Abstract
Background
Lung adenocarcinoma (LUAD) patients have a dismal survival rate because of cancer metastasis and drug resistance. The study aims to identify the genes that concurrently modulate EMT, metastasis and EGFR-TKI resistance, and to investigate the underlying regulatory mechanisms.
Methods
Cox regression and Kaplan–Meier analyses were applied to identify prognostic oncogenes in LUAD. Gene set enrichment analysis (GSEA) was used to indicate the biological functions of the gene. Wound-healing and Transwell assays were used to detect migratory and invasive ability. EGFR-TKI sensitivity was evaluated by assessing the proliferation, clonogenic survival and metastatic capability of cancer cells with treatment with gefitinib. Methylated RNA immunoprecipitation (MeRIP) and RNA immunoprecipitation (RIP) analyses established the level of m6A modification present on the target gene and the protein’s capability to interact with RNA, respectively. Single-sample gene set enrichment (ssGSEA) algorithm used to investigate levels of immune cell infiltration.
Results
Our study identified dual-specificity phosphatase 5 (DUSP5) as a novel and powerful predictor of adverse outcomes for LUAD by using public datasets. Functional enrichment analysis found that DUSP5 was positively enriched in EMT and transforming growth factor-beta (TGF-β) signaling pathway, a prevailing pathway involved in the induction of EMT. As expected, DUSP5 knockdown suppressed EMT via inhibiting the canonical TGF-β/Smad signaling pathway in in vitro experiments. Consistently, knockdown of DUSP5 was first found to inhibit migratory ability and invasiveness of LUAD cells in in vitro and prevent lung metastasis in in vivo. DUSP5 knockdown re-sensitized gefitinib-resistant LUAD cells to gefitinib, accompanying reversion of EMT progress. In LUAD tissue samples, we found 14 cytosine-phosphate-guanine (CpG) sites of DUSP5 that were negatively associated with DUSP5 gene expression. Importantly, 5′Azacytidine (AZA), an FDA-approved DNA methyltransferase inhibitor, restored DUSP5 expression. Moreover, RIP experiments confirmed that YTH N6-methyladenosine RNA binding protein 1 (YTHDF1), a m6A reader protein, could bind DUSP5 mRNA. YTHDF1 promoted DUSP5 expression and the malignant phenotype of LUAD cells. In addition, the DUSP5-derived genomic model revealed the two clusters with distinguishable immune features and tumor mutational burden (TMB).
Conclusions
Briefly, our study discovered DUSP5 which was regulated by epigenetic modification, might be a potential therapeutic target, especially in LUAD patients with acquired EGFR-TKI resistance.
Graphical Abstract

Similar content being viewed by others
Introduction
Lung cancer is the deadliest type of tumor globally, remaining first in terms of mortality [1], and non-small cell lung cancer (NSCLC) is a primary type of lung cancer, constituting approximately 85% of all cases. As a predominant type of NSCLC, the Lung adenocarcinoma (LUAD) has a high propensity to metastasize to the distant organs and develop resistance to treatment, resulting in a grim five-year overall survival rate [2, 3]. Epidermal growth factor receptor tyrosine kinase inhibitors (EGFR-TKIs), such as gefitinib and erlotinib, are the conventional first-line therapy for advanced LUAD patients with sensitive epidermal growth factor receptor (EGFR) mutations and have demonstrated significant efficacy for these patients [4]. Unfortunately, the majority of LUAD patients experience incomplete responses, and eventually develop resistance to EGFR-TKIs within 9 to 13 months, which leads to the progression of the fatal disease [5]. The discovered acquired mechanisms of EGFR-TKI resistance have been largely classified as EGFR secondary mutations (e.g. EGFR-T790M mutation), bypass signaling activations, and phenotypic changes such as EMT [6]. This diversity and uncertainty in resistance mechanisms pose challenges to developing new therapeutic targets and predicting the effectiveness of EGFR-TKI treatment [7]. In conclusion, it is essential and attractive to discover and validate precise biomarkers for coupling modules of metastasis and EGFR-TKI resistance, offering a strategy to therapeutically combat the malignant progression of LUAD.
Epithelial-mesenchymal transition (EMT), a hallmark of tumorigenic transformation, is a cellular program where tumor cells lose their well-differentiated epithelial phenotypes and acquire invasive mesenchymal, fibroblast-like ones [8, 9]. Increasing evidence suggest that EMT is a critical contributor to cancer metastasis and drug resistance, including that to EGFR-TKIs [10]. During the sophisticated metastatic cascade of cancer, EMT progress is a prerequisite [11]. The drastically altered cancer cell surface during EMT prompts the re-localization or degradation of intercellular attachments, leading to the “leaky” tight junctions (TJs) and the destabilized adherence junctions (AJs) [12]. EMT lends metastatic cancer stem cells to survive, escaping from the primary site, and to invade and migrate to secondary areas [13]. Additionally, a growing number of studies disclosed that EMT or mesenchymal phenotype is correlated with both intrinsic and acquired resistance to EGFR-TKIs [14,15,16]. The molecular mechanisms by which the mesenchymal phenotype-mediated the EGFR-TKI resistance are still unknown [16]. Recent research suggested that mesenchymal cells might inherently exhibit reduced sensitivity to the inhibited intracellular signaling pathway, indicating a potential vulnerability to EGFR-TKIs [16,17,18]. For this dynamic and reversible process, EMT is strictly controlled by internal and external cues, including various regulators, effectors, and signaling pathways [19]. The transforming growth factor-beta (TGF-β) signaling pathway, in particular, has been presented to promote EMT, EMT-mediated metastasis and EGFR-TKI resistance [20]. Nevertheless, the mechanisms and factors that concurrently govern EMT-mediated metastasis and EGFR-TKI resistance remain largely unknown.
The rapid advancement of next-generation high throughput RNA sequencing technologies has resulted in the production of tremendous amounts of high-dimensional omics data [21]. Due to vast and reliable data sharing efforts by the research communities, such as the cancer genome atlas (TCGA), extensive and various omics data with unprecedented details are available [22, 23]. TCGA database and many datasets from the gene expression omnibus (GEO) provide detailed clinical information [24, 25]. In LUAD, according to the presence or absence of lymph node metastasis in the early pathologic T stage, Dong et al. identified macrophage-related secreted phosphoprotein 1 (SPP1) as a risk indicator for early lymph node metastasis across multiple cohorts, suggesting that SPP1 might function as a putative biomarker for the early detection of lymph node metastasis [26]. Gu et al. established a risk model comprising three differentially methylated genes, which possesses the ability to accurately predict prognosis in LUAD, based on the pathologic tumor node metastasis (TNM) stage [27]. However, the considerable volume of valuable omics data for LUAD metastasis and EMT had not been fully utilized and analyzed.
Herein, we first applied a novel combination of two public datasets (TCGA-LUAD and GEO: GSE11117) to identify distant metastasis-associated genes, and the seven differentially up-regulated genes were obtained. Among them, dual-specificity phosphatase 5 (DUSP5) exhibited an excellent ability to predict prognosis in multiple independent data sets. As indicated by functional enrichment analysis performed with GSEA, these experimental results validated that DUSP5 knockdown reversed EMT and inhibited the canonical TGF-β/Smad signaling pathway. Suppression of DUSP5 impeded tumor metastasis and EGFR-TKI resistance in LUAD cells. Meanwhile, YTHDF1 could bind DUSP5 mRNA to regulate its expression. Additionally, our clusters based on the DUSP5-originated genomic model exhibited distinct immune microenvironments, tumor mutational burden (TMB), and prognostic characteristics in LUAD. In short, the results revealed that DUSP5 acted as an influential mediator of EMT and could become a potential biomarker of prognosis and antitumor therapy of LUAD patients. Our results aimed to deliver a novel insight into the biological complexity related to metastasis and the resistance to targeted therapy, contributing to the tailored precision therapy for LUAD.
Methods
Data downloading and preprocessing
Transcriptome RNA-sequencing and clinical follow-up data were obtained from patients with TCGA-LUAD (https://portal.gdc.cancer.gov/projects). These following steps screened 482 TCGA-LUAD patients: (1) Retaining patients with clinical characteristics and OS time; (2) Keeping genes with Fragments Per Kilobase of exon model per Million mapped fragments (FPKM) greater than 1 in over 50% of the individuals. Furthermore, the data of GSE11117, GSE30219, GSE3141, GSE31210, GSE41271, GSE50081 and GSE72094 datasets were gained from GEO (https://www.ncbi.nlm.nih.gov/geo/).
The analysis of differentially expressed genes (DEGs)
The “edgeR” R package was used to obtain DEGs from TCGA, with the threshold criteria consisting of a |Log2 fold change (FC)| greater than 0.585, false-discovery rate (FDR), and P < 0.05. The DEGs from GSE11117 were filtered using GEO2R (http://www.ncbi.nlm.nih.gov/geo/geo2r/) with a cutoff criteria of P < 0.05 and |Log2FC| greater than 0.585. DEGs were plotted as volcano plots based on bioinformatics (http://www.bioinformatics.com.cn/). PCA was used to visualize the discrimination between 2 groups (with distant metastasis and without distant metastasis) in terms of the R package “edgeR”, “ggplot2”, “ggrepel” and “FactoMineR”. A Venn diagram (https://hiplot-academic.com/basic/venn) was carried out to create interesting overlaps between two sets for identifying key targets.
