Abstract
Background
The Living Atlas is an open source platform used to collect, visualise and analyse biodiversity data from multiple sources, and serves as the national biodiversity data hub in many countries. Although powerful, the Living Atlas has had limited functionality for species occurrence data derived from DNA sequences. As a step toward integrating this fast-growing data source into the platform, we developed the Amplicon Sequence Variant (ASV) portal: a web interface to sequence-based biodiversity observations in the Living Atlas.
Results
The ASV portal allows data providers to submit denoised metabarcoding output to the Living Atlas platform via an intermediary ASV database. It also enables users to search for existing ASVs and associated Living Atlas records using the Basic Local Alignment Search Tool, or via filters on taxonomy and sequencing details. The ASV portal is a Python-Flask/jQuery web interface, implemented as a multi-container docker service, and is an integral part of the Swedish Biodiversity Data Infrastructure.
Conclusion
The ASV portal is a web interface that effectively integrates biodiversity data derived from DNA sequences into the Living Atlas platform.
Similar content being viewed by others
Background
Biodiversity research is rapidly developing into big-data science, enabling researchers to model processes that affect entire biotas and to predict ecosystem-wide effects of environmental change. To facilitate this, infrastructures that provide open access to species observation data for all types of life are crucial. The Living Atlas (LA) is an infrastructure for integration of biodiversity data from multiple sources with environmental and contextual information. It was originally developed by the Atlas of Living Australia, in response to growing demands of the biodiversity research community for open access to extensive databases and analysis tools [1]. It is, however, also supported by the Global Biodiversity Information Facility [2], and now serves as the main biodiversity data hub in 27 countries and regions [3]. The software is developed in open collaboration, and more than 100 developers have contributed to the codebase.
Although the LA accommodates less traditional data types such as images, or output from animal tracking devices, it has so far offered limited functionality for DNA sequence-based observations. Meanwhile, molecular methods for species observation, in particular metabarcoding (amplicon sequencing of taxonomic marker genes) of environmental DNA (eDNA) and bulk samples, are becoming increasingly important tools for documenting the diversity of life [4], especially in the microscopic realm (prokaryotes, protists and fungi; see e.g., [5]).
We identified three features that would make the LA platform more useful for handling occurrence data derived from metabarcoding: (1) the option to store processed barcode sequences in the form of Amplicon Sequence Variants (ASVs), underlying occurrences in the atlas, and to use the Basic Local Alignment Search Tool (BLAST; [6]) to find such occurrences, (2) the possibility of searching for ASVs and occurrence records based on sequencing details, such as target genes and primers, and (3) a dynamic approach to taxonomic annotation of observed ASVs, allowing for easy updates as reference databases develop. Below, we present an application that provides these features, and functions as a semi-integrated LA module.
Implementation
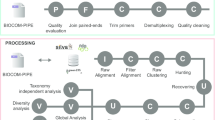
The ASV portal is a web interface to sequence-based biodiversity observations in the LA platform, and is implemented as five separate microservices that are defined and orchestrated with Docker Compose ([7]; Fig. 1). The main application includes a Python-Flask [8] backend, a jQuery [9] frontend, and a uWSGI [10] application server that forwards requests to Flask from the NGINX reverse proxy server [11]. Flask, in turn, retrieves ASV and occurrence records from a PostgreSQL [12] database, turned into a RESTful API by the PostgREST [13] server. In addition, the main application delegates BLAST jobs to a worker, spawning additional worker processes when needed. Finally, the service configuration includes volumes for persistent storage of e.g. file uploads, BLAST and ASV database records.
Results
The ASV portal provides options to submit and search for denoised metabarcoding data and associated occurrence records via intermediary ASV and BLAST databases (Fig. 1).
Data providers submit their data using a spreadsheet template based on the Darwin Core (DwC) standard for biodiversity data [14]. Specifically, the template corresponds to a DwC event core with associated contextual (‘extended Measurement or Facts’) and sequence-related (‘DNA derived data’ [15]) extensions. Each event is also associated with occurrences reported in ASV table format, i.e. as read counts given per sample (row) and ASV (column), rather than in the typical DwC occurrence format.
