Abstract
Background
Glioma is a deadly and immunosuppressive brain tumour. Complement C1r subcomponent like (C1RL), a prognostic biomarker in several kinds of tumours, has attracted increasing attention from oncologists. However, the role of C1RL in glioma remains unclear.
Methods
Through analysis of 2120 glioma patients from 5 public datasets, the relationships between C1RL expression and clinicopathological characteristics were evaluated. Furthermore, the C1RL-associated genes were screened, and Gene Ontology (GO) analysis was conducted to investigate biological process enrichment. In addition, tumour purity, leukocyte infiltration and overall survival were evaluated based on C1RL expression.
Results
We found that C1RL expression was upregulated in glioblastoma (GBM), especially mesenchymal GBM and primary GBM. Increased C1RL expression accompanied the IDH1-wt phenotype in both lower grade glioma (LGG) and GBM. C1RL- associated genes were mainly enriched in biological processes related to the immune response. C1RL expression was also correlated with reduced tumour purity and increased M2 macrophage infiltration. Higher C1RL expression predicted unfavourable survival in patients with glioma and therapeutic resistance in GBM.
Conclusions
Our results imply that C1RL is involved in immunological activities and is an independent unfavourable prognostic biomarker in patients with glioma. C1RL is a potential clinical immunotherapeutic target for glioma treatment in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Glioblastoma (GBM; WHO grade IV) and lower grade glioma (LGG; WHO grade II and III) are incurable brain tumour. Existing therapeutic strategies only prolong the survival of glioma patients to a limited extent. Patients with glioma eventually die from tumour recurrence, even with aggressive treatment. Novel therapies that have been successful in other tumours, such as PD-1 inhibition [1] and bevacizumab administration [2, 3], have failed to extend the overall survival time of patients with glioma. Tumour treating fields (TTF), a novel therapy that was recently approved for GBM treatment by the Food and Drug Administration (FDA), is not widely used in clinical practice because of its high price and difficult process [4, 5]. The current poor situation pushes us to explore the mechanism of glioma development and identify novel therapies.
The immunosuppressive microenvironment significantly contributes to the progression and therapeutic resistance of glioma. On the one hand, glioma cells induce a relatively weak immune response and enhance immunosuppression. Compared to other malignancies, glioma exhibits a lower mutational burden and fewer infiltrating T cells [6]. GBM cells block T cell activation and proliferation in response to T cell receptor stimulation by generating extracellular vesicles carrying PD-L1 [7]. Glioma cells promote the expression of PD-L1 on macrophages derived from healthy donors [8, 9]. Intratumoural immunosuppressive education by glioma also contributes to the rise of systemic immunosuppressive myeloid-derived suppressor cells (MDSCs) [10]. On the other hand, the brain provides an immunosuppressive environment for glioma. Compared tomelanoma in the flank, melanoma in the brain contains fewer CD8 T cells [11]. Moreover, antigen-specific cytotoxicity is systemically impaired in mice with brain melanoma [11]. Naïve T cells are sequestered in large numbers in the bone marrow in cancer patients. This phenomenon characterizes a variety of tumours only when the tumours are located in the intracranial compartment [12].
Complement C1r subcomponent like (C1RL) was found to be a prognostic marker in hepatocellular carcinoma [13] and renal cell cancer [14]. A gene-based analysis showed significant associations between non-Hodgkin lymphoma or diffuse large B-cell lymphoma and the C1RL gene [15]. C1RL also mediates the progression of Burkitt’s lymphoma [16]. C1RL is a protein-coding gene associated with ovarian adenocarcinoma and leucorrhea. In terms of molecular function, the C1RL protein, which is homologous to C1r, is identified as the active form of serine hydrolase [17]. The C1RL protein cleaves prohaptoglobin in the endoplasmic reticulum [18]. In addition, pro-C1s is proteolytically cleaved into two fragments with sizes identical to those of the two chains of active C1s by the C1RL protein [19]. However, the immunological and clinicopathological characteristics of C1RL in glioma remain unclear.
