, **P < 0.01, and ***P < 0.001). These analyses were repeated twice with similar results. DAH, days after heading; NS, not significant
OsbHLH079 represses the transcription of OsABF2, OsABF4, OsABI5, and OsNAP. A Description of the constructs used in the dual-luciferase reporter assay, including the effectors, reporters, and internal control. LUC was fused to proOsABF2 (-1,553 to -1), proOsABF4 (-1,709 to + 7), proOsABI5 (-1,971 to + 58), or proOsNAP (-1,542 to + 84). LUC, luciferase; NOS, NOS terminator; pUbi, promoter of Ubiquitin; Rluc, renilla luciferase. B The repression of proOsABF2, proOsABF4, proOsABI5, and proOsNAP by OsbHLH079-MYC expression in the dual-luciferase reporter assay. MYC was used as a negative control. The LUC activity of each sample was represented relative to that of the negative control, set as 1. Averages and standard deviations were obtained from five independent samples. Significant differences between means were determined using a two-tailed Student’s t-test (**P < 0.01, and ***P < 0.001). LUC, luciferase; pUbi, promoter of Ubiquitin
Discussion
OsbHLH079 Negatively Regulates ABA-Induced Leaf Senescence
Basic helix-loop-helix proteins are a large family of transcription factors that are widely distributed in fungi, plants, and animals (Carretero-Paulet et al. 2010; Pires and Dolan 2010). Within the plant kingdom, bHLH TFs play crucial roles in numerous biological processes, including responses to biotic and abiotic stresses, reproductive organ development, embryo growth, and hormonal signaling (Friedrichsen et al. 2002; Gremski et al. 2007; Kondou et al. 2008; Ariyarathne and Wone 2022). For example, overexpression of AhHLH112 in peanuts enhances drought tolerance by increasing endogenous ABA levels (Li et al. 2021). In Arabidopsis, a mutation in DYSFUNCTIONAL TAPETUM1 (DYT1), which encodes a bHLH TF, leads to male sterility by downregulating several tapetum-preferential genes (Zhang et al. 2006). Recently, bHLH TFs have been reported to modulate leaf senescence in Arabidopsis: MYC2, MYC3, and MYC4 promote jasmonic acid-mediated senescence (Qi et al. 2015), whereas ATBS1-INTERACTING FACTOR 2 (AIF2) delays brassinosteroid (BR)-mediated leaf senescence (Kim et al. 2020). However, the role of bHLH TFs in rice leaf senescence has not been extensively investigated. This study shows that OsbHLH079 acts to delay both natural and dark-induced leaf senescence. Overexpression of OsbHLH079 delays leaf senescence, while osbhlh079 knockout mutants exhibit early leaf senescence (Figs. 2 and 3). In addition, OsbHLH079 represses the expression of ABA signaling genes, including OsABF2, OsABF4, OsABI5, and OsNAP (Figs. 8 and 9). This suggests that OsbHLH079 suppresses ABA signaling during leaf senescence. Consistent with these data, OsbHLH079 was shown to delay ABA-induced leaf senescence (Fig. 7B-D). Therefore, we propose that OsbHLH079 functions to inhibit leaf senescence by desensitizing ABA signaling.
OsbHLH079 Functions as a Negative Regulator of Leaf Senescence
Senescence-associated transcription factors (Sen-TFs), whose expressions are elevated during leaf senescence, are categorized into senescence-promoting and senescence-inhibiting types (Lee and Masclaux-Daubresse 2021). In this study, we observed that OsbHLH079-overexpressing lines exhibited a stay-green phenotype, whereas osbhlh079 knock-out mutant lines displayed premature leaf yellowing (Figs. 2 and 3), indicating that OsbHLH079 functions as a senescence-inhibiting sen-TFs. Recently, several sen-TFs have been found to play roles in delaying leaf senescence. For example, ONAC106, a senescence-associated NAC TF in rice, negatively regulates leaf senescence (Sakuraba et al. 2015). Similarly, OsMYB102, a MYB-type TF in rice, retards both natural senescence and dark-induced senescence: the expression of OsMYB102 was increased during leaf senescence, and OsMYB102-overexpressing plants exhibited delayed leaf senescence phenotypes (Piao et al. 2019). These types of TFs are believed to function antagonistically to senescence-promoting sen-TFs in regulating leaf senescence (Sakuraba et al. 2015; Cao et al. 2023). For instance, MYC2, MYC3, and MYC4 redundantly promote JA-mediated leaf senescence by directly up-regulating SAG29; meanwhile, bHLH03, bHLH13, bHLH14, and bHLH17 attenuate MYC2/MYC3/MYC4-mediated senescence by directly repressing SAG29 in Arabidopsis (Qi et al. 2015). Therefore, we propose that OsbHLH079 antagonizes senescence-promoting sen-TFs, ensuring that leaf senescence initiates at the proper time for survival under fluctuating environmental conditions.
OsbHLH079 Retards Leaf Senescence by Dampening ABA Signaling, Not by Reducing ABA Accumulation
To date, several sen-TFs that govern ABA-mediated leaf senescence have been identified (Park et al. 2018; Piao et al. 2019; Sakuraba et al. 2020; Xie et al. 2022). In many cases, these sen-TFs regulate both ABA metabolism and ABA signaling. For example, OsMYB102, a MYB-type TF in rice, retards leaf senescence by inhibiting ABA accumulation and attenuating ABA signaling under senescing conditions (Piao et al. 2019). In addition, OsWRKY53, a WRKY TF in rice, increases endogenous ABA levels to promote leaf senescence; the expression levels of several ABA signaling genes were also higher in OsWRKY53-overexpressing plants compared to the WT (Xie et al. 2022). Some sen-TFs, however, have been found to regulate leaf senescence by modulating ABA signaling, rather than by influencing ABA metabolism. For instance, ONAC054, a NAC TF in rice, promotes ABA-induced leaf senescence by activating ABA signaling genes, including OsABF4 and OsABI5; conversely, the endogenous ABA contents in onac054 mutants were nearly identical to those in the WT (Sakuraba et al. 2020). Similarly, OsRL3, a MYB-related TF in rice, up-regulates ABA signaling genes, such as OsRK1, OsRAB16C, and OsRAB16D, to accelerate dark-induced senescence; meanwhile, the expression levels of ABA biosynthetic genes were unchanged in osrl3 mutants compared to the WT (Park et al. 2018). These findings indicate that sen-TFs controlling ABA-induced leaf senescence do not necessarily modulate ABA metabolism under senescing conditions. In this study, we found that OsbHLH079 delays ABA-mediated leaf senescence (Fig. 7B). Subsequent RT-qPCR and dual-luciferase reporter assays revealed that OsbHLH079 suppresses the transcriptions of downstream ABA signaling genes, including OsABF2, OsABF4, OsABI5, and OsNAP (Figs. 8 and 9). Interestingly, the endogenous ABA levels in flag leaves of osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 plants were almost the same as the WT (Additional file 1: Fig. S11). Therefore, we propose that OsbHLH079 governs ABA-mediated leaf senescence by dampening ABA signaling, rather than by reducing ABA contents, as observed in the cases of ONAC054 and OsRL3.
