Abstract
Background
The maintenance of mitochondrial homeostasis is critical for tumor initiation and malignant progression because it increases tumor cell survival and growth. The molecular events controlling mitochondrial integrity that facilitate the development of hepatocellular carcinoma (HCC) remain unclear. Here, we report that UBX domain-containing protein 1 (UBXN1) hyperactivation is essential for mitochondrial homeostasis and liver tumorigenesis.
Methods
Oncogene-induced mouse liver tumor models were generated with the Sleeping Beauty (SB) transposon delivery system. Assessment of HCC cell growth in vivo and in vitro, including tumour formation, colony formation, TUNEL and FACS assays, was conducted to determine the effects of UBXN1 on HCC cells, as well as the involvement of the UBXN1-prohibitin (PHB) interaction in mitochondrial function. Coimmunoprecipitation (Co-IP) was used to assess the interaction between UBXN1 and PHB. Liver hepatocellular carcinoma (LIHC) datasets and HCC patient samples were used to assess the expression of UBXN1.
Results
UBXN1 expression is commonly upregulated in human HCCs and mouse liver tumors and is associated with poor overall survival in HCC patients. UBXN1 facilitates the growth of human HCC cells and promotes mouse liver tumorigenesis driven by the NRas/c-Myc or c-Myc/shp53 combination. UBXN1 interacts with the inner mitochondrial membrane protein PHB and sustains PHB expression. UBXN1 inhibition triggers mitochondrial damage and liver tumor cell apoptosis.
Conclusions
UBXN1 interacts with PHB and promotes mitochondrial homeostasis during liver tumorigenesis.
Similar content being viewed by others
Background
HCC is the third leading cause of cancer-related mortality worldwide, with a 5-year survival rate of approximately 18% [1, 2]. The incidence of genetic alterations in HCCs is relatively low compared to that in other cancer types, and alterations only occur in some subgroups of HCCs; aberrant molecular events regulating the processes of apoptosis and mitochondrial homeostasis are commonly observed in HCC and drive hepatocarcinogenesis [3,4,5,6].
The maintenance of mitochondrial homeostasis in tumor cells is critical for cellular energy production and the survival of tumor cells under genetic or environmental stresses [3, 4, 7]. Oncogenic driver mutations, such as those affecting KRas and c-Myc, maintain proper mitochondrial bioenergetics and ATP levels by modulating the expression of mitochondrial proteins, including MFN2, OPA1 and DRP1, in transformed cells [4, 8,9,10]. Loss of these mitochondrial proteins damages mitochondria and shifts mitochondrial dynamics, which alters mitochondrial membrane potential and oxidative phosphorylation, resulting in excessive production of reactive oxygen species (ROS) and induction of mitochondria-associated apoptosis, which are detrimental to liver tumorigenesis [3,4,5, 11, 12]. Notably, the integrity of mitochondria is tightly regulated by mitochondrial inner membrane proteins, such as PHB, TIMM13 and VDAC1, which are involved in membrane architecture and dynamics [11, 13,14,15,16]. Emerging evidence has shown that PHB is highly expressed in multiple types of cancer, including pancreatic cancer (PDAC), breast carcinoma (BC) and gastric cancer (GC), and participates in biological processes related to tumorigenesis [17,18,19]. For instance, PHB is phosphorylated by AKT at Thr258 and thereby facilitates bladder cancer cell proliferation [20]. Additionally, PHB promotes the growth and metastasis of cervical cancer through binding to Raf-1 and enhances the Ras-mediated activation of the Raf/MEK/ERK pathway [21]. Targeting PHB using genetic or pharmacological approaches decrease the expression of Bcl-2, activates caspase(s) and induces mitochondria-dependent apoptosis [22, 23]. However, the molecular events controlling PHB expression in tumor development and progression remain elusive.
UBXN1, a member of the ubiquitin regulatory X domain-containing protein (UBXN) family, comprises 13 members that share similar functional ubiquitin-associated domain (UBA) or ubiquitin regulatory X (UBX) domains and is implicated in regulating multiple cellular processes, including the innate immune response, replisome disassembly and mitophagy [24,25,26,27,28]. UBXN1 has dual effects on tumor growth in different types of cancer [29,30,31]. YTHDF2 recognizes METTL3-mediated m6A on UBXN1 mRNA and decreases its expression, which in turn promotes NF-κB signaling and the malignant progression of gliomas [31]. In contrast, the inhibition of UBXN1 attenuated prostate cancer cell proliferation by inducing mitochondria-associated apoptosis [29]. Furthermore, UBXN1 recognizes and binds to autoubiquitinated BRCA1, a tumor suppressor for breast cancer [32], and inhibits its enzymatic function, suggesting that UBXN1 plays an oncogenic role in cancer.
In this study, we found that the expression of UBXN1 was upregulated in NRas G12V/c-Myc-driven and c-Myc/shp53-driven mouse HCC tumors and human HCCs. UBXN1 maintained PHB expression by binding to it, which promoted mitochondrial homeostasis. Aberrant upregulation of UBXN1 inhibited mitochondria-associated cellular dysfunction and promoted liver tumorigenesis.
Materials and methods
Cell lines and tissue specimens
PLC/PRF/5, Hepa1-6 and HEK-293T cell lines were purchased from American Type Culture Collection (ATCC, Manassas VA, USA). HCC cell line MHCC97H was supplied by Liver Cancer Institute, Fudan University (Shanghai, China). Mouse LPC-HRas cells were previously generated in our laboratory [33]. All cells were cultured in DMEM (Gibco) containing 10% FBS, 50 U/ml of penicillin and streptomycin (Yeasen, China) at 37 °C under 5% CO2. The absence of mycoplasma contamination was determined by PCR assay. Human 21 paired HCC samples were obtained from HCC patients who received curative surgery in the Department of Liver Surgery, Renji Hospital, School of Medicine, Shanghai Jiaotong University (Shanghai, China). The tissue microarray LVC-1605 is from Shanghai Outdo Biotech Company (Shanghai, China). The clinical information of HCC patients is presented in supplementary Tables 4 or 5.
Reagents and antibodies
Puromycin (CL13900) was purchased from MCE. CCCP (C2759) and MTT (Thiazolyl Blue Tetrazolium Bromide, M5655) were purchased from Sigma. Lipofectamine 2000 Transfection Reagent was purchased from Invitrogen. Polyethyleneimine transfection reagent (23966-1) was purchased from Polysciences. JC-1 fluorescent probe (IJ0300) was purchased from Solarbio. Caspase Inhibitor Z-VAD-FMK (C1202) and TUNEL Apoptosis Assay Kit (C1089) were obtained from Beyotime. MitoTracker™ Deep Red FM (M22426) and MitoTracker™ Green FM (M7514) Fluorescent Dyes were from ThermoFisher. DMSO was from SantaCruz. All antibodies used in this study are listed in Supplementary Table 1.
