Abstract
Background
Aminoglycosides have been a cornerstone of the treatment of nosocomial infections caused by Pseudomonas aeruginosa for over 80 years. However, escalating emergence of resistance poses a significant challenge. Therefore, this study aimed to investigate the prevailing patterns of aminoglycoside resistance among clinical isolates of P. aeruginosa in Iran; as well as the underlying resistance mechanisms observed in patients referred to Ardabil hospitals.
Methods
A total of 200 isolates from five hospitals were evaluated. The resistance profiles of P. aeruginosa isolates to tobramycin, amikacin, and netilmicin were determined using the disk diffusion method. The capacity of aminoglycoside-resistant isolates to form biofilms was assessed through a phenotypic assay, and the results were confirmed using the gene amplification technique. The presence of genes associated with aminoglycoside resistance was detected using polymerase chain reaction (PCR). Quantitative reverse transcription PCR (qRT-PCR) was performed to measure the expression levels of genes encoding the MexXY-OprM efflux pump and PhoPQ two-component system (TCS).
Results
The prevalence of aminoglycoside-resistant P. aeruginosa isolates was 48%, with 94.7% demonstrating multidrug resistance (MDR). All aminoglycoside-resistant P. aeruginosa strains exhibited biofilm-forming capabilities and harbored all the genes associated with biofilm production. Among the nine genes encoding 16S rRNA methylase and aminoglycoside-modifying enzymes, three genes were detected in these isolates: aac(6’)-Ib (85.4%), ant(2’’)-Ia (18.7%), and aph(3’)-VI (3.1%). Additionally, all aminoglycoside-resistant P. aeruginosa isolates carried mexY and phoP genes, although the expression levels of mexY and phoP were 75% and 87.5%, respectively.
Conclusion
Given the considerably high prevalence of aminoglycoside-resistant P. aeruginosa strains, urgent measures are warranted to transition towards the use of novel aminoglycosides and to uphold vigilant surveillance of resistance patterns.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Hospitalized patients face a heightened risk of acquiring diverse hospital-acquired infections, among which Pseudomonas aeruginosa stands out as a prominent causative agent, particularly in intensive care units (ICUs) [1,2,3,4]. Commonly prescribed antibiotics for combating P. aeruginosa infections in hospitals settings encompass β-lactams, fluoroquinolones, and aminoglycosides [5, 6]. However, effective eradication of this nosocomial pathogen from hospital environments and curbing mortality rates in severe infections present considerable challenges because of its propensity to develop resistance against a broad spectrum of antiseptics, disinfectants, and antibiotics. This has led to increased costs and prolonged hospital stay [3, 7, 8].
Aminoglycosides, known for their bactericidal properties and synergistic effects with β-lactams, are commonly employed in the treatment of various P. aeruginosa infections, notably pulmonary infections in patients with cystic fibrosis (CF) [8]. However, the emergence of aminoglycoside-resistant P. aeruginosa strains has become a global concern since its initial recognition in the 1960s [8]. The resistance of P. aeruginosa to aminoglycosides can be attributed to several mechanisms, including (1) alterations in outer membrane permeability facilitated by lipopolysaccharide (LPS) modifications mediated by the PhoPQ two-component system (TCS); (2) efflux systems such as MexXY-OprM; (3) ribosomal changes caused by 16S rRNA ribosomal methyltransferases; (4) production of antibiotic-inactivating enzymes such as aminoglycoside phosphotransferases (APH), aminoglycoside acetyltransferases (AAC), and aminoglycoside nucleotidyltransferases (ANT); and (5) biofilm formation [8, 9].
Previous investigations conducted in Ardabil city have underscored elevated rates of resistance and elucidated resistance mechanisms in clinical isolates of P. aeruginosa against penicillins, β-lactam combination agents, cephems, monobactams, carbapenems, fluoroquinolones, and lipopeptides [1, 2, 4,5,6, 10]. Nevertheless, comprehensive data regarding aminoglycoside-resistant strains and their underlying resistance mechanisms in this region remain elusive.