Submitted data files are curated and imported into the ASV database by portal administrators. A standard taxonomic annotation is then applied to each ASV, using current versions of selected classification algorithms and reference databases. The database schema also allows for successive re-annotations, enabling improved taxonomic accuracy and resolution as reference databases develop. Each DwC occurrence is, however, also assigned a unique taxon ID, based on the MD5 checksum of the underlying ASV sequence. This ensures that identification is consistent between data providers, and unaffected by changes in the mapping of ASVs to different taxon concepts.
Imported datasets are shared with GBIF and LA via the Integrated Publishing Toolkit [16]. The ASV database schema includes linked DwC views that can be accessed and filtered to create a new data resource in the IPT. The portal administrator then invites the data provider to fill in dataset-level metadata in the IPT form, before the dataset is formally published and made available to LA users.
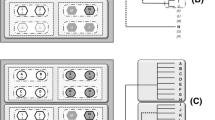
The ASV portal provides two options for finding ASVs and published LA records: BLAST or FILTER search. In the BLAST form, users can paste in FASTA sequences, and set the minimum identity and query coverage of returned hits. Sequences are then aligned against a BLAST database that portal administrators rebuild when new data are imported into the ASV database. The FILTER form lets the user filter out ASVs based on sequencing details (e.g. target gene) and taxonomy. Search results are presented in similar, paginated tables in which users can select specific ASV records. Users can download these directly, in Excel or delimited text format, or choose to explore associated occurrence records in the LA platform. An illustrated use case for ASV portal search is given in Fig. 2, and a video tutorial covering both data submission and searching is available on YouTube [17].
Use-case: Searching for Acinetobacter sequences and occurrence records in the ASV portal. A user interested in finding denoised sequences and associated occurrence records of a specific taxon, is presented with two search options in the start page of the ASV portal: BLAST and filter search (a). Filtering for ASVs derived from the 16S rRNA target gene in the genus Acinetobacter (b), produces a list of six ASVs, available for direct download (c). Alternatively, BLAST:ing against a known marker sequence from the targeted taxon (d), results in a corresponding list of ASVs (e). The user then opts for showing associated occurrence records in the main atlas platform (f), where data can be visualised and analysed together with other species observations as well as environmental and contextual data layers
Future development
The ASV portal is currently an integral part of the Swedish LA instance [18], but given the rate at which sequence-based biodiversity data are being collected around the world, we envision that the LA community at large will benefit from our initiative to integrate this data source. We aim to keep the portal up to date, and welcome user requests, as well as contributions from biodiversity informatics programmers that want to join this open source project. The application will likely need to be optimised to support larger amounts of data in the future, and possible development includes adding an option for direct API access to data, by providing custom R and Python client libraries for this.
Conclusion
The ASV portal is a Python-Flask web interface that integrates DNA sequence-based biodiversity data into the Living Atlas platform, where they can be combined with a multitude of other data sources to e.g. model processes that affect entire biotas, and to predict system-wide effects of environmental change. Most importantly, the portal provides straightforward options to submit data from metabarcoding studies in a convenient (ASV table) format, and to search for ASVs and associated occurrence records using sequence alignment (BLAST), as well as filters on e.g. target genes or primers. The application is developed in open collaboration, and containerized for easy deployment on any platform.
Availability and requirements
Project name: ASV portal.
Project home page: https://asv-portal.biodiversitydata.se (running instance), https://github.com/biodiversitydata-se/mol-mod (development repository).
Archived version: https://zenodo.org/record/6394275.
Operating systems: Platform independent.
Programming language: Python, jQuery.
Other requirements: Docker and Docker Compose.
License: CC0 1.0 Universal (jQuery, DataTables and select2 components: MIT license).
Any restrictions to use by non-academics: None.
Availability of data and materials
The dataset supporting the conclusions of this article is available from the db-backup folder of the development repository (https://github.com/biodiversitydata-se/mol-mod) and the archived resource (https://zenodo.org/record/6394275). A video tutorial of the application is available on YouTube (https://www.youtube.com/watch?v=9P1qcJqZQtA).