OsbHLH079 -mediated Attenuation of ABA Signaling during Leaf Senescence may be Attributed to the Antagonistic Interaction between BR and ABA
BR, a class of plant-specific polyhydroxylated steroid hormones, play an important role in the regulation of various physiological processes, including cell elongation, cell wall regeneration, fruit ripening, pollen development, and root growth (Hacham et al. 2011; Vogler et al. 2014; Peres et al. 2019). In recent decades, there has been increasing evidence that BR also affects leaf senescence. Mutants with increased BR levels or enhanced BR signaling tend to undergo accelerated leaf senescence, whereas mutants with reduced BR levels or suppressed BR signaling show a delay in leaf senescence (Li et al. 1996; Li and Chory 1997; Yin et al. 2002; Husar et al. 2011). In addition, application of exogenous epibrassinolide, an active form of BR, leads to premature leaf senescence in a dose-dependent manner (Sağlam-Çağ 2007). These studies highlight the importance of BRs as essential phytohormones that promote senescence in plants.
In our previous study (Seo et al. 2020), we found that OsbHLH079 regulates leaf angle and kernel length in a BR-dependent manner. OsbHLH079 overexpressors exhibited exaggerated leaf angles and elongated grains, whereas transgenic lines with suppressed OsbHLH079 expression displayed upright leaves and shorter grains, similar to mutants with altered BR signaling. BR signaling genes were upregulated in the OsbHLH079 overexpressors and downregulated in the OsbHLH079-targeted RNAi lines. Consistent with these data, overexpression of OsbHLH079 resulted in increased responsiveness to epibrassinolide in a BR-induced lamina joint (LJ) tilt assay, highlighting the role of OsbHLH079 in amplifying BR signaling and consequently increasing leaf angle and kernel length. In this study, we demonstrated the critical role of OsbHLH079 in leaf senescence. Interestingly, it appeared that BR do not serve as the primary phytohormone regulating OsbHLH079-mediated leaf senescence, since OsbHLH079, a positive regulator of the BR pathway, actually induced a delay in leaf senescence (Figs. 2 and 3). Instead, OsbHLH079 was found to attenuate ABA signaling, thereby delaying the process of leaf senescence (Figs. 7 and 8).
To date, several molecular and physiological studies have provided substantial evidence that ABA and BR function in an antagonistic manner to control a variety of biological processes, such as seed germination, stomatal movement, and root growth (Xue et al. 2009; Hu and Yu 2014; Clouse 2016). For instance, BIN2-overexpressing lines, which are defective in BR signaling, were hypersensitive to ABA during seed germination in Arabidopsis (Hu and Yu 2014). Similarly, the BR-deficient Arabidopsis mutant det2 displayed enhanced responsiveness to ABA-induced suppression of root elongation (Xue et al. 2009). More recently, a molecular link between ABA and BR signaling pathways was uncovered in Arabidopsis: BZR1, a master TF in BR signaling, directly binds to G-box cis-elements within the promoter of ABA INSENSITIVE 5 (ABI5) and inhibits its expression (Yang et al. 2016b). Interestingly, our study demonstrated that OsbHLH079 stimulated the transcription of OsBZR1 (Seo et al. 2020), the closest counterpart of Arabidopsis BZR1 in rice, whereas downregulating the expression of OsABI5, a functional homolog of Arabidopsis ABI5 (Fig. 8C). Based on these findings, we speculate that the increased expression of OsBZR1 driven by OsbHLH079 may contribute to OsbHLH079-mediated dampening of ABA signaling, although we cannot exclude the possibility that OsbHLH079 influences ABA signaling through a mechanism independent of BR. Taken together, our results provide insights into the interplay between ABA and BR signaling pathways in the context of leaf senescence.
OsbHLH079 -mediated Suppression of ABA Signaling Alters Expression of SAGs and CDGs
Numerous ABA signaling genes, including OsABF2, OsABF4, OsABI5, OsbZIP23, OsEEL, and OsNAP, have been found in the rice genome, and their involvement in abiotic stress response has been extensively elucidated (Xiang et al. 2008; Lu et al. 2009; Hossain et al. 2010a, b; Chen et al. 2014; Yang et al. 2019). For example, the transgenic rice overexpressing OsABF4 exhibited enhanced sensitivity to ABA, resulting in increased drought tolerance (Lu et al. 2009). In contrast, T-DNA insertional knockout mutants of OsABF2 or OsABI5 displayed susceptibility to drought and salinity due to the impaired ABA response (Hossain et al. 2010a, b). Interestingly, recent investigations have unveiled a number of sen-TFs capable of influencing the expression of these ABA signaling genes, thereby controlling leaf senescence (Kang et al. 2019; Piao et al. 2019; Sakuraba et al. 2020). For instance, ONAC054 acts as a positive regulator of leaf senescence by driving the transcription of OsABF4 and OsABI5 (Sakuraba et al. 2020). In addition, OsMYB102 delays both natural senescence and DIS by repressing OsNAP and OsABF4 (Piao et al. 2019). In parallel, the function of OsbHLH079 was confirmed in inhibiting the expression of various ABA signaling genes, including OsABF2, OsABF4, OsABI5, and OsNAP, ultimately contributing to the delay of leaf senescence (Figs. 8 and 9). Apparently, all these sen-TFs play a central role in the modulation of ABA-mediated leaf senescence.
To date, there is increasing evidence that ABA signaling genes themselves are capable of orchestrating the expression of SAGs and/or CDGs (Chen et al. 2014; Piao et al. 2019; Sakuraba et al. 2020). For example, OsABF4 directly induces the expression of OsNYC1 and OsSGR (Piao et al. 2019). In addition, OsABI5 was also shown to directly transactivate OsNYC1 and OsSGR (Sakuraba et al. 2020). Furthermore, chromatin immunoprecipitation (ChIP) assays revealed that OsNAP binds to the promoters of several SAGs and CDGs, such as Osl57, Osh69, OsNYC1, OsSGR, OsNYC3, and OsRCCR1 (Chen et al. 2014). Therefore, we propose that OsbHLH079-mediated alterations in the expression of ABA signaling genes could consequently induce the downregulation of SAGs and CDGs during the process of leaf senescence, although we cannot exclude the alternative possibility that OsbHLH079 may directly inhibit the expression of SAGs and CDGs through various feed-forward regulatory loops.
Conclusions
In this study, we show that OsbHLH079, a bHLH TF in rice, is involved in the process of leaf senescence. Similar to other sen-TFs, OsbHLH079 exhibited a progressive increase in its expression during leaf senescence. Overexpression of OsbHLH079 delayed leaf senescence, whereas the loss-of-function mutation in OsbHLH079 induced premature leaf senescence. RT-qPCR analysis revealed that OsbHLH079 negatively regulates the expression of SAGs and CDGs during leaf senescence. Furthermore, we found that OsbHLH079 retards ABA-induced leaf senescence and substantially suppresses the expression of key ABA signaling genes, including OsABF2, OsABF4, OsABI5, and OsNAP. Hence, we propose that OsbHLH079 acts to attenuate ABA signaling, ultimately leading to a delay in the progression of leaf senescence in rice.
Materials and Methods
Plant Materials and Growth Conditions
The activation-tagged T-DNA insertion line of OsbHLH079, designated as osbhlh079-D (PFG_3A-01275), was obtained from the Salk Institute Genomics Analysis Laboratory (http://signal.salk.edu/cgi-bin/RiceGE) (Jeon et al. 2000; Jeong et al. 2002). The genetic information of osbhlh079-D has been described previously (Seo et al. 2020). For this study, the rice (Oryza sativa) plants, including osbhlh079-D, OsbHLH079-OE, osbhlh079-1, osbhlh079-2, and their parental japonica cultivar ‘Dongjin’ (referred to as wild type; WT), were grown under natural long-day conditions (approximately 14 h of light per day) in a paddy field located in Suwon, South Korea (37ºN latitude). The seeds were sown on seedbeds and grown in a greenhouse for one month before transplanting in the paddy field. Rice cultivation followed common agricultural practices adapted to Korean rice varieties.