The aims of this study encompassed the investigation of the following aspects: (1) the resistance patterns of P. aeruginosa strains to aminoglycosides, (2) assessing the prevalence of genes encoding 16S rRNA methylase (including rmtA, rmtB, rmtC, rmtD, and armA genes) and aminoglycoside-modifying enzymes (such as aac(6’)-Ib, aac(6’)-IIa, aph(3’)-VI, and ant(2’’)-Ia genes), (3) evaluating both phenotypic and genotypic biofilm formation through the analysis of genes algD, pslD, pelF, Ppgl, and PAPI-1, and (4) investigating the expression levels of genes encoding the MexXY-OprM efflux pump (mexY gene) and the PhoPQ TCS (phoP gene) among aminoglycoside-resistant P. aeruginosa strains isolated from various patient specimens in Ardabil hospitals.
Materials and methods
Bacterial isolates
In this study, glycerol stocks of P. aeruginosa clinical isolates (n = 200) were utilized, sourced from five affiliated hospitals of Ardabil Medical University between June 2019 and May 2023. These isolates were previously identified through standard phenotypic and genotypic testing protocols. P. aeruginosa clinical isolates had been collected from inpatients and outpatients and duplicate isolates were excluded from the study.
Antimicrobial susceptibility testing
The resistance profiles of P. aeruginosa to three aminoglycosides (i.e., tobramycin, amikacin, and netilmicin) were assessed using the disk diffusion method, following the guidelines outlined by the Clinical and Laboratory Standards Institute (CLSI, 2024) [11]. The disk diffusion procedure was conducted according to established protocols, with P. aeruginosa (ATCC 27,853) utilized as a control strain for consistency [4]. Colistin resistance was previously done using the colistin agar test [10].
Biofilm formation assay
Biofilm formation ability was determined for aminoglycoside-resistance P. aeruginosa clinical isolates using the colorimetric microtiter plate, following the method previously described by Jabalameli et al. [12]. In this procedure, the optical density (OD) of each distinct bacterial strain was measured at 590 nm using an ELISA reader and the results were recorded as ODs (the OD of a strain) ≤ ODc (the OD of the negative control) (no biofilm producer), ODc < ODs < 2× ODc (weak biofilm producer), 2× ODc < ODs < 4× ODc (moderate biofilm producer), and 4× ODc < ODs (strong biofilm producer).
Amplification of aminoglycosides resistance genes
The genes responsible for biofilm formation (i.e., algD, pslD, pelF, Ppgl, and PAPI-1 genes), 16S rRNA methylase (i.e., rmtA, rmtB, rmtC, rmtD, and armA genes), aminoglycoside-modifying enzymes (i.e., aac(6’)-Ib, aac(6’)-IIa, aph(3’)-VI, and ant(2’’)-Ia genes), efflux pump (mexY gene) and TCS (phoP gene) were detected using the polymerase chain reaction (PCR) method. Deoxyribonucleic acid (DNA) extraction from aminoglycoside-resistance P. aeruginosa strains was carried out using the boiling method. Each PCR reaction had a final volume of 25 µL and followed a standardized thermal cycling protocol: 1 cycle of initial denaturation at 95 °C for 5 min, followed by 30 to 34 cycles consisting of denaturation at 94 °C for 1 min, annealing at various temperatures (as specified in Table 1) for 1 min, and extension at 72 °C for 1 min. The oligonucleotide sequences of the primers used and the corresponding annealing temperatures for each gene are provided in Table 1. Sequencing of the PCR product of each amplified gene was conducted, and the sequences were utilized as positive control.
The accession numbers for these sequences are available in the NCBI GenBank repository (OR855380, OR855383 to OR855385, PP468580 to PP468582, ON920997, and OR855381).