Abbreviations
- API:
-
Application programming interface
- ASV:
-
Amplicon sequence variant
- BLAST:
-
Basic local alignment search tool
- DwC(A):
-
Darwin core (archive)
- eDNA:
-
Environmental DNA
- GBIF:
-
Global biodiversity information facility
- IPT:
-
Integrated publishing toolkit
- LA:
-
Living Atlas
- SBDI:
-
Swedish biodiversity data infrastructure
References
Belbin L, Wallis E, Hobern D, Zerger A. The Atlas of Living Australia: History, current state and future directions. Biodiv Data J. 2021;9:1–35.
What is GBIF? https://www.gbif.org/what-is-gbif. Accessed 24 Jan 2022.
Living atlases. https://living-atlases.gbif.org. Accessed 24 Jan 2022.
Deiner K, Bik HM, Mächler E, Seymour M, Lacoursière-Roussel A, Altermatt F, et al. Environmental DNA metabarcoding: transforming how we survey animal and plant communities. Mol Ecol. 2017;26:5872–95.
Hugerth LW, Andersson AF. Analysing microbial community composition through amplicon sequencing: from sampling to hypothesis testing. Front Microbiol. 2017;8(SEP):1–22.
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–402.
Overview of Docker compose. 2022. https://docs.docker.com/compose/. Accessed 11 May 2022.
Welcome to flask—flask documentation (2.1.X). https://flask.palletsprojects.com/en/2.1.x/. Accessed 11 May 2022.
JS Foundation-js. Foundation. JQuery. https://jquery.com/. Accessed 11 May 2022.
The uWSGI project—uWSGI 2.0 documentation. https://uwsgi-docs.readthedocs.io/en/latest/. Accessed 11 May 2022.
NGINX reverse proxy. https://docs.nginx.com/nginx/admin-guide/web-server/reverse-proxy/. Accessed 11 May 2022.
PostgreSQL Global Development Group. PostgreSQL. PostgreSQL. 2022. https://www.postgresql.org/. Accessed 11 May 2022.
PostgREST documentation—PostgREST 9.0.0 documentation. https://postgrest.org/en/stable/. Accessed 11 May 2022.
Wieczorek J, Bloom D, Guralnick R, Blum S, Doring M, Giovanni R, et al. Darwin core: an evolving community-developed biodiversity data standard. PLoS ONE. 2012;7: e29715.
Andersson AF, Bissett A, Finstad AG, Fossøy F, Grosjean M, Hope M, et al. Publishing DNA-derived data through biodiversity data platforms. Version 1.0. 2020. https://docs.gbif.org/publishing-dna-derived-data/1.0/en/. Accessed 24 Jan 2022.
IPT: the integrated publishing toolkit. https://www.gbif.org/ipt. Accessed 24 Jan 2022.
Prager M. The Swedish ASV portal. 2021. https://www.youtube.com/watch?v=9P1qcJqZQtA. Accessed 4 Nov 2022.
Swedish Biodiversity Data Infrastructure (SBDI). 2021. https://biodiversitydata.se/. Accessed 4 May 2022.
Acknowledgements
We thank Martin Norling, Andreas Kusalananda Kähäri and Pontus Freyhult at National Bioinformatics Infrastructure Sweden (NBIS) for advice and coding contributions, as well as Manash Shah, Anna Rosling, Jeanette Tångrot and two anonymous reviewers for useful feedback.
Funding
Open access funding provided by Royal Institute of Technology. This work is part of the Swedish Biodiversity Data Infrastructure (SBDI), funded by its partner organizations, and the Swedish Research Council VR through grant no. 2019-00242.
Author information
Authors and Affiliations
Contributions
AA, DL and FR identified the problem and conceptualised the solution. MP designed and implemented the application (with support from consultants at NBIS), under supervision by AA and DL. MP drafted the initial manuscript. All authors read, edited and approved the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Prager, M., Lundin, D., Ronquist, F. et al. ASV portal: an interface to DNA-based biodiversity data in the Living Atlas. BMC Bioinformatics 24, 6 (2023). https://doi.org/10.1186/s12859-022-05120-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12859-022-05120-z