Animal studies
Male C57BL/6 mice (6–8 weeks old) were purchased from Lingchang Biotechnology Co. (Shanghai, China). BALB/c nude mice were purchased from Shanghai SLAC Laboratory Animal Company (Shanghai, China). C57BL/6 mice were subjected to hydrodynamic tail vein injection (HDTVi) with the endotoxin-free plasmid combination [28]: (1) 2.5 µg SB100, 12 µg pT-Caggs-NRas G12V, 12 µg pT3-EF1α-c-Myc; (2) 2.5 µg SB100, 12 µg pT2-shp53, 12 µg pT3-EF1α-c-Myc; (3) 2.5 µg SB100, 12 µg pT-Caggs-NRasG12V, 12 µg pT3-EF1α-c-Myc, 12 µg plentiV2- Cas9-sgUbxn1 (or sgScramble); (4) 2.5 µg SB100, 12 µg pT2-shp53, 12 µg pT3-EF1α-c-Myc, 12 µg plentiV2- Cas9-sgUbxn1 (or sgScramble); (5) 2.5 µg SB100, 12 µg pT-Caggs-NRasG12V, 12 µg pT3-EF1α-c-Myc-IRES-PHB (or Control), 12 µg plentiV2- Cas9-sgUbxn1 (or sgScramble). All mice were sacrificed when they developed abdominal swelling and liver weights and other data were recorded, and the livers were removed for further study. Six-week-old BALB/c nude mice were subcutaneously injected with MHCC97H cells (7 × 106) utilizing shRNA knockdown of UBXN1 or control. Mice were subcutaneously injected with MHCC97H cells (5 × 106) expressing UBXN1 or vector control. Once the tumors were formed, the tumor size of the mice was measured and recorded every two days. After the mice reached a tumor size of 1000 mm3, the mice were sacrificed to remove the tumors for further study. Mice were housed in a SPF condition. All experiments were conducted in accordance with National Institutes of Health guidelines and were agreed by the Institutional Animal Care and Use Committee of Shanghai Jiao Tong University and Shanghai Cancer Institute.
Plasmids and stable cell line construction
V5-tagged UBXN1 (full length), UBXN1 with the deletion of N-terminal UBA domain (ΔUBA: 1-42aa) and the deletion of C-terminal UBX domain (ΔUBX: 205-313aa) were cloned into PLVX-ZSgreen plasmid. The overexpression plasmids (UBXN1-Flag, PHB-His, PHB2-Flag and GFP-UBXN1) were cloned into PLVX-IRES-Puro. The shRNA sequence was inserted into pLKO.1. The UBXN1 gDNA sequence was inserted into LentiV2-CRISPR plasmid. HEK293T cells and packaging plasmids (psPAX2, pMD2.G) were used for lentivirus production. HCC cells were infected with lentivirus adding polybrene. Cells overexpressing the PLVX-ZSgreen plasmid were sorted by flow cytometry. SB100, pT-Caggs-NRas G12V, pT3-EF1α-c-Myc and pT2-shp53 plasmids were from Addgene. pT3-EF1α-c-Myc-IRES-PHB palasmid was self-designed and constructed by homologous recombination using pT3-EF1α-c-Myc as a template. All plasmids injected into the tail vein of mice were extracted using an endotoxin-free plasmid extraction kit (DP117, TIANGEN). All sequences of designed shRNA or guide DNA in this study are listed in Supplementary Table 2.
Western blotting and Co-IP
Western blotting and IP assay were conducted following the protocols [34]. In brief, the cells or tissues were lysed in RIPA lysis buffer (Beyotime, P0013B) with Protease Inhibitor Cocktail (Roche, 05892970001) and Phosphatase Inhibitor (Roche, 04906845001) for 1 h. After 12,000 rpm centrifugation for 20 min at 4 °C, the supernatant was taken. Then, the protein supernatants were separated using SDS-PAGE and transformed onto nitrocellulose filter membranes (Pall Corporation, 66485). After blocking by 5% BSA for 1 h, the membranes were incubated with primary antibodies at 4 ℃ overnight. The blots were measured by laser scanner (Odyssey, Licor, USA). Cells were lysed in western blot & IP lysis buffer (Beyotime, P0013). The lysates were incubated with protein A/G beads (Santa Cruz, sc-2002) and primary antibody for 12 h at 4 °C. The precipitates were collected by centrifugation at 1500 rpm for 5 min at 4 °C and washed three times with the cold 1×TBS. The protein samples were subjected to western blotting analysis.
Mass spectrometry analysis
The protein samples were preprocessed with reference to protocols in the published article [34]. Briefly, cell lysates were immunoprecipitated and separated by SDS-PAGE, then stained with Caumas Brilliant Blue. LC-MS/MS analysis and identification of peptide mixtures were performed by Hangzhou PTMBIO Biotechnology Co.
Quantitative real-time PCR
The qRT-PCR assay referenced the protocols of the published article [34]. The mRNA from the cells were extracted by RNAiso Plus (Takara, 9108) based on the manufacturer’s manual. Then, the mRNA was reverse transcribed into cDNA by Hifair™ III Super Mix plus (yeasen, 11141C). qRT-PCR was conducted using Hieff® qPCR SYBR Green Master Mix (yeasen, 11202ES03) and the Applied Biosystems 7500 (Life Technologies, USA). The gene expression was normalized with GAPDH. All sequences of designed primers in this study are listed in Supplementary Table 3.
RNA-sequencing (RNA-seq) and bioinformatics analysis
RNA from tumor cell lines was extracted with TRIZOL Reagent (Life Technologies, USA) following the manufacturer’s instructions. Briefly, transcriptome library was built by the TruSeq™ RNA preparation kit (Illumina, CA). Paired-end library was conducted by Illumina NovaSeq 6000 system. The reads were mapped to the human reference genome (GRCh38) for human cells, using Hisat2 to produce the BAM files. The expression level of transcripts was calculated according to the per Transcripts Per Million reads (TPM) method. Differentially expressed genes were estimated by limma algorithm. The pathway enrichment analysis, Kyoto Encyclopedia of Genes and Genomes (KEGG), was performed using the Cluster Profiler package through R programming language.
The GSE14520 transcriptome dataset was downloaded from the GEO database. The TCGA-LIHC gene expression dataset was downloaded from the TCGA data portal. The ICGC-LIHC dataset was obtained from the ICGC data portal. The survival and survminer packages were used to generate Kaplan‒Meier plots. The cut-off value for UBXN1 expression was determined by quartile. The tumor samples from the TCGA-LIHC database were classified into two groups based on the transcript levels of UBXN1: the top 25% were classified as the high expression group, and the bottom 25% were classified as the low expression group. GSEA was conducted using GSEA version 4.2.3 of the Broad Institute. A false discovery rate (FDR) < 0.05 and p value < 0.05 were considered to indicate a statistically significant difference. The raw RNA-seq data and detailed experimental design are deposited under GSE247383.
Using the RNA-seq data of MHCC97H cells, the genes regulated by shUBXN1 (shUBXN1 vs. shRFP, fold change ≥ 2 or ≤ 0.5, p value < 0.05.) were identified and subjected to KEGG enrichment analysis via the clusterProfiler package.