Expression of the MexXY-OprM efflux pump and PhoPQ TCS
The quantitative reverse transcription PCR (qRT-PCR) technique was employed to quantify the expression levels of the genes encoding MexXY-OprM efflux pump (mexY gene) and PhoPQ TCS (phoP gene). Sixteen multidrug-resistant (MDR) P. aeruginosa clinical isolates were selected for this analysis. These MDR isolates also were extensively drug-resistant (XDR) or pandrug-resistant (PDR) or colistin-resistant bacteria. Total RNA extraction and cDNA synthesis were conducted using TRIzol™ reagent (Bio Basic, Canada) and cDNA synthesis kit (Yekta Tajhiz Azma, Iran), respectively [13]. Each qRT-PCR reaction had a final volume of 15 µL and all reactions followed a standardized thermal cycling protocol: pre-incubation at 95 °C for 600 s, followed by 40 cycles of three amplification steps at 95 °C for 20 s, 64 °C, 62 °C, and 59 °C (for rpsL, mexY, and phoP genes, respectively) for 20 s, and 72 °C for 30 s. The rpsL housekeeping gene along with P. aeruginosa ATCC 27,853 were served as the reference gene and the reference strain, respectively, and changes in gene expression levels of the target genes were calculated using the comparative Ct (2-ΔΔCt) method. Each experiment was replicated twice, and the interpretation of the results was conducted based on previous studies [13, 14].
Result
In this study, a comprehensive examination of clinical strains of 200 P. aeruginosa revealed a prevalence rate of 48% for aminoglycoside-resistant isolates, as determined by the disk diffusion method (n = 96). The breakdown of resistance rates for individual aminoglycoside antibiotics was as follows: tobramycin 45.5% (91/200), amikacin 43% (86/200), and netilmicin 39.2% (33/84). Table 2 provides detailed demographic information for the 96 aminoglycoside-resistant P. aeruginosa isolates, including data on the hospital, specimen type, patient sex, and age.
Furthermore, the antibiotic resistance profiles of these aminoglycoside-resistant P. aeruginosa isolates to a range of antibiotics were investigated. The results indicated high resistance rates, including: piperacillin 88.5% (85/96), piperacillin-tazobactam 62.5% (60/96), ceftazidime 86.4% (83/96), cefepime 91.6% (88/96), aztreonam 19.7% (19/96), imipenem 95.8% (92/96), meropenem 90.6% (87/96), ciprofloxacin 93.7% (90/96), levofloxacin 93.7% (90/96), norfloxacin 91.6% (88/96), ofloxacin 60.4% (58/96), and colistin 3.1% (3/96). Remarkably, 91 of the 96 (94.7%) aminoglycoside-resistant P. aeruginosa isolates exhibited multi-drug resistance (MDR).
Phenotypic and genotypic analyses demonstrated that all aminoglycoside-resistant P. aeruginosa isolates possessed the ability to produce biofilms. Among these isolates, 70.8% (68/96) were categorized as weak biofilm producers, 17.7% (17/96) as moderate biofilm producers, and 6.2% (6/96) as strong biofilm producers. These isolates carried all genes associated with biofilm production, including algD, pslD, pelF, Ppgl, and PAPI-1 genes.
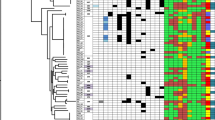
Based on PCR analysis, 86.4% (83/96) of the tested isolates were positive for genes encoding aminoglycoside-modifying enzymes. The most prevalent gene detected was aac(6’)-Ib (82/96, 85.4%), followed by ant(2’’)-Ia (18/96, 18.7%), and aph(3’)-VI (3/96, 3.1%). Notably, the aac(6’)-IIa gene was not identified in our study. Interestingly, among the 96 aminoglycoside-resistant P. aeruginosa isolates, 14 (14.5%) did not harbor any known aminoglycoside resistance genes. The genotypic and phenotypic profiles of aminoglycoside resistance among the 96 P. aeruginosa isolates are presented in Tables 3 and 4. Additionally, the genes responsible for methylation of the 16S rRNA in the 30 S ribosomal subunit were not found to be associated with the emergence of aminoglycoside resistance in P. aeruginosa isolates. Details of genotypic detection of aminoglycoside resistance-associated genes in P. aeruginosa clinical strains are depicted in Supplementary Figure S1.