Vector Construction and Rice Transformation
To generate the OsbHLH079-OE transgenic plant, the full-length coding region of OsbHLH079 was amplified by polymerase chain reaction (PCR) using cDNA obtained from leaves of the WT as a template and gene-specific primers (see Additional file 2: Table S2). The PCR product was subcloned into the pCRTM8/GW/TOPO® entry vector (Invitrogen, Carlsbad, CA, USA) and transferred into the pMDC32 Gateway-compatible binary destination vector (Curtis and Grossniklaus 2003) through an LR recombination reaction using the Gateway™ LR Clonase™ II Enzyme Mix (Invitrogen). To generate the osbhlh079-1 and osbhlh079-2 mutants, a specific 20-nt spacer sequence, GACGTTTCACGACACCGGAA, was designed using the CRISPRdirect software (https://crispr.dbcls.jp/) (Naito et al. 2015) and subcloned into a guide RNA expression cassette in the pOs-sgRNA entry vector (Miao et al. 2013). The resulting cassette was then transferred into the pH-Ubi-cas9-7 destination vector containing a Cas9 expression cassette (Miao et al. 2013) via the LR reaction.
The Agrobacterium tumefaciens strain LBA4404 (Ooms et al. 1982) was transformed with the resulting constructs, respectively, using the freeze-thaw method (Höfgen and Willmitzer 1988), and the calli generated from mature WT seed embryos were subjected to the Agrobacterium-mediated transformation of rice according to the previously described protocol (Jeon et al. 2000). The transformed calli were selected on 2N6 medium containing 50 mg L− 1 hygromycin (Duchefa Biochemie, Haarlem, The Netherlands) and regenerated into transgenic rice plants, including OsbHLH079-OE, osbhlh079-1, and osbhlh079-2, respectively.
Reverse Transcription and Quantitative Real-Time PCR (RT-qPCR) Analysis
Total RNAs were isolated from rice leaves using the MG Total RNA Extraction Kit (MGmed, Seoul, Republic of Korea) according to the manufacturer’s instructions. The extracted RNAs were reverse transcribed to generate first-strand cDNAs using the Oligo(dT)15 Primer (Promega, Madison, WI, USA) and M-MLV Reverse Transcriptase (Promega). The resulting product mixtures were diluted four-fold with distilled water. Quantitative real-time PCR (qPCR) was performed using the GoTaq® qPCR Master Mix (Promega) and a LightCycler® 480 system (Roche, Basel, Switzerland). The qPCR reaction mix, with a final volume of 20 µl, was prepared by combining 2 µl first-strand cDNA mixture, 0.4 µl 10 µM forward primer, 0.4 µl 10 µM reverse primer, 10 µl GoTaq® qPCR Master Mix, and 7.2 µl nuclease-free water. Gene-specific primers are listed in Additional file 2: Table S2. The qPCR conditions consisted of an initial denaturation step at 95 °C for 2 min followed by 50 cycles of 95 °C for 15 s and 60 °C for 1 min. Data obtained by qPCR were analyzed using the 2−ΔΔCT method (Livak and Schmittgen 2001) with OsGAPDH as the reference gene for normalization (Jain et al. 2006).
Dark Treatments
For the dark treatment, leaf discs were harvested from the flag leaves of rice plants at the heading stage, which were grown under natural day-night conditions in the paddy field. The leaf discs were then carefully placed abaxial side up on a 3 mM MES buffer (pH 5.8) and incubated for the indicated periods in complete darkness in an artificial growth chamber maintained at 30 °C.
Phytohormone Treatments
To examine the expression level of OsbHLH079 under different phytohormone treatments, WT seeds were sterilized in 70% (v/v) aqueous ethanol for 10 min and in 2% (w/v) sodium hypochlorite for 20 min. The sterilized seeds were then rinsed three times with sterile water and germinated on half strength Murashige and Skoog (MS) solid medium (pH 5.7) under continuous light conditions (100 µmol m− 2 s− 1) at 30 °C in an artificial growth chamber. After 10 days of growth, the seedlings were transferred to a half-strength MS liquid medium containing 100 µM ABA (Duchefa Biochemie), 10 mM ACC (Sigma-Aldrich, Saint Louis, MO, USA), 100 µM MeJA (Sigma-Aldrich), or 100 µM SA (Sigma-Aldrich). Seedlings in a half-strength MS liquid medium without additional phytohormones were used as a mock. Leaves were harvested at 3 and 6 h after treatment, and their total RNA was isolated for further analysis.
To evaluate the senescence phenotype in response to the senescence-associated phytohormone treatments, leaf discs were collected from flag leaves of WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 plants at the heading stage, grown under natural long-day conditions in the rice field. Leaf discs were then floated, abaxial side up, on a 3 mM MES buffer (pH 5.8) containing 50 µM ABA (Duchefa Biochemie), 20 mM ACC (Sigma-Aldrich), 50 µM MeJA (Sigma-Aldrich), or 100 µM SA (Sigma-Aldrich), followed by incubation for the indicated periods under continuous light conditions (100 µmol m− 2 s− 1) at 30 °C in an artificial growth chamber. Leaf discs that were incubated on a 3 mM MES buffer (pH 5.8) without any phytohormones were used as a control.
Total Chlorophyll Quantification
To determine the total chlorophyll content, approximately 10 mg of flag leaves were weighed into a 2 ml microcentrifuge tube and homogenized in liquid nitrogen using a TissueLyser II (Qiagen, Hilden, Germany). The ground leaves were then dissolved in 500 µl of 80% (v/v) ice-cold acetone, followed by centrifugation at 12,000 rpm for 15 min at 10 °C. The absorbance of the supernatant was measured at wavelengths of 663 nm and 647 nm using a UV/VIS spectrophotometer (PowerWave X, BioTek, Winooski, VT, USA). Finally, total chlorophyll concentrations were calculated according to Porra et al. (1989).
Measurement of Ion Leakage Rates
Ion leakage rates were determined as previously described by Fan et al. (1997) with minor adjustments. Five rice leaf discs, each approximately 1 cm2 in size, were placed in 6 ml of 0.4 M mannitol (Duchefa Biochemie) with gentle rotation for 3 h at room temperature. The initial conductivity of the solution was then measured with a conductivity meter (CON 6, LaMotte, Maryland, USA). After incubation at 90 °C for 30 min, the total conductivity of the solution was measured with the same conductivity meter (CON 6, LaMotte). Finally, the ion leakage rate was calculated as the percentage of the initial conductivity divided by the total conductivity.
Determination of Fv / Fm Ratios
The Fv/Fm ratios, which represent the maximum quantum efficiency, were measured using a chlorophyll fluorometer (OS30p+, Opti-Sciences Inc., New Hampshire, USA). To ensure the complete oxidation of QA, a bound plastoquinone, the middle part of each flag leaf of rice plants grown in the paddy field was dark adapted for 15 min before measuring the Fv/Fm ratio. Each plant was subjected to three experimental replicates.
