Abstract
Candidiasis is the most incessant fungal disease affecting human population worldwide. In this research work, we explored the characteristics contributing to pathogenicity of Candida infections, from serious bloodstream conditions like candidemia. Candida spp. were isolated from the blood of 120 suspected patients and assessed their susceptibility to antifungal treatments via MIC measurements. Enzyme activities (phospholipase, proteinase, hemolysin, and esterase) were examined alongside biofilm formation, with surface analysis conducted via SEM. Statistical analysis, facilitated by GraphPad Prism 8.4.2, identified correlations among data points. Results reveal that non-albicans spp. like Candida auris heightened resistance to antifungal treatments compared to Candida albicans. Candida tropicalis exhibited no phospholipase activity but shows activity to proteinase and esterase. Conversely, C. albicans show a case of elevated hemolysin activity. Non-albicans Candida i.e. C. tropicalis and Candida krusei were isolated as high biofilm formers in comparison to C. albicans. Statistical analysis yielded a highly significant p-value (< 0.001), affirming the robustness of our results. In conclusion, non-albicans species exhibit a broader spectrum of resistance mechanisms compared to C. albicans in pathogenicity. This investigation sheds light on diverse virulence strategies employed by different Candida species, informing future treatment approaches.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Candida species are omnipresent yeasts that colonize the skin, oral, gastrointestinal, and reproductive systems asymptomatically and are significantly found in the human population [1]. Genus Candida is made up of a diverse group of microorganisms, with over 17 distinct Candida species to cause infection in humans. Of these species, opportunistic infections are most frequently caused by C. albicans. Furthermore, drug-resistant strains of Candida glabrata, along with the emerging global cases of C. auris, alongside other species such as Candida parapsilosis, C. tropicalis, and C. krusei, collectively pose significant public health challenges [2, 3]. The WHO recognizes their importance and has categorized them into different priority groups based on their impact. The WHO inaugural fungal priority pathogens list categorizes species into specific priority groups: C. auris and C. albicans are classified under the critical priority group, while C. krusei falls under medium priority group. The high priority group includes C. glabrata, C. parapsilosis and C. tropicalis. Several virulence characteristics of Candida species allow them to successfully spread illness and evade the effects of antifungal host defenses and pharmaceutical interventions. These characteristics help them to flourish in a variety of conditions inside the human body, which can result in a variety of illnesses [4]. Comprehending these pathways is essential for formulating efficacy of antifungal treatment and enhancing patient results. There is growing worry about the uptick in infections attributed to non-albicans fungal species, coupled with the development of resistance to antifungal medications [5].
At present, antifungal drugs used against Candida species fall into four main categories: azoles, polyenes, echinocandins, and 5-flucytosine. These classes are characterized by distinct mechanisms of action and a varied spectrum of activity [6]. The increased use of antifungal therapy for prophylactic or empirical treatment has led to the emergence of various Candida species with decreased susceptibility or resistance to antifungal drugs, as it selects for microorganisms that can survive and reproduce in the presence of these drugs [7]. Due to these changes, developing a standardized in vitro antifungal susceptibility assay has become essential for optimizing antifungal therapy and predicting clinical outcomes. Thus, given the rise of non-albicans and rare Candida species with diverse resistance patterns, proper antifungal testing and species-level identification are crucial for managing Candida infections [8].
Another important aspect of studying Candida species infections is examining the unique virulence factors they possess, which contribute to their pathogenicity [9]. One significant virulence factor is biofilm formation, which enables Candida species to overcome the body's defense mechanisms [10]. Candida species display variation in their capacities for forming biofilms, particularly concerning their physical structure, the properties of the extracellular matrix and their ability to develop resistance against antifungal treatments. These distinctions underline the diversity among Candida species in terms of biofilm formation making it a complex and multifaceted phenomenon to study [11]. The diversity in these characteristics heightens the complexity of addressing Candida biofilms as a distinct challenge, thereby amplifying the difficulty in deciphering effective solutions to combat the associated threats. The ECM work as a supporting frame for a cell to attach to various surfaces. It serves as a protective shield, separating the cells within the biofilms from their external environment [12]. These disparities underscore the intricate nature of the mechanism driving biofilm formation, thus highlighting the challenge of identifying a universal approach for eradicating all forms of Candida biofilms. Notably, in each scenario, the development of biofilms undermines the effectiveness of antifungal therapies. When these biofilms form on implanted medical devices, such as central venous catheters, it frequently necessitates the replacement of the implant as a solution [13]. The formation of biofilm by Candida spp., therefore, possess a significant clinical challenge, that necessitates their evaluation.
Another factor that contributes to their pathogenicity is the secretion of hydrolytic enzymes that allows adherence, tissue damage, and changes in the immune response, and hence enhancing their activity leading to acute candidiasis [14]. Various species of Candida produce a range of hydrolytic enzymes, including proteinase, phospholipase, esterase, and hemolysin, which contribute to tissue invasion by disrupting cell membranes [15]. These organisms possess a mechanism to break down hemoglobin and liberate iron by the production of a lytic protein known as hemolysin [16]. The significant role of hemolysins to aid in the long-term survival of pathogens, helping them acquire iron, and promoting hyphal invasion in systemic or disseminated candidiasis cases, underscores its crucial importance as a virulence factor [17]. Candida species were reported to have 30–100% production of phospholipase [18]. Along with this, a multifunctional molecule SAP play a role in Candida pathogenesis by enhancing the colonization and penetration of host tissues [19]. C. albicans and non-albicans species naturally produce SAP genes. Among highly pathogenic Candida species, such as C. glabrata, C. tropicalis, C. parapsilosis, C. dubliniensis, and C. albicans, the expression of these genes is highly observed [20]. Esterase is another hydrolytic enzyme that breaks down cell membrane ester bonds and promotes tissue invasion [9]. Given this, studying the hydrolytic enzymes associated with Candida species is crucial for monitoring their infections.