Analysis of cox regression and nomogram model
Exploring independent prognostic genes based on Cox regression analyses followed by using R, and these results showed as forest plots with the “survminer” R package. A nomogram prediction model was designed from these significant characteristics and genes obtained from multivariate Cox regression analysis. Then, we depicted the calibration plots to access the nomogram’s prognostic value.
Survival analysis
Patients with TCGA-LUAD were categorized based on various clinical characteristics, which encompassed TNM stage (I/II and III/IV), T stage (T1/T2 and T3/T4), N stage (N0 and N1/N2/N3), M stage (M0 and M1), age (< 55 and ≥ 55 years old), and gender (male and female). In terms of optimal cutoff expression values, R tool was used to investigate OS in different cohorts by the Kaplan–Meier analysis. GEPIA analysis (http://gepia.cancer-pku.cn/) was used to assess the presence of DUSP5 in invasive breast carcinoma, cholangiocarcinoma, thyroid carcinoma, and esophageal carcinoma. The DUSP5 expression in various clinical features of TCGA-LUAD patients was calculated by Prism 8.0 (GraphPad, USA).
LUAD cell culture, siRNA transfection and lentivirus infection
A549, H1299, HCC827, PC9, HCC827GR, and PC9GR cells were grown in RPMI1640 or DMEM medium supplemented with fetal bovine serum (FBS). The manufacturer’s instructions were followed for transient transfection with siRNA (Ribobio, Guangzhou, China) using Lipofectamine 3000 (Invitrogen). The following sequence: si1: GAGACTTTCTACTCGGAAT; si2: GTGGTAAATGTCAGCTACA. Lentiviral YTHDF1 shRNA-expressing (shYT-1 and shYT-2) vectors and control shRNA-expressing (NC) vectors were infected and selected by adding 2 µg/ml puromycin into medium.
RNA immunoprecipitation (RIP) and methylated RNA immunoprecipitation (MeRIP)
A RIP Kit (Bes5101, BersinBio, China) was utilized to conduct the RIP assay. Input and coimmunoprecipitated RNAs were detected using qPCR.
Initially, the entire RNA was isolated using TRIzol reagent from TAKARA, a Japanese company. For the MeRIP assay, we utilized the riboMeRIPTM m6A Transcriptome Profiling Kit (C11051-1, Ribobio, China). Then the m6A-modified RNAs were analyzed for qPCR. SRAMP (http://www.cuilab.cn/sramp) was used to predict the MeRIP-qPCR primers for the target genes.
Biological enrichment analysis
Genes with a correlation coefficient ≥ 0.3 were chosen to annotate using gene ontology (GO) enrichment analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses in R software. Meanwhile, we visualized biological enrichment in bioinformatics (http://www.bioinformatics.com.cn/). The gene set named “h.all.v7.5.1.symbols.gmt” and gene set enrichment analysis (GSEA) (version 4.1.0) were utilized to identify hallmark pathways. Pathways with normalized P < 0.05 and FDR q value < 0.25 were considered significant. Additionally, the top 10 pathways were displayed based on the ranking of normalized enrichment scores (NESs).
Obtainment of EMT-related genes
The Molecular Signatures Database v7.5.1 (MSigDB) (http://www.gsea-msigdb.org/gsea/msigdb) and dbEMT2 (http://www.dbemt.bioinfo-minzhao.org/browser.cgi#tsgene) provided access to EMT-associated genes. MSigDB contained 200 genes associated with EMT in the module labeled “HALLMARK_EPITHELIAL_MESENCHYMAL_TRANSITION”, while dbEMT2 contained 191 EMT-related genes.
Single-cell analysis and Gene set cancer analysis (GSCALite)
In the CancerSEA (http://biocc.hrbmu.edu.cn/CancerSEA/), we also evaluated the functional conditions of DUSP5 in LUAD, including angiogenesis, apoptosis, cell cycle, differentiation, DNA damage, DNA repair, EMT, hypoxia, inflammation, invasion, metastasis, proliferation, quiescence, and stemness [28].
GSCALite is a foundational tool for cancer genomics analysis [29]. We examined the activation or inhibition of various pathways, including apoptosis, cell cycle, DNA damage response, EMT, hormone AR, hormone ER, PI3K/AKT, RAS/MAPK, RTK, and TSC/mTOR pathways, among the tools explored.
Assessment of DNA methylation
TCGA has identified links between DNA methylation and gene expression through the use of MEXPRESS (https://mexpress.be/) [30] and SMART (http://www.bioinfo-zs.com/smartapp/) [31]. The MethPrimer software [32] was used to predict cytosine-phosphate-guanine (CpG) islands (CGIs) in the sequences of gene promoters. Meanwhile, for analyses of tumor vs. normal DNA methylation, we used the TCGA database. 5′Azacytidine (AZA) as a DNA methyltransferase inhibitor was purchased from MedChemExpress (HY-10,586).
Western blotting
The proteins of cells were separated and transferred to polyvinylidene fluoride membranes using SDS-PAGE. After that, the primary antibodies DUSP5 (Abcam, ab200708), E-cadherin (Proteintech, 60335-1-lg), N-cadherin (Proteintech, 22018-1-AP), Vimentin (Abcam, ab92547), Snail (Cell Signaling Technology, #3879), Smad2 (Cell Signaling Technology, #5339), Smad3 (Cell Signaling Technology, #9523), P-smad2 (Cell Signaling Technology, #3108), P-smad3 (Cell Signaling Technology, #9520), YTHDF1 (Proteintech, 26787-1-AP) and GAPDH (Proteintech, 60004-1-lg) followed by, a secondary antibody was applied. The bands were imaged by a Tanon 5200 Imaging System (Tanon, China).
Wound-healing and transwell assays
The cells were grown in 6-well dishes containing 4 × 105 cells/ per well and then subjected to a scratch to create a wound resembling a crossroad. The serum-free 1640 was used to culture the cells after the scratch, and the migration of cells was imaged after 24, 36, or 48 h. ImageJ software was used to estimate wound closure rate.
Using Transwell assays to measure migratory and invasive capabilities via adding or not adding Matrigel coating. 2 × 104 cells/well were placed in each well of the upper chamber using RPMI-1640. Additionally, 700 µL of RPMI1640 with FBS was introduced into the lower chambers. The enclosure was placed in an environment for a period of 24, 36, or 48 h. Following that, the cells were immobilized and colored using paraformaldehyde and crystal violet consecutively. Inverted microscopy was used to capture images.
Cell counting kit (CCK-8), colony formation assays and EdU incorporation assay
In each well of 96-well plate, 3000 cells were cultured for 1–4 days. After incubating the CCK-8 reagent for 2 h, the absorption intensity at 450 nm was measured.
In the presence or absence of gefitinib therapy, a total of 1000 cells were grown in each well and then cultured in complete media for 12 days. Subsequently, the cells were fixed and stained sequentially with paraformaldehyde and crystal violet. Finally, they were adequately photographed and counted.
A total of 5000 cells were grown in a 96-well dish, followed by fixed and permeabilized using paraformaldehyde and Triton X-100. Subsequently, the cells were treated with EdU during incubation. Lastly, they were photographed by an inverted fluorescent microscope.
Animal models
All animal experiments were approved by the Ethics Committee of Harbin Medical University. To establish the metastasis model in nude mice, six nude mice (Beijing Vitalstar Biotechnology, China) were randomly divided into two groups: NC and shDUSP5. Six nude mice were injected via the tail vein with 5 × 106 A549-derived LUAD cells (NC and shDUSP5 cells). After 48 days, D-luciferin potassium salt was injected intraperitoneally at a dosage of 100 mg/kg to measure the bioluminescence signal from the tumors in the nude mice. The lungs of mice with metastatic tumors were promptly excised, treated with D-luciferin potassium salt, and the bioluminescence signal was measured again.
To establish EGFR-TKI sensitivity models in nude mice, ten mice were randomly divided into two groups: NC + Gefitinib and shDUSP5 + Gefitinib. We injected 1 × 107 tumor cells subcutaneously into 4-week-old BALB/c mice. Once detectable tumors formed, the mice were administered gefitinib daily at a dosage of 100 mg/kg. After 28 days, the mice were euthanized. Tumor dimensions (long and short diameters) were measured using vernier calipers, and tumor weights were also recorded.
Immune cell infiltration estimation
The ssGSEA analysis was conducted to quantify 29 different immune cell types in tissues based on normalized gene expression profiles by R. Wilcoxon test was used to examine the variations in infiltration levels. The TMB was assessed by the “maftools” package. We obtained 32 joint immune checkpoints in the literature [33], 20 (FPKM > 1 in more than 50% of the patients) of which were analyzed differentially.
Statistical analysis
We analyzed our data with R language (version 4.1.3) and Prism 8.0 (GraphPad, USA). Differences were tested using the Student’s t-test and one-way ANOVA. P < 0.05 indicated statistically significant. The correlation was identified by Pearson’s correlation analysis in the bioinformatics website (http://www.bioinformatics.com.cn/) and Hiplot (https://hiplot-academic.com/basic/).