The MexXY-OprM efflux pump and PhoPQ TCS genes were identified in all aminoglycoside-resistant P. aeruginosa isolates using the PCR analysis. However, as outlined in Table 5, the expression levels of the mexY and phoP genes, assessed via qRT-PCR, among 16 selected aminoglycoside-resistant P. aeruginosa isolates were 75% (12/16) and 87.5% (14/16), respectively. Details of amplification curves and melting peaks for each target in qRT-PCR are presented in Supplementary Figure S2.
Discussion
Since the introduction of broad-spectrum aminoglycosides in the 1940s, alongside beta-lactams and fluoroquinolones, they have remained crucial as antipseudomonal agents [13, 23]. However, a comprehensive analysis of our current and previous studies reveals a significant prevalence of aminoglycoside resistance among P. aeruginosa strains isolated from hospitals in Ardabil (48%), which is comparable to the resistance rates observed for beta-lactams such as penicillins (46.4% to 94%), carbapenems (33.3% to 66.7%), monobactams (42.9%), cephems (46.5% to 50%), and fluoroquinolones (52.4% to 76.2%) [4]. This is a matter of concern and underscores the potential threat posed to the health of individuals receiving medical care at Ardabil hospitals. An important contributing factor to the high levels of aminoglycoside resistance in Ardabil hospitals is their extensive use in the treatment of life-threatening infections caused by various organisms, including urinary tract infections, sepsis, and pneumonia [23]. Therefore, it is imperative to develop strategies to optimize the use of antimicrobial agents, such as combination therapy, rather than relying solely on individual agents. Among the aminoglycosides, amikacin, gentamicin, and tobramycin are the most commonly used in clinical practice [23]. Amikacin serves as an indicator antibiotic for the treatment of P. aeruginosa infections, and amikacin-resistant strains often display cross-resistance to other aminoglycosides [17]. In this study, among the 86 tested amikacin-resistant P. aeruginosa strains, resistance rates to tobramycin and netilmicin were 95.3%, and 100%, respectively. It is worth noting that the average prevalence of amikacin resistance among P. aeruginosa strains in Iran is 50.6% [24]. The resistance to amikacin observed in this study was higher than the rates reported in Urmia (30.7%), Hamadan (30.2%), and Zanjan (21.7%), while being lower than the rates reported in Isfahan (95.5%), Tehran (80%), Ahvaz (55.2%), and Guilan (48.8%) [24]. The prevalence of amikacin-resistant P. aeruginosa strains in other countries was as follows: China (85%), the USA (6%), and 11 European countries (12.9%) [25,26,27]. Differences in these results may be attributed to geographic variations, overuse of aminoglycosides in hospitals, self-medication practices without prescription, and differences in the overall health status of the populations studied. Amikacin and gentamicin, when combined with other antibiotic classes, are recommended for the treatment of infections caused by MDR Gram-negative organisms [23]. Gentamicin disk diffusion and MIC breakpoints for P. aeruginosa was deleted in new CLSI breakpoint revision [11]. Based on the previous version, the resistance rate to gentamicin observed in this study was similar to the national average (46% vs. 46.9%) [24]. Interestingly, the prevalence of MDR strains among the 96 aminoglycoside-resistant P. aeruginosa strains was higher than the national average (94.7%) [24]. In addition, it was higher than Ahvaz (91.9%), Hamadan (88.7%), Tehran (81.3%), Tabriz (68%), Zanjan (65%), Isfahan (63.1%), Urmia (56.9%), Guilan (45.5%), and Zahedan (16.4%) [24]. The identification of genes encoding resistance to aminoglycosides is crucial for managing drug-resistant infections and preventing treatment failure [17]. The predominant mechanism of aminoglycoside resistance involves aminoglycoside modifying enzymes [28], which was corroborated in our study, with 86.4% of the strains exhibiting this type of resistance. Among these enzymes, the aac(6')-Ib gene emerged as the most