MTT and Colony formation assay
Cells were spread on a 96-well plate at a density of 1000 cells per well. At the indicated time points, 15 µL of MTT (0.5 mg/ml) was added to each well, followed by incubation at 37 °C for 3 h. Absorbance values at 493 nm were detected using a Fluorescence Microplate Reader (Infinite TM M200, TECAN). Cells were seeded in 6-well plates at a density of 2000 cells per well. After 10 days, cells were fixed in methanol and stained with crystal violet (Beyotime, C0121).
Flow cytometric analysis
Apoptotic cells were detected by using APC-Annexin V (Biolegend, 640920) and 7-AAD (Biolegend, 640926) Apoptosis Detection Kit. The fluorescent probe MitoTracker Deep Red (50nM) was used to label mitochondria. In addition, mitochondrial membrane potential changes were detected using the JC-1(5µM) Assay Kit. All these experiments were performed according to the corresponding manufacturer’s instructions. At least 5000 live cells were detected by flow cytometric analysis on a FACS flow cytometer (Becton Dickinson, LSRFortessa). Experimental data were analyzed using Flow Jo software v10.2.
Immunohistochemistry
Tumors or liver tissues were fixed and embedded in paraffin. Tissue sections were deparaffinized with xylene and rehydrated in decreasing concentrations of alcohol, followed by immersion in 3% hydrogen peroxide for 10 min at room temperature. Then, the sections were boiled in 0.01 mol/L citrate buffer for 20 min on low heat for antigen extraction. The tissue sections were then cooled to room temperature and blocked with immunostaining blocking solution for 1 h. The tissue sections were then incubated with primary antibody at 4 °C overnight. The sections were rinsed three times with PBS and incubated with secondary antibody at room temperature for 1 h. After rinsing three times with PBS, the sections were stained with hematoxylin for nuclear staining and dehydrated with different concentrations of ethanol.
Immunofluorescence
Cells were fixed with 4% paraformaldehyde for 20 min at room temperature, and permeabilized with 0.1% Triton X-100 for 20 min. Cells were then blocked with immunostaining blocking solution (Beyotime, P0023A) for 1 h and incubated with primary antibody at 4 °C overnight. The cells were washed three times with PBS and incubated with secondary antibody at room temperature for 1 h. After washing three times with PBS, the cells were stained with DAPI for 15 min. The fluorescent probe Mito Tracker Green (50nM) was used to label mitochondria of living cells. The experiment was performed according to the corresponding manufacturer’s instructions. Pictures of cells at 488 nm were taken using a fluorescence microscope (TCS SP8, Leica).
TUNEL assay
Fragmented DNA in cells was labelled using a TUNEL kit with red fluorescence according to the manufacturer’s instructions. In brief, cells were fixed with 4% paraformaldehyde for 20 min at room temperature. The DNA fragments of cells were labelled with dUTP by the terminal transferase reaction. Cell nuclei were stained with DAPI. Pictures of cells or tissues at 550 nm were taken using a fluorescence microscope (TCS SP8, Leica).
Mitochondrial DNA quantification
The mtDNA extraction kit (Abcam, ab65321) was used to extract DNA from the cells according to the corresponding manufacturer’s instructions. The DNA then analyzed by qPCR using primers targeting mtDNA (tRNA-Leu). The mtDNA primers were synthesized to detect tRNA-Leu (Forward: CACCCAAGAACAGGGTTTGT; Reverse: TGGCCATGGGTATGTT GTTA) and GAPDH.
Statistical analysis
All experiments were repeated at least three times and data are expressed as mean ± SD. Statistical analysis was performed using GraphPad Prism7. Paired and unpaired and two-tailed Student’s t-tests were used to compare the data between the two groups. Survival analysis was performed using the log-rank test. Pearson’s test was used to measure correlation between the two groups. *p < 0.05; **p < 0.01, and ***p < 0.001. All results with p values < 0.05 were statistically significant.
Results
UBXN1 is aberrantly upregulated in HCC and associated with poor outcomes
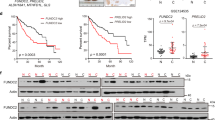
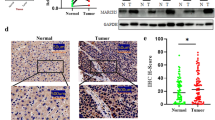
To address the potential involvement of UBXN proteins in the development and progression of HCC, we first analyzed the mRNA expression patterns of UBXN proteins in the tumor and nontumor tissues of HCC patients and found that the transcript level of UBXN1 was consistently and significantly elevated in tumor tissues compared to nontumor tissues from the TCGA-LIHC, ICGC-LIHC and GEO14520 cohorts (Fig. 1A and Supplementary 1A–C). Furthermore, survival analysis revealed that patients with high UBXN1 levels had shorter overall survival than those with low UBXN1 levels (Fig. 1B). To support the in silico results, we assessed the expression profiles of UBXN1 in 21 HCC clinical specimens and observed that UBXN1 protein expression was robustly upregulated in ∼ 61.9% of the tumor tissue samples, as evidenced by the increased protein and mRNA levels in the tumor tissue samples compared with nontumor tissue specimens. (Fig. 1C-D). Additionally, IHC analysis of UBXN1 expression in 80 paired HCC patient samples revealed that the UBXN1 protein was predominantly located in the cytoplasm and highly expressed in HCC cells compared with adjacent cells (Fig. 1E). Next, we measured expression of UBXN1 in mouse liver tumors by using two set of oncogenic-driven primary liver cancer model, which were generated by the SB transposon delivery of the plasmids expressing NRas G12V (NRas)/c-Myc and c-Myc/shp53, and developed HCC approximately 2 or 5 weeks post-HDTVi, respectively (Supplementary 1D). After confirming the efficiency of NRas- and c-Myc-overexpression as well as p53-knockdown (Supplementary 1E), we found that the protein abundance of UBXN1 was aberrantly increased in mouse liver tumors induced by NRas G12V/c-Myc or c-Myc/shp53 (Fig. 1F–G). Collectively, these results suggest that UBXN1 may contribute to the development and progression of HCC.