Results
DUSP5 is a novel independent prognostic predictor for LUAD
To identify distant metastasis-associated genes, we employed a new combination of two public datasets (TCGA-LUAD and GEO: GSE11117). Initially, we stratified patients from the TCGA-LUAD and GSE11117 datasets into cohorts with (DM) and without (N-DM) distant metastasis, based on their M stage. Using principal components analysis (PCA), we found apparent differences between DM and N-DM (Fig. 1A). Compared with N-DM, 210 genes were up-regulated in DM compared to N-DM, whereas 231 genes were down-regulated, based on the TCGA-LUAD database. 672 up-regulated genes and 604 down-regulated genes were discovered based on GSE11117 (Fig. 1B). As shown in Fig. 1C, Venn diagrams indicated seven up-regulated and six downregulated genes as the intersected genes. Next, univariate and multivariate cox-regression analyses were utilized to investigate the index-pendent tumor-promoting factor in the TCGA-LUAD dataset. In LUAD, DUSP5 and T stage were found to be independent risk biomarkers for poor prognosis (Fig. 1D, E).
Identification of DUSP5 as an independent and poor prognostic indicator. (A) PCA for transcriptome profiles of 2 groups (DM: distant metastases and N-DM: no distant metastases), performing a difference on transcriptome between DM group and N-DM group. (B) Volcano plots exhibited these DEGs between the DM group and the N-DM group in TCGA and GSE11117. (C) Venn plots indicated 7 upregulated and 6 downregulated genes as intersected genes. (D, E) Univariate and Multivariate Cox regression analysis of genes and clinical features using the forest map. (F-H) Kaplan–Meier overall survival plot of DUSP5 based on TCGA, GSE30219 and GSE3141. FC: fold change; T stage: primary tumor stage; N stage: regional lymph node stage; M stage: distant metastasis stage
To reliability, with put-back 100 times in advance, the median was used to randomly split 482 patients from the LUAD-TCGA dataset into two appropriate validation sets, preventing the random allocation bias. The clinical characteristics of these two groups were homogenously distributed (Table S1). The high DUSP5 expression could strongly predict dismal outcomes of LUAD patients in the two random TCGA validation sets, GSE30219 and GSE3141 datasets according to Kaplan–Meier overall survival (OS) curves (Fig. 1F-H, Figure S1A, B). In addition, using a clinical parameters-stratified analysis, such as T, N, M, TNM stage, gender and age, high DUSP5 expression also predicted poor prognosis in LUAD patient cohorts (Figure S2, S3). It is the first time that DUSP5 may contribute to the progression of LUAD.
Expression levels of DUSP5 exhibit a positive correlation with T, M, and TNM stages
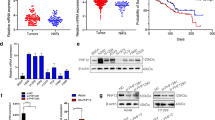
DUSP5 expression was significantly elevated in tumor tissue samples than in normal lung tissue samples based on the TCGA-LUAD dataset (Fig. 2A). Moreover, we unearthed that the DUSP5 was upregulated in various cancers, including invasive breast carcinoma, cholangiocarcinoma, thyroid carcinoma, and esophageal carcinoma (Figure S4A). High DUSP5 expression was relative to the advanced pathologic T stage, M stage, and TNM stage (Fig. 2B-D). There was no relevance between the expression of DUSP5 and N stage, age, and gender (Fig. S4B-D).
Increased DUSP5 expression is positively associated with clinicopathologic characteristics. (A) The levels of DUSP5 expression were examined between tumor and normal tissues according to data taken from the TCGA database. (B-D) Expression of DUSP5 in T stage, M stage and TNM stage based on TCGA database. (E) The nomogram predicted the probability of OS. (F) The calibration plot accessed the actual survival and predicted survival probability of OS. *P < 0.05, **P < 0.01, ***P < 0.001
For a good predictive model for clinical application, a nomogram was established according to total risk scores of DUSP5, T, N, M, and TNM stage (Fig. 2E). Through evaluation of the calibration curves which had the consistency of actual survival and predicted probability of 1-year, 2-year, and 3-year overall survival, we found that this model had excellent predictive accuracy (Fig. 2F).
DUSP5 exhibited a positive correlation with metastasis, EMT, and TGF-β signaling pathway
KEGG pathway analysis exhibited that focal adhesion, TNF, MAPK, and NF-kappa B signaling pathways that were reported to govern cancer metastasis were listed as top 10 (Figure S5A) [34,35,36,37]. GO enrichment showed metastasis-associated biological processes, such as wound healing, epithelial cell migration, focal adhesion, and cell-cell junction enriched (Figure S5B) [38,39,40,41,42].
Notably, GSEA was performed on Hallmark gene sets, EMT, and TGF-β signaling, the most prominent EMT inducer [43], as the top 10 Hallmark pathways were positively interrelated and remarkably enriched (Fig. 3A). In the two LUAD single-cell sequencing datasets, DUSP5 was also positively related to metastasis and EMT (Fig. 3B). Using GSCALite, DUSP5, CDH2, SNAIL, TWIST1, and VIM were associated with activated EMT progress, whereas CDH1 was involved in inhibited EMT progress (Fig. 3C, Figure S5C). As expected, 19 EMT-related genes from “MsigDB” and 8 oncogenic EMT-related genes from “dbEMT” correlated with DUSP5 expression with coefficients greater than 0.3 (Figure S6, S7A). In addition, EMT-related markers (VIM, FN1, SNAIL, SNAI2, TWIST1, and TWIST2) demonstrated elevated expression in the cohort characterized by high DUSP5 expression relative to the group with diminished DUSP5 expression levels (Figure S7B-C). Shortly, these data indicated that DUSP5 facilitated tumorigenesis and metastasis by inducing the EMT biological process in LUAD.
DUAP5 depletion inhibited EMT and TGF-β signaling pathways. (A) Hallmark analysis of DUSP5-related genes was performed by GSEA. (B) The correlation analysis between metastasis and DUSP5, and EMT and DUSP5 in the single-cell sequencing datasets (EXP0066 and EXP0067) of LUAD was based on the CancerSEA website. (C) The pie was used to represent the strength of genes (CDH1, CDH2, DUSP5, SNAIL, TWIST1 and VIM) of LUAD using GSCALite, red: activity; green: inhibition; grey: non-significant. (D) The RNA levels of DUSP5 expression in LUAD cells. (E)The protein levels of DUSP5 expression in LUAD cells were examined by Western blotting. (F) The levels of vimentin, N-cadherin and E-cadherin expression were assessed in UT, siNC, si1 and si2 cell lines. (G) The levels of P-smad2, Smad2, P-smad3, Smad3 and snail expression were assessed in UT, siNC, si1 and si2 cell lines. (H) Cell migratory capability was elucidated by a Wound-healing assay. (I) The invasive and migratory abilities were evaluated by Transwell assays. (J) The images display the bioluminescence signal intensities in each BALB/c nude mouse on day 48 after tail vein injection with NC or shDUSP5 cells (left panel). The bar charts present the results of quantitative and statistical analyses of the bioluminescence signal intensities for the NC and shDUSP5 groups (right panel). *P < 0.05, **P < 0.01, ***P < 0.001
DUAP5 depletion inhibited metastasis, EMT and TGF-β signaling pathway
To confirm the above bioinformatics results, we conducted loss-of-function experiments in which we exploited A549 and H1299 cell lines to silence DUSP5 expression using DUSP5-specific siRNAs (Fig. 3D-E). We found that the knockdown of DUSP5 enhanced E-cadherin expression and suppressed N-cadherin and vimentin expression in LUAD cells (Fig. 3F). In H1299 and A549 cells, the activation of the p-Smad2/3 and EMT-associated transcription factors snail were dramatically decreased by DUSP5 knockdown (Fig. 3G). According to Wound healing and Transwell assays, we verified that DUSP5 silence reduced the migratory and invasive ability of LUAD cells (Fig. 3H-I, Figure S8A-B). In the in vivo metastasis assays, knockdown of DUSP5 reduced the lung metastatic capability of LUAD cells in nude mice following tail vein injection, as indicated by lower luciferase signals. (Figure S8C, Fig. 3J). To verify whether DUSP5 regulated the EMT process and metastasis in LUAD through the TGF-β/Smad signaling pathway, we used SRI-011381, an administered agonist of the TGF-β signaling pathway. As shown in WB, SRI-011381 reversed the effects of the knockdown of DUSP5 resulting in the inhibition of P-smad2/3 and EMT (Fig. 4A, B). In addition, when treated with SRI-011381, DUSP5 knockdown failed to inhibit the migratory and invasive ability of LUAD cells (Fig. 4C, D). Together, our data indicated that DUSP5 is a driver for the EMT progression by activating the canonical TGF-β/Smad signaling pathway in LUAD.
