Abstract
Dorema (Apiaceae) includes 12 species distributed in the Irano-Turanian region. The genus differs from other genera of subfamily Apioideae in having paniculate inflorescences with simple umbels. Recently, molecular phylogenetic analyses placed six species of Dorema, including the generitype, in Ferula, thus subsuming the former into the synonymy of the latter. The remaining six congeners have not been yet included in molecular analyses. Here, we fill this gap by determining the phylogenetic position of these species. Because of a low resolution in some parts of the Ferula tree in earlier studies, alongside standard set of markers—nuclear nrDNA ITS and plastid rps16 and rpoC1 introns and the rpoB-trnC intergenic spacer—we additionally assessed the utility of nrDNA ETS sequences in resolving infrageneric relationships in the genus. We included 43 ingroup species representing major infrageneric divisions recognised in the most recent taxonomic revision. Although we were not successful in obtaining all markers for all newly studied species, they were placed with high support in one clade within section Peucedanoides together with formerly studied congeners in all maximum likelihood and Bayesian analyses of nuclear, plastid, and combined data sets. The addition of the ETS marker to the combined phylogenetic analyses confirmed the most recent classification of Ferula with increased branch support in some parts of the tree. However, many branches still remained unresolved. In particular, the monophyly of former Dorema within section Peucedanoides as suggested by morphology has not been definitely confirmed. A new combination and two nomina nova are proposed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dorema D.Don (Apiaceae Lindl.) includes 12 species distributed in semi-deserts, steppes and dry mountain slopes in the Irano-Turanian region (Pimenov 1988). The genus is well distinguished from other genera of the subfamily Apioideae by the presence of paniculate inflorescences with simple umbels. Due to dorsally flattened mericarps (single-seeded parts of a fruit constituting a dispersal unit), Dorema was traditionally classified in the tribe Peucedaneae, while its affinity to Ferula L. (Drude 1898; Pimenov & Leonov 1993) was supported due to the presence of fibrous or prosenchymatic lignified hypendocarp, i.e. the inner layer of the pericarp (Drude 1898; Korovin 1939; Pimenov 1988). The close relationship between Dorema and Ferula was subsequently confirmed by immunological studies, although the placement of these two genera in Peucedaneae Dumort. was questioned (Shneyer et al. 1995). Recent phylogenetic studies, based on nuclear and plastid markers, firmly established the position of both genera far from Peucedanum L. (currently a member of Selineae Spreng.) in tribe Scandiceae Spreng. and clearly showed the position of Dorema within Ferula (Ajani et al. 2008; Kurzyna-Młynik et al. 2008; Panahi et al. 2015; Piwczyński et al. 2018). In a new classification system of the genus Ferula (Panahi et al. 2018), the studied species of Dorema were placed in the section Peucedanoides Boiss. of subgenus Narthex (Falc.) Drude together with some species of Ferula (F. caspica M.Bieb., F. feruloides (Steud.) Korovin) having unusual arrangement of compound umbels: a central umbellet is replaced by successive compound umbel resulting therefore in pseudoverticillate, ‘proliferating’ inflorescence. Interestingly, the inflorescence morphology was considered one of the most important criteria in distinguishing infrageneric units in Ferula in its traditional taxonomic system (Korovin 1939, 1947). However, low resolution within the section Peucedanoides in molecular analyses allowed neither to confirm whether members of Dorema form a clade nor to indicate their closest relatives. Moreover, only six species of Dorema were molecularly analysed (Panahi et al. 2018). The remaining six were either included in Ferula based solely on the morphology with two species synonymised under a single species name (F. namanganica Sennikov ≡ D. microcarpum Korovin, F. sabulosa (Litv.) Sennikov ≡ D. sabulosum Litv. and = D. karataviense Korovin; Sennikov and Tojibaev 2021) or were left as incertae sedis (D. badhysi Pimenov, D. balchanorum Pimenov and D. kopetdaghense Pimenov).