In summary, virulence factors expression and in vitro antifungal susceptibility profiles of Candida spp., may differ based on the infecting species, geographical origin, kind of infection, the site and stage of infection [21]. Unfortunately, to the best of my knowledge, no surveillance study has been seen in the northern region of India which can monitor the changes in susceptibility of antifungal drugs and their virulence factor of Candida species. In this study, we assessed the activity of aforementioned virulence factors among Candida species isolated from blood and also find their anti-fungal susceptibility against different antifungal drugs.
2 Materials and methods
2.1 Materials
Antifungal pharmaceutical compounds were procured from Sigma-Aldrich, USA. Fungal culture media, including SDA, BHI, YPD, Agar, and RPMI 1640 without sodium bicarbonate, were collected from Hi-media Laboratories Ltd., India. Plastic wares, such as sterile 50 ml Falcon tubes, 1.5 ml MCT, and 96-well flat-bottom microtiter plates, were collected from Tarson product Pvt. Ltd., India. Solvents like BSA, NaCl, CaCl2, MgSO4, K2HPO4, Peptone, Tween 80, and Crystal violet were obtained from Merck Life Science Pvt. Ltd., India. For this research, all the chemicals were used are of analytical grade quality.
2.2 Ethics approval and participants
The study approval was confirmed by the Institutional Ethical Committee (ECR/526/Inst/UP/2019), at the Institute of Medical Sciences, B.H.U, Varanasi, India. All patients were provided with detailed informed consent form for the study. The study adhere to Helsinki Declaration. Participants have the right to ask questions and receive satisfactory answers before signing informed consent form. For unconscious patients, their family members signed the informed consent form on their behalf ensuring all ethical standard legal requirement was met before participating in the study. No patients were influenced to participate. They were informed about their rights to withdraw from study at any point without any consequences. Informed Consent forms were provided in both English and Hindi languages.
2.3 Yeast isolates
A total of 120 Candida species were isolated from blood cultures of various age group patients. Blood samples were collected from the Microbiological laboratory, Institute of Medical Sciences, Varanasi in the year 2022–23. For the identification of Candida species we performed phenotypic and genotypic methods. Phenotypic identification were assessed through colony characteristics on SDA media followed by Gram’s staining and germ tube method. Colony color was also evaluated by utilizing a special agar called Hi-chrome Candida agar. Corn-Meal tween-80 was used to study the chlamydospores production. These isolates were further identified by Biochemical method according to the standard operating procedure for mycology laboratories. Confirmatory identification was done by MALDI-TOF MS a sophisticated technique that provides scores. A score of at least 2 log was required for species identification, 1.7–1.99 for genus identification, and scores below 1.7 were considered as unreliable. This final step of identification was conducted at PGIMER Chandigarh using equipment from Bruker Daltonik GmbH in Bremen, Germany. Stock cultures of the isolates were maintained in 15% glycerol broth at – 20 ℃ for future use. Before subjecting them to test, all isolates underwent sub-culturing on SDA under aerobic conditions at 37 ℃ for 24 h to confirm their viability and purity. C. albicans ATCC 90028, C. tropicalis ATCC 750, C. krusei ATCC 6258, and C. auris CDC B11903 were taken as standard reference strains in this experiment.
2.4 Anti-fungal drug susceptibility profile
Antifungal susceptibility of Candida isolates were assessed against antifungal drugs in a controlled laboratory setting by following CLSI M27-A3 guidelines [22]. The antifungal agents were used are Fluconazole, Itraconazole, Amphotericin B, Voriconazole, Posaconazole, Caspofungin, Anidulafungin, and Micafungin. We performed serial dilutions of the antifungal agents, utilizing RPMI-1640 medium distributing them into a 96-well round bottom microtiter plate containing 100 µl of each diluted drugs. The harvested cells were suspended in RPMI-1640 medium to achieve a concentration of approximately 5.0 × 102 to 2.5 × 103 cells/ml and 100 µl of the inoculum were added to each microdiluted well so that total volume of each well become 200 µl. Following the 24 h incubation period at 35 °C, we interpreted the susceptibility patterns of the tested isolates based on the CLSI M27-A3. This systematic approach allowed us to evaluate how the Candida isolates responded to different antifungal drugs under standardized conditions.
2.5 Enzymatic activity testing
The Extracellular enzymatic activities of Candida isolates were screened by plate methods, where 5 µl of Candida inoculum, derived from overnight growth on SDA media, were applied to each test medium. Following that, the plates were placed in an incubator at 37 ℃ and evaluated on different days for different enzymatic activity with the inoculum suspension of concentration equal to 107 CFU/ml. Enzymatic activity was assessed utilizing the Pz index, a semi quantitative measurement derived from both colony appearance and precipitation zone measurement. The Pz index values were classified into three categories: Strongly positive (Pz ≤ 0.64), Positive (0.64 < Pz < 0.99) and Negative activity (Pz = 1) [23]. This approach provided a systematic and standardized method for assessing the enzymatic activity of the Candida isolates in the study. The experiment was replicated three times on distinct occasions for every isolate, with C. albicans ATCC were employed as a positive control.