prevalent among aminoglycoside-resistant P. aeruginosa strains [28]. Our findings revealed the presence of the aac(6')-Ib gene in 85.4% of the strains, consistent with studies by El-Far et al. (94.4%) [15], Dubois et al. (36.5%) [16], Ahmadian et al. (60.4%) [29], and Jafari et al. (74%) [30]. The aac(6')-Ib gene known to confer resistance to tobramycin and amikacin [31]. In our study, 86.8% (79/91) of isolates harboring the aac(6')-Ib gene were resistant to tobramycin, and 88.3% (76/86) were resistant to amikacin. This underscores the significance of the aac(6')-Ib gene as a key determinant of tobramycin and amikacin resistance in clinical isolates of P. aeruginosa in Ardabil hospitals. Furthermore, other prevalent modifying enzymes identified in P. aeruginosa include ant(2'')-I, aac(6')-II, and aph(3')-VI genes [31]. In our investigation, we observed the presence of ant(2'')-Ia (18.5%) and aph(3')-VI (3.1%) genes among aminoglycoside-resistant P. aeruginosa strains, while the aac(6')-IIa gene was not detected. In a study conducted by Kim et al., the aph(3')-VI gene was reported as the most commonly encountered (77%) [17]. The ant(2'')-I and aac(6')-II genes are associated with resistance to gentamicin and tobramycin [31]. Based on the previous CLSI version, 94.4% of isolates harboring the ant(2'')-Ia gene were resistant to both gentamicin and tobramycin. The aph(3')-VI gene mediates resistance to amikacin [31]. However, in our analysis, no significant correlation was observed between tobramycin resistance and the presence of aph(3')-VI (Table 4). Considering the high prevalence of aminoglycoside resistance among P. aeruginosa strains in Ardabil hospitals, largely attributed to aminoglycoside-modifying enzymes, the utilization of semisynthetic aminoglycosides to overcome common resistance mechanisms is recommended [23]. One notable example of a semisynthetic aminoglycoside is plazomicin, which was approved by the FDA (Food and Drug Administration) in June 2018 for the treatment of urinary tract infections caused by certain susceptible bacteria [23]. Plazomicin was specifically engineered to circumvent aminoglycoside-modifying enzymes [23]. Fourteen aminoglycoside-resistant P. aeruginosa strains exhibited positive phenotypic tests but did not show the presence of genes encoding aminoglycoside-modifying enzymes. There are two possible explanations for this observation: 1) the genes encoding aminoglycoside-modifying enzymes are typically carried on plasmids [28], and 2) other resistance mechanisms, such as 16S rRNA methylases, biofilm formation, MexXY-OprM efflux pump, and TCS, may be involved. In line with a report by Kim et al., none of the P. aeruginosa strains in our study harbored the 16S rRNA methylases gene. We speculate that this is because the 16S rRNA methylases gene is encoded on the same plasmid as the aminoglycoside-modifying enzymes [28]. Decreased drug accumulation via overexpression of the MexXY-OprM efflux pump gene confers low-level intrinsic resistance to aminoglycosides in P. aeruginosa [28]. As shown in Table 3, aminoglycoside-resistant P. aeruginosa strains with resistance mechanisms independent of aminoglycoside-modifying enzymes and 16S rRNA methylase exhibited biofilm production as well as overproduction of TCS and efflux pump. Studies have indicated that extracellular DNA (eDNA), a component of the biofilm matrix, is involved in aminoglycoside resistance by inducing the expression of genes regulated by the PhoPQ TCS [9]. In the current study, 87.5% of aminoglycoside-resistant P. aeruginosa strains exhibited expression of the PhoPQ TCS. Some additional experiments were beyond the scope of this study and could be acknowledged as limitations but also provide opportunities for future research. These include evaluation of: 1) gene expression levels of the MexXY-OprM efflux pump and PhoPQ TCS in all aminoglycoside-resistant P. aeruginosa strains; 2) mutations in the PhoPQ TCS genes; and 3) the genetic relationship between bacterial strains isolated from different hospitals using a molecular typing method.