UBXN1 expression is upregulated in HCC and correlated with poor clinical prognosis. (A) The expression level of UBXN1 (FPKM) in HCC (T) and nontumor (NT) tissues from TCGA-LIHC (NT = 50, T = 374), ICGC-LIHC (NT = 202, T = 243) or GSE14520 (NT = 241, T = 247) databases. (B) Kaplan–Meier survival curves showing overall survival of HCC patients stratified by UBXN1 mRNA expression level (TCGA-LIHC (n = 364), ICGC-LIHC (n = 243) or GSE14520 (n = 242). (C) Western blotting analysis of UBXN1 in 21 paired clinical HCC tissues (T) and adjacent normal tissues (NT). The quantitative measurement was conducted to analyze grayscale value (right, UBXN1/Tubulin, paired Mann Whitney test). (D) qRT-PCR analysis of UBXN1 in 21 paired clinical HCC tissues (T) and adjacent normal tissues (NT). (E) Representative IHC images (left) and IHC score (right) of UBXN1 expression in paired clinical HCC tissues (T) and adjacent normal tissues (NT), n = 80. Scale bar: 100 μm. (F-G) The C57BL/6 mice were hydrodynamically injected with indicated plasmids (n = 5 or 6). IHC images (F, left) and IHC score (F, right) of mouse livers at 4 or 7 weeks post hydrodynamic injection. Scale bar: 100 μm. Western blotting analysis of UBXN1 expression in liver tissues (G). The data are presented as mean ± SD. **, p < 0.01; ***, p < 0.001; by two-tailed Student t test
UBXN1 promotes oncogene-driven liver tumorigenesis
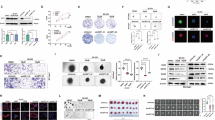
To explore the requirement of UBXN1 for liver tumorigenesis, we transduced CRISPR/Cas9 vectors targeting Ubxn1 together with transposon plasmids expressing NRas/c-Myc or c-Myc/shp53 into C57BL/6 mice by HDTVi (Fig. 2A–H). The knockout efficiency of Ubxn1 with the designed sgRNA was confirmed prior to the experiments (Supplementary 2A). As expected, knockout of Ubxn1 significantly inhibited tumor development in these two sets of autochthonous liver tumor models, as evidenced by a decreased liver/weight ratio and increased survival in the Ubxn1-KO (sgUbxn1) group compared with those in the control (sgCon) group (Fig. 2B–D and F–H). Consistent with these findings, xenograft experiments revealed substantial decreases in the volume and weight of UBXN1-knockdown human HCC tumors compared with control tumors (Fig. 2I–K). Notably, UBXN1 overexpression had the opposite effect on the growth of HCC tumors (Fig. 2L–N). Next, we investigated whether UBXN1 regulates tumor proliferation in vitro. The results showed that overexpression of UBXN1 strongly promoted the proliferation potential of tumor cells cultured under serum starvation conditions, and that knockdown of UBXN1 suppressed the proliferation of cultured PLC/PRF/5, MHCC97 H and LPC-HRas cells (Fig. 2O-P). A colony formation assay revealed similar results: knockdown of UBXN1 inhibited clonogenicity, whereas ectopic UBXN1 expression increased clonogenicity in cells cultured in serum-insufficient conditions (Supplementary 2B-C), suggesting that UBXN1 is required for liver tumor growth.
UBXN1 promotes HCC cell proliferation and tumorigenicity. (A–D) The C57BL/6 mice were hydrodynamic co-injected transposon-based vectors expressing NRas G12V, c-Myc, along with vector expressing Cas9 and a CRISPR guide RNA targeting UBXN1 or scramble RNA (n = 6). Schematic representation of the HDTVi (A), morphology of livers (B), ratio of liver/body weight (C) or Kaplan-Meier overall survival curves (D) were shown. (E–H) The indicated vectors were hydrodynamic injected into C57BL/6 mice (n = 5). Schematic of vectors (E), morphology (F), ratio of liver/body weight (G) or Kaplan-Meier overall survival curves (H) were shown. (I–K) BALB/C nude mice were subcutaneously implanted with 7 × 106 UBXN1-knockdown or control MHCC97H cells (n = 6). Tumors were harvested at day 28 post implantation. Representative pictures (I), tumor volumes (J) and tumor mass (K) were shown. (L–N) BALB/C nude mice were subcutaneously implanted with 5 × 106 UBXN1-ovexpressed and control MHCC97H cells (n = 5). Tumors were harvested at day 35 post implantation. Representative pictures (L), tumor volumes (M) and tumor mass (N) were shown. (O) The indicated cells were cultured with medium containing 0.1% FBS with starvation for one day or two, and then switched to complete medium for the following days. The growth capacity was tested by MTT assay. (P) MTT assay was used for determine the growth capacity of indicated cells. The data are presented as mean ± SD. **, p < 0.01; ***, p < 0.001 by two-tailed Student t test
UBXN1 promotes tumor cell survival and growth by inhibiting cell apoptosis
We next performed GSEA of the genes differentially expressed in TCGA-LIHC tumors with a high abundance of UBXN1 transcripts (UBXN1high) compared with those in the UBXN1low group and revealed a significant correlation between the mRNA levels of UBXN1 and genes involved in the regulation of apoptosis (Fig. 3A). We further used a TUNEL assay to examine tumor cell apoptosis in human HCC xenografts, and the results showed that UBXN1-knockdown greatly increased the number of TUNEL+ cells (Fig. 3B). Additionally, the number of TUNEL+ apoptotic cells was increased in cultured PLC/PRF/5 and MHCC97H cells upon UBXN1-knockdown (Fig. 3C). FACS analysis also revealed that the percentages of apoptotic cells resulting from UBXN1-knockdown were robustly greater than those of control cells (PLC/PRF/5, MHCC97H, and LPC-HRas, Fig. 3D). These results were further corroborated by increased levels of BAX and cleaved caspase-3 and decreased expression of Bcl-xL in UBXN1-knockdown HCC cells (Fig. 3E). Furthermore, the induction of cell apoptosis by UBXN1-knockdown was efficiently reversed by the pan-caspase inhibitor Z-VAD (Fig. 3F). Notably, UBXN1 overexpression markedly attenuated tumor cell apoptosis induced by serum starvation (Fig. 3G). These results suggest that UBXN1 promotes HCC cell survival and tumor growth by inhibiting caspase-mediated cellular apoptosis.