In the present study, we employed 2120 glioma specimens and 23 non-tumour brain tissues from 5 datasets to explore the clinicopathological and biological characteristics of C1RL in glioma. The clinicopathological features evaluated included WHO grade, histology, GBM status, IDH mutation status, GBM subtype, overall survival and therapeutic resistance. The biological process enrichment of C1RL-associated genes was analysed to explore the biological characteristics of C1RL. Moreover, the relationships between C1RL expression and tumour purity or leukocyte infiltration were analysed.
Methods data collection
Five datasets including transcriptomic files and corresponding clinicopathological information for patients who were diagnosed with glioma (WHO II-IV) were downloaded. A microarray dataset containing 539 samples (TCGAmic) and an RNA sequencing dataset containing 702 samples (TCGAseq) were downloaded from The Cancer Genome Atlas (TCGA; https://xenabrowser.net). A microarray dataset containing 301 samples (CGGAmic) and an RNA-sequencing dataset containing 325 samples (CGGAseq) were downloaded from The Chinese Glioma Genome Atlas (CGGA; http://www.cgga.org.cn/). A microarray dataset containing 276 samples (GSE16011mic) was downloaded from Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/).
Statistics of C1RL expression patterns
An unpaired t test was used in comparisons of C1RL expression between two groups. Ordinary one-way ANOVA (multiple comparisons) was applied to compare C1RL expression among three or more groups. P < 0.05 was considered significant.
C1RL-associated gene siftings and gene ontology (GO) analyses
Pearson correlation coefficients between C1RL and all other genes were calculated in RStudio 1.1.453 with the cor.test algorithm. C1RL-associated genes were defined as genes with an r value > 0.4 in the GBM dataset (TCGAmic) and r > 0.5 in the glioma datasets (TCGAseq, CGGAmic, and CGGAseq). All the C1RL-associated genes were introduced into DAVID (https://david.ncifcrf.gov/) for further GO analyses. The top 10 biological process terms of the GO analysis results are listed in Fig. 2.
Moreover, the detailed correlations between C1RL and immunosuppressive genes (CD86, LGALS9, and TGFB1) are shown in Fig. 3.
Tumour purity and leukocyte infiltration
The ESTIMATE algorithm package was used to analyse tumour purity. The CIBERSORT tool (https://cibersort.stanford.edu/) was used to evaluate leukocyte infiltration. Heatmaps were produced in MORPHEUS (https://software.broadinstitute.org/morpheus/) online. The colour shows the Z score (subtract mean, divided by standard deviation) of all the expression data. The samples were ordered according to the expression of C1RL.
Survival analyses
The log-rank test and Kaplan-Meier survival curves were used to describe survival differences between two groups. The survival analysis of the GSE16011 dataset was conducted in R2 (https://hgserver1.amc.nl/cgi-bin/r2/main.cgi).
Results
C1RL expression was upregulated in GBM, especially mesenchymal GBM, primary GBM and IDH1-wt GBM
In this study, we employed 2120 glioma specimens and 23 non-tumour brain tissues from 5 datasets. The characteristics and clinical information of the 5 datasets were summarized in Table S1.
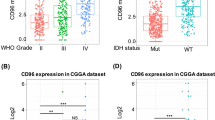
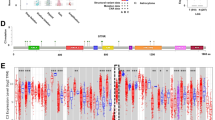
C1RL expression was analysed according to the WHO classification, GBM subtype, GBM status and IDH1 mutation status. First, the expression of C1RL was always highest in GBM in the 5 datasets according to both the grading system and the histology system (Fig. 1.A-H). However, the expression levels of C1RL in GBM samples varied greatly. Furthermore, C1RL expression among different subgroups of GBM was analysed. Among the four transcriptomic subgroups of GBM, C1RL expression was always highest in mesenchymal GBM (Fig. 1.I-L). Secondary GBM was developed from lower grade glioma and exhibited lower C1RL expression than primary GBM (Fig. 1.M-N). The IDH mutation status is a well-accepted marker for glioma classification. C1RL expression was higher in IDH1-wt GBM than in IDH1-mt GBM (Fig. 1.O-Q). In addition, C1RL expression was higher in IDH1-wt LGG than in IDH1-mt LGG (Fig. 1 P and Q). These results suggested that higher C1RL expression accompanies more advaced malignancy in glioma, especially in GBM.