TGF-β signaling pathway is essential for DUSP5-mediated EMT and metastasis. (A, B) Western blot analysis validated the effect of treatment with SRI-011381 (10 µM) or DMSO for 24 h on the TGF-β signaling pathway and EMT after DUSP5 knockdown in LUAD cells. (C, D) The migratory and invasive ability of H1299 and A549 cells, with or without SRI-011381 treatment, was detected by Wound-healing assay and Transwell assays
DUSP5 knockdown reversed EGFR-TKI resistance in gefitinib-resistant LUAD cells by inhibiting EMT
EMT is recognized to facilitate EGFR-TKI resistance [44, 45]. To investigate if DUSP5 could induce drug resistance to EGFR-TKI, we transfected DUSP5-specific siRNAs into gefitinib-resistant cell lines (HCC827GR and PC9GR) to achieve DUSP5 knockdown (Figure S8E-F). Knockdown of DUSP5 re-sensitized gefitinib-resistant cell lines at different concentrations and treatment times of gefitinib (Fig. 5A-B). Interestingly, EGFR-TKI-resistant cell lines showed resistance to gefitinib, however, DUSP5 knockdown caused a significant decrease in cellular activity and inhibited the colony-forming ability of resistant cell lines (Fig. 5B-C, Figure S8G). Furthermore, similar results were obtained using the EdU incorporation experiments (Fig. 5D). In the in vivo gefitinib resistance experiment, xenografts (NC and shDUSP5) derived from PC9GR cells were established in nude athymic mice (Figure S8D). DUSP5 knockdown attenuated the gefitinib resistance in these cells, as evidenced by reduced tumor volume (Fig. 5E-F) and tumor weight (Fig. 5G). When treated with gefitinib, DUSP5 silence attenuated the metastatic abilities of EGFR-TKI-resistant LUAD cells; these effects were further enhanced when the combination of DUSP5 knockdown (Fig. 6A-B).
EGFR-TKI resistant LUAD cells could be re-sensitized by knocking down DUSP5 in vitro and in vivo. (A, B) CCK-8 analysis results performed the proliferation and viability of DUSP5 knockdown cell lines and EGFR-TKI resistant cell lines. (C, D) The proliferation of gefitinib-treated or untreated cells was detected by colony formation assay and EdU incorporation assay. (E-G) Panel E shows images of xenograft tumors from the two designated groups. Panels F and G depict the results of t-tests used to compare the mean tumor volumes (F) and weights (G) between the groups. *P < 0.05, **P < 0.01, ***P < 0.001
DUSP5 knockdown reverses EGFR-TKI resistance in gefitinib-resistant LUAD cells by inhibiting EMT. (A) The effect of DUSP5 knockdown on the motility of EGFR-TKI resistant cells was shown by Wound-healing assay. (B) The invasive and migratory abilities of EGFR-TKI resistant cells were assessed by Transwell assays after DUSP5 knockdown. (C, D) The levels of vimentin, N-cadherin and E-cadherin expression after DUSP5 knockdown. *P < 0.05, **P < 0.01, ***P < 0.001
Of interest, examined by Western blotting, EGFR-TKI resistant cell lines displayed significantly lower abundance of E-cadherin, but higher expression of Vimentin and N-cadherin than parental cells (Fig. 6C-D). After DUSP5 knockdown, the mesenchymal phenotype in resistant cell lines was reversed, as indicated by increased E-cadherin and diminished N-cadherin and VIM expression (Fig. 6C-D). Collectively, our data indicated that DUSP5 decreased sensitivity to TKI treatment by inducing EMT.
DUSP5 was epigenetically regulated by DNA methylation and m6A modification
Methylation of DNA involves the dysregulation of gene expression [46]. Within the MEXPRESS database, fourteen CpG sites have been identified that exhibit an inverse correlation with the levels of DUSP5 expression (Fig. 7A). Using MethPrimer software, we obtained the CGI at the 1417–2446 sites in the promoter region of DUSP5, which meets the CGI criteria, including island size greater than 100 bp, GC percentage higher than 50.0, CpG dinucleotide Obs/Exp ratio greater than 0.6 [32] (Fig. 7B).
DUSP5 was epigenetically regulated by DNA methylation in LUAD. (A) The relationship between DNA methylation of the DUSP5 promoter region and DUSP5 expression with MEXPRESS. r: correlation coefficient. (B) Schematic representation of the CpG islands in the promoter region of DUSP5. (C) The beta value of CpG sites in tumor and normal tissue tissues. (D, E) The levels of DUSP5 expression were assessed with or without 1 µM AZA treatment. *P < 0.05, **P < 0.01, and ***P < 0.001
Next, we used the beta value to quantify methylation status. In comparison with normal tissue, the beta values in tumor tissue are high, indicating hypomethylation utilizing data from the TCGA-LUAD database (Fig. 7C). Further, the SMART website revealed that there were 11 significant correlations between the 22 CpG sites and the expression of DUSP5, including one positive correlation and ten negative correlations (|R| > 0.2, P < 0.05) (Figure S9). Consistently, AZA, a DNA methylation inhibitor, stimulated DUSP5 expression in LUAD cells, with or without depletion of DUSP5 (Fig. 7D, E). Our data suggest that DUSP5 was regulated by DNA methylation, and this regulation is negatively controlled by the degree of methylation of the DUSP5 promoter.
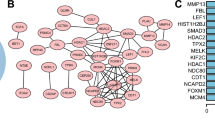
In addition to the exploration of DNA modification of DUSP5, we further investigated its mRNA modification. The most common modification of mRNA is m6A [47]. Using the online bioinformatic tools, SRAMP and MEME, the typical m6A motif RRACH (D = A, G or U; R = A or G; H = A, U or C) was identified in DUSP5 mRNA based on GSE136433 (Fig. 8A-C, Figure S10A-B). MeRIP-qPCR experiments revealed that DUSP5 underwent significant m6A methylation in LUAD cells (Fig. 8D). Using RM2Target [48] and RMVar [49] online databases, we predicted six m6A regulators that might modify DUSP5 mRNA m6A (Fig. 8E). Among them, YTHDF1 was most significantly overexpressed in LUAD tissues at the optimal P-value (Fig. 8F, Figure S10C).
YTHDF1 is involved in the regulation of DUSP5 through the recognition of m6A methylation. (A) SRAMP website showed the potential sites of DUSP5 mRNA with m6A modification. The horizontal axis represented the position of the m6A modification sites, the vertical axis represented the combined score with m6A. (B) A graphical representation of the secondary structure context around the m6A modification sites of DUSP5 mRNA in SRAMP. The red dots represented sites of m6A modification. (C) m6A motif RRACH (D = A, G or U; R = A or G; H = A, U or C) was predicted by MEME. (D) MeRIP-qPCR showed that m6A levels of DUSP5 mRNA in H299 and A549 cells. (E) Venn plots displayed 6 intersected genes associated with m6A based on RM2target and RMVar. (F) Differential significance of six m6A-related genes between carcinoma and paracancer. (G) The interaction probability for DUSP5 and YTHDF1 predicted by RPISeq. (H) RIP-PCR showed that DUSP5 mRNA was immunoprecipitated by YTHDF1 protein. (I) The RNA levels of DUSP5 expression in LUAD cells. (J) Western blotting showed the DUSP5, Vimentin, N-cadherin and E-cadherin expression after YTHDF1 knockdown. (K-L) Cell migratory capability was elucidated by a Wound-healing assay. (M-N) The invasive and migratory abilities were evaluated by Transwell assays. **P < 0.01, and ***P < 0.001
Furthermore, we anticipated the robust interactions between YTHDF1 and DUSP5 (probability score > 0.5 was considered positive) via RPISeq approaches (Fig. 8G). Then, RIP experiments revealed that DUSP5 mRNA was immunoprecipitated by the YTHDF1 protein (Fig. 8H). We successfully knocked down DUSP5 (Fig. 8I). Western blotting confirmed that YTHDF1 silence inhibited DUSP5 expression and EMT progress (Fig. 8J). Consistently, the knockdown of DUSP5 resulted in decreased migration and invasion abilities of LUAD cells (Fig. 8K-N). These data suggested that YTHDF1 could facilitate the DUSP5-mediated EMT process and metastasis through m6A modification.
Construction of the DUSP5-originated genomic model
We identified 111 DUSP5-originated genes in LUAD patients using the limits of |FC| >2 and FDR < 0.05 (Figure S11A). GO annotations data and KEGG analysis showed that DUSP5-originated genes were linked with many malignant biological behaviors and oncogenic signaling pathways, such as epithelial cell migration and MAPK signaling pathway (Figure S11B, C). Using univariate-Cox regression models, we discovered that 69 DUSP5-originated genes can predict the prognosis of LUAD patients (Table S2). Then 69 genes were taken into the least absolute shrinkage and selection operator (LASSO) regression analysis to establish a DUSP5-originated genomic model for LUAD patients. Three prognosis-related genes were identified, including, Gap junction protein, beta 3 (GJB3), family with sequence similarity 83 member A (FAM83A), and fascin actin-bundling protein 1 (FSCN1) (Figure S11D, E). The LUAD patients were stratified into two cohorts predicated upon the median value of the risk score (Figure S11F). The three genes were all strongly expressed in the high-risk group (Figure S11G). LUAD patients who belonged to the high-risk group had a remarkably worse OS (Figure S11H). As observed in Figure S11I, the DUSP5-originated genomic model had the highest C-index among these prognostic factors including risk score, and T, N, M, and TNM stage.
Verification of prognostic significance of DUSP5-originated genomic model
The predictive value of this DUSP5-originated genomic model was verified among external datasets. The high-risk group could forecast a worse outcome in the GSE30219, GSE31210, GSE41271, GSE50081, and GSE72094 datasets (Figure S12A-E). Furthermore, the risk score of the DUSP5-originated genomic model could predict poor prognosis following cox-regression analysis in the TCGA-LUAD dataset (Figure S12F, G). To produce an accurate predictive model for therapeutic application, a nomogram based on the risk score, T, N, M, and TNM stage was built (Figure S12H). The calibration curves revealed that this model had excellent prediction accuracy (Figure S12I).