2.5.1 Phospholipase assay
Phospholipase activity in the isolated sample was assessed through the plate method. The SDA medium was enriched with 1 M NaCl, 0.005 M CaCl2, 3% glucose, and 10% sterile egg yolk. First, the components without the egg yolk were mixed and sterilized [24]. Then egg yolk was centrifuged at 500 × g for 10 min at room temperature and 20 ml of supernatant was added to a sterilized medium [25]. 5 µl of standard suspension of the test and control Candida isolates were deposited on the egg yolk agar medium and left to dry at room temperature. Each inoculated culture plate was then incubated at 37℃ for 48 h, after which precipitation zone around the colony was calculated.
2.5.2 Proteinase assay
Testing for proteinase production was conducted utilizing bovine serum albumin, following a previously described method [26]. In summary, the test medium was formulated by combining 0.1% K2HPO4, 0.05% MgSO4, 0.2% BSA, 1% Dextrose, and 0.01% yeast extract while maintaining a pH of 5. This mixture was then blended with molten agar to attain a final concentration of 2% (w/v) BSA agar. 5 µl of each Candida suspension was spot inoculated on the BSA plate and incubated at 30℃ for 48 h.
2.5.3 Hemolysin assay
The SDA medium was enriched with glucose concentrations of 7% and 3% (w/v), and was adjusted to 5.6 pH. 5 µl of this suspension was spot inoculated on prepared medium to yield a circular inoculation site of about 5 mm in diameter. The plates were incubated for 2 days at 30 ℃ in the presence of 5% CO2. The presence of a distinct translucent halo around the inoculum site, viewed with transmitted light, is indicative of hemolytic activity [27].
2.5.4 Esterase assay
Esterase Activity was performed on a Tween 80 opacity medium. The medium was prepared using 10gm of bacteriological peptone, 5 gm of NaCl, 0.1 gm of CaCl2, and 15 gm of Agar dissolved in 1000 ml of distilled water and autoclaved. Then 5 ml of autoclaved Tween 80 was added to the medium cooled at 45 ℃ and distributed in sterile 90 mm petri dishes. Overnight grown culture was taken for suspension preparation. Each Candida suspension, in volumes of 5 µl was spot inoculated onto plates and then placed in incubation at 30 ℃. Daily observations were conducted over 10 days [27].
2.6 Biofilm formation assay
The method employed for quantitatively measuring biofilm production involved initial culturing of Candida isolates on SDA at 37 °C for 24 h. Afterward, individual colonies were suspended into physiological saline (0.89% NaCl); the suspension was achieved to a 107 CFU/ml (0.5-Mcfarland turbidity). Then 20 µl sample of Candida cell suspension, was inoculated into a 96-well flat bottom microtiter plate containing 180 µl of YPD supplemented with 2% glucose, followed by an incubation period at 37 °C for 24 h. After period of incubation, the planktonic cells were eliminated, and the biofilm underwent two washes with 200 µl of PBS (pH: 7.4). The biofilm was then heat fixed at 60 °C for 20 min and stained with 175 µl of 0.5% crystal violet dye for 5 min. After rinsing off the excess dye with running tap water, bound crystal violet was solubilized with 150 µl of mixture of 80% ethanol and 20% acetone. The solubilized biofilm was resuspended in new 96-well microtiter plate to measure absorbance signal at 595 nm wavelength in ELISA plate reader (Thermo Scientific, USA), utilizing media blanks as negative controls. Each isolate underwent four independent replicate experiments. Biofilm density categorization into four groups was determined by established cut-off values (ODc), which were calculated from the mean ODnc of negative controls plus three standard deviations (3xSDnc). These categories were: OD ≤ ODc (no biofilm formation), ODc < OD ≤ 2 × ODc (low biofilm formation), 2 × ODc < OD ≤ 4 × ODc (moderate biofilm formation), and 4 × ODc < OD (high biofilm formation) [28].
2.7 Field-emission scanning electron microscopy
In this study, biofilm formation by various Candida species was investigated by FE-SEM. The experimental procedure involved seeding 2 ml of a standardized cell suspension in YPD media containing 1 × 107 CFU/ml into individual wells of a 12-well culture plate, followed by incubation for 24 h at 37 °C. Subsequently, after 12 h of incubation, 1 ml of the YPD medium was replaced with an equal volume of fresh YPD to sustain biofilm growth. After a total incubation period of 24 h, the culture medium was aspirated, and non-adherent cells were eliminated by washing the biofilm with PBS (pH: 7.4). The formed biofilm on coverslip was then subjected to fixation by immersion in a 2.5% paraformaldehyde solution. Post-fixation procedures were conducted using 1% osmium tetroxide for 1 h, followed by treatment with 1% tannic acid for an additional hour. Following a brief rinse in distilled water, the samples were subjected to dehydration by being immersed sequentially in ethanol solutions with ascending concentrations (30%, 50%, 70%, 90% and 100%), each for 10 min. Following dehydration, the samples were air-dried and bottom of the wells was cut for observation involved mounting the base of the dehydrated biofilm material onto aluminum stubs, followed by sputter-coating with gold. The samples were examined using a field emission scanning electron microscope (EVO 18, Zeiss, Germany). This experimental protocol ensures the preservation of biofilm morphology and structure, allowing for comprehensive SEM-based analysis of Candida biofilms [29].
2.8 Statistical analysis
Statistical analysis was performed utilizing Graph pad prism statistical software 8.4.2. Due to the non-normal distribution of the data, non-parametric tests were utilized. Descriptive statistics for the variables were reported as median, minimum, and maximum values. The degree of association between paired data was assessed utilizing Spearman’s correlation coefficient (rs), where values nearer to ± 1 suggest a more robust relationship. Variations in median values of quantitative variables across distinct groups were examined employing Mann–Whitney’s test and Kruskal–Wallis’s test. Statistical significance was defined at p < 0.05 for all analyses.