Histopathological characteristics of C1RL in glioma. a. C1RL expression in primary GBM (p-GBM) and nontumour brain tissues in TCGAmic. C1RL expression in distinct WHO grades of glioma in CGGAseq (b), CGGAmic (c), and GSE16011mic (d). C1RL expression in distinct histological types of glioma in TCGAseq (e), CGGAseq (f), CGGAmic (g), and GSE16011mic (h). C1RL expression in four GBM subtypes in TCGAmic (i), TCGAseq (j), CGGAseq (k), and CGGAmic (l). C1RL expression in samples with different GBM statuses in CGGAseq (m) and CGGAmic (n). C1RL expression in IDH1-mutant or IDH1-wild-type LGG or GBM in TCGAmic (o), CGGAseq (p), and CGGAmic (q). O, oligodendroglioma; AO, anaplastic oligodendroglioma; OA, oligoastrocytoma; AOA, anaplastic oligoastrocytoma; A, astrocytoma; AA, anaplastic astrocytoma. Mut, mutant; wt, wild-type. *, p < 0.05; **, p < 0.01; ***, p < 0.001; ****, p < 0.0001
C1RL-associated genes were enriched in the biological processes of the immune response
The biological function of C1RL, especially in tumours, has not been clarified thoroughly. Therefore, we aimed to identify the possible biological function of C1RL through analysis of the biological functions of C1RL-associated genes. C1RL-associated genes were defined as genes with expression trends similar to those of C1RL in glioma samples. All the C1RL genes from each dataset were listed in Table S2 and were evaluated by GO analysis. The biological processes are listed in reverse order of their p values. The GO analyses showed that C1RL-associated genes were mainly enriched in the biological processes of the immune response, inflammatory response, IFN-γ mediated signalling pathway, and innate immune response (Fig. 2.A-D).
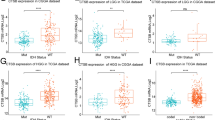
To determine whether C1RL plays a positive role in the anti-glioma immune response, the expression relationships between C1RL and existing biomarkers were analysed. The CD86 protein is the receptor of CTLA4 and is mainly expressed on dendritic cells and monocytes. Galectin-9, which is encoded by LGALS9, was identified as the ligand of Tim-3 and plays a key role in T cell apoptosis. TGFB1 encodes a secreted ligand in the transforming growth factor-beta (TGF-beta) superfamily of proteins. CD86 [20], LGALS9 [21], and TGFB1 [22] play immunosuppressive roles in glioma. Our results showed that C1RL expression exhibited positive relations with CD86, LGALS9, and TGFB1 (Fig. 3.A-L).
C1RL expression was correlated with reduced tumour purity and increased M2 macrophage infiltration
The immune response is based on the migration of immune cells. Both tumour purity and the infiltration of 22 types of leukocytes were assessed for each sample in the TCGA datasets. The samples are displayed in order of their C1RL expression level. Both the immune score and the stromal score exhibited a positive correlation with C1RL expression trends (Fig. 4.A and B, top panels). In addition, tumour purity showed an inverse correlation with C1RL expression trends (Fig. 4.A and B, middle panels). Moreover, C1RL expression was mostly related to the infiltration of M2 macrophages among the 22 types of leukocytes (Fig. 4.A and B, bottom panels).
The correlations of C1RL with tumour purity and leukocyte infiltration. The correlation of C1RL with nontumour cell infiltration in TCGAmic (a, GBM) and TCGAseq (b, glioma). Samples are listed from left to right according to their C1RL expression level from low to high. Both the immune score and the stromal score are shown in the top panels. Tumour purity is shown in the middle panels. Leukocyte infiltration levels are listed in the bottom panels
High expression of C1RL predicted unfavourable survival and therapeutic resistance in glioma
The median expression value of C1RL was used to separate samples into two subgroups. We evaluated the prognostic value of C1RL in the four glioma datasets. The patients with glioma exhibiting higher C1RL expression had significantly shorter survival times than their counterparts in the GSE16011mic, TCGAseq, CGGAmic, and CGGAseq datasets (Fig. 5.A-D). However, the histopathology characteristics in the two subgroups are significantly different (Table S3). The histopathology characteristics may contribute to the survival differences. Next, we compared survival times among GBM patients with different C1RL expression levels in the five datasets. All the survival curves exhibited significant differences (Fig. 5.E-I). Moreover, the effectiveness of well-accepted treatment was evaluated in different C1RL expression groups. The primary GBM patients with lower C1RL expression showed better responses to resection, radiochemotherapy (temozolomide), and standard therapy (Fig. 5.J-L). These results indicate that C1RL may contribute to therapeutic resistance.