Transmission Electron Microscopy (TEM)
Flag leaves were collected from rice plants grown in the paddy field and prepared for TEM analysis. Sample preparation followed the conventional method described by Inada et al. (1998) using a microwave tissue processor (PELCO BioWave® Pro+, Ted Pella, Redding, CA, USA) with microwave irradiation, as previously reported (Mowery and Bauchan 2018), with some modifications. Briefly, samples were vacuum-infiltrated in modified Karnovsky’s fixative (2% [w/v] paraformaldehyde and 2% [w/v] glutaraldehyde in 50 mM sodium cacodylate buffer, pH 7.2) for 60 min, followed by overnight incubation at 4 °C in complete darkness. The specimens were then washed three times with 50 mM sodium cacodylate buffer, pH 7.2, for 10 min each at 4 °C before postfixation with 1% (w/v) osmium tetroxide in 50 mM sodium cacodylate buffer (pH 7.2) using a microwave tissue processor. The postfixed specimens were rinsed twice with distilled water at room temperature and stained en bloc with 0.5% (w/v) uranyl acetate using the microwave tissue processor. To dehydrate the samples, the microwave-assisted dehydration method was used with a graded series of increasing concentrations of ethanol (one change each in 30% [v/v], 50% [v/v], 70% [v/v], and 90% [v/v] aqueous ethanol, followed by three changes in 100% [v/v] ethanol). After dehydration, the specimens were treated twice with propylene oxide under microwave irradiation and infiltrated gradually with increasing concentrations of Spurr’s resin (Spurr 1969) in propylene oxide: one change each in 20% (v/v), 40% (v/v), 60% (v/v), and 80% (v/v) Spurr’s resin, followed by two changes in 100% (v/v) Spurr’s resin, all performed using the microwave tissue processor. The specimens were then embedded in 100% (v/v) Spurr’s resin, polymerized at 70 °C for 24 h in an oven, and sectioned into 70 nm sections using an ultramicrotome (EM UC7, Leica Microsystems, Wetzlar, Germany) equipped with a Diatome diamond knife. The resulting sections were mounted on Formvar-coated copper grids (EMS, Hatfield, PA, USA), and stained with 2% (w/v) uranyl acetate and Reynolds’ lead citrate (Reynolds 1963) for 7 min each at room temperature. Finally, the chloroplast structure in each sample was observed under a transmission electron microscope (Talos L120C, FEI, Czech Republic) operating at 120 kV. Additional information on the detailed microwave procedures used for sample preparation can be found in Additional file 2: Table S3.
Determination of ABA Contents
Flag leaves at the heading stage were collected from WT, osbhlh079-D, OsbHLH079-OE, osbhlh079-1, and osbhlh079-2 grown in the natural rice field. The flag leaves were pulverized in liquid nitrogen using a mortar and pestle, and then subjected to lyophilization for 48 h in a freeze dryer (Bondiro, ilShin® Lab Co. Ltd., Yang-Ju, Republic of Korea). Approximately 150 mg of the freeze-dried sample was carefully weighed and placed in a 5 ml snap-cap centrifuge tube. The sample was dissolved in 1.5 ml of 80% (v/v) methanol containing 1 mM butylated hydroxytoluene (Sigma-Aldrich) as an antioxidant using an ultrasonic bath (Powersonic 420, Hwashin Tech Co. Ltd., Gwangju, Republic of Korea) at 0 °C for 15 min, followed by overnight rotation at 4 °C in complete darkness. After centrifugation at 4,000 g for 15 min at 4 °C, the supernatant was transferred to a 1.5 ml light-proof centrifuge tube and analyzed by enzyme-linked immunosorbent assay (ELISA) using an ABA ELISA kit (MyBioSource, San Diego, CA, USA) according to the manufacturer’s instructions. Absorbance at 450 nm was measured using a UV/VIS spectrophotometer (PowerWave X, BioTek), and ABA levels were determined using a standard curve.
Dual-luciferase Reporter Assay
To construct the reporter plasmids containing the LUC reporter gene under the control of various promoters, promoter fragments of OsABF2 (-1,553 to -1), OsABF4 (-1,709 to + 7), OsABI5 (-1,971 to + 58), or OsNAP (-1,542 to + 84) were cloned, respectively, into the pJD301 vector (Luehrsen et al. 1992). For the effector plasmids, the cDNA of OsbHLH079 was cloned upstream of a sequence encoding six copies of a MYC epitope tag in the pGA3817 vector (Kim et al. 2009). The reporter (2 µg), effector (4 µg), and internal control (1 µg) plasmids were co-transfected into 5 × 104 rice protoplasts using the PEG-mediated transfection method (Yoo et al. 2007). The transfected protoplasts were subsequently suspended in protoplast culture medium (0.4 mM mannitol, 4 mM MES, 15 mM MgCl2, pH 5.8), followed by overnight incubation in complete darkness for 12 h at room temperature. The LUC activity of each cell lysate was determined using the LUC reporter assay system kit (Promega).
Statistical Analysis
Statistical analyses were performed using the two-tailed Student’s t-test with Microsoft Excel 2016. Significant differences between means are indicated by asterisks (*P < 0.05, **P < 0.01, and ***P < 0.001).
Accession Numbers
The sequence data from this article can be found in the Rice Genome Annotation Project (http://rice.uga.edu/analyses_search_locus.shtml), GenBank (https://www.ncbi.nlm.nih.gov/genbank/), and EMBL’s European Bioinformatics Institute (EMBL-EBI; https://www.ebi.ac.uk/) databases under the accession numbers listed in Additional file 2: Table S1.
Data availability
All data supporting the findings of this study are available within the paper and within its supplementary materials published online.
Abbreviations
- ABA:
-
Abscisic acid
- ACC:
-
1-Aminocyclopropane-1-carboxylic acid
- bHLH:
-
basic Helix-Loop-Helix
- BR:
-
Brassinosteroid
- CDG:
-
Chlorophyll degradation gene
- Chl:
-
Chlorophyll
- DAH:
-
Days after heading
- DDI:
-
Day(s) of dark incubation
- DIS:
-
Dark-induced senescence
- DT:
-
Day(s) of treatment
- DW:
-
Dry weight
- ELISA:
-
Enzyme-linked immunosorbent assay
- ET:
-
Ethylene
- FW:
-
Fresh weight
- JA:
-
Jasmonic acid
- LJ:
-
Lamina joint
- MeJA:
-
Methyl jasmonate
- NAC:
-
NAM/ATAF1/2/CUC2
- NS:
-
Not significant
- ORF:
-
Open reading frame
- PCR:
-
Polymerase chain reaction
- RT-qPCR:
-
Reverse transcription quantitative real-time PCR
- SA:
-
Salicylic acid
- SAG:
-
Senescence-associated gene
- Sen-TF:
-
Senescence-associated transcription factor
- TEM:
-
Transmission electron microscopy
- TF:
-
Transcription factor
- WT:
-
Wild-type
References