3 Results
3.1 Candida species identification
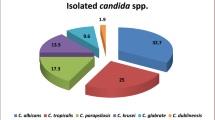
This research revealed that among 120 clinical isolates of Candida obtained from blood cultures, C. albicans was the most prevalent species, accounting for 40% (n = 48), followed by C. tropicalis at 39% (n = 47), C. krusei at 20% (n = 12), and C. auris at 20% (n = 13), which were examined by MALDI-TOF MS Notably, C. tropicalis emerged as the most common non-albicans species. Furthermore, the study demonstrated that non-albicans Candida collectively represented 60% (n = 72) of the isolates, surpassing the proportion of C. albicans, which accounted for 40% (n = 48) as shown in (Fig. 1). The phenotypic identification of Candida spp. were done by Gram staining, growth on CHROMagar, germ tube test (Fig. 2) and morphology detection was performed on corn meal agar (Fig. 3).
3.2 Antifungal drug susceptibility test
This study reveals the antifungal susceptibility profiles of various Candida species, including C. albicans, C. tropicalis, C. auris, and C. krusei was determined according to CLSI recommended guidelines. A comprehensive analysis was conducted, focusing on 48 isolates of C. albicans, revealing a wide range of MIC50 and MIC90 values across eight different antifungal agents. Remarkably, fluconazole showed a MIC50 of 0.25 μg/ml and a MIC90 of 8 μg/ml, with a mode of 0.12 μg/ml for C. albicans. Similar patterns were observed with other antifungal medications, such as itraconazole and caspofungin show MIC90 value 0.5 μg/ml, posaconazole and anidulafungin show MIC90 value 0.12 μg/ml, voriconazole and micafungin show MIC90 value 0.06 μg/ml, and for amptotericin B MIC90 value 2 μg/ml was observed for C. albicans. Out of the 47 isolates representing C. tropicalis, we found a similar range of variability regarding antifungal susceptibility. Specifically, fluconazole showed a MIC50 value of 0.12 μg/ml and a MIC90 value of 0.5 μg/ml. Interestingly, for C. tropicalis the most common MIC90 value was 0.12 μg/ml for most of antifungal drugs except fluconazole, amphotericin B and anidulafungin as shown in Table 1. We also examined 13 strains of C. auris, finding that fluconazole showed higher MIC50 value 16 μg/ml and MIC90 value 64 μg/ml. Surprisingly, amphotericin B had a mode of 2 μg/ml and MIC90 value of 4 μg/ml. The effectiveness of other antifungal treatments differed against C. auris, with unique MIC90 and mode value for each. A noteworthy finding emerged when studying C. krusei, with 12 isolates displaying distinct antifungal susceptibility patterns. Specifically, fluconazole demonstrated a high MIC50, MIC90 value and mode of 16 μg/ml. Further examination of the remaining antifungal drugs revealed unique MIC values, providing invaluable insight into C. krusei susceptibility (Fig. 4). These findings underline the significance of considering MIC50 and MIC90 values, along with mode, when evaluating antifungal susceptibility against various Candida species. The observed variations highlight the intricate nature of antifungal resistance patterns among Candida isolates, underscoring the necessity for tailored therapeutic approaches in clinical settings.
3.3 Evaluation of enzymatic activity
The phospholipase, proteinase, hemolysin, and esterase activity (Pz value) was measured by taking the ratio of the diameter of the colony plus the precipitation zone in mm shown in Fig. 5. Enzyme activity precipitation zone (Pz) of 1 mm was evaluated as negative, 0.99–0.90 mm as weak, 0.89–0.8 mm as mild, 0.79–0.7 as relatively strong, and < 0.69 as a very strong activity.
In our study, all isolates of C. tropicalis was phospholipase-negative. Maximum phospholipase activity was shown by C. albicans followed by C. krusei and C. auris. None of the isolates showed weak or mild activity. Very few percent which means 4% of C. albicans showed positive activity whereas about 81% of C. albicans showed very strong positive activity shown in Table 2. C. tropicalis followed by C. albicans showed maximum proteinase activity. Half of the C. krusei showed very strong activity and half of them showed negative activity. About 76% of C. auris showed relatively strong activity and 7% of isolates were showing negative activity as depicted in Table 3. C. albicans and C. tropicalis showed the maximum hemolytic activity which was nearly equal followed by C. auris whereas C. krusei isolate showed comparatively less hemolytic activity. Thus, for C. albicans, C. tropicalis, and C. auris, hemolysin is a strong virulence factor as mentioned in Table 4. Maximum esterase activity was shown by C. tropicalis followed by C. albicans. 76% of the C. auris isolates showed very strong activity and 15% of isolates showed relatively strong activity as shown in Table 5. The isolates of C. krusei showed 58% of very strong activity, and 41% were depicting negative activity against esterase. Thus, C. tropicalis is the most virulent, and C. krusei is the least virulent.
3.4 Evaluation of biofilm
This assessment depicted the percentage of biofilm formation among Candida isolates. Based on CV staining, out of 120 isolates, 116 were producing biofilms and 4 were non-biofilm producers, as inferred by OD values (Table S1 of supplementary material). According to the cut off OD (ODc) of the CV staining assay, clinical isolates of C. tropicalis (n = 14), C. krusei (n = 8), C. albicans (n = 1) and C. auris (n = 1) exhibited the highest biofilm production when grown on an YPD medium supplemented with 2% glucose. Results of the high, moderate, and weak biofilm former shown that non-albicans Candida strains had a high percentage of biofilm production shown in Fig. 6. There was significant variability in biofilm formation observed both among different species and within the same species, as illustrated in Table 6. The highest levels of biomass production were noted in C. tropicalis and C. krusei. Biofilm comparisons between different Candida species were analyzed by using SEM at two different Magnifications shown in Fig. 7
3.5 Statistical correlation analyses
The statistical analysis was conducted on the enzymatic activity against various clinical isolates of Candida spp. Results revealed significant (p < 0.05) when employing three different tests: the Friedman test, the Kruskal–Wallis test, and the Two-Way ANOVA for non-parametric data as shown in Table 7. Furthermore, a significant Spearman correlation was found between MIC50 and MIC90 values against antifungals across all Candida isolates shown in Table 8.