Survival differences according to C1RL expression in glioma. High C1RL expression predicted shortened overall survival for glioma patients (a) and primary glioma patients (b-d). High C1RL expression predicted shortened overall survival for GBM patients (e) and primary GBM patients (f-i). Primary GBM with high C1RL expression showed increased resistance to radiochemotherapy (j), resection (k), and standard therapy (l)
Discussion
We analysed the characteristics of C1RL in gliomas from various angles. According to the 2016 WHO classification of glioma, the current diagnosis of glioma is mainly based on the WHO grades and pathology and IDH mutation [23]. Besides, the GBM can be classified into Proneural, Neural, Classical, and Mesenchymal subtypes [24], or primary and secondary GBM [25]. The results showed that the subgroups with worse prognosis always have higher levels of C1RL (Fig. 1). Based on the perspectives that C1RL should have similar biological functions with C1RL-associated genes, we tried to explore the biological function of C1RL with C1RL-associated genes. The GO analyses showed that C1RL-associated genes enriched in immune related biological functions. But we still unsure whether C1RL promotes anti-tumour immune response or suppress it. Given that CD86 [20], LGALS9 [21], and TGFB1 [22] play immunosuppressive roles in glioma, we further investigated the expressing relationship between C1RL and these immunosuppressive genes. Besides, low tumour purity [26] and high M2 macrophages infiltration [27] were reported to promotes glioma progression. So, we also analysed the relationships between C1RL and glioma purity and leukocyte infiltration.
C1RL is a negative biomarker for glioma prognosis. C1RL had not been mentioned in cancer until a report indicating significant associations of non-Hodgkin lymphoma and diffuse large B-cell lymphoma with the C1RL gene in 2012 [15]. In recent years, C1RL has been reported to be a prognostic biomarker in hepatocellular carcinoma [13] and renal cell cancer [14]. Our results showed evidence that C1RL is highly expressed in glioma samples and predicts a poor prognosis. GBM is WHO grade IV glioma and has the worst prognosis of glioma types, with a median overall survival time of 14.6 months [28]. C1RL expression was always higher in GBM than in LGG (Fig. 1.A-H). Unsupervised transcriptomic analysis revealed that of the GBM subtypes, mesenchymal GBM has the worst survival [29]. C1RL expression was higher in mesenchymal GBM than in other GBM subgroups (Fig. 1.I-L). Secondary GBM progresses from LGG within 5–10 years of diagnosis and is accompanied by a better prognosis than primary GBM [25]. Secondary GBM exhibited less C1RL expression than primary GBM (Fig. 1.M and N). Patients with IDH1-mut glioma have a better outcome than those with IDH1-wt glioma [30]. Relatively low C1RL expression was found in both IDH1-mut LGG and IDH1-mut GBM (Fig. 1.O-Q). C1RL not only predicts more advanced malignancy but also worse overall survival in glioma. Due to the distinct outcomes of the glioma subgroups, the differences in C1RL expression in different histopathological subgroups (Fig. 1.A-H) may contribute to the observed differences in survival. We further investigated survival differences between the high C1RL expression group and the low C1RL expression group in GBM and even primary GBM. The results confirmed that the expression level of C1RL was a survival indicator in primary GBM (Fig. 5.F-I). Resection following chemoradiation is a well-accepted strategy for primary GBM patients. Considering the effects of variant therapies, C1RL may play a role in therapeutic resistance (Fig. 5.J-L). Overall, our results indicate that C1RL is a biomarker of poor outcomes in glioma patients.