The DUSP5-originated genomic model revealed two clusters with distinct immune features
We then investigated the variations in levels of immune cells between various groups to comprehend the relationships between the DUSP5-originated genomic model and the immune microenvironment (TME). ssGSEA algorithm revealed that 16 of the 28 immune infiltrating cells varied between two groups, with the high-risk group showing higher infiltration of immune cells, indicating an immunological “hot” phenotype. On the contrary, the low-risk group displayed an immunological phenotype known as “cold” (Figure S13A). Additionally, we found the expression of immune checkpoint genes, including PDCD1 (PD-1), CD274 (PD-L1), IL23A, JAK1, LDHA, LAMA3, PDCD1LG2, PVR, TNFRSF18, TNFRSF4, and TNFRSF9, were higher in the high-risk group compared with that in the low-risk group (Figure S13B, Figure S14).
As shown in Figure S13C, the 15 genes, the most frequently mutated proto-oncogene or tumor suppressor gene, had a high rate of mutations (93.19% and 87.29%) in the two groups, respectively. TMB is an accurate predictive biomarker for immune checkpoint blockade (ICB) therapy, and patients with higher TMB may suggest better immunotherapy efficiency [50]. Our data revealed disclosed an augmented tumor mutational burden (TMB) within the high-risk cohort, suggesting an enhanced potential responsiveness to immunotherapeutic interventions (Figure S13D). Additionally, survival analysis elucidated that the patients within the high-TMB cohort exhibited a superior OS compared to their counterparts in the low-TMB cohort (Figure S13E). Further, the Kaplan-Meier survival curve demonstrated that the cohort characterized by a high TMB concomitant with a low-risk score manifested the most favorable five-year survival prognosis, whereas the group with a low TMB and high-risk score presented the least favorable prognosis (Figure S13F). Consequently, patients in the high-risk group exhibited an increased likelihood of deriving clinical benefit from ICB therapy.
Discussion
The majority of LUAD patients succumb to tumor metastasis, therapeutic resistance including EGFR-TKI treatment, and relapse of disease, rather than their primary tumor [51, 52]. The relevant players and mechanisms have not been sufficiently investigated. We first combined the two public datasets (TCGA-LUAD and GEO: GSE11117) and performed an integrative analysis, identifying DUSP5 with robust prognostic significance and its functions in the EMT and TGF-β signaling pathway. Experimentally, we found that suppression of DUSP5 significantly inhibited the TGF-β signaling pathway-mediated EMT, thus impeding tumor metastasis and EGFR-TKI resistance. The DUSP5 expression was first postulated and then validated to be regulated by DNA methylation. Meanwhile, we have discovered that YTHDF1 could bind DUSP5 mRNA and regulate DUSP5 expression. Then, the two clusters with distinct immune features and TMB were revealed by the DUSP5-derived genomic model (Fig. 9).
For the first time, DUSP5 was demonstrated to result in LUAD progression in this study. The kinase-interacting motif of DUSP5 facilitates its capacity to catalyze the dephosphorylation of threonine/serine as well as tyrosine residues within its substrate molecules [53]. DUSP5 dephosphorylate toward extracellular signal-regulated kinase (ERK), contributing to nuclear translocation and inactivation of ERK [54]. It is reasonable that both DUSP5 and ERK are “double-edged sword” [55, 56]. DUSP5 is overexpressed in human papillary thyroid carcinomas (PTCs), and DUSP5 silencing suppresses the migratory ability and invasiveness of PTCs cells [57]. Consistently, Olaia et al. disclose that DUSP5 was a potential biomarker of poor prognosis, and they speculate that DUSP5 may prevent ERK1/2-mediated tumor development in human neuroblastoma [58]. In line with these findings, our data suggested a pro-tumorigenic role of DUSP5 by both experimental and bioinformatic approaches in LUAD. On the contrary, DUSP5 is found to be a tumor-suppressive factor by inactivation of ERK in gastric cancer, colorectal cancer and so on [59,60,61,62,63]. The reason that DUSP5 functions as an oncogene in LUAD is not investigated here but warrants further exploration.
Our findings indicate that DUSP5 augments resistance to EGFR-TKI and fosters the EMT process by mediating the activation of the TGF-β/Smad signaling pathway in LUAD. TGF-β signaling pathway is the most well-known pathway that controls the EMT process [64]. The canonical TGF-β signaling pathway involves TGF-receptor I kinase directly phosphorylating Smad2/3 before activated Smad2/3 binds with Smad4 to create a complex that translocates into the nucleus and augments the expression of downstream target genes [65]. Non-canonical TGF-β signaling pathways involve JNK, Ras-ERK, PI3K-Akt pathway, and so on [66]. Whether DUSP5 regulates non-canonical TGF-β signaling pathways to modulate EMT in LUAD remains unknown and further exploration is needed. So far, several inhibitors of the TGF-β signaling pathway have been used in preclinical and clinical stages, such as TβRII-Fc (ligand traps), AP12009 (antisense oligonucleotides), SB-431,542 (small molecule receptor kinase inhibitors), Trx-SARA (peptide aptamers) [67, 68]. Importantly, it has been hypothesized that neoplastic cells are induced to initiate EMT as a mechanism to acquire resistance to EGFR-TKIs [69]. Our research outcomes have the potential to inform the advancement of enhanced precision therapeutic strategies for patients with LUAD, particularly those who are most likely to benefit from interventions targeting the TGF-β signaling axis or EGFR-TKIs.
Herein, DNA methylation can control DUSP5 gene expression. DNA methylation, histone modification, nucleosome remodeling, and RNA interference represent exemplars of epigenetic regulatory mechanisms [70]. DNA methylation is the most well-characterized and reversible epigenetic modification to date [71, 72]. Generally, it encompasses the enzymatic conveyance of a methyl moiety to the fifth carbon atom of the cytosine nucleotide, resulting in the formation of 5-methylcytosine in genomic DNA by DNA methyltransferases (DNMTs) [73]. This change modifies chromatin structure, interactions with DNA proteins, DNA structure, stability, and gene expression [74]. In accordance with our study, we observed hypomethylation of DUSP5 in LUAD cells, which resulted in the upregulation of DUSP5 expression. AZA is a well-known, highly effective anticancer medication that is used to treat acute myeloid leukemia, melanoma, breast cancer, and colon cancer [75,76,77,78,79,80]. AZA promotes DNA demethylation by inhibiting DNMTs during replication, inducing silence of gene expression [81, 82]. According to our results, DUSP5 may function as a putative biomarker for directing the stratification of patients with a heightened propensity for responsiveness to AZA.
Here, our findings delineate, for the inaugural instance, that DUSP5 is subject to regulation via YTHDF1-mediated m6A modification. Through a variety of molecular mechanisms, including enhancing protein translation and modifying mRNA stability, YTHDF1 controls the expression of target genes [83]. According to the previous study, YTHDF1 is responsible for promoting translation initiation and subsequent protein translation [84]. Our results of RIP experiments suggested that YTHDF1 modulates DUSP5 expression by altering mRNA stability. Similar mechanisms have previously been reported [85,86,87]. Zhao et al. elucidated that YTHDF1 enhances the stability of c-Myc mRNA catalyzed by METTL3, thereby augmenting the expression of c-Myc [85]. . Additionally, YTHDF1 knockdown dramatically reduces the half-life of HK2 mRNA, suggesting that YTHDF1 may play a role in maintaining HK2 RNA stability [86]. YTHDF1, through its interaction with the m6A-modified 5’UTR of the mRNA, enhances the translation elongation and mRNA stability of PDK4 by recruiting eukaryotic translation elongation factor 2 (eEF-2) and insulin-like growth factor 2 mRNA-binding protein 3 (IGF2BP3) [87]. The specific mechanisms by which YTHDF1 governs DUSP5 expression require further exploration.
Tumor-infiltrating immune cell levels, the expressions of immune checkpoints including PD-1, PD-L1 and CTLA-4, and TMB were found to be elevated in the high-risk group relative to the low-risk group, as classified by the genomic model-derived from DUSP5. Immunotherapy, including ICB therapy, has brought revolutionary changes to the management of LUAD [88, 89]. Immune infiltrations are the primary targets of the immunotherapy [90]. Immune “hot” tumors with more infiltrating immune cells are more likely to be responsive to immunotherapy than immune “cold” tumors with a lower density of infiltrating immune cells [91]. Moreover, immune checkpoint molecules and TMB have been proven to be reliable biomarkers to predict the response of immune checkpoint blockade therapy [92, 93]. Thus, our DUSP5-originated genomic model for LUAD could help to appraise a predictor of response to ICB treatment strategies and display robust performance on survival prediction.
Conclusions
In summary, we first demonstrated that DUSP5 is a novel prognostic biomarker and pro-oncogenic gene for LUAD. Our data indicated DNA hypomethylation and YTHDF1-mediated aberrant increased expression of DUSP5 orchestrates metastasis and development of EGFR-TKI resistance in LUAD by inducing TGF-β/Smad signaling pathway-mediated EMT. Our results suggested that the involvement of the DUSP5 and DUSP5-derived genes might be utilized to guide a personalized approach to metastatic patients and patients with target therapy and immunotherapy in LUAD.
Data availability
Data is available on reasonable request. All data generated or analyzed during this study are available from either the supplementary information or the corresponding author.