The infrageneric relationships within Ferula have usually been inferred using ITS sequence variation and/or three plastid markers: rps16 and rpoC1 introns and the rpoB-trnC intergenic spacer. However, even the combined analyses left many internal parts of the phylogenetic tree of Ferula unresolved. Recently, Piwczyński et al. (2018) examined the utility of three potentially highly variable pDNA regions, trnH-psbA, trnS-trnG, and atpB-rbcL intergenic spacers, for resolving relationships within the genus Ferula. They found that combined pDNA markers provided a low number of parsimony informative sites, even lower than ITS alone. Moreover, pDNA and ITS phylogenetic trees were incongruent. The cause of this incongruence might be manifold, including hybridisation, a lack of a phylogenetic signal, or homoplastic substitutions (Panahi et al. 2015, 2018). These examples show that highly variable molecular markers are necessary to increase the resolution of the phylogenetic tree of the genus. One of the markers potentially having strong phylogenetic signal is nuclear ribosomal DNA external transcribed spacer (ETS) (Logacheva et al. 2010; Yi et al. 2015). This marker, likewise as ITS, is a part of the rDNA cistron, which occurs in high-copy numbers in the genome and is subject to homogenisation (Poczai and Hyvönen 2010). In Apiaceae, the ETS region has been shown to evolve faster and to have higher number of parsimony informative sites than ITS (Logacheva et al. 2010). Therefore, this marker is the most prospective candidate that may help to increase the resolution of the phylogenetic tree of Ferula.
The aim of our study was (1) to determine molecular phylogenetic position of all species of Dorema and to ascertain whether their paniculate inflorescence, a unique trait among apioid umbellifers, is synapomorphic and (2) to assess the utility of ETS sequences in resolving infrageneric relationships in Ferula.
Materials and methods
Taxon sampling
The genus Dorema was subsumed into Ferula based on molecular analysis of six species (Panahi et al. 2015) out of 12 recognised by Pimenov (1988). In spite of the morphological similarity of the unsampled species, Panahi et al. (2015) refrained from their formal inclusion in Ferula until their placement is confirmed with molecular data. Three of these species were recently included into Ferula based exclusively on their morphology (Sennikov and Tojibaev 2021). Here, we sampled herbarium material for all six Dorema species that have never been included in the molecular analyses: D. badhysi, D. balchanorum, D. kopedaghense, D. microcarpum (≡ F. namanganica in Sennikov and Tojibaev 2021), D. karataviense (= F. sabulosa in Sennikov and Tojibaev 2021) and D. sabulosum (≡ F. sabulosa in Sennikov and Tojibaev 2021). For these samples, we attempted to obtain nuclear rDNA ITS and ETS, plastid rpoB-trnC intergenic spacer, and two plastid introns: rps16 and rpoC1, which were used in previous studies of the genus (Panahi et al. 2015, 2018). We supplemented these sequences with ITS and plastid data for 36 species of Ferula (39 accessions) and a representative of Leutea Pimenov from GenBank representing all infrageneric groups recognised by Panahi et al. (2018) and additionally sequenced ETS marker for these accessions. Finally, our matrix consisted of 43 ingroup taxa (46 accessions) and two outgroup species from subtribe Daucinae Dumort.: Laserpitium gallicum subsp. paradoxum (O.Bolòs & Font Quer) P.Monts. and Thapsia garganica L. (Table S1).