C1RL probably plays an important role in glioma immunosuppression. The C1RL protein is confirmed to be an active form of serine hydrolase [17] and cleaves prohaptoglobin and pro-C1s into their active forms [18, 19]. On the one hand, due to the suppression of lymphocyte function by haptoglobin [31], C1RL may modulate immunosuppression in glioma by releasing active haptoglobin. On the other hand, the association of C1s with C1r and C1q, following ligand recognition, triggers the activation of the classical complement pathway [32]. C1q plays a fundamental role in the pathogenesis of glioma [33]. C1RL may trigger the classical complement pathway by activating C1s and thus contribute to the pathogenesis of glioma. In addition, accumulated evidence shows that C1RL expression is upregulated during inflammation [34, 35]. GO analyses of C1RL-associated genes revealed that they were mainly enriched in the biological processes of the immune response, inflammatory response, IFN-γ mediated signalling pathway, and innate immune response (Fig. 2.A-D). Furthermore, C1RL exhibited positive correlations with immunosuppressive markers (Fig. 3.A-L).
C1RL expression was correlated with leukocyte infiltration, especially M2 macrophage infiltration. Tumour purity was proposed as an important factor in glioma. Low purity cases were independently associated with poor prognosis [26]. Glioma evolution is associated with immunological changes in the microenvironment [29]. M2 macrophages promotes glioma growth [27]. The ESTIMATE algorithm is a well-accepted method to predict the tumour purity in genomic and transcriptomic studies [36, 37]. Besides, CIBERSORT algorithm, also known as in silico flow cytometry, was developed to accurately assess the infiltration of many leukocyte subsets in bulk tumour samples, along with a signature genes file that enumerates the genes that define the signature expression profile for each immune cell [38]. The CIBERSORT algorithm can be access online (https://cibersort.stanford.edu/) to characterize cell composition of complex tissues from their gene expression profiles. In this study, both ESTIMATE algorithm and CIBERSORT algorithm were used to further assess the relationships between C1RL mRNA expression and 22 different immune cell populations. Increased amounts of immune cells, especially M2 macrophages, migrated into glioma tumours with relatively high C1RL expression (Fig. 4.A and B). All these results are consistent with the hypothesis that C1RL palys an immunosuppressive role in glioma.
Conclusions
In conclusion, we analysed the immunological and clinicopathological characteristics of C1RL in 2120 glioma patients from five datasets. The results indicate that C1RL is a negative biomarker for the patients with glioma. Furthermore, C1RL probably plays an immunosuppressive role in the pathogenesis of glioma by triggering the activation of haptoglobin and C1s.
Availability of data and materials
The microarray dataset of 539 samples (TCGAmic) and the RNA sequencing dataset of 702 samples (TCGAseq) were downloaded from The Cancer Genome Atlas (TCGA, https://xenabrowser.net). The microarray dataset of 301 samples (CGGAmic) and the RNA sequencing dataset of 325 samples (CGGAseq) were downloaded from The Chinese Glioma Genome Atlas (CGGA, http://www.cgga.org.cn/). The microarray dataset of 276 samples (GSE16011mic) was downloaded from Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/). The survival analysis of GSE16011 dataset was conducted in R2 (https://hgserver1.amc.nl/cgi-bin/r2/main.cgi).
Abbreviations
- C1RL:
-
Complement C1r subcomponent like
- CGGA:
-
The Chinese Glioma Genome Atlas
- FDA:
-
The Food and Drug Administration
- GBM:
-
Glioblastoma
- GO:
-
Gene Ontology
- LGG:
-
Lower grade glioma
- MDSCs:
-
Myeloid-derived suppressor cells
- TCGA:
-
The Cancer Genome Atlas
- TGF-beta:
-
The transforming growth factor-beta
- TTF:
-
Tumour treating fields
References
Filley AC, Henriquez M, Dey M. Recurrent glioma clinical trial, CheckMate-143: the game is not over yet. Oncotarget. 2017;8:91779–94.
Wick W, Gorlia T, Bendszus M, Taphoorn M, Sahm F, Harting I, Brandes AA, Taal W, Domont J, Idbaih A, Campone M, Clement PM, Stupp R, Fabbro M, Le RE, Dubois F, Weller M. von DA, Golfinopoulos V, Bromberg JC, Platten M, Klein M, van den Bent MJ. Lomustine and Bevacizumab in Progressive Glioblastoma. N Engl J Med. 2017;377:1954–63.