Conclusion
Considering the high prevalence of aminoglycoside-resistant P. aeruginosa strains with diverse resistance mechanisms in Ardabil hospitals, the following strategies are suggested to combat bacterial resistance to aminoglycosides: 1) enhancing public awareness regarding antibiotic resistance and advocating for judicious antibiotic use, 2) tailoring antibiotic prescriptions based on local antimicrobial resistance patterns and considering combination therapy when appropriate, 3) mitigating the occurrence of hospital-acquired infections through stringent adherence to infection control protocols, 4) implementing ongoing surveillance and research initiatives to monitor the prevalence and mechanisms of aminoglycoside resistance, given its plasmid-mediated nature, and 5) transitioning towards the utilization of novel antipseudomonal antibiotics, including emerging aminoglycosides, within clinical settings.
Availability of data and materials
The datasets generated and analyzed during the current study are available in the NCBI GenBank repository, under the accession numbers: OR855380, OR855383 to OR855385, PP468580 to PP468582, ON920997, and OR855381.
Abbreviations
- P. aeruginosa :
-
Pseudomonas aeruginosa
- PCR:
-
Polymerase chain reaction
- qRT-PCR:
-
Quantitative reverse transcription PCR
- TCSs:
-
Two-component systems
- MDR:
-
Multidrug-resistant
- ICUs:
-
Intensive care units
- CF:
-
Cystic fibrosis
- LPS:
-
Lipopolysaccharide
- APH:
-
Aminoglycoside phosphotransferase
- AAC:
-
Aminoglycoside acetyltransferase
- ANT:
-
Aminoglycoside nucleotidyltransferase
- CLSI:
-
Clinical and laboratory standards institute
- FDA:
-
Food and drug administration
- ATCC:
-
American type culture collection
- OD:
-
Optical density
- DNA:
-
Deoxyribonucleic acid
- RNA:
-
Ribonucleic acid
- TOB:
-
Tobramycin
- AMK:
-
Amikacin
- NET:
-
Netilmicin
- PIP:
-
Piperacillin
- TZP:
-
Piperacillin-tazobactam
- CAZ:
-
Ceftazidime
- FEP:
-
Cefepime
- ATM::
-
Aztreonam
- IMP:
-
Imipenem
- MEM:
-
Meropenem
- CIP:
-
Ciprofloxacin
- LVX:
-
Levofloxacin
- NOR:
-
Norfloxacin
- OFX:
-
Ofloxacin
- CST:
-
Colistin
- R:
-
Resistant
- S:
-
Susceptible
- I:
-
Intermediate
References
Nazari M, Ahmadi H, Hosseinzadeh S, Sahebkar A, Khademi F. Imipenem resistance associated with amino acid alterations of the OprD porin in Pseudomonas aeruginosa clinical isolates. Acta Microbiol Immunol Hung. 2023;70(3):206–12.
Hasanpour F, Ataei N, Sahebkar A, Khademi F. Distribution of class A extended-spectrum β-Lactamases among Pseudomonas aeruginosa clinical strains isolated from Ardabil hospitals. Jundishapur J Microbiol. 2023;16(4):e135726.
Namaki M, Habibzadeh S, Vaez H, Arzanlou M, Safarirad S, Bazghandi SA, Sahebkar A, Khademi F. Prevalence of resistance genes to biocides in antibiotic-resistant Pseudomonas aeruginosa clinical isolates. Mol Biol Rep. 2022;49(3):2149–55.
Bazghandi SA, Arzanlou M, Peeridogaheh H, Vaez H, Sahebkar A, Khademi F. Prevalence of virulence genes and drug resistance profiles of Pseudomonas aeruginosa isolated from clinical specimens. Jundishapur J Microbiol. 2021;14(8):e118452.
Khademi F, Maarofi K, Arzanlou M, Peeri-Dogaheh H, Sahebkar A. Which missense mutations associated with DNA gyrase and topoisomerase IV are involved in Pseudomonas aeruginosa clinical isolates resistance to ciprofloxacin in Ardabil? Gene Rep. 2021;24:101211.
Safarirad S, Arzanlou M, Mohammadshahi J, Vaez H, Sahebkar A, Khademi F. Prevalence and characteristics of metallo-beta-lactamase-positive and high-risk clone ST235 Pseudomonas aeruginosa at Ardabil hospitals. Jundishapur J Microbiol. 2021;14(3):e115819.