4 Discussion
A comprehensive study was conducted in 2022–23 at Sir Sunderlal Hospital, BHU, Varanasi, India. In this study 120 common species of Candida, isolated from bloodstream infections were analyzed for anti-fungal susceptibility and virulence factors. This epidemiological investigation is pivotal for understanding and treating invasive candidiasis, shedding light on the mechanisms of Candida infection [30]. In India, there is limited research on epidemiology of Candidemia. This study elucidated that 60% of the isolated species were non-albicans Candida (n = 72), rest isolates were C. albicans (n = 48). Previous studies also showed similar species distribution where non-albicans Candida has higher proportion than C. albicans as main etiological agent that causes candidiasis [23].
Susceptibility profiles of various Candida species to different antifungal agents provides essential insights into treatment efficacy [31]. Notably, in our study, C. albicans and C. tropicalis exhibited MIC90 values of 8 µg/ml and 0.5 µg/ml respectively for fluconazole as shown in Fig. 4. In compared to another azole groups drugs (itraconazole, voriconazole and posaconazole), fluconazole displayed less susceptibility towards all isolated Candida species. C. auris and C. krusei demonstrated varying degrees of resistance to fluconazole with reduced efficacy in both species and Berkow et al. also reported similar type of results for these species [32]. Fluconazole is still considered an acceptable alternative for non-albicans Candida infections due to its low cost and toxicity as compared to echinocandins. It is widely available and used orally in non-critically unwell individuals [33]. In our study, for amphotericin B, C. albicans and C. krusei show MIC90 value 2 µg/ml for both species whereas C. tropicalis and C. auris show MIC90 value 1 µg/ml and 4 µg/ml respectively. The epidemiological cutoff values (ECVs) of C. tropicalis, C. krusei and C. albicans for amphotericin B was 1 µg/ml, 4 µg/ml and 0.5 µg/ml respectively. After comparing the MIC90 and epidemiological cutoff value, result shows less MIC90 value than ECV. For echinocandin, according to CLSI antifungal breakpoint guideline, all isolates of C. tropicalis, C. krusei and C. albicans show MIC90 value within susceptible range except C. albicans MIC90 value of 0.5 µg/ml for caspofungin which falls under intermediate range of the guidelines [34]. Echinocandins work by inhibiting (1,3)-β-d-glucan synthase [35], azole blocks the synthesis of ergosterol by targeting Erg11p or Cyp51p [36], while polyenes binds to ergosterol, causing pore formation, and ion leakage with fungicidal effect [37]. The possible mechanism of antifungal resistance to echinocandins is attributed by a mutation in the FKS gene. Azole drug resistance was mediated by sequential acquisition under drug pressure. This causes mutation in genes ABC or the MFS genes and the second mechanism is overproduction of target enzymes Erg11p. The mechanism of polyenes resistance remains unclear, but in some literature, it was reported with mutation in genes that are involved in the ergosterol biosynthesis pathway [38, 39].
Furthermore, several research have extensively investigated for enzymatic activity of Candida species to understand their pathogenicity and interactions with the host [40, 41]. Enzymes such as phospholipase and proteinase play crucial roles in Candida infections. This study reveals that C. albicans have a highest phospholipase activity while C. tropicalis exhibiting the highest proteinase activity which are also in concordance with the work demonstrated by Sriphannam et al. [9] (Fig. 5). These enzymes aid in tissue invasion and spread of the fungus within the host [42, 43]. Moreover, the virulence of Candida species enables them to break down hemoglobin from the host to obtain iron, facilitating their persistence and survival within the host environment. In this study, C. albicans and C. tropicalis had the highest hemolysin activity, which contributed to their pathogenicity. Similar results were reported in other research works conducted earlier, with C. albicans and C. tropicalis having higher hemolytic activity [44, 45]. Esterase enzyme activity, which is essential for tissue invasion, was notably high in C. tropicalis and C. albicans, suggesting their potential virulence which was concordant with previous studies [9, 44].
Biofilm formation is a significant factor contributing to Candida’s pathogenicity and resistance to antifungal treatments. C. tropicalis exhibited the highest biofilm formation, followed by C. krusei and C. albicans as shown in Fig. 6. Similar results were also revealed by Sriphannam et al. where C. tropicalis isolates show high biofilm formation [9, 10]. The ability to form robust biofilms is particularly concerning with their resistance to antifungal treatments and increased incidence of hospital-acquired infections [46,47,48]. Candida species, like other microorganisms, can form biofilms with different shapes and thicknesses. These biofilms have a protective layer around the cells called an extracellular matrix. This matrix acts like a shield, guarding the cells within the biofilm. [49,50,51]. All Candida strains were examined by scanning electron microscopy also to address the surface analysis and adherence of yeast cells to the substrate (Fig. 7) [52,53,54].