Gilbert MR, Dignam JJ, Armstrong TS, Wefel JS, Blumenthal DT, Vogelbaum MA, Colman H, Chakravarti A, Pugh S, Won M, Jeraj R, Brown PD, Jaeckle KA, Schiff D, Stieber VW, Brachman DG, Werner-Wasik M, Tremont-Lukats IW, Sulman EP, Aldape KD, Curran WJ, Mehta MP. A randomized trial of bevacizumab for newly diagnosed glioblastoma. N Engl J Med. 2014;370:699–708.
Stupp R, Taillibert S, Kanner A, Read W, Steinberg D, Lhermitte B, Toms S, Idbaih A, Ahluwalia MS, Fink K, Di MF, Lieberman F, Zhu JJ, Stragliotto G, Tran D, Brem S, Hottinger A, Kirson ED, Lavy-Shahaf G, Weinberg U, Kim CY, Paek SH, Nicholas G, Bruna J, Hirte H, Weller M, Palti Y, Hegi ME, Ram Z. Effect of tumor-treating fields plus maintenance Temozolomide vs maintenance Temozolomide alone on survival in patients with Glioblastoma: a randomized clinical trial. JAMA. 2017;318:2306–16.
Palmer JD, Bhamidipati D, Mehta M, Williams NL, Dicker AP, Werner-Wasik M, Shi W. Treatment recommendations for elderly patients with newly diagnosed glioblastoma lack worldwide consensus. J Neuro-Oncol. 2018;140:421–6.
Li B, Severson E, Pignon JC, Zhao H, Li T, Novak J, Jiang P, Shen H, Aster JC, Rodig S, Signoretti S, Liu JS, Liu XS. Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome Biol. 2016;17:174.
Ricklefs FL, Alayo Q, Krenzlin H, Mahmoud AB, Speranza MC, Nakashima H, Hayes JL, Lee K, Balaj L, Passaro C, Rooj AK, Krasemann S, Carter BS, Chen CC, Steed T, Treiber J, Rodig S, Yang K, Nakano I, Lee H, Weissleder R, Breakefield XO, Godlewski J, Westphal M, Lamszus K, Freeman GJ, Bronisz A, Lawler SE, Chiocca EA. Immune evasion mediated by PD-L1 on glioblastoma-derived extracellular vesicles. Sci Adv. 2018;4:eaar2766.
Tong L, Li J, Choi J, Pant A, Xia Y, Jackson C, Liu P, Yi L, Boussouf E, Lim M, Yang X. CLEC5A expressed on myeloid cells as a M2 biomarker relates to immunosuppression and decreased survival in patients with glioma. Cancer Gene Ther. 2019.
Bloch O, Crane CA, Kaur R, Safaee M, Rutkowski MJ, Parsa AT. Gliomas promote immunosuppression through induction of B7-H1 expression in tumor-associated macrophages. Clin Cancer Res. 2013;19:3165–75.
Chae M, Peterson TE, Balgeman A, Chen S, Zhang L, Renner DN, Johnson AJ, Parney IF. Increasing glioma-associated monocytes leads to increased intratumoral and systemic myeloid-derived suppressor cells in a murine model. Neuro-Oncology. 2015;17:978–91.
Jackson CM, Kochel CM, Nirschl CJ, Durham NM, Ruzevick J, Alme A, Francica BJ, Elias J, Daniels A, Dubensky TW, Lauer P, Brockstedt DG, Baxi EG, Calabresi PA, Taube JM, Pardo CA, Brem H, Pardoll DM, Lim M, Drake CG. Systemic tolerance mediated by melanoma brain tumors is reversible by radiotherapy and vaccination. Clin Cancer Res. 2016;22:1161–72.
Chongsathidkiet P, Jackson C, Koyama S, Loebel F, Cui X, Farber SH, Woroniecka K, Elsamadicy AA, Dechant CA, Kemeny HR, Sanchez-Perez L, Cheema TA, Souders NC, Herndon JE, Coumans JV, Everitt JI, Nahed BV, Sampson JH, Gunn MD, Martuza RL, Dranoff G, Curry WT, Fecci PE. Sequestration of T cells in bone marrow in the setting of glioblastoma and other intracranial tumors. Nat Med. 2018;24:1459–68.
Xu B, Lv W, Li X, Zhang L, Lin J. Prognostic genes of hepatocellular carcinoma based on gene coexpression network analysis. J Cell Biochem. 2019.