Abbreviations
- EMT:
-
Epithelial-mesenchymal transition
- EGFR:
-
Epidermal growth factor receptor
- EGFR-TKIs:
-
Epidermal growth factor receptor tyrosine kinase inhibitors
- LUAD:
-
Lung adenocarcinoma
- TCGA:
-
The Cancer Genome Atlas
- GEO:
-
Gene Expression Omnibus
- GSEA:
-
Gene set enrichment analysis
- MeRIP:
-
methylated RNA immunoprecipitation
- RIP:
-
RNA immunoprecipitation
- ssGSEA:
-
Single-sample gene set enrichment
- DUSP5:
-
Dual-specificity phosphatase 5
- YTHDF1:
-
YTH N6-methyladenosine RNA binding protein 1
- AZA:
-
5′Azacytidine
- TMB:
-
Tumor mutational burden
- NSCLC:
-
Non-small cell lung cancer
- TJs:
-
Tight junctions
- AJs:
-
Adherence junctions
- SPP1:
-
Secreted phosphoprotein 1
- m6A:
-
N6-methyladenosine
- FPKM:
-
Fragments Per Kilobase of exon model per Million mapped fragments
- FC:
-
Fold change
- FDR:
-
False-discovery rate
- FBS:
-
Fetal bovine serum
- NESs:
-
Normalized enrichment scores
- MSigDB:
-
Molecular Signatures Database v7.5.1
- GSCALite:
-
Gene Set Cancer Analysis
- CGIs:
-
CpG islands
- DM:
-
With distant metastasis
- N-DM:
-
Without distant metastasis
- PCA:
-
Principal components analysis
- FAM83A:
-
Family with sequence similarity 83 member A
- GJB3:
-
Gap junction protein, beta 3
- FSCN1:
-
Fascin actin-bundling protein 1
- TME:
-
The immune microenvironment
- ICB:
-
Immune checkpoint blockade
- ERK:
-
Extracellular signal-regulated kinase
- PTCs:
-
Papillary thyroid carcinomas
- DNMTs:
-
DNA methyltransferases
References
Zhang HT, Peng R, Chen S, Shen A, Zhao L, Tang W, Wang XH, Li ZY, Zha ZG, Yi M, et al. Versatile Nano-PROTAC-Induced Epigenetic Reader degradation for efficient Lung Cancer Therapy. Adv Sci (Weinh). 2022;9:e2202039.
Qiu H, Cao S, Xu R. Cancer incidence, mortality, and burden in China: a time-trend analysis and comparison with the United States and United Kingdom based on the global epidemiological data released in 2020. Cancer Commun (Lond). 2021;41:1037–48.
Zhang LX, Gao J, Long X, Zhang PF, Yang X, Zhu SQ, Pei X, Qiu BQ, Chen SW, Lu F, et al. The circular RNA circHMGB2 drives immunosuppression and anti-PD-1 resistance in lung adenocarcinomas and squamous cell carcinomas via the miR-181a-5p/CARM1 axis. Mol Cancer. 2022;21:110.
Wang S, Gao A, Liu J, Sun Y. First-line therapy for advanced non-small cell lung cancer with activating EGFR mutation: is combined EGFR-TKIs and chemotherapy a better choice? Cancer Chemother Pharmacol. 2018;81:443–53.
Suda K, Rivard CJ, Mitsudomi T, Hirsch FR. Overcoming resistance to EGFR tyrosine kinase inhibitors in lung cancer, focusing on non-T790M mechanisms. Expert Rev Anticancer Ther. 2017;17:779–86.
Lei HM, Zhang KR, Wang CH, Wang Y, Zhuang GL, Lu LM, Zhang J, Shen Y, Chen HZ, Zhu L. Aldehyde dehydrogenase 1A1 confers erlotinib resistance via facilitating the reactive oxygen species-reactive carbonyl species metabolic pathway in lung adenocarcinomas. Theranostics. 2019;9:7122–39.
Jacobsen K, Bertran-Alamillo J, Molina MA, Teixido C, Karachaliou N, Pedersen MH, Castellvi J, Garzon M, Codony-Servat C, Codony-Servat J, et al. Convergent akt activation drives acquired EGFR inhibitor resistance in lung cancer. Nat Commun. 2017;8:410.
Wang J, Zhang N, Han Q, Lu W, Wang L, Yang D, Zheng M, Zhang Z, Liu H, Lee TH, et al. Pin1 inhibition reverses the acquired resistance of human hepatocellular carcinoma cells to Regorafenib via the Gli1/Snail/E-cadherin pathway. Cancer Lett. 2019;444:82–93.
Yuan K, Xie K, Lan T, Xu L, Chen X, Li X, Liao M, Li J, Huang J, Zeng Y, et al. TXNDC12 promotes EMT and metastasis of hepatocellular carcinoma cells via activation of beta-catenin. Cell Death Differ. 2020;27:1355–68.
Tan TZ, Miow QH, Miki Y, Noda T, Mori S, Huang RY, Thiery JP. Epithelial-mesenchymal transition spectrum quantification and its efficacy in deciphering survival and drug responses of cancer patients. EMBO Mol Med. 2014;6:1279–93.
Hatami R, Sieuwerts AM, Izadmehr S, Yao Z, Qiao RF, Papa L, Look MP, Smid M, Ohlssen J, Levine AC, et al. KLF6-SV1 drives breast cancer metastasis and is associated with poor survival. Sci Transl Med. 2013;5:169ra12.
Lamouille S, Xu J, Derynck R. Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol. 2014;15:178–96.
Singh M, Yelle N, Venugopal C, Singh SK. EMT: mechanisms and therapeutic implications. Pharmacol Ther. 2018;182:80–94.
Suda K, Tomizawa K, Fujii M, Murakami H, Osada H, Maehara Y, Yatabe Y, Sekido Y, Mitsudomi T. Epithelial to mesenchymal transition in an epidermal growth factor receptor-mutant lung cancer cell line with acquired resistance to erlotinib. J Thorac Oncol. 2011;6:1152–61.
Witta SE, Gemmill RM, Hirsch FR, Coldren CD, Hedman K, Ravdel L, Helfrich B, Dziadziuszko R, Chan DC, Sugita M, et al. Restoring E-cadherin expression increases sensitivity to epidermal growth factor receptor inhibitors in lung cancer cell lines. Cancer Res. 2006;66:944–50.
Sequist LV, Waltman BA, Dias-Santagata D, Digumarthy S, Turke AB, Fidias P, Bergethon K, Shaw AT, Gettinger S, Cosper AK, et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci Transl Med. 2011;3:75ra26.
Singh A, Greninger P, Rhodes D, Koopman L, Violette S, Bardeesy N, Settleman J. A gene expression signature associated with K-Ras addiction reveals regulators of EMT and tumor cell survival. Cancer Cell. 2009;15:489–500.
Carretero J, Shimamura T, Rikova K, Jackson AL, Wilkerson MD, Borgman CL, Buttarazzi MS, Sanofsky BA, McNamara KL, Brandstetter KA, et al. Integrative genomic and proteomic analyses identify targets for Lkb1-deficient metastatic lung tumors. Cancer Cell. 2010;17:547–59.
Kondratyeva L, Chernov I, Kopantzev E, Didych D, Kuzmich A, Alekseenko I, Kostrov S, Sverdlov E. Pancreatic lineage specifier PDX1 increases adhesion and decreases motility of Cancer cells. Cancers (Basel). 2021; 13.
Soucheray M, Capelletti M, Pulido I, Kuang Y, Paweletz CP, Becker JH, Kikuchi E, Xu C, Patel TB, Al-Shahrour F, et al. Intratumoral Heterogeneity in EGFR-Mutant NSCLC results in Divergent Resistance mechanisms in response to EGFR tyrosine kinase inhibition. Cancer Res. 2015;75:4372–83.
Guo Y, Peng D, Zhou J, Lin S, Wang C, Ning W, Xu H, Deng W, Xue Y. iEKPD 2.0: an update with rich annotations for eukaryotic protein kinases, protein phosphatases and proteins containing phosphoprotein-binding domains. Nucleic Acids Res. 2019;47:D344–50.
Zhu Y, Cang S, Chen B, Gu Y, Jiang M, Yan J, Shao F, Huang X. Patient stratification of Clear Cell Renal Cell Carcinoma using the global transcription factor activity Landscape Derived from RNA-Seq Data. Front Oncol. 2020;10:526577.
Nersisyan L, Hopp L, Loeffler-Wirth H, Galle J, Loeffler M, Arakelyan A, Binder H. Telomere Length Maintenance and its transcriptional regulation in Lynch Syndrome and sporadic colorectal carcinoma. Front Oncol. 2019;9:1172.
Tran AN, Walker K, Harrison DG, Chen W, Mobley J, Hocevar L, Hackney JR, Sedaka RS, Pollock JS, Goldberg MS, et al. Reactive species balance via GTP cyclohydrolase I regulates glioblastoma growth and tumor initiating cell maintenance. Neuro Oncol. 2018;20:1055–67.
Xie Y, Liu Y, Li Q, Chen J. Polo-like kinase 2 promotes chemoresistance and predicts limited survival benefit from adjuvant chemotherapy in colorectal cancer. Int J Oncol. 2018;52:1401–14.
Dong B, Wu C, Huang L, Qi Y. Macrophage-related SPP1 as a potential biomarker for early Lymph Node Metastasis in Lung Adenocarcinoma. Front Cell Dev Biol. 2021;9:739358.