Inhibition of UBXN1 induces HCC cell apoptosis. (A) GSEA of TCGA-LIHC dataset with UBXN1 transcript levels and REACTOME _REGULATION_OF_APOPTOSIS signature. (B) UBXN1 staining (left) and TUNEL staining (right) of indicated samples derived from MHCC97H xenograft tumor. Scale bar: 100 μm. (C) IF staining of TUNEL in indicated PLC/PRF/5 and MHCC97H cells. Scale bar: 100 μm. (D) Flow cytometry analysis of apoptotic ratio in PLC/PRF/5, MHCC97H and LPC-HRas cells. (E) Western blotting analysis of indicated proteins in UBXN1-knockdown PLC/PRF/5 and MHCC97H cells. (F) Flow cytometry analysis of apoptosis ratio in indicated cells treated with or without Z-VAD-FMK (20µM) for 48 h. (G) Flow cytometry analysis of apoptosis ratio in indicated PLC/PRF/5 and MHCC97H cells coculturing with or without medium containing 0.1%FBS for 48 h serum starvation. The data are presented as mean ± SD. ***, p < 0.001 by two-tailed Student t test
UBXN1 alleviates mitochondrial damage in HCC cells
To investigate the detailed mechanisms underlying the pro-survival effects of UBXN1, we conducted Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis of the differentially expressed genes (DEGs) in UBXN1-knockdown MHCC97H cells compared with control cells and found that the DEGs were notably enriched in signaling pathways related to mitochondrial homeostasis and NF-κB signaling (Fig. 4A). Accordingly, GSEA of the transcriptional profiles of the TCGA-LIHC cohort revealed that the mitophagy-related signatures were upregulated in the UBXN1high tumor group (Fig. 4B). We also found a strong correlation between the mitophagy signature and the apoptosis signature in liver tumors from the TCGA-LIHC cohort (Supplementary 3 A-B). Next, we observed mitochondrial phenotypes and found that the knockdown of UBXN1 resulted in a significant increase in the number of HCC cells with abnormal mitochondrial aggregation (Fig. 4C). The MitoTracker Deep Red probe was then used to label intact mitochondria, and FACS analysis revealed that damaged mitochondria were robustly more abundant in UBXN1-knockdown cells than in control cells (Fig. 4D). In addition, a noticeable reduction in mtDNA copy number, accompanied by severe loss of membrane potential, was observed in PLC/PRF/5 and MHCC97H cells with UBXN1-knockdown (Fig. 4E-F). In contrast, upon treatment with CCCP, which blocks electron flow from PS II to PS I, ectopic expression of UBXN1 effectively inhibited mitochondrial damage, membrane potential reduction and cellular apoptosis in PLC/PRF/5 and MHCC97H cells (Fig. 4G-H, Supplementary 3 C-D), suggesting that UBXN1 is required for the maintenance of mitochondrial integrity under stress conditions. Next, to explore whether NF-κB signaling pathway was involved in UBXN1-regulated mitochondrial homeostasis, we first incubated HCC cells with CCCP, which induced mitochondria damage within 24 h in HCC cells (Fig. 4G), and found that the expression of active form of NF-κB (p65), p-p65, was comparable in HCC cells treated with CCCP for 0–24 h (supplementary Fig. 3E). Of note, UBXN1-knockdown was not capable to induce the expression of p-p65 in PLC/PRF/5 and LPC-HRas cells (supplementary Fig. 3F). These results therefore imply that UBXN1 inhibits the damage and loss of mitochondria in HCC cells independently of regulating NF-κB signaling pathway.
Inhibiting UBXN1 promotes mitochondrial damage in HCC cells. (A) KEGG enrichment analysis of UBXN1-upregulated genes in MHCC97H cells using compilation C5 (MSigDB). Upregulated genes were determined with the criteria: p<0.05 and log2 (fold change) ≥ 1. (B) GSEA of TCGA-LIHC dataset with UBXN1 transcript levels and REACTOME_MITOPHAGY signature. (C) Representative pictures of mitochondrial morphology stained with MitoTracker Green in indicated PLC/PRF/5 and MHCC97H cells. Scale bar: 100 μm. (D) The indicated cells were stained with MitoTracker Deep Red (50 nM) for 15 min. The percentage of damaged mitochondria in HCC cells was measured by FACS. (E) mtDNA copy numbers in PLC/PRF/5 and MHCC97H cells were analyzed by qPCR. (F) The indicated cells were stained with JC-1 (5µM) for 15 min. The analysis of mitochondrial membrane potential loss was measured by FACS. (G) The indicated cells were treated with CCCP (20µM) for 12–24 h and stained with MitoTracker Deep Red (50 nM) for 15 min. The percentage of damaged mitochondria in HCC cells was measured by FACS. (H) mtDNA copy numbers analyzed by qPCR with tRNA-Leu to GAPDH in PLC/PRF/5 and MHCC97H cells with or without UBXN1 overexpression, CCCP (20µM) for 12–24 h. The data are presented as mean ± SD. *, p < 0.05; **, p < 0.01; ***, p < 0.001 by two-tailed Student t test
UBXN1 interacts with PHB and regulates its expression
To investigate proteins involved in UBXN1-regulated mitochondrial homeostasis, we coimmunoprecipitated UBXN1 from human MHCC97H cells and mouse Hepa1-6 cells and performed mass spectrometry analysis to screen for UBXN1-interacting proteins. PHB and PHB2, which are present in the inner mitochondrial membrane, were identified as candidates (Fig. 5A). We further explored the subcellular localization of UBXN1 by transfecting HCC cells with a plasmid encoding the UBXN1-GFP fusion protein and observed that UBXN1 was mainly located and dispersed in the cytoplasm of cells cultured under normal conditions (Fig. 5B). Notably, the localization patterns of the UBXN1 protein changed, and the UBXN1 protein became more focally aggregated and colocalized with TOM20, a mitochondrial marker, in cells treated with CCCP (Fig. 5B), emphasizing the involvement of UBXN1 in the regulation of mitochondrial function. Then, we co-expressed UBXN1 with PHB or PHB2 in HEK293T cells and found that UBXN1 interacted with both PHB and PHB2 (Fig. 5C, Supplementary 4 A). Consistently, we also confirmed this interaction in the PLC/PRF/5 and MHCC97H HCC cell lines (Fig. 5D). Remarkably, we found that the level of PHB, but not the PHB2 protein, was decreased in UBXN1-knockdown HCC cells vs. control cells (Fig. 5E). Notably, knockdown of UBXN1 had little effect on the mRNA expression of both PHB and PHB2 (Fig. 5F). In addition, increased levels of PHB protein were detected in UBXN1-overexpressing cells under serum starvation conditions (Fig. 5G).
UBXN1 interacts with PHB and maintains its expression. (A) Venn diagram illustrated UBXN1 interactors (overlap based on IP-MS of MHCC97H and Hepa1-6, unique peptides > 2) with mitophagy regulator (GOBP AUTOPHAGY OF MITOCHONDRION). (B) GFP-UBXN1-expressed MHCC97H cells were treated with or without CCCP for 2 h MHCC97H cells. Representative pictures of IF staining of indicated cells was shown. (C) HEK293T cells were transient transfected with PHB-His together with UBXN1-Flag or control plasmid. Total lysates were immunoprecipitated with Flag M2 affinity gel after 48 h and the indicated proteins were detected by western blotting. (D) Total lysates from indicated PLC/PRF5 and MHCC97H cells were immunoprecipitated with Flag M2 affinity gel and then indicated proteins were detected by western blotting. (E) Western blotting analysis of indicted proteins in MHCC97H and PLC/PRF5 cells transduced with lentivirus UBXN1 shRNA or shRNA-control plasmid. (F) qRT-PCR analysis of PHB and PHB2 in MHCC97H and PLC/PRF5 cells transduced with lentivirus UBXN1 shRNA or shRNA-control plasmid. (G) Indicated PLC/PRF/5 and MHCC97H cells coculturing with or without medium containing 0.1% FBS for 24 h serum starvation. The expression of indicated proteins was measured by western blotting. (H) HEK293T cells were transient transfected with PHB-His together with UBXN1-V5, UBXN1-ΔUBA-V5, UBXN1-ΔUBX-V5 or control plasmid. Total lysates were immunoprecipitated with His affinity gel after 48 h and the indicated proteins were detected by western blotting. (I) Endogenous UBXN1 was knocked down by transfecting lentiviral shRNA targeting the 3′UTR of UBXN1 in UBXN1-, UBXN1-ΔUBA- and UBXN1-ΔUBX-overexpressed MHCC97H cells. The indicated proteins were detected by western blotting. (J) Flow cytometry analysis of apoptotic ratio in indicated MHCC97H cells. The data are presented as mean ± SD. **, p < 0.01; ***, p < 0.001 by two-tailed Student t test
To explore the functional domain of UBXN1 in regulating PHB expression, we generated two truncated mutants of UBXN1, the ΔUBA mutant (with a 1–42 aa deletion) and the ΔUBX mutant (with a 205–313 aa deletion), to measure the ability of these mutants to bind PHB. We found that the UBX domain is essential for this interaction (Fig. 5H). Functionally, the overexpression of wild-type UBXN1 or the ΔUBA mutant, but not the ΔUBX mutant, robustly increased PHB expression and inhibited the cellular apoptosis induced by UBXN1-knockdown (Fig. 5I-J). In summary, these data suggest that a functional interaction between PHB and UBXN1 regulates tumor cell survival.