The statistical analysis of this work also shows significant value after applying a non-parametric test for enzymatic activity and correlation with MIC50 and MIC90. While this study provides valuable insights into the virulence factors and biofilm formation of common Candida species isolated from bloodstream infections, it has some limitations also. One major limitation is the relatively small sample size, which may not fully represent the diversity of Candida strains and their virulence profiles. Additionally, our study focused on a specific demography (ICU patients), limiting the generalization of our findings to other patient populations. Future research could address these limitations by conducting larger-scale epidemiological studies encompassing a broader range of patient demographics and healthcare settings. Furthermore, investigations into the molecular mechanisms underlying virulence factor expression and biofilm formation in Candida species could provide deeper insights into their pathogenicity and aid in the development of more effective treatment strategies. Additionally, exploring the impact of environmental factors on Candida virulence could enhance our understanding of host–pathogen interactions and provide infection prevention measures.
5 Conclusion
In summary, our findings demonstrate how several factors influence the potency and survival of Candida species seen in bloodstream infections. The study reveals that non-albicans Candida is more likely to cause candidemia than C. albicans in Northern India. The albicans species has the highest antifungal sensitivity, whereas non-albicans exhibit resistance to fluconazole and other antifungal medications. Candida species like C. auris and C. krusei demonstrate the strongest resistance to fluconazole and also found that caspofungin, anidulafungin, and micafungin are effective against several Candida species. C. tropicalis demonstrated the highest enzymatic activity for proteinase and esterase, but no activity against phospholipase. While C. albicans has the highest hemolysin activity. C. tropicalis developed more biofilm than C. albicans. In conclusion, the increased incidence of non-albicans Candida in blood stream infections and high rates of fluconazole resistance underline the importance of ongoing candidemia surveillance, antifungal stewardship for retaining antifungal efficacy and risk factor analysis related with switch to candidemia caused by non-albicans Candida. These findings emphasize the need for continued monitoring and personalized treatment approaches to effectively manage serious Candida infections.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- MIC:
-
Minimum inhibitory concentration
- FE-SEM:
-
Field emission scanning electron microscopy
- ECM:
-
Extracellular matrix
- SAP:
-
Secretory aspartyl proteinase
- ICU:
-
Intensive care unit
- SDA:
-
Sabouraud dextrose agar
- CFU:
-
Colony forming unit
- BSA:
-
Bovine serum albumin
- PBS:
-
Phosphate buffer saline
- YPD:
-
Yeast extract peptone dextrose
- MCT:
-
Micro centrifuge tubes
References
Chow BDW, Reardon JR, Perry EO, Tucker R, Bliss JM. Expressed breast milk as a predictor of neonatal yeast colonization in an intensive care setting. J Pediatr Infect Dis Soc. 2014;3(3):213–20. https://doi.org/10.1093/jpids/pit090.
Lamoth F, Lockhart SR, Berkow EL, Calandra T. Changes in the epidemiological landscape of invasive candidiasis. J Antimicrob Chemother. 2018;73:i4–13. https://doi.org/10.1093/jac/dkx444.
Pappas PG, Lionakis MS, Arendrup MC, Ostrosky-zeichner L, Kullberg BJ. Invasive candidiasis. Nat Publ Gr. 2018;4:1–20. https://doi.org/10.1038/nrdp.2018.26.
Lopes JP, Lionakis MS. Pathogenesis and virulence of Candida albicans ABSTRACT. Virulence. 2022;13(1):89–121. https://doi.org/10.1080/21505594.2021.2019950.
Pereira GH, Müller PR, Szeszs MW, Levin AS, Melhem MSC. Five-year evaluation of bloodstream yeast infections in a tertiary hospital: the predominance of non-C. albicans Candida species. Med Mycol. 2010;48(6):839–42. https://doi.org/10.3109/13693780903580121.
Bhattacharya S, Sae-tia S, Fries BC. Candidiasis and mechanisms of antifungal resistance. Antibiotics. 2020. https://doi.org/10.3390/antibiotics9060312.
Morace G, Perdoni F, Borghi E. Antifungal drug resistance in Candida species. J Glob Antimicrob Resist. 2014;2:254–9.
Pfaller MA. Antifungal drug resistance: mechanisms, epidemiology, and consequences for treatment. Am J Med. 2012;125(1 Suppl):S3–13. https://doi.org/10.1016/j.amjmed.2011.11.001.
Sriphannam C, Nuanmuang N, Saengsawang K, Amornthipayawong D, Kummasook A. Anti-fungal susceptibility and virulence factors of Candida spp. isolated from blood cultures. J Mycol Med. 2019;29(4):325–30. https://doi.org/10.1016/j.mycmed.2019.08.001.
Marcos-Zambrano LJ, Escribano P, Bouza E, Guinea J. Production of biofilm by Candida and non-Candida spp. isolates causing fungemia: comparison of biomass production and metabolic activity and development of cut-off points. Int J Med Microbiol. 2014;304(8):1192–8. https://doi.org/10.1016/j.ijmm.2014.08.012.
Seneviratne CJ, Jin L, Samaranayake LP. Biofilm lifestyle of Candida: a mini review. Oral Dis. 2008;14(7):582–90. https://doi.org/10.1111/j.1601-0825.2007.01424.x.
Mitchell KF, Zarnowski R, Andes DR. The extracellular matrix of fungal biofilms. Adv Exp Med Biol. 2016;931:21–35. https://doi.org/10.1007/5584_2016_6.
Mermel LA, et al. Guidelines for the management of intravascular catheter-related infections. Clin Infect Dis. 2001;32:1249–72.
Jafarian H, Gharaghani M, Seyedian SS, Mahmoudabadi AZ. Genotyping, antifungal susceptibility, enzymatic activity, and phenotypic variation in Candida albicans from esophageal candidiasis. J Clin Lab Anal. 2021;35(7):1–10. https://doi.org/10.1002/jcla.23826.