Chinello C, Cazzaniga M, De Sio G, Smith AJ, Grasso A, Rocco B, Signorini S, Grasso M, Bosari S, Zoppis I, Mauri G, Magni F. Tumor size, stage and grade alterations of urinary peptidome in RCC. J Transl Med. 2015;13:332.
Bassig BA, Zheng T, Zhang Y, Berndt SI, Holford TR, Hosgood HD, Hu W, Leaderer B, Yeager M, Menashe I, Boyle P, Xu J, Zou K, Zhu Y, Chanock S, Rothman N, Lan Q. Polymorphisms in complement system genes and risk of non-Hodgkin lymphoma. Environ Mol Mutagen. 2012;53:145–51.
Han B, Wang S, Zhao H. MicroRNA-21 and microRNA-155 promote the progression of Burkitt's lymphoma by the PI3K/AKT signaling pathway. Int J Clin Exp Pathol. 2020;13:89–98.
Navarrete M, Ho J, Krokhin O, Ezzati P, Rigatto C, Reslerova M, Rush DN, Nickerson P, Wilkins JA. Proteomic characterization of serine hydrolase activity and composition in normal urine. Clin Proteomics. 2013;10:17.
Wicher KB, Fries E. Prohaptoglobin is proteolytically cleaved in the endoplasmic reticulum by the complement C1r-like protein. Proc Natl Acad Sci U S A. 2004;101:14390–5.
Ligoudistianou C, Xu Y, Garnier G, Circolo A, Volanakis JE. A novel human complement-related protein, C1r-like protease (C1r-LP), specifically cleaves pro-C1s. Biochem J. 2005;387:165–73.
Tumangelova-Yuzeir K, Naydenov E, Ivanova-Todorova E, Krasimirova E, Vasilev G, Nachev S, Kyurkchiev D. Mesenchymal stem cells derived and cultured from Glioblastoma Multiforme increase Tregs, Downregulate Th17, and induce the Tolerogenic phenotype of monocyte-derived cells. Stem Cells Int. 2019;2019:6904638.
Yuan F, Ming H, Wang Y, Yang Y, Yi L, Li T, Ma H, Tong L, Zhang L, Liu P, Li J, Lin Y, Yu S, Ren B, Yang X. Molecular and clinical characterization of Galectin-9 in glioma through 1,027 samples. J Cell Physiol. 2020;235:4326–34.
Han J, Chen X, Chu J, Xu B, Meisen WH, Chen L, Zhang L, Zhang J, He X, Wang QE, Chiocca EA, Kaur B, Caligiuri MA, Yu J. TGFβ treatment enhances Glioblastoma Virotherapy by inhibiting the innate immune response. Cancer Res. 2015;75:5273–82.
Louis DN, Perry A, Reifenberger G, von DA F-BD, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW. The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 2016;131:803–20.
Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP, Alexe G, Lawrence M, O'Kelly M, Tamayo P, Weir BA, Gabriel S, Winckler W, Gupta S, Jakkula L, Feiler HS, Hodgson JG, James CD, Sarkaria JN, Brennan C, Kahn A, Spellman PT, Wilson RK, Speed TP, Gray JW, Meyerson M, Getz G, Perou CM, Hayes DN. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell. 2010;17:98–110.
Hu H, Mu Q, Bao Z, Chen Y, Liu Y, Chen J, Wang K, Wang Z, Nam Y, Jiang B, Sa JK, Cho HJ, Her NG, Zhang C, Zhao Z, Zhang Y, Zeng F, Wu F, Kang X, Liu Y, Qian Z, Wang Z, Huang R, Wang Q, Zhang W, Qiu X, Li W, Nam DH, Fan X, Wang J, Jiang T. Mutational Landscape of Secondary Glioblastoma Guides MET-Targeted Trial in Brain Tumor. Cell. 2018;175:1665–78 e18.
Zhang C, Cheng W, Ren X, Wang Z, Liu X, Li G, Han S, Jiang T, Wu A. Tumor purity as an underlying key factor in Glioma. Clin Cancer Res. 2017;23:6279–91.
Zhou W, Ke SQ, Huang Z, Flavahan W, Fang X, Paul J, Wu L, Sloan AE, McLendon RE, Li X, Rich JN, Bao S. Periostin secreted by glioblastoma stem cells recruits M2 tumour-associated macrophages and promotes malignant growth. Nat Cell Biol. 2015;17:170–82.
Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, Taphoorn MJ, Belanger K, Brandes AA, Marosi C, Bogdahn U, Curschmann J, Janzer RC, Ludwin SK, Gorlia T, Allgeier A, Lacombe D, Cairncross JG, Eisenhauer E, Mirimanoff RO. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med. 2005;352:987–96.
Wang Q, Hu B, Hu X, Kim H, Squatrito M, Scarpace L, de Carvalho AC, Lyu S, Li P, Li Y, Barthel F, Cho HJ, Lin YH, Satani N, Martinez-Ledesma E, Zheng S, Chang E, SCE G, Olar A, Lan ZD, Finocchiaro G, Phillips JJ, Berger MS, Gabrusiewicz KR, Wang G, Eskilsson E, Hu J, Mikkelsen T, RA DP, Muller F, Heimberger AB, Sulman EP, Nam DH, RGW V. Tumor Evolution of Glioma-Intrinsic Gene Expression Subtypes Associates with Immunological Changes in the Microenvironment. Cancer Cell. 2018;33:152.
Yan H, Parsons DW, Jin G, McLendon R, Rasheed BA, Yuan W, Kos I, Batinic-Haberle I, Jones S, Riggins GJ, Friedman H, Friedman A, Reardon D, Herndon J, Kinzler KW, Velculescu VE, Vogelstein B, Bigner DD. IDH1 and IDH2 mutations in gliomas. N Engl J Med. 2009;360:765–73.
Sadrzadeh SM, Bozorgmehr J. Haptoglobin phenotypes in health and disorders. Am J Clin Pathol. 2004;121(Suppl):S97–104.
Lu J, Kishore U. C1 complex: an adaptable Proteolytic module for complement and non-complement functions. Front Immunol. 2017;8:592.
Mangogna A, Belmonte B, Agostinis C, Zacchi P, Iacopino DG, Martorana A, Rodolico V, Bonazza D, Zanconati F, Kishore U, Bulla R. Prognostic implications of the complement protein C1q in Gliomas. Front Immunol. 2019;10:2366.
Severino P, Ariga SK, Barbeiro HV, de Lima TM, de Paula SE, Barbeiro DF, MCC M, Nizet V, dSF P, et al. J Mol Med (Berl). 2017;95:995–1003.
Shi L, Zhu B, Xu M, Wang X. Selection of AECOPD-specific immunomodulatory biomarkers by integrating genomics and proteomics with clinical informatics. Cell Biol Toxicol. 2018;34:109–23.
Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW, Levine DA, Carter SL, Getz G, Stemke-Hale K, Mills GB, Verhaak RG. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612.
Cao JY, Guo Q, Guan GF, Zhu C, Zou CY, Zhang LY, Cheng W, Wang GL, Cheng P, Wu AH, Li GY. Elevated lymphocyte specific protein 1 expression is involved in the regulation of leukocyte migration and immunosuppressive microenvironment in glioblastoma. Aging (Albany NY). 2020;12:1656–84.
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, Hoang CD, Diehn M, Alizadeh AA. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12:453–7.
Acknowledgements
Not Applicable.
Funding
This work was supported by grants from the Beijing-Tianjin-Hebei Basic Research Cooperation Project (No. 19JCZDJC64200) and the State Scholarship Fund from China Scholarship Council (No. 201806940031).
Author information
Authors and Affiliations
Contributions
XY and JW participated the design of this study. JW and LT performed the statistical analysis and drafted the manuscript. GL and HW carried out data acquisition. LZ edited the tables, the figures, and the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was approved by the Ethics Committee of Tianjin Medical University General Hospital.
Consent for publication
Informed consent was obtained from all participants for publication.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Additional file 1 Table S1
. Clinical information of 2143 patients from the different datasets.
Additional file 2 Table S2
. C1RL associated genes.
Additional file 3 Table S3
. C1RL and WHO grade and glioma histopathology.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Wang, J., Tong, L., Lin, G. et al. Immunological and clinicopathological characteristics of C1RL in 2120 glioma patients. BMC Cancer 20, 931 (2020). https://doi.org/10.1186/s12885-020-07436-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12885-020-07436-6