UBXN1 function in promoting mitochondrial homeostasis in tumors depends on PHB
We next investigated the functional requirement of the PHB-UBXN1 interaction in liver tumorigenesis. We found that knockdown of PHB induced cell arrest, apoptosis, and severe mitochondrial damage (Supplementary 5 A–D), which reflects the importance of the PHB-UBXN1 interaction in maintaining mitochondrial homeostasis. Accordingly, ectopic expression of PHB significantly attenuated the cell proliferation arrest induced by UBXN1-knockdown (Fig. 6A-B, Supplementary 5E) and alleviated mitochondrial damage and the induction of cell apoptosis (Fig. 6C-D). More importantly, we integrated a CRISPR/Cas9 deletion of Ubxn1 into an NRas/c-Myc-driven liver tumor model with or without PHB overexpression (Fig. 6E). Although Ubxn1 knockout effectively attenuated NRas/c-Myc-induced liver tumor growth (sg Ubxn1/NRas/c-Myc-IRES-Con vs. sg Con/NRas/c-Myc-IRES-Con), PHB expression significantly promoted liver tumorigenesis in the Ubxn1-knockout groups of mice (sg Ubxn1/NRas/c-Myc-IRES-PHB vs. sg Ubxn1/NRas/c-Myc-IRES-Con, Fig. 6F–H). Collectively, these results indicate that PHB is required for the UBXN1-mediated maintenance of mitochondrial homeostasis and liver tumorigenesis.
PHB counteracts UBXN1-knockdown-mediated HCC cell apoptosis and mitochondrial damage. (A–B) MTT assay (A) and colony formation assay (B) were used for measured proliferation of indicated cells. (C–D) Percentage of apoptotic cells (C) and damaged mitochondria (D) were measured by FACS. (E–H) The indicated vectors were hydrodynamic injected into C57BL/6 mice (n = 5). Schematic of vectors (E), morphology (F), ratio of liver/body weight (G) or Kaplan-Meier overall survival curves (H) were shown. Student’s t-test, ***p < 0.001
Discussion
Tumor cells are adept at achieving a survival advantage by maintaining mitochondrial homeostasis upon exposure to endogenous or exogenous stresses [35,36,37]. The detailed mechanism by which mitochondrial homeostasis is maintained in tumor cells, however, remains elusive. Here, we identified UBXN1 as a tumor promoter in HCC via the maintenance of mitochondrial integrity. The expression of UBXN1 is aberrantly upregulated in human HCCs and mouse liver tumors, and high UBXN1 expression is associated with poor overall survival in patients with HCC. Mechanistically, the interaction between UBXN1 and PHB contributes to mitochondrial integrity, thereby promoting HCC cell survival and liver tumorigenesis.
UBXN1 is aberrantly expressed in several types of cancer and is implicated in the regulation of multiple biological processes through interactions with the multifunctional AAA-ATPase p97/VCP protein or ubiquitinated proteins via its UBX or UBA domain [38, 39]. Recent studies have demonstrated that the expression of UBXN1 is heterogeneous across tumors but generally decreased in gliomas [30]. Our results showed that the expression levels of UBXN1 are generally increased in the HCC samples examined. Interestingly, some of these samples exhibited marginal alterations or even decreased expression of UBXN1. Since the intertumor (tumor by tumor) heterogeneity in genetic background or biological context substantially exists in HCC [40], the compensatory mechanisms consisting of overlapping molecular events or alteration of the surrogates may be present and account for this divergence in UBXN1 expression. Nevertheless, our findings are the first to reveal that UBXN1 is highly expressed in HCC tissues and mouse liver tumors and contributes to HCC growth and malignant progression.
UBXN1 has diverse and sometimes opposite functions in regulating tumor progression due to differences in its substrates and cancer types [29,30,31]. YTHDF2 and PRADX decrease UBXN1 expression at the posttranscriptional level and thereby facilitate NF-κB signaling activity in glioblastoma and colon adenocarcinoma progression [30, 31]. In contrast, UBXN1 promotes AKT signaling and cell proliferation, migration and invasion in prostate cancer cells [29]. Our RNA-seq data revealed an increase in NF-κB signaling in UBXN1-knockdown MHCC97H cells, but silencing UBXN1 did not promote the expression of the active form of NF-κB in human PLC/PRF/5 or murine LPC-HRas HCC cells. Furthermore, mitochondrial damage induced by CCCP could not be sufficient to rapidly activate NF-κB signaling. Therefore, the promotion of liver tumorigenesis by UBXN1 is mainly due to the regulation of mitochondrial homeostasis but not NF-κB signaling.
Maintenance of mitochondrial homeostasis depends on mitochondrial quality control mechanisms that control mitochondrial integrity and function [41, 42]. A recent study showed that the inhibition of UBXN1 attenuates PINK1-mediated mitophagy via the recruitment of MFN2 to damaged mitochondria in cells treated with CCCP [27], emphasizing the role of UBXN1 in the clearance of damaged mitochondria. Similar to the findings of the present study, knockdown of UBXN1 triggered mitochondrial damage, resulting in excessive accumulation of dysfunctional mitochondria. Notably, we revealed that UBXN1 interacts with the mitochondrial inner membrane protein PHB and maintains its expression, which is required for mitochondrial membrane integrity. PHB and PHB2 are evolutionarily conserved scaffold proteins that form a multimeric ring complex at the inner mitochondrial membrane that acts as a protein and lipid scaffold to ensure the function of the inner mitochondrial membrane [43,44,45]. Consistent with our observation, PHB deficiency disrupts the integrity of the mitochondrial membrane and triggers NOXA-mediated apoptosis in multiple myeloma cells [22, 46]. Intriguingly, pharmacologically inhibiting PHB induces mitochondrial-dependent apoptosis and decreases the progression and development of gastric tumors [23, 47]. These findings suggest that PHB plays a prominent role in regulating UBXN1-mediated mitochondrial quality control and that PHB is a tumor-promoting factor in certain types of cancers.