Silva S, Negri M, Henriques M, Oliveira R, Williams DW, Azeredo J. Candida glabrata, Candida parapsilosis and Candida tropicalis: biology, epidemiology, pathogenicity and antifungal resistance. FEMS Microbiol Rev. 2012;36(2):288–305. https://doi.org/10.1111/j.1574-6976.2011.00278.x.
Linares CEB, et al. Enzymatic and hemolytic activities of Candida dubliniensis strains. Rev Inst Med Trop Sao Paulo. 2007;49(4):203–6. https://doi.org/10.1590/s0036-46652007000400001.
Pendrak ML, Roberts DD. Hemoglobin is an effective inducer of hyphal differentiation in Candida albicans. Med Mycol. 2007;45(1):61–71. https://doi.org/10.1080/13693780601028691.
Rossoni RD, Barbosa JO, Godinho Vilela SF, Cardoso Jorge AO, Junqueira JC. Comparison of the hemolytic activity between C. albicans and non-albicans Candida species. Braz Oral Res. 2013;27(6):484–9. https://doi.org/10.1590/S1806-83242013000600007.
Pandey N, Gupta MK, Tilak R. Extracellular hydrolytic enzyme activities of the different Candida spp. isolated from the blood of the Intensive Care Unit-admitted patients. J Lab Phys. 2018;10(04):392–6. https://doi.org/10.4103/jlp.jlp_81_18.
Meenambiga SS, Venkataraghavan R, Abhishek Biswal R. In silico analysis of plant phytochemicals against secreted aspartic proteinase enzyme of Candida albicans. J Appl Pharm Sci. 2018;8(11):140–50. https://doi.org/10.7324/JAPS.2018.81120.
Deorukhkar SC, Saini S, Mathew S. Virulence factors contributing to pathogenicity of candida tropicalis and its antifungal susceptibility profile. Int J Microbiol. 2014. https://doi.org/10.1155/2014/456878.
Alexander BD. Document M27-A4. Ref Method Broth Dilution Antifung. Suscetibility Test. Yeasts. 2012.
Maria H, Canela S, Coelho HC, Cardoso B, Martinez R, Eliana M. Prevalence, virulence factors and antifungal susceptibility of Candida spp. isolated from bloodstream infections in a tertiary care hospital in Brazil. Mycoses. 2017. https://doi.org/10.1111/myc.12695.
Price MF, Wilkinson ID, Gentry LO. Plate method for detection of phospholipase activity in Candida albicans. Med Mycol. 1982. https://doi.org/10.1080/00362178285380031.
Costa CR, Passos XS, Kioko L, Lucena PDA, De Fátima O, Fernandes L. Differences in exoenzyme production and adherence ability of Candida spp. isolates from catheter, blood and oral cavity. Rev Inst Med Trop S Paulo. 2010;52(3):139–43. https://doi.org/10.1590/S0036-46652010000300005.
Sacristan B, Blanco MT, Galan-Ladero MA, Blanco J. “Short communications aspartyl proteinase, phospholipase, hemolytic activities and biofilm production of Candida albicans isolated from bronchial aspirates of ICU patients. Med Mycol. 2011. https://doi.org/10.3109/13693786.2010.482947.
Fatahinia M, Poormohamadi F, Mahmoudabadi AZ. Comparative study of esterase and hemolytic activities in clinically important Candida species, isolated from oral cavity of diabetic and non-diabetic individuals. Jundishapur J Microbiol. 2015. https://doi.org/10.5812/jjm.20893.
Singh AK, Prakash P, Achra A, Singh GP, Das A, Singh RK. Standardization and classification of in vitro biofilm formation by clinical isolates of Staphylococcus aureus. J Glob Infect Dis. 2017. https://doi.org/10.4103/jgid.jgid.
Relucenti M, et al. Microscopy methods for biofilm imaging: focus on SEM and VP-SEM pros and cons. Biology. 2021. https://doi.org/10.3390/biology10010051.
Kothavade RJ, Kura MM, Valand AG, Panthaki MH. Candida tropicalis: its prevalence, pathogenicity and increasing resistance to fluconazole. J Med Microbiol. 2010;59(8):873–80. https://doi.org/10.1099/jmm.0.013227-0.
Nicholls S, MacCallum DM, Kaffarnik FAR, Selway L, Peck SC, Brown AJP. Activation of the heat shock transcription factor Hsf1 is essential for the full virulence of the fungal pathogen Candida albicans. Fungal Genet Biol. 2011;48(3):297–305. https://doi.org/10.1016/j.fgb.2010.08.010.
Berkow EL, Lockhart SR. Fluconazole resistance in Candida species: a current perspective. Infect Drug Resist. 2017;10:237–45. https://doi.org/10.2147/IDR.S118892.
Ou HT, Lee TY, Chen YC, Charbonneau C. Pharmacoeconomic analysis of antifungal therapy for primary treatment of invasive candidiasis caused by Candida albicans and non-albicans Candida species. BMC Infect Dis. 2017;17(1):1–9. https://doi.org/10.1186/s12879-017-2573-8.
Wiederhold NP. Antifungal susceptibility testing: a primer for clinicians. Open Forum Infect Dis. 2021;8(11):1–13. https://doi.org/10.1093/ofid/ofab444.
Perlin DS. Current perspectives on echinocandin class drugs. Future Microbiol. 2011;6(4):441–57. https://doi.org/10.2217/fmb.11.19.
Odds FC, Brown AJP, Gow NAR. Antifungal agents: mechanisms of action. Trends Microbiol. 2003;11(6):272–9. https://doi.org/10.1016/S0966-842X(03)00117-3.
Gray KC, et al. Amphotericin primarily kills yeast by simply binding ergosterol. Proc Natl Acad Sci USA. 2012;109(7):2234–9. https://doi.org/10.1073/pnas.1117280109.