Gu X, Huang X, Zhang X, Wang C. Development and validation of a DNA methylation-related classifier of circulating Tumour cells to Predict Prognosis and to provide a therapeutic strategy in Lung Adenocarcinoma. Int J Biol Sci. 2022;18:4984–5000.
Yuan H, Yan M, Zhang G, Liu W, Deng C, Liao G, Xu L, Luo T, Yan H, Long Z, et al. CancerSEA: a cancer single-cell state atlas. Nucleic Acids Res. 2019;47:D900–8.
Liu CJ, Hu FF, Xia MX, Han L, Zhang Q, Guo AY. GSCALite: a web server for gene set cancer analysis. Bioinformatics. 2018;34:3771–2.
Koch A, Jeschke J, Van Criekinge W, van Engeland M, De Meyer T. MEXPRESS update 2019. Nucleic Acids Res. 2019;47:W561–5.
Caruso FP, D’Andrea MR, Coppola L, Landriscina M, Condelli V, Cerulo L, Giordano G, Porras A, Pancione M. Lymphocyte antigen 6G6D-mediated modulation through p38alpha MAPK and DNA methylation in colorectal cancer. Cancer Cell Int. 2022;22:253.
Li LC, Dahiya R. MethPrimer: designing primers for methylation PCRs. Bioinformatics. 2002;18:1427–31.
He R, Man C, Huang J, He L, Wang X, Lang Y, Fan Y. Identification of RNA methylation-related lncRNAs signature for Predicting Hot and Cold tumors and prognosis in Colon cancer. Front Genet. 2022;13:870945.
Banerjee P, Xiao GY, Tan X, Zheng VJ, Shi L, Rabassedas MNB, Guo HF, Liu X, Yu J, Diao L, et al. The EMT activator ZEB1 accelerates endosomal trafficking to establish a polarity axis in lung adenocarcinoma cells. Nat Commun. 2021;12:6354.
Karamanou K, Franchi M, Vynios D, Brezillon S. Epithelial-to-mesenchymal transition and invadopodia markers in breast cancer: Lumican a key regulator. Semin Cancer Biol. 2020;62:125–33.
Tang X, Sui X, Weng L, Liu Y. SNAIL1: linking Tumor Metastasis to Immune Evasion. Front Immunol. 2021;12:724200.
Cruceriu D, Baldasici O, Balacescu O, Berindan-Neagoe I. The dual role of tumor necrosis factor-alpha (TNF-alpha) in breast cancer: molecular insights and therapeutic approaches. Cell Oncol (Dordr). 2020;43:1–18.
Massague J. TGFbeta Cancer Cell. 2008;134:215–30.
Loie E, Charrier LE, Sollier K, Masson JY, Laprise P. CRB3A controls the morphology and cohesion of Cancer cells through Ehm2/p114RhoGEF-Dependent signaling. Mol Cell Biol. 2015;35:3423–35.
Parikh BH, Liu Z, Blakeley P, Lin Q, Singh M, Ong JY, Ho KH, Lai JW, Bogireddi H, Tran KC, et al. A bio-functional polymer that prevents retinal scarring through modulation of NRF2 signalling pathway. Nat Commun. 2022;13:2796.
Mackenzie NC, Raz E. Found in translation: a new player in EMT. Dev Cell. 2006;11:434–6.
Tiwari N, Gheldof A, Tatari M, Christofori G. EMT as the ultimate survival mechanism of cancer cells. Semin Cancer Biol. 2012;22:194–207.
Li Q, Sodroski C, Lowey B, Schweitzer CJ, Cha H, Zhang F, Liang TJ. Hepatitis C virus depends on E-cadherin as an entry factor and regulates its expression in epithelial-to-mesenchymal transition. Proc Natl Acad Sci U S A. 2016;113:7620–5.
Thomson S, Buck E, Petti F, Griffin G, Brown E, Ramnarine N, Iwata KK, Gibson N, Haley JD. Epithelial to mesenchymal transition is a determinant of sensitivity of non-small-cell lung carcinoma cell lines and xenografts to epidermal growth factor receptor inhibition. Cancer Res. 2005;65:9455–62.
Park KS, Raffeld M, Moon YW, Xi L, Bianco C, Pham T, Lee LC, Mitsudomi T, Yatabe Y, Okamoto I, et al. CRIPTO1 expression in EGFR-mutant NSCLC elicits intrinsic EGFR-inhibitor resistance. J Clin Invest. 2014;124:3003–15.
Lue NZ, Garcia EM, Ngan KC, Lee C, Doench JG, Liau BB. Base editor scanning charts the DNMT3A activity landscape. Nat Chem Biol. 2022.
Chen Y, Wan R, Zou Z, Lao L, Shao G, Zheng Y, Tang L, Yuan Y, Ge Y, He C, et al. O-GlcNAcylation determines the translational regulation and phase separation of YTHDF proteins. Nat Cell Biol. 2023;25:1676–90.
Bao X, Zhang Y, Li H, Teng Y, Ma L, Chen Z, Luo X, Zheng J, Zhao A, Ren J, et al. RM2Target: a comprehensive database for targets of writers, erasers and readers of RNA modifications. Nucleic Acids Res. 2023;51:D269–79.
Luo X, Li H, Liang J, Zhao Q, Xie Y, Ren J, Zuo Z. RMVar: an updated database of functional variants involved in RNA modifications. Nucleic Acids Res. 2021;49:D1405–12.
Li X, Dong H, Chen L, Wang Y, Hao Z, Zhang Y, Jiao Y, Zhao Z, Peng X, Zhan X. Identification of N7-methylguanosine related subtypes and construction of prognostic model in gastric cancer. Front Immunol. 2022;13:984149.
Choi J, Yoon S, Kim D, Moon YW, Lee CH, Seo S, Cheon J, Gho YS, Kim C, Lee ER, et al. Transglutaminase 2 induces intrinsic EGFR-TKI resistance in NSCLC harboring EGFR sensitive mutations. Am J Cancer Res. 2019;9:1708–21.
Zhou Y, Hong T, Tong L, Liu W, Yang X, Luo J, Wang F, Li J, Yan L. Astragalus polysaccharide combined with 10-hydroxycamptothecin inhibits metastasis in non-small cell lung carcinoma cell lines via the MAP4K3/mTOR signaling pathway. Int J Mol Med. 2018;42:3093–104.
Chen HF, Chuang HC, Tan TH. Regulation of dual-specificity phosphatase (DUSP) Ubiquitination and Protein Stability. Int J Mol Sci. 2019; 20.
Mandl M, Slack DN, Keyse SM. Specific inactivation and nuclear anchoring of extracellular signal-regulated kinase 2 by the inducible dual-specificity protein phosphatase DUSP5. Mol Cell Biol. 2005;25:1830–45.
Sugiura R, Satoh R, Takasaki T. ERK: a double-edged Sword in Cancer. ERK-Dependent apoptosis as a potential therapeutic strategy for Cancer. Cells. 2021; 10.
Kutty RG, Talipov MR, Bongard RD, Lipinski RAJ, Sweeney NL, Sem DS, Rathore R, Ramchandran R. Dual specificity phosphatase 5-Substrate Interaction: a mechanistic perspective. Compr Physiol. 2017;7:1449–61.
Buffet C, Hecale-Perlemoine K, Bricaire L, Dumont F, Baudry C, Tissier F, Bertherat J, Cochand-Priollet B, Raffin-Sanson ML, Cormier F, et al. DUSP5 and DUSP6, two ERK specific phosphatases, are markers of a higher MAPK signaling activation in BRAF mutated thyroid cancers. PLoS ONE. 2017;12:e0184861.
Aurtenetxe O, Zaldumbide L, Erramuzpe A, Lopez R, Lopez JI, Cortes JM, Pulido R, Nunes-Xavier CE. DUSP5 expression associates with poor prognosis in human neuroblastoma. Exp Mol Pathol. 2018;105:272–8.
Wang R, Bao HB, Du WZ, Chen XF, Liu HL, Han DY, Wang LG, Wu JN, Wang CL, Yang MC, et al. P68 RNA helicase promotes invasion of glioma cells through negatively regulating DUSP5. Cancer Sci. 2019;110:107–17.
Yang Z, Huang D, Meng M, Wang W, Feng J, Fang L, Chen H, Zou S. BAF53A drives colorectal cancer development by regulating DUSP5-mediated ERK phosphorylation. Cell Death Dis. 2022;13:1049.
Ma Z, Gao X, Shuai Y, Wu X, Yan Y, Xing X, Ji J. EGR1-mediated linc01503 promotes cell cycle progression and tumorigenesis in gastric cancer. Cell Prolif. 2021;54:e12922.
Nokin MJ, Bellier J, Durieux F, Peulen O, Rademaker G, Gabriel M, Monseur C, Charloteaux B, Verbeke L, van Laere S, et al. Methylglyoxal, a glycolysis metabolite, triggers metastasis through MEK/ERK/SMAD1 pathway activation in breast cancer. Breast Cancer Res. 2019;21:11.
Kidger AM, Saville MK, Rushworth LK, Davidson J, Stellzig J, Ono M, Kuebelsbeck LA, Janssen KP, Holzmann B, Morton JP, et al. Suppression of mutant Kirsten-RAS (KRAS(G12D))-driven pancreatic carcinogenesis by dual-specificity MAP kinase phosphatases 5 and 6. Oncogene. 2022;41:2811–23.