Taken together, our findings revealed that UBXN1 hyperactivation frequently occurs and is crucial for liver tumorigenesis and that the tumor-promoting function of UBXN1 occurs through the UBXN1-PHB interaction, which regulates mitochondrial homeostasis.
Data availability
The RNA-seq data were deposited in the NCBI Gene Expression Omnibus under GEO: 247383.
Abbreviations
- HCC:
-
Hepatocellular carcinoma
- PHB:
-
Prohibitin
- mtDNA:
-
mitochondrial DNA
- ROS:
-
reactive oxygen species
- UBX:
-
ubiquitin regulatory X
- UBA:
-
ubiquitin-associated domain
- CCCP:
-
Carbonyl Cyanide3-ChloroPhenylhydrazone
- qRT-PCR:
-
quantitative Real-Time Polymerase Chain Reaction
- LC-MS/MS:
-
Liquid chromatography-tandem mass spectrometry
- GO:
-
Gene Ontology
- GSEA:
-
Gene Set Enrichment Analysis
- LIHC:
-
Liver hepatocellular carcinoma
- HDTVi:
-
hydrodynamic tail-vein injection
- SB:
-
Sleeping Beauty
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- Co-IP:
-
Coimmunoprecipitation
References
Llovet JM, Kelley RK, Villanueva A, Singal AG, Pikarsky E, Roayaie S, et al. Hepatocellular carcinoma. Nat Reviews Disease Primers. 2021;7(1):6.
Vogel A, Meyer T, Sapisochin G, Salem R, Saborowski A. Hepatocellular carcinoma. Lancet (London England). 2022;400(10360):1345–62.
Chan DC. Mitochondrial dynamics and its involvement in Disease. Annu Rev Pathol. 2020;15:235–59.
Zong WX, Rabinowitz JD, White E. Mitochondria and Cancer. Mol Cell. 2016;61(5):667–76.
Chen Y, Chen HN, Wang K, Zhang L, Huang Z, Liu J, et al. Ketoconazole exacerbates mitophagy to induce apoptosis by downregulating cyclooxygenase-2 in hepatocellular carcinoma. J Hepatol. 2019;70(1):66–77.
Wang Q, Yu P, Liu C, He X, Wang G. Mitochondrial fragmentation in liver cancer: emerging player and promising therapeutic opportunities. Cancer Lett. 2022;549:215912.
Sarwar A, Zhu M, Su Q, Zhu Z, Yang T, Chen Y, et al. Targeting mitochondrial dysfunctions in pancreatic cancer evokes new therapeutic opportunities. Crit Rev Oncol Hematol. 2022;180:103858.
Zhang Y, Yang B, Tu C, Ping Y, Chen S, Wu T, et al. Mitochondrial impairment and downregulation of Drp1 phosphorylation underlie the antiproliferative and proapoptotic effects of alantolactone on oral squamous cell carcinoma cells. J Transl Med. 2023;21(1):328.
Bryant KL, Stalnecker CA, Zeitouni D, Klomp JE, Peng S, Tikunov AP, et al. Combination of ERK and autophagy inhibition as a treatment approach for pancreatic cancer. Nat Med. 2019;25(4):628–40.
Crowell PD, Giafaglione JM, Jones AE, Nunley NM, Hashimoto T, Delcourt AML, et al. MYC is a regulator of androgen receptor inhibition-induced metabolic requirements in prostate cancer. Cell Rep. 2023;42(10):113221.
Liu Y, Zhang H, Liu Y, Zhang S, Su P, Wang L, et al. Hypoxia-induced GPCPD1 depalmitoylation triggers mitophagy via regulating PRKN-mediated ubiquitination of VDAC1. Autophagy. 2023;19(9):2443–63.
Giordano F, Paolì A, Forastiero M, Marsico S, De Amicis F, Marrelli M, et al. Valproic acid inhibits cell growth in both MCF-7 and MDA-MB231 cells by triggering different responses in a cell type-specific manner. J Transl Med. 2023;21(1):165.
Han Q, Yan P, Song R, Liu F, Tian Q. HOXC13-driven TIMM13 overexpression promotes osteosarcoma cell growth. Cell Death Dis. 2023;14(7):398.
Liu H, Fan H, He P, Zhuang H, Liu X, Chen M, et al. Prohibitin 1 regulates mtDNA release and downstream inflammatory responses. Embo j. 2022;41(24):e111173.
Chen X, Liu Q, Chen Y, Wang L, Yang R, Zhang W, et al. Carboxylesterase 2 induces mitochondrial dysfunction via disrupting lipid homeostasis in oral squamous cell carcinoma. Mol Metab. 2022;65:101600.
Ploeger C, Huth T, Sugiyanto RN, Pusch S, Goeppert B, Singer S, et al. Prohibitin, STAT3 and SH2D4A physically and functionally interact in tumor cell mitochondria. Cell Death Dis. 2020;11(11):1023.
Canevari RA, Marchi FA, Domingues MA, de Andrade VP, Caldeira JR, Verjovski-Almeida S, et al. Identification of novel biomarkers associated with poor patient outcomes in invasive breast carcinoma. Tumour Biol. 2016;37(10):13855–70.
Wang W, Xu L, Yang Y, Dong L, Zhao B, Lu J, et al. A novel prognostic marker and immunogenic membrane antigen: prohibitin (PHB) in pancreatic cancer. Clin Transl Gastroenterol. 2018;9(9):178.
Kang X, Zhang L, Sun J, Ni Z, Ma Y, Chen X, et al. Prohibitin: a potential biomarker for tissue-based detection of gastric cancer. J Gastroenterol. 2008;43(8):618–25.
Jiang L, Dong P, Zhang Z, Li C, Li Y, Liao Y, et al. Akt phosphorylates Prohibitin 1 to mediate its mitochondrial localization and promote proliferation of bladder cancer cells. Cell Death Dis. 2015;6(2):e1660.
Chiu CF, Ho MY, Peng JM, Hung SW, Lee WH, Liang CM, et al. Raf activation by Ras and promotion of cellular metastasis require phosphorylation of prohibitin in the raft domain of the plasma membrane. Oncogene. 2013;32(6):777–87.
Núñez-Vázquez S, Sánchez-Vera I, Saura-Esteller J, Cosialls AM, Noisier AFM, Albericio F, et al. NOXA upregulation by the prohibitin-binding compound fluorizoline is transcriptionally regulated by integrated stress response-induced ATF3 and ATF4. Febs j. 2021;288(4):1271–85.
Moncunill-Massaguer C, Saura-Esteller J, Pérez-Perarnau A, Palmeri CM, Núñez-Vázquez S, Cosialls AM, et al. A novel prohibitin-binding compound induces the mitochondrial apoptotic pathway through NOXA and BIM upregulation. Oncotarget. 2015;6(39):41750–65.