Ariyarathne MA, Wone BWM (2022) Overexpression of the Selaginella lepidophylla bHLH transcription factor enhances water-use efficiency, growth, and development in Arabidopsis. Plant Sci 315:111129. https://doi.org/10.1016/j.plantsci.2021.111129
Balazadeh S, Kwasniewski M, Caldana C, Mehrnia M, Zanor MI, Xue GP, Mueller-Roeber B (2011) ORS1, an H2O2-responsive NAC transcription factor, controls senescence in Arabidopsis thaliana. Mol Plant 4(2):346–360. https://doi.org/10.1093/mp/ssq080
Bengoa Luoni S, Astigueta FH, Nicosia S, Moschen S, Fernandez P, Heinz R (2019) Transcription factors associated with leaf senescence in crops. Plants 8(10):411. https://doi.org/10.3390/plants8100411
Besagni C, Kessler F (2013) A mechanism implicating plastoglobules in thylakoid disassembly during senescence and nitrogen Starvation. Planta 237:463–470. https://doi.org/10.1007/s00425-012-1813-9
Breeze E, Harrison E, McHattie S, Hughes L, Hickman R, Hill C, Kiddle S, Kim YS, Penfold CA, Jenkins D, Zhang C, Morris K, Jenner C, Jackson S, Thomas B, Tabrett A, League R, Moore JD, Wild DL, Ott S, Rand D, Beynon J, Denby K, Mead A, Buchanan-Wollaston V (2011) High-resolution temporal profiling of transcripts during Arabidopsis leaf senescence reveals a distinct chronology of processes and regulation. Plant Cell 23(3):873–894. https://doi.org/10.1105/tpc.111.083345
Buchanan-Wollaston V (1997) The molecular biology of leaf senescence. J Exp Bot 48(2):181–199. https://doi.org/10.1093/jxb/48.2.181
Buchanan-Wollaston V, Page T, Harrison E, Breeze E, Lim PO, Nam HG, Lin JF, Wu SH, Swidzinski J, Ishizaki K, Leaver CJ (2005) Comparative transcriptome analysis reveals significant differences in gene expression and signalling pathways between developmental and dark/starvation-induced senescence in Arabidopsis. Plant J 42(4):567–585. https://doi.org/10.1111/j.1365-313X.2005.02399.x
Cao J, Liu H, Tan S, Li Z (2023) Transcription factors-regulated leaf senescence: current knowledge, challenges, and approaches. Int J Mol Sci 24(11):9245. https://doi.org/10.3390/ijms24119245
Carmo-Silva E, Andralojc PJ, Scales JC, Driever SM, Mead A, Lawson T, Raines CA, Parry MAJ (2017) Phenotyping of field-grown wheat in the UK highlights contribution of light response of photosynthesis and flag leaf longevity to grain yield. J Exp Bot 68(13):3473–3486. https://doi.org/10.1093/jxb/erx169
Carretero-Paulet L, Galstyan A, Roig-Villanova I, Martínez-García JF, Bilbao-Castro JR, Robertson DL (2010) Genome-wide classification and evolutionary analysis of the bHLH family of transcription factors in Arabidopsis, Poplar, rice, moss, and algae. Plant Physiol 153(3):1398–1412. https://doi.org/10.1104/pp.110.153593
Chen X, Wang Y, Lv B, Li J, Luo L, Lu S, Zhang X, Ma H, Ming F (2014) The NAC family transcription factor OsNAP confers abiotic stress response through the ABA pathway. Plant Cell Physiol 55(3):604–619. https://doi.org/10.1093/pcp/pct204
Clouse SD (2016) Brassinosteroid/abscisic acid antagonism in balancing growth and stress. Dev Cell 38(2):118–120. https://doi.org/10.1016/j.devcel.2016.07.005
Curtis MD, Grossniklaus U (2003) A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol 133(2):462–469. https://doi.org/10.1104/pp.103.027979
Fan L, Zheng S, Wang X (1997) Antisense suppression of phospholipase D alpha retards abscisic acid- and ethylene-promoted senescence of Postharvest Arabidopsis leaves. Plant Cell 9(12):2183–2196. https://doi.org/10.1105/tpc.9.12.2183
Friedrichsen DM, Nemhauser J, Muramitsu T, Maloof JN, Alonso J, Ecker JR, Furuya M, Chory J (2002) Three redundant brassinosteroid early response genes encode putative bHLH transcription factors required for normal growth. Genetics 162(3):1445–1456. https://doi.org/10.1093/genetics/162.3.1445
Gan SS, Hörtensteiner S (2013) Frontiers in plant senescence research: from bench to bank. Plant Mol Biol 82:503–504. https://doi.org/10.1007/s11103-013-0098-0
Gao S, Gao J, Zhu X, Song Y, Li Z, Ren G, Zhou X, Kuai B (2016) ABF2, ABF3, and ABF4 promote ABA-mediated chlorophyll degradation and leaf senescence by transcriptional activation of chlorophyll catabolic genes and senescence-associated genes in Arabidopsis. Mol Plant 9(9):1272–1285. https://doi.org/10.1016/j.molp.2016.06.006
Gepstein S, Glick BR (2013) Strategies to ameliorate abiotic stress-induced plant senescence. Plant Mol Biol 82:623–633. https://doi.org/10.1007/s11103-013-0038-z
Gregersen PL, Culetic A, Boschian L, Krupinska K (2013) Plant senescence and crop productivity. Plant Mol Biol 82:603–622. https://doi.org/10.1007/s11103-013-0013-8
Gremski K, Ditta G, Yanofsky MF (2007) The HECATE genes regulate female reproductive tract development in Arabidopsis thaliana. Development 134(20):3593–3601. https://doi.org/10.1242/dev.011510
Guo Y, Gan S (2005) Leaf senescence: signals, Execution, and regulation. Curr Top Dev Biol 71:83–112. https://doi.org/10.1016/S0070-2153(05)71003-6
Guo Y, Cai Z, Gan S (2004) Transcriptome of Arabidopsis leaf senescence. Plant Cell Environ 27(5):521–549. https://doi.org/10.1111/j.1365-3040.2003.01158.x
Guo Y, Ren G, Zhang K, Li Z, Miao Y, Guo H (2021) Leaf senescence: progression, regulation, and application. Mol Hort 1:5. https://doi.org/10.1186/s43897-021-00006-9
Hacham Y, Holland N, Butterfield C, Ubeda-Tomas S, Bennett MJ, Chory J, Savaldi-Goldstein S (2011) Brassinosteroid perception in the epidermis controls root meristem size. Development 138(5):839–848. https://doi.org/10.1242/dev.061804
Havé M, Marmagne A, Chardon F, Masclaux-Daubresse C (2017) Nitrogen remobilization during leaf senescence: lessons from Arabidopsis to crops. J Exp Bot 68(10):2513–2529. https://doi.org/10.1093/jxb/erw365
Höfgen R, Willmitzer L (1988) Storage of competent cells for Agrobacterium transformation. Nucleic Acids Res 16(20):9877. https://doi.org/10.1093/nar/16.20.9877
Hossain MA, Cho JI, Han M, Ahn CH, Jeon JS, An G, Park PB (2010a) The ABRE-binding bZIP transcription factor OsABF2 is a positive regulator of abiotic stress and ABA signaling in rice. J Plant Physiol 167(17):1512–1520. https://doi.org/10.1016/j.jplph.2010.05.008
Hossain MA, Lee Y, Cho JI, Ahn CH, Lee SK, Jeon JS, Kang H, Lee CH, An G, Park PB (2010b) The bZIP transcription factor OsABF1 is an ABA responsive element binding factor that enhances abiotic stress signaling in rice. Plant Mol Biol 72:557–566. https://doi.org/10.1007/s11103-009-9592-9