Morehead MS, Scarbrough C. Emergence of global antibiotic resistance. Prim care. 2018;45(3):467–84.
Poole K. Aminoglycoside resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother. 2005;49(2):479–87.
Pang Z, Raudonis R, Glick BR, Lin TJ, Cheng Z. Antibiotic resistance in Pseudomonas aeruginosa: mechanisms and alternative therapeutic strategies. Biotechnol Adv. 2019;37(1):177–92.
Jafari-Ramedani S, Nazari M, Arzanlou M, Peeri-Dogaheh H, Sahebkar A, Khademi F. Prevalence and molecular characterization of colistin resistance in Pseudomonas aeruginosa isolates: insights from a study in Ardabil hospitals. BMC Microbiol. 2024;24(1):152.
CLSI. Performance standards for antimicrobial susceptibility testing. 31st ed. Wayne, USA: Clinical and Laboratory Standards Institute; 2024.
Jabalameli F, Mirsalehian A, Khoramian B, Aligholi M, Khoramrooz SS, Asadollahi P, Taherikalani M, Emaneini M. Evaluation of biofilm production and characterization of genes encoding type III secretion system among Pseudomonas aeruginosa isolated from burn patients. Burns. 2012;38(8):1192–7.
Yousefi S, Nazari M, Ramazanzadeh R, Sahebkar A, Safarzadeh E, Khademi F. Association of carbapenem and multidrug resistance with the expression of efflux pump-encoding genes in Pseudomonas aeruginosa clinical isolates. Acta Microbiol Immunol Hung. 2023;70(2):161–6.
Lin J, Xu C, Fang R, Cao J, Zhang X, Zhao Y, Dong G, Sun Y, Zhou T. Resistance and heteroresistance to colistin in Pseudomonas aeruginosa isolates from Wenzhou, China. Antimicrob Agents Chemother. 2019;63(10):10–128.
El-Far SW, Abukhatwah MW. Prevalence of aminoglycoside resistance genes in clinical isolates of Pseudomonas aeruginosa from Taif, Saudi Arabia—An emergence indicative study. Microorganisms. 2023;11(9): 2293.
Dubois V, Arpin C, Dupart V, Scavelli A, Coulange L, André C, Fischer I, Grobost F, Brochet JP, Lagrange I, Dutilh B. β-Lactam and aminoglycoside resistance rates and mechanisms among Pseudomonas aeruginosa in French general practice (community and private healthcare centres). J Antimicrob Chemother. 2008;62(2):316–23.
Kim JY, Park YJ, Kwon HJ, Han K, Kang MW, Woo GJ. Occurrence and mechanisms of amikacin resistance and its association with β-lactamases in Pseudomonas aeruginosa: a Korean nationwide study. J Antimicrob Chemother. 2008;62(3):479–83.
Yamane K, Wachino JI, Doi Y, Kurokawa H, Arakawa Y. Global spread of multiple aminoglycoside resistance genes. Emerg Infect Dis. 2005;11(6):951–3.
Fritsche TR, Castanheira M, Miller GH, Jones RN, Armstrong ES. Detection of methyltransferases conferring high-level resistance to aminoglycosides in Enterobacteriaceae from Europe, North America, and Latin America. Antimicrob Agents Chemother. 2008;52(5):1843–5.
Lee H, Yong D, Yum JH, Roh KH, Lee K, Yamane K, Arakawa Y, Chong Y. Dissemination of 16S rRNA methylase-mediated highly amikacin-resistant isolates of Klebsiella pneumoniae and Acinetobacter baumannii in Korea. Diagn Microbiol Infect Dis. 2006;56(3):305–12.
Castanheira M, Fritsche TR, Sader HS, Jones RN. RmtD 16S RNA methylase in epidemiologically unrelated spm-1-producing Pseudomonas aeruginosa isolates from Brazil. Antimicrob Agents Chemother. 2008;52(4):1587–8.
Rajabi H, Salimizand H, Khodabandehloo M, Fayyazi A, Ramazanzadeh R. Prevalence of algD, pslD, pelF, Ppgl, and PAPI-1 genes involved in biofilm formation in clinical Pseudomonas aeruginosa strains. BioMed Res Int. 2022;2022:1–7.