Sonneville R, Moreno SP, Knebel A, Johnson C, Hastie CJ, Gartner A, et al. CUL-2(LRR-1) and UBXN-3 drive replisome disassembly during DNA replication termination and mitosis. Nat Cell Biol. 2017;19(5):468–79.
Wang P, Yang L, Cheng G, Yang G, Xu Z, You F, et al. UBXN1 interferes with Rig-I-like receptor-mediated antiviral immune response by targeting MAVS. Cell Rep. 2013;3(4):1057–70.
Wang Y-B, Tan B, Mu R, Chang Y, Wu M, Tu H-Q, et al. Ubiquitin-associated domain-containing ubiquitin regulatory X (UBX) protein UBXN1 is a negative regulator of nuclear factor κB (NF-κB) signaling. J Biol Chem. 2015;290(16):10395–405.
Mengus C, Neutzner M, Bento ACPF, Bippes CC, Kohler C, Decembrini S, et al. VCP/p97 cofactor UBXN1/SAKS1 regulates mitophagy by modulating MFN2 removal from mitochondria. Autophagy. 2022;18(1):171–90.
Zhang L, Jiao K, Liu Y, Xu G, Yang Z, Xiang L et al. UBXN9 inhibits the RNA exosome function to promote T cell control of liver tumorigenesis. Hepatology. 2023.
Oh JJ, Ho JN, Byun SS. ARRDC4 and UBXN1: Novel Target genes correlated with prostate Cancer Gleason Score. Cancers (Basel). 2021;13:20.
Li Y, Liu X, Cui X, Tan Y, Wang Q, Wang Y, et al. LncRNA PRADX-mediated recruitment of PRC2/DDX5 complex suppresses UBXN1 expression and activates NF-κB activity, promoting tumorigenesis. Theranostics. 2021;11(9):4516–30.
Chai RC, Chang YZ, Chang X, Pang B, An SY, Zhang KN, et al. YTHDF2 facilitates UBXN1 mRNA decay by recognizing METTL3-mediated m(6)a modification to activate NF-κB and promote the malignant progression of glioma. J Hematol Oncol. 2021;14(1):109.
Wu-Baer F, Ludwig T, Baer R. The UBXN1 protein associates with autoubiquitinated forms of the BRCA1 tumor suppressor and inhibits its enzymatic function. Mol Cell Biol. 2010;30(11):2787–98.
Zhang L, Yang Z, Ma A, Qu Y, Xia S, Xu D, et al. Growth arrest and DNA damage 45G down-regulation contributes to Janus kinase/signal transducer and activator of transcription 3 activation and cellular senescence evasion in hepatocellular carcinoma. Hepatology. 2014;59(1):178–89.
Zhang L, Liu Y, Wang B, Xu G, Yang Z, Tang M, et al. POH1 deubiquitinates pro-interleukin-1β and restricts inflammasome activity. Nat Commun. 2018;9(1):4225.
Hu Q, Xu J, Wang L, Yuan Y, Luo R, Gan M et al. SUCLG2 regulates mitochondrial dysfunction through Succinylation in Lung Adenocarcinoma. Adv Sci (Weinh). 2023:e2303535.
Ng MYW, Wai T, Simonsen A. Quality control of the mitochondrion. Dev Cell. 2021;56(7):881–905.
Ma Z, Han H, Zhao Y. Mitochondrial dysfunction-targeted nanosystems for precise tumor therapeutics. Biomaterials. 2023;293:121947.
Rezvani KUBXD, Proteins. A family of proteins with diverse functions in Cancer. Int J Mol Sci. 2016;17(10).
LaLonde DP, Bretscher A. The UBX protein SAKS1 negatively regulates endoplasmic reticulum-associated degradation and p97-dependent degradation. J Biol Chem. 2011;286(6):4892–901.
Craig AJ, von Felden J, Garcia-Lezana T, Sarcognato S, Villanueva A. Tumour evolution in hepatocellular carcinoma. Nat Rev Gastroenterol Hepatol. 2020;17(3):139–52.
Ruan L, Wang Y, Zhang X, Tomaszewski A, McNamara JT, Li R. Mitochondria-Associated Proteostasis. Annu Rev Biophys. 2020;49:41–67.
Zhang J, Qiao W, Luo Y. Mitochondrial quality control proteases and their modulation for cancer therapy. Med Res Rev. 2023;43(2):399–436.
Hernando-Rodríguez B, Artal-Sanz M. Mitochondrial Quality Control mechanisms and the PHB (Prohibitin) Complex. Cells. 2018;7(12).
Xia L, Liu Y, Zhang S, Yang Y, Zhou Z, Tu J. Can Prohibitin 1 be a safeguard against liver disease? Ann Hepatol. 2019;18(6):790–5.
Fernandez-Abascal J, Artal-Sanz M. Prohibitins in neurodegeneration and mitochondrial homeostasis. Front Aging. 2022;3:1043300.
Ross JA, Robles-Escajeda E, Oaxaca DM, Padilla DL, Kirken RA. The prohibitin protein complex promotes mitochondrial stabilization and cell survival in hematologic malignancies. Oncotarget. 2017;8(39):65445–56.
Tao L, Yin Z, Ni T, Chu Z, Hao S, Wang Z, et al. The Ethyl acetate Extract from Celastrus orbiculatus promotes apoptosis of gastric Cancer cells through Mitochondria Regulation by PHB. Front Pharmacol. 2021;12:635467.
Acknowledgements
We appreciate all the participants who provide supports for the study.
Funding
This work was supported by grants from National Natural Science Foundation of China (No. 82273002, No. 82273005, No. 81972579 and No. 81902373), Shanghai Rising-Star Program (21QA1408400), Natural Science Foundation of Shanghai (20ZR1454100), the Grants from State Key Laboratory of Systems Medicine for Cancer (No. zz-94-2304, No. zz-RCPY-23-05, No. JY2193 and No. JY2224) and Innovative Research Team of High-level Local Universities in Shanghai.
Author information
Authors and Affiliations
Contributions
All authors have approved the final version and agreed to publish this paper. YZ.L., K.J. and L.Z. defined the project and designed the experiments. K.J., G.X. and L.Z. carried out most of the experiments and analyzed data. Y.L., Z.Y., L.X., Z.C. C.X., Y.Z., Z.W., W.X. and N.Z. participated in experiment conduction. YZ.L., K.J. and L.Z. wrote and edited the paper.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All experiments on mice were conducted through guidelines approved by the Animal Protection Committee of Shanghai Jiao Tong University. The use of human tissues was authorized by the Ethics Committee of Renji Hospital, and this study was conducted following ethical guidelines (Declaration of Helsinki).
Consent for publication
This manuscript is approved by all authors for publication.
Conflict of interests
We declare that we have no potential financial conflict of interest related to this manuscript. None of the material in this paper has been published or is under consideration for publication elsewhere.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Jiao, K., Xu, G., Liu, Y. et al. UBXN1 promotes liver tumorigenesis by regulating mitochondrial homeostasis. J Transl Med 22, 485 (2024). https://doi.org/10.1186/s12967-024-05208-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12967-024-05208-5