Hu Y, Yu D (2014) BRASSINOSTEROID INSENSITIVE2 interacts with ABSCISIC ACID INSENSITIVE5 to mediate the antagonism of brassinosteroids to abscisic acid during seed germination in Arabidopsis. Plant Cell 26(11):4394–4408. https://doi.org/10.1105/tpc.114.130849
Huang Y, Guo Y, Liu Y, Zhang F, Wang Z, Wang H, Wang F, Li D, Mao D, Luan S, Liang M, Chen L (2018) 9-cis-epoxycarotenoid dioxygenase 3 regulates plant growth and enhances multi-abiotic stress tolerance in rice. Front Plant Sci 9:162. https://doi.org/10.3389/fpls.2018.00162
Huang P, Li Z, Guo H (2022) New advances in the regulation of leaf senescence by classical and peptide hormones. Front Plant Sci 13:923136. https://doi.org/10.3389/fpls.2022.923136
Husar S, Berthiller F, Fujioka S, Rozhon W, Khan M, Kalaivanan F, Elias L, Higgins GS, Li Y, Schuhmacher R, Krska R, Seto H, Vaistij FE, Bowles D, Poppenberger B (2011) Overexpression of the UGT73C6 alters brassinosteroid glucoside formation in Arabidopsis thaliana. BMC Plant Biol 11:51. https://doi.org/10.1186/1471-2229-11-51
Inada N, Sakai A, Kuroiwa H, Kuroiwa T (1998) Three-dimensional analysis of the senescence program in rice (Oryza sativa L.) coleoptiles. Planta 205:153–164. https://doi.org/10.1007/s004250050307
Jain M, Nijhawan A, Tyagi AK, Khurana JP (2006) Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem Biophys Res Commun 345(2):646–651. https://doi.org/10.1016/j.bbrc.2006.04.140
Jan S, Abbas N, Ashraf M, Ahmad P (2019) Roles of potential plant hormones and transcription factors in controlling leaf senescence and drought tolerance. Protoplasma 256:313–329. https://doi.org/10.1007/s00709-018-1310-5
Jeon JS, Lee S, Jung KH, Jun SH, Jeong DH, Lee J, Kim C, Jang S, Lee S, Yang K, Nam J, An K, Han MJ, Sung RJ, Choi HS, Yu JH, Choi JH, Cho SY, Cha SS, Kim SI, An G (2000) T-DNA insertional mutagenesis for functional genomics in rice. Plant J 22(6):561–570. https://doi.org/10.1046/j.1365-313x.2000.00767.x
Jeong DH, An S, Kang HG, Moon S, Han JJ, Park S, Lee HS, An K, An G (2002) T-DNA insertional mutagenesis for activation tagging in rice. Plant Physiol 130(4):1636–1644. https://doi.org/10.1104/pp.014357
Jiao BB, Wang JJ, Zhu XD, Zeng LJ, Li Q, He ZH (2012) A novel protein RLS1 with NB-ARM domains is involved in chloroplast degradation during leaf senescence in rice. Mol Plant 5(1):205–217. https://doi.org/10.1093/mp/ssr081
Jing HC, Sturre MJG, Hille J, Dijkwel PP (2002) Arabidopsis onset of leaf death mutants identify a regulatory pathway controlling leaf senescence. Plant J 32(1):51–63. https://doi.org/10.1046/j.1365-313X.2002.01400.x
Kang K, Shim Y, Gi E, An G, Paek NC (2019) Mutation of ONAC096 enhances grain yield by increasing panicle number and delaying leaf senescence during grain filling in rice. Int J Mol Sci 20(20):5241. https://doi.org/10.3390/ijms20205241
Kim SR, Lee DY, Yang JI, Moon S, An G (2009) Cloning vectors for rice. J Plant Biol 52:73–78. https://doi.org/10.1007/s12374-008-9008-4
Kim T, Kang K, Kim SH, An G, Paek NC (2019) OsWRKY5 promotes rice leaf senescence via senescence-associated NAC and abscisic acid biosynthesis pathway. Int J Mol Biol 20(18):4437. https://doi.org/10.3390/ijms20184437
Kim Y, Park SU, Shin DM, Pham G, Jeong YS, Kim SH (2020) ATBS1-INTERACTING FACTOR 2 negatively regulates dark- and brassinosteroid-induced leaf senescence through interactions with INDUCER OF CBF EXPRESSION 1. J Exp Bot 71(4):1475–1490. https://doi.org/10.1093/jxb/erz533
Kondou Y, Nakazawa M, Kawashima M, Ichikawa T, Yoshizumi T, Suzuki K, Ishikawa A, Koshi T, Matsui R, Muto S, Matsui M (2008) RETARDED GROWTH OF EMBRYO1, a new basic helix-loop-helix protein, expresses in endosperm to control embryo growth. Plant Physiol 147(4):1924–1935. https://doi.org/10.1104/pp.108.118364
Kong Z, Li M, Yang W, Xu W, Xue Y (2006) A novel nuclear-localized CCCH-type zinc finger protein, OsDOS, is involved in delaying leaf senescence in rice. Plant Physiol 141(4):1376–1388. https://doi.org/10.1104/pp.106.082941
Kusaba M, Ito H, Morita R, Iida S, Sato Y, Fujimoto M, Kawasaki S, Tanaka R, Hirochika H, Nishimura M, Tanaka A (2007) Rice NON-YELLOW COLORING1 is involved in light-harvesting complex II and grana degradation during leaf senescence. Plant Cell 19(4):1362–1375. https://doi.org/10.1105/tpc.106.042911
Lee S, Masclaux-Daubresse C (2021) Current understanding of leaf senescence in rice. Int J Mol Sci 22(9):4515. https://doi.org/10.3390/ijms22094515
Lee RH, Wang CH, Huang LT, Chen SCG (2001) Leaf senescence in rice plants: cloning and characterization of senescence up-regulated genes. J Exp Bot 52(358):1117–1121. https://doi.org/10.1093/jexbot/52.358.1117
Li J, Chory J (1997) A putative leucine-rich repeat receptor kinase involved in brassinosteroid signal transduction. Cell 90(5):929–938. https://doi.org/10.1016/S0092-8674(00)80357-8
Li J, Nagpal P, Vitart V, McMorris TC, Chory J (1996) A role for brassinosteroids in light-dependent development of Arabidopsis. Science 272(5260):398–401. https://doi.org/10.1126/science.272.5260.398
Li X, Duan X, Jiang H, Sun Y, Tang Y, Yuan Z, Guo J, Liang W, Chen L, Yin J, Ma H, Wang J, Zhang D (2006) Genome-wide analysis of basic/helix-loop-helix transcription factor family in rice and Arabidopsis. Plant Physiol 141(4):1167–1184. https://doi.org/10.1104/pp.106.080580
Li Z, Peng J, Wen X, Guo H (2012) Gene network analysis and functional studies of senescence-associated genes reveal novel regulators of Arabidopsis leaf senescence. J Integr Plant Biol 54(8):526–539. https://doi.org/10.1111/j.1744-7909.2012.01136.x
Li Z, Zhang Y, Zou D, Zhao Y, Wang HL, Zhang Y, Xia X, Luo J, Guo H, Zhang Z (2020) LSD 3.0: a comprehensive resource for the leaf senescence research community. Nucleic Acids Res 48(D1):D1069–D1075. https://doi.org/10.1093/nar/gkz898
Li C, Yan C, Sun Q, Wang J, Yuan C, Mou Y, Shan S, Zhao X (2021) The bHLH transcription factor AhbHLH112 improves the drought tolerance of peanut. BMC Plant Biol 21:540. https://doi.org/10.1186/s12870-021-03318-6
Liang C, Wang Y, Zhu Y, Tang J, Hu B, Liu L, Ou S, Wu H, Sun X, Chu J, Chu C (2014) OsNAP connects abscisic acid and leaf senescence by fine-tuning abscisic acid biosynthesis and directly targeting senescence-associated genes in rice. Proc Natl Acad Sci USA 111(27):10013–10018. https://doi.org/10.1073/pnas.1321568111
Lim PO, Kim HJ, Nam HG (2007) Leaf senescence. Annu Rev Plant Biol 58:115–136. https://doi.org/10.1146/annurev.arplant.57.032905.105316