Serio AW, Keepers T, Andrews L, Krause KM. Aminoglycoside revival: review of a historically important class of antimicrobials undergoing rejuvenation. EcoSal Plus. 2018;8(1):10–128.
Vaez H, Salehi-Abargouei A, Ghalehnoo ZR, Khademi F. Multidrug resistant Pseudomonas aeruginosa in Iran: a systematic review and metaanalysis. J Glob Infect Dis. 2018;10(4):212–7.
Dou Y, Huan J, Guo F, Zhou Z, Shi Y. Pseudomonas aeruginosa prevalence, antibiotic resistance and antimicrobial use in Chinese burn wards from 2007 to 2014. J Int Med Res. 2017;45(3):1124–37.
Atassi G, Medernach R, Scheetz M, Nozick S, Rhodes NJ, Murphy-Belcaster M, Murphy KR, Alisoltani A, Ozer EA, Hauser AR. Genomics of aminoglycoside resistance in Pseudomonas aeruginosa bloodstream infections at a United States Academic Hospital. Microbiol Spectr. 2023;11(3):e05087-05022.
Torrens G, van der Schalk TE, Cortes-Lara S, Timbermont L, del Barrio-Tofiño E, Xavier BB, Zamorano L, Lammens C, Ali O, Ruzin A, Goossens H. Susceptibility profiles and resistance genomics of Pseudomonas aeruginosa isolates from European ICUs participating in the ASPIRE-ICU trial. J Antimicrob Chemother. 2022;77(7):1862–72.
Wang N, Luo J, Deng F, Huang Y, Zhou H. Antibiotic combination therapy: a strategy to overcome bacterial resistance to aminoglycoside antibiotics. Front Pharmacol. 2022;13: 839808.
Ahmadian L, Norouzi Bazgir Z, Ahanjan M, Valadan R, Goli HR. Role of Aminoglycoside-Modifying Enzymes (AMEs) in resistance to aminoglycosides among clinical isolates of pseudomonas aeruginosa in the North of Iran. Biomed Res Int. 2021;2021:7077344.
Jafari M, Fallah F, Borhan RS, Navidinia M, Rafiei Tabatabaei S, Karimi A, et al. The first report of CMY, Aac (6′)-Ib and 16S rRNA methylase genes among Pseudomonas aeruginosa isolates from Iran. Arch Pediatr Infect Dis. 2013;1(3):109–12.
Vaziri F, Peerayeh SN, Nejad QB, Farhadian A. The prevalence of aminoglycoside-modifying enzyme genes (aac (6′)-I, aac (6′)-II, ant (2 ″)-I, aph (3′)-VI) in Pseudomonas aeruginosa. Clinics. 2011;66(9):1519–22.
Acknowledgements
Not applicable.
Funding
This research has been financially supported by the Vice Chancellor for Research (grant number:402000378), Ardabil University of Medical Sciences, Iran.
Author information
Authors and Affiliations
Contributions
NS: Methodology, Investigation, and Formal analysis. MN: Methodology, and Investigation. RR: Conceptualization, Review, and Editing. SJR: Methodology, and Investigation. AS: Review, and Editing. FK: Conceptualization, Supervision, Project administration, and Original draft preparation.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This research has been approved by the Regional Research Ethics Committee (approval ID: IR.ARUMS.MEDICINE.REC.1402.157). All methods were carried out according to relevant guidelines and regulations. Clinical isolates were collected from the hospital’s bacterial repository solely for research purposes, and neither patient samples nor patient data were utilized in this study. Therefore, the requirement for informed consent from participants was waived by the Regional Research Ethics Committee of Ardabil University of Medical Sciences.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Saeli, N., Jafari-Ramedani, S., Ramazanzadeh, R. et al. Prevalence and mechanisms of aminoglycoside resistance among drug-resistant Pseudomonas aeruginosa clinical isolates in Iran. BMC Infect Dis 24, 680 (2024). https://doi.org/10.1186/s12879-024-09585-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12879-024-09585-6