Coste A, et al. Genotypic evolution of azole resistance mechanisms in sequential Candida albicans isolates. Eukaryot Cell. 2007;6(10):1889–904. https://doi.org/10.1128/EC.00151-07.
Sanglard D, Coste A, Ferrari S. Antifungal drug resistance mechanisms in fungal pathogens from the perspective of transcriptional gene regulation. FEMS Yeast Res. 2009;9(7):1029–50. https://doi.org/10.1111/j.1567-1364.2009.00578.x.
Tsang CSP, Chu FCS, Leung WK, Jin LJ, Samaranayake LP, Siu SC. Phospholipase, proteinase and haemolytic activities of Candida albicans isolated from oral cavities of patients with type 2 diabetes mellitus. J Med Microbiol. 2007;56(10):1393–8. https://doi.org/10.1099/jmm.0.47303-0.
Kuriyama T, Dw W, Mao L. Kuriyama2003. 2003;(5):405–7.
Seifi Z, Mahmoudabadi AZ, Zarrin M. Extracellular enzymes and susceptibility to fluconazole in Candida strains isolated from patients with vaginitis and healthy individuals. Jundishapur J Microbiol. 2015. https://doi.org/10.5812/jjm.20162.
de Riceto ÉBM, de Menezes RP, Penatti MPA, dos Pedroso RS. Enzymatic and hemolytic activity in different Candida species. Rev Iberoam Micol. 2015;32(2):79–82. https://doi.org/10.1016/j.riam.2013.11.003.
Sachin C, Ruchi K, Santosh S. In vitro evaluation of proteinase, phospholipase and haemolysin activities of Candida species isolated from clinical specimens. Int J Med Biomed Res. 2012;1(2):153–7. https://doi.org/10.14194/ijmbr.1211.
Galán-Ladero MA, Blanco MT, Sacristán B, Fernández-Calderón MC, Pérez-Giraldo C, Gómez-García AC. Enzymatic activities of Candida tropicalis isolated from hospitalized patients. Med Mycol. 2010;48(1):207–10. https://doi.org/10.3109/13693780902801242.
Williams D, Lewis M. Pathogenesis and treatment of oral candidosis. J Oral Microbiol. 2011;3(2011):1–12. https://doi.org/10.3402/jom.v3i0.5771.
Nobile CJ, Johnson AD. Candida albicans biofilms and human disease. Annu Rev Microbiol. 2015;69(1):71–92. https://doi.org/10.1146/annurev-micro-091014-104330.
Soll DR, Daniels KJ. Plasticity of Candida albicans biofilms. Microbiol Mol Biol Rev. 2016;80(3):565–95. https://doi.org/10.1128/mmbr.00068-15.
Hall CW, Mah TF. Molecular mechanisms of biofilm-based antibiotic resistance and tolerance in pathogenic bacteria. FEMS Microbiol Rev. 2017;41(3):276–301. https://doi.org/10.1093/femsre/fux010.
Silva S, Henriques M, Martins A, Oliveira R, Williams D, Azeredo J. Biofilms of non-Candida albicans Candida species: quantification, structure and matrix composition. Med Mycol. 2009;47(7):681–9. https://doi.org/10.3109/13693780802549594.
Atriwal T, et al. Mechanistic understanding of Candida albicans biofilm formation and approaches for its inhibition. Front Microbiol. 2021. https://doi.org/10.3389/fmicb.2021.638609.
Marak MB, Dhanashree B. Antifungal susceptibility and biofilm production of Candida spp. isolated from clinical samples. Int J Microbiol. 2018;2018:6–10. https://doi.org/10.1155/2018/7495218.
Kulig K, et al. Insight Into the properties and immunoregulatory effect of extracellular vesicles produced by Candida glabrata, Candida parapsilosis, and Candida tropicalis biofilms. Front Cell Infect Microbiol. 2022;12:1–13. https://doi.org/10.3389/fcimb.2022.879237.
Handorf O, et al. Nonthermal plasma jet treatment negatively affects the viability and structure of Candida albicans SC5314 biofilms. Appl Environ Microbiol. 2018;84(21):1–15. https://doi.org/10.1128/AEM.01163-18.
Acknowledgements
We acknowledge SATHI-BHU, Varanasi, India for providing the SEM facilities. Prof. Ragini Tilak expresses her gratitude to the IoE, BHU, Varanasi, India, for the financial backing received.
Funding
The two funding bodies financially supported this research work. First, Joint CSIR-UGC New Delhi, India (JRF Grants no. 1055/(CSIR-UGC NET JUNE 2019) for Punit Tiwari, and Second, Institute of Eminence (IoE), BHU, Varanasi, India, for Prof. Ragini tilak.
Author information
Authors and Affiliations
Contributions
PT: Conceptualization, and execution of the work; KT and TP performed experiments; AN: Biostatistics, Editing, and draft writing. The study was supervised by RT and MKG; PT and AN analyzed the data; RT, MKG, PT and AN revised the manuscript; PT edited the final version of the manuscript; All authors read and approved the submitted version.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The study was granted approval from the Ethical Committee of the Institute under the reference number ECR/526/Inst/UP/2019.
Competing interests
The authors declare that they have no conflict of interest. Author Agreement: Authors certify that they have seen and approved the final version of the manuscript being submitted. Submission declaration and verification: The authors declare that the work described has not been published previously, it is not under consideration for publication elsewhere.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tiwari, P., Nikhil, A., Tiwari, K. et al. In vitro determination of antifungal susceptibility and virulence factors in Candida species causing candidemia in North India Region. Discov Public Health 21, 50 (2024). https://doi.org/10.1186/s12982-024-00175-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12982-024-00175-0