Lin JF, Wu SH (2004) Molecular events in senescing Arabidopsis leaves. Plant J 39(4):612–628. https://doi.org/10.1111/j.1365-313X.2004.02160.x
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2–∆∆CT method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Lu G, Gao C, Zheng X, Han B (2009) Identification of OsbZIP72 as a positive regulator of ABA response and drought tolerance in rice. Planta 229:605–615. https://doi.org/10.1007/s00425-008-0857-3
Luehrsen KR, de Wet JR, Walbot V (1992) Transient expression analysis in plants using firefly luciferase reporter gene. Methods Enzymol 216:397–414. https://doi.org/10.1016/0076-6879(92)16037-k
Mao C, Lu S, Lv B, Zhang B, Shen J, He J, Luo L, Xi D, Chen X, Ming F (2017) A rice NAC transcription factor promotes leaf senescence via ABA biosynthesis. Plant Physiol 174(3):1747–1763. https://doi.org/10.1104/pp.17.00542
Masclaux-Daubresse C, Reisdorf-Cren M, Orsel M (2008) Leaf nitrogen remobilisation for plant development and grain filling. Plant Biol 10(s1):23–36. https://doi.org/10.1111/j.1438-8677.2008.00097.x
Miao Y, Smykowski A, Zentgraf U (2008) A novel upstream regulator of WRKY53 transcription during leaf senescence in Arabidopsis thaliana. Plant Biol 10(s1):110–120. https://doi.org/10.1111/j.1438-8677.2008.00083.x
Miao J, Guo D, Zhang J, Huang Q, Qin G, Zhang X, Wan J, Gu H, Qu LJ (2013) Targeted mutagenesis in rice using CRISPR-Cas system. Cell Res 23(10):1233–1236. https://doi.org/10.1038/cr.2013.123
Moore B, Zhou L, Rolland F, Hall Q, Cheng WH, Liu YX, Hwang I, Jones T, Sheen J (2003) Role of the Arabidopsis glucose sensor HXK1 in nutrient, light, and hormonal signaling. Science 300(5617):332–336. https://doi.org/10.1126/science.1080585
Morita R, Sato Y, Masuda Y, Nishimura M, Kusaba M (2009) Defect in non-yellow coloring 3, an α/β hydrolase-fold family protein, causes a stay-green phenotype during leaf senescence in rice. Plant J 59(6):940–952. https://doi.org/10.1111/j.1365-313X.2009.03919.x
Mowery J, Bauchan G (2018) Optimization of rapid microwave processing of botanical samples for transmission electron microscopy. Microsc Microanal 24(S1):1202–1203. https://doi.org/10.1017/S1431927618006499
Naito Y, Hino K, Bono H, Ui-Tei K (2015) CRISPRdirect: software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 31(7):1120–1123. https://doi.org/10.1093/bioinformatics/btu743
Ooms G, Hooykaas PJJ, Van Veen RJM, Van Beelen PV, Regensburg-Tuïnk TJG, Schilperoort RA (1982) Octopine Ti-plasmid deletion mutants of Agrobacterium tumefaciens with emphasis on the right side of the T-region. Plasmid 7(1):15–29. https://doi.org/10.1016/0147-619X(82)90023-3
Park SY, Yu JW, Park JS, Li J, Yoo SC, Lee NY, Lee SK, Jeong SW, Seo HS, Koh HJ, Jeon JS, Park YI, Paek NC (2007) The senescence-induced staygreen protein regulates chlorophyll degradation. Plant Cell 19(5):1649–1664. https://doi.org/10.1105/tpc.106.044891
Park DY, Shim Y, Gi E, Lee BD, An G, Kang K, Paek NC (2018) The MYB-related transcription factor RADIALIS-LIKE3 (OsRL3) functions in ABA-induced leaf senescence and salt sensitivity in rice. Environ Exp Bot 156:86–95. https://doi.org/10.1016/j.envexpbot.2018.08.033
Peng Z, Zhang Y (2017) Methyl jasmonate induces the apoptosis of human Colorectal cancer cells via downregulation of EZH2 expression by microRNA-101. Mol Med Rep 15(2):957–962. https://doi.org/10.3892/mmr.2016.6061
Pennell RI, Lamb C (1997) Programmed cell death in plants. Plant Cell 9(7):1157–1168. https://doi.org/10.1105/tpc.9.7.1157
Peres ALGL, Soares JS, Tavares RG, Righetto G, Zullo MA, Mandava NB, Menossi M (2019) Brassinosteroids, the sixth class of phytohormones: a molecular view from the discovery to hormonal interactions in plant development and stress adaptation. Int J Mol Sci 20(2):331. https://doi.org/10.3390/ijms20020331
Piao W, Han SH, Sakuraba Y, Paek NC (2017) Rice 7-hydroxymethyl chlorophyll a reductase is involved in the promotion of chlorophyll degradation and modulates cell death signaling. Mol Cells 40(10):773–786. https://doi.org/10.14348/molcells.2017.0127
Piao W, Kim SH, Lee BD, An G, Sakuraba Y, Paek NC (2019) Rice transcription factor OsMYB102 delays leaf senescence by down-regulating abscisic acid accumulation and signaling. J Exp Bot 70(10):2699–2715. https://doi.org/10.1093/jxb/erz095
Pires N, Dolan L (2010) Origin and diversification of basic-helix-loop-helix proteins in plants. Mol Biol Evol 27(4):862–874. https://doi.org/10.1093/molbev/msp288
Porra RJ, Thompson WA, Kriedemann PE (1989) Determination of accurate extinction coefficients and simultaneous equations for assaying chlorophylls a and b extracted with four different solvents: verification of the concentration of chlorophyll standards by atomic absorption spectroscopy. Biochim Biophys Acta - Bioenerg 975(3):384–394. https://doi.org/10.1016/S0005-2728(89)80347-0
Pružinská A, Tanner G, Aubry S, Anders I, Moser S, Müller T, Ongania KH, Kräutler B, Youn JY, Liljegren SJ, Hörtensteiner S (2005) Chlorophyll breakdown in senescent Arabidopsis leaves. Characterization of chlorophyll catabolites and of chlorophyll catabolic enzymes involved in the degreening reaction. Plant Physiol 139(1):52–63. https://doi.org/10.1104/pp.105.065870
Qi T, Wang J, Huang H, Liu B, Gao H, Liu Y, Song S, Xie D (2015) Regulation of jasmonate-induced leaf senescence by antagonism between bHLH subgroup IIIe and IIId factors in Arabidopsis. Plant Cell 27(6):1634–1649. https://doi.org/10.1105/tpc.15.00110
Quiles MJ, García C, Cuello J (1995) Differential effects of abscisic acid and methyl jasmonate on endoproteinases in senescing barley leaves. Plant Growth Regul 16:197–204. https://doi.org/10.1007/BF00029541
Reynolds ES (1963) The use of lead citrate at high pH as an electron-opaque stain in electron microscopy. J Cell Biol 17(1):208–121. https://doi.org/10.1083/jcb.17.1.208
Sağlam-Çağ S (2007) The effect of epibrassinolide on senescence in wheat leaves. Biotechnol Biotechnol Equip 21(1):63–65. https://doi.org/10.1080/13102818.2007.10817415
Sakuraba Y, Piao W, Lim JH, Han SH, Kim YS, An G, Paek NC (2015) Rice ONAC106 inhibits leaf senescence and increases salt tolerance and tiller angle. Plant Cell Physiol 56(12):2325–2339. https://doi.org/10.1093/pcp/pcv144
Sakuraba Y, Kim D, Han SH, Kim SH, Piao W, Yanagisawa S, An G, Paek NC (2020) Multilayered regulation of membrane-bound ONAC054 is essential for abscisic acid-induced leaf senescence in rice. Plant Cell 32(3):630–649. https://doi.org/10.1105/tpc.19.00569