Gay D, Ghinatti G, Guerrero-Juarez CF, Ferrer RA, Ferri F, Lim CH, Murakami S, Gault N, Barroca V, Rombeau I, et al. Phagocytosis of wnt inhibitor SFRP4 by late wound macrophages drives chronic wnt activity for fibrotic skin healing. Sci Adv. 2020;6:eaay3704.
Chan SC, Zhang Y, Shao A, Avdulov S, Herrera J, Aboudehen K, Pontoglio M, Igarashi P. Mechanism of fibrosis in HNF1B-Related autosomal Dominant Tubulointerstitial kidney disease. J Am Soc Nephrol. 2018;29:2493–509.
Ikushima H, Miyazono K. TGFbeta signalling: a complex web in cancer progression. Nat Rev Cancer. 2010;10:415–24.
Connolly EC, Freimuth J, Akhurst RJ. Complexities of TGF-beta targeted cancer therapy. Int J Biol Sci. 2012;8:964–78.
Hu Q, Hisamatsu T, Haemmerle M, Cho MS, Pradeep S, Rupaimoole R, Rodriguez-Aguayo C, Lopez-Berestein G, Wong STC, Sood AK, et al. Role of platelet-derived Tgfbeta1 in the progression of Ovarian Cancer. Clin Cancer Res. 2017;23:5611–21.
Lu Y, Liu Y, Oeck S, Zhang GJ, Schramm A, Glazer PM. Hypoxia induces resistance to EGFR inhibitors in Lung Cancer cells via Upregulation of FGFR1 and the MAPK pathway. Cancer Res. 2020;80:4655–67.
Liu C, Xiong Q, Li Q, Lin W, Jiang S, Zhang D, Wang Y, Duan X, Gong P, Kang N. CHD7 regulates bone-fat balance by suppressing PPAR-gamma signaling. Nat Commun. 2022; 13:1989.
Nabais MF, Gadd DA, Hannon E, Mill J, McRae AF, Wray NR. An overview of DNA methylation-derived trait score methods and applications. Genome Biol. 2023;24:28.
Wang M, Wang P, Tu L, Zhu S, Zhang L, Li Z, Zhang Q, Yuan D, Zhang X. Multi-omics maps of cotton fibre reveal epigenetic basis for staged single-cell differentiation. Nucleic Acids Res. 2016;44:4067–79.
Yuan H, Du S, Chen L, Xu X, Wang Y, Ji F. Hypomethylation of nerve growth factor (NGF) promotes binding of C/EBPalpha and contributes to inflammatory hyperalgesia in rats. J Neuroinflammation. 2020;17:34.
Jie C, Li R, Cheng Y, Wang Z, Wu Q, Xie C. Prospects and feasibility of synergistic therapy with radiotherapy, immunotherapy, and DNA methyltransferase inhibitors in non-small cell lung cancer. Front Immunol. 2023;14:1122352.
El Baroudi M, La Sala D, Cinti C, Capobianco E. Pathway landscapes and epigenetic regulation in breast cancer and melanoma cell lines. Theor Biol Med Model. 2014;11(Suppl 1):S8.
Momparler RL. Epigenetic therapy of cancer with 5-aza-2’-deoxycytidine (decitabine). Semin Oncol. 2005;32:443–51.
Lemaire M, Momparler LF, Bernstein ML, Marquez VE, Momparler RL. Enhancement of antineoplastic action of 5-aza-2’-deoxycytidine by zebularine on L1210 leukemia. Anticancer Drugs. 2005;16:301–8.
Momparler RL, Bovenzi V. DNA methylation and cancer. J Cell Physiol. 2000;183:145–54.
Maurillo L, Venditti A, Spagnoli A, Gaidano G, Ferrero D, Oliva E, Lunghi M, D’Arco AM, Levis A, Pastore D, et al. Azacitidine for the treatment of patients with acute myeloid leukemia: report of 82 patients enrolled in an Italian compassionate program. Cancer. 2012;118:1014–22.
Li W, Wu D, Niu Z, Jiang D, Ma H, He H, Zuo X, Xie X, He Y. 5-Azacytidine suppresses EC9706 cell proliferation and metastasis by upregulating the expression of SOX17 and CDH1. Int J Mol Med. 2016;38:1047–54.
Wei H, Jiang S, Chen L, He C, Wu S, Peng H. Characterization of cytosine methylation and the DNA methyltransferases of Toxoplasma Gondii. Int J Biol Sci. 2017;13:458–70.
Deng X, Hu Z. Generation of cochlear hair cells from Sox2 positive supporting cells via DNA demethylation. Int J Mol Sci. 2020; 21.
Ren W, Yuan Y, Li Y, Mutti L, Peng J, Jiang X. The function and clinical implication of YTHDF1 in the human system development and cancer. Biomark Res. 2023;11:5.
Nettersheim D, Berger D, Jostes S, Kristiansen G, Lochnit G, Schorle H. N6-Methyladenosine detected in RNA of testicular germ cell tumors is controlled by METTL3, ALKBH5, YTHDC1/F1/F2, and HNRNPC as writers, erasers, and readers. Andrology. 2019;7:498–506.
Zhao W, Cui Y, Liu L, Ma X, Qi X, Wang Y, Liu Z, Ma S, Liu J, Wu J. METTL3 facilitates oral squamous cell Carcinoma Tumorigenesis by enhancing c-Myc Stability via YTHDF1-Mediated m(6)a modification. Mol Ther Nucleic Acids. 2020;20:1–12.
Wang Q, Guo X, Li L, Gao Z, Su X, Ji M, Liu J. N(6)-methyladenosine METTL3 promotes cervical cancer tumorigenesis and Warburg effect through YTHDF1/HK2 modification. Cell Death Dis. 2020;11:911.
Li Z, Peng Y, Li J, Chen Z, Chen F, Tu J, Lin S, Wang H. N(6)-methyladenosine regulates glycolysis of cancer cells through PDK4. Nat Commun. 2020;11:2578.
Park C, Na KJ, Choi H, Ock CY, Ha S, Kim M, Park S, Keam B, Kim TM, Paeng JC, et al. Tumor immune profiles noninvasively estimated by FDG PET with deep learning correlate with immunotherapy response in lung adenocarcinoma. Theranostics. 2020;10:10838–48.
Wang Z, Chen L, Ma Y, Li X, Hu A, Wang H, Wang W, Li X, Tian B, Dong J. Peptide vaccine-conjugated mesoporous carriers synergize with immunogenic cell death and PD-L1 blockade for amplified immunotherapy of metastatic spinal. J Nanobiotechnol. 2021;19:243.
Tang L, Wei R, Chen R, Fan G, Zhou J, Qi Z, Wang K, Wei Q, Wei X, Xu X. Establishment and validation of a cholesterol metabolism-related prognostic signature for hepatocellular carcinoma. Comput Struct Biotechnol J. 2022;20:4402–14.
Galon J, Bruni D. Approaches to treat immune hot, altered and cold tumours with combination immunotherapies. Nat Rev Drug Discov. 2019;18:197–218.
Chen Y, Wang X, Deng X, Zhang Y, Liao R, Li Y, Yang H, Chen K. DNA damage repair Status predicts Opposite Clinical Prognosis Immunotherapy and Non-immunotherapy in Hepatocellular Carcinoma. Front Immunol. 2021;12:676922.
Park S, Joung JG, Min YW, Nam JY, Ryu D, Oh D, Park WY, Lee SH, Choi Y, Ahn JS, et al. Paired whole exome and transcriptome analyses for the immunogenomic changes during concurrent chemoradiotherapy in esophageal squamous cell carcinoma. J Immunother Cancer. 2019;7:128.
Acknowledgements
Not applicable.
Funding
Project funded by the National Natural Science Foundation of China (81772474, 82072563 to LC, 81803023, 82172587 to YX, 82103519 to JH), China Postdoctoral Science Foundation (2017M621307 to YX, 2021M693826 to JH), Heilongjiang Provincial Postdoctoral Science Foundation (LBH-Z17182 to YX, LBH-Z21187 to JH), Haiyan Foundation of Harbin Medical University Cancer Hospital (JJQN2018-02 to SY, JJZD2021-07 to YX), the Top-Notch Youth Fund from Harbin Medical University Cancer Hospital (BJQN2019-07 to YX, BJQN 2021-02 to JH), the Outstanding Youth Project of Haiyan Fund (YX), Climing program of Harbin Medical University Cancer Hospital (PDYS2024-17 to LC), 2023 Basic Research Operating Expenses of Provincial Higher Education Institutions in Heilongjiang Province for Scientific Research Projects (MRC).
Author information
Authors and Affiliations
Contributions
LC, JH, MRC and WNF designed the study. WNF, YX, SY, WL, JFN and FLT carried development of methodology. WNF, YX, XW, YNZ and LXL supported analysis and interpretation of data. WNF and YX conducted experiments. WNF, YX and SY wrote, reviewed, and edited the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All animal work was approved by the Ethics Committee of Harbin Medical University Cancer Hospital .
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Fan, W., Xing, Y., Yan, S. et al. DUSP5 regulated by YTHDF1-mediated m6A modification promotes epithelial-mesenchymal transition and EGFR-TKI resistance via the TGF-β/Smad signaling pathway in lung adenocarcinoma. Cancer Cell Int 24, 208 (2024). https://doi.org/10.1186/s12935-024-03382-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12935-024-03382-6