Sato Y, Morita R, Katsuma S, Nishimura M, Tanaka A, Kusaba M (2009) Two short-chain dehydrogenase/reductases, NON-YELLOW COLORING 1 and NYC1-LIKE, are required for chlorophyll b and light-harvesting complex II degradation during senescence in rice. Plant J 57(1):120–131. https://doi.org/10.1111/j.1365-313X.2008.03670.x
Schippers JHM, Schmidt R, Wagstaff C, Jing HC (2015) Living to die and dying to live: the survival strategy behind leaf senescence. Plant Physiol 169(2):914–930. https://doi.org/10.1104/pp.15.00498
Seo H, Kim SH, Lee BD, Lim JH, Lee SJ, An G, Paek NC (2020) The rice basic Helix-Loop-Helix 79 (OsbHLH079) determines leaf angle and grain shape. Int J Mol Sci 21(6):2090. https://doi.org/10.3390/ijms21062090
Singh S, Singh A, Nandi AK (2016) The rice OsSAG12-2 gene codes for a functional protease that negatively regulates stress-induced cell death. J Biosci 41(3):445–453. https://doi.org/10.1007/s12038-016-9626-9
Smart CM (1994) Gene expression during leaf senescence. New Phytol 126(3):419–448. https://doi.org/10.1111/j.1469-8137.1994.tb04243.x
Spurr AR (1969) A low-viscosity epoxy resin embedding medium for electron microscopy. J Ultrastruct Res 26(1–2):31–43. https://doi.org/10.1016/S0022-5320(69)90033-1
Su Y, Hu S, Zhang B, Ye W, Niu Y, Guo L, Qian Q (2017) Characterization and fine mapping of a new early leaf senescence mutant es3(t) in rice. Plant Growth Regul 81:419–431. https://doi.org/10.1007/s10725-016-0219-2
Tang Y, Li M, Chen Y, Wu P, Wu G, Jiang H (2011) Knockdown of OsPAO and OsRCCR1 cause different plant death phenotypes in rice. J Plant Physiol 168(16):1952–1959. https://doi.org/10.1016/j.jplph.2011.05.026
van der Graaff E, Schwacke R, Schneider A, Desimone M, Flügge UI, Kunze R (2006) Transcription analysis of Arabidopsis membrane transporters and hormone pathways during developmental and induced leaf senescence. Plant Physiol 141(2):776–792. https://doi.org/10.1104/pp.106.079293
Ververidis P, John P (1991) Complete recovery in vitro of ethylene-forming enzyme activity. Phytochemistry 30(3):725–727. https://doi.org/10.1016/0031-9422(91)85241-Q
Vogler F, Schmalzl C, Englhart M, Bircheneder M, Sprunck S (2014) Brassinosteroids promote Arabidopsis pollen germination and growth. Plant Reprod 27:153–167. https://doi.org/10.1007/s00497-014-0247-x
Woo HR, Kim HJ, Lim PO, Nam HG (2019) Leaf senescence: systems and dynamics aspects. Annu Rev Plant Biol 70:347–376. https://doi.org/10.1146/annurev-arplant-050718-095859
Xiang Y, Tang N, Du H, Ye H, Xiong L (2008) Characterization of OsbZIP23 as a key player of the basic leucine zipper transcription factor family for conferring abscisic acid sensitivity and salinity and drought tolerance in rice. Plant Physiol 148(4):1938–1952. https://doi.org/10.1104/pp.108.128199
Xie W, Li X, Wang S, Yuan M (2022) OsWRKY53 promotes abscisic acid accumulation to accelerate leaf senescence and inhibit seed germination by downregulating abscisic acid catabolic genes in rice. Front Plant Sci 12:816156. https://doi.org/10.3389/fpls.2021.816156
Xue LW, Du JB, Yang H, Xu F, Yuan S, Lin HH (2009) Brassinosteroids counteract abscisic acid in germination and growth of Arabidopsis. Z Naturforsch C 64(3–4):225–230. https://doi.org/10.1515/znc-2009-3-413
Yang Y, Xu J, Huang L, Leng Y, Dai L, Rao Y, Chen L, Wang Y, Tu Z, Hu J, Ren D, Zhang G, Zhu L, Guo L, Qian Q, Zeng D (2016a) PGL, encoding chlorophyllide a oxygenase 1, impacts leaf senescence and indirectly affects grain yield and quality. J Exp Bot 67(5):1297–1310. https://doi.org/10.1093/jxb/erv529
Yang X, Bai Y, Shang J, Xin R, Tang W (2016b) The antagonistic regulation of abscisic acid-inhibited root growth by brassinosteroids is partially mediated via direct suppression of ABSCISIC ACID INSENSITIVE 5 expression by BRASSINAZOLE RESISTANT 1. Plant Cell Environ 39(9):1994–2003. https://doi.org/10.1111/pce.12763
Yang S, Xu K, Chen S, Li T, Xia H, Chen L, Liu H, Luo L (2019) A stress-responsive bZIP transcription factor OsbZIP62 improves drought and oxidative tolerance in rice. BMC Plant Biol 19:260. https://doi.org/10.1186/s12870-019-1872-1
Yin Y, Wang ZY, Mora-Garcia S, Li J, Yoshida S, Asami T, Chory J (2002) BES1 accumulates in the nucleus in response to brassinosteroids to regulate gene expression and promote stem elongation. Cell 109(2):181–191. https://doi.org/10.1016/S0092-8674(02)00721-3
Yoo SD, Cho YH, Sheen J (2007) Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat Protoc 2:1565–1572. https://doi.org/10.1038/nprot.2007.199
Zakari SA, Asad MAU, Han Z, Guan X, Zaidi SHR, Gang P, Cheng F (2020) Senescence-related translocation of nonstructural carbohydrate in rice leaf sheaths under different nitrogen supply. Agron J 112(3):1601–1616. https://doi.org/10.1002/agj2.20163
Zhang W, Sun Y, Timofejeva L, Chen C, Grossniklaus U, Ma H (2006) Regulation of Arabidopsis tapetum development and function by DYSFUNCTIONAL TAPETUM1 (DYT1) encoding a putative bHLH transcription factor. Development 133(16):3085–3095. https://doi.org/10.1242/dev.02463
Zhang L, Zhao G, Jia J, Liu X, Kong X (2012) Molecular characterization of 60 isolated wheat MYB genes and analysis of their expression during abiotic stress. J Exp Bot 63(1):203–214. https://doi.org/10.1093/jxb/err264
Zhao D, Derkx AP, Liu DC, Buchner P, Hawkesford MJ (2015) Overexpression of a NAC transcription factor delays leaf senescence and increases grain nitrogen concentration in wheat. Plant Biol 17(4):904–913. https://doi.org/10.1111/plb.12296
Acknowledgements
We thank Gynheung An (Kyung Hee University, South Korea) for the donation of osbhlh079-D and Li-Jia Qu (Beijing University, Beijing, China) for providing the pOs-sgRNA and pH-Ubi-cas9-7 vectors.
Funding
This research was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (NRF-2022R1A2C1091553).
Author information
Authors and Affiliations
Contributions
N-CP conceived and supervised the project. S-HK and JY performed the research with the help of HK. S-JL analyzed the data. N-CP, S-HK, JY, HK, and S-JL wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, SH., Yoon, J., Kim, H. et al. Rice Basic Helix-Loop-Helix 079 (OsbHLH079) Delays Leaf Senescence by Attenuating ABA Signaling. Rice 16, 60 (2023). https://doi.org/10.1186/s12284-023-00673-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12284-023-00673-w