DNA isolation and sequencing
Total genomic DNA was extracted from ca. 20 mg of dried plant material using the DNeasy Plant Mini Kit (Qiagen, Valencia, California, U.S.A.) following the manufacturer’s instructions. The PCR reactions, primers description and sequencing protocol for obtaining ITS and three plastid markers for newly sampled Dorema species are provided in details elsewhere (Panahi et al. 2015). To PCR-amplify the ETS marker, we used the universal primers for Apiaceae, 18S-ETS and Umb-ETS (Logacheva et al. 2010). Each 20-μL polymerase chain reaction (PCR) was prepared using 1 × PCR buffer, 0.2 mM of deoxynucleoside triphosphates (dNTPs), 0.2 μM of each primer, 2.5 mM of MgCl2, 0.2 U of Taq DNA polymerase (ThermoScientific) and 1 μL of DNA template. The PCR included the initial denaturation at 95 °C for 5 min, followed by 30 cycles comprising 1 min of denaturation at 95 °C, 40 s of annealing at 64 °C and 45 s of extension at 72 °C. The final extension at 72 °C lasted for 5 min. Each PCR product was subsequently electrophoresed using 1% agarose gel and stained with GelRed (Biotium) to determine whether the amplification was successful. We considered a PCR product appropriate for sequencing if no obvious polymorphism (multiple bands from a single PCR product) was observed. The PCR product was then purified using Ampure (1:1.8) and sequenced using fluorescent Big Dye terminators (Applied Biosystems, Foster City, CA, USA). The sequencing products were precipitated using 0.3 M sodium acetate, washed with 80% ethanol, dried, and then resolved using an automated DNA sequencer at the Laboratory of Molecular Biology Techniques, Adam Mickiewicz University (Poznań, Poland) or Genomed (Warsaw, Poland). The sequences were assembled and edited using SeqMan II v. 4.0 (DNASTAR, Madison, WI, USA). All newly obtained sequences were deposited in GenBank (Table S1).
Phylogenetic analyses
DNA sequences were aligned using MUSCLE v.3.8.31 (Edgar 2004) using a graphical interface in Seaview v.4.6.1 (Gouy et al. 2010), separately for each marker. The resulting matrices were corrected manually if necessary. In the case of plastid markers, we also manually excluded long autapomorphic indels and parts of the alignment with ambiguous site homologies (e.g. long mono- or dinucleotide repeats). Subsequently, we concatenated the alignments into five data sets: nuclear markers only (ITS and ETS), two data sets containing plastid markers only (rpoB-trnC, rpoC1 and rps16) with and without manual elimination of ambiguous sites (full and trimmed thereafter), and two concatenated alignments including nuclear and full or trimmed plastid data sets. With these five data sets, we tested the influence of our alignment manipulation on phylogenetic relationships as well as the putative influence of incongruence between nuclear and plastid data sets, recognised in earlier studies (Panahi et al. 2018), on the placement of analysed Dorema species.
Phylogenetic analyses were performed using the maximum likelihood (ML) method implemented in RAxML v.8.2.12 (Stamatakis 2014) and the Bayesian inference (BI) implemented in MrBayes v.3.2.7 (Ronquist et al. 2012). Optimal partitioning schemes and substitution models were inferred with PartitionFinder v.1.1.1 (Lanfear et al. 2012, 2014) using the corrected Akaike information criterion (AICc). The search algorithm was set to ‘all’ to choose among all possible partitioning combinations. In the case of ML analyses, we fixed model substitution to GTR + G (GTR is the only model implemented in RAxML) and searched only for the best partitioning scheme with PartitionFinder (models = GTR + G). Although GTR + G + I model, with the proportion of invariable sites, is also available in RAxML, GTR + G is preferred due to the redundancy of the I parameter, which is not independent of the alpha parameter of the gamma distribution. MrBayes supports more models than RAxML, although it is also restricted to the subset of all possible general time-reversible models. To analyse this particular subset, we set models = mrbayes in PartitionFinder. All analyses were run using models with branch lengths linked among partitions (branchlengths = linked).
All ML analyses consisted of 2000 independent searches starting from distinct randomised maximum parsimony trees, and the branch support (BS) was evaluated based on 2000 standard non-parametric bootstrap replicates. In Bayesian analyses, two independent runs were executed simultaneously, each with four Monte Carlo Markov chains with 10 000 000 generations, a sampling frequency of 1000 generations, and all priors set to default. The initial 25% of saved trees were discarded as burn-in and the results were summarised on the 50% majority rule consensus tree. The effective sample size (ESS) for the estimated parameters and the convergence of the independent runs were checked using Tracer v.1.7.1 (Rambaut et al. 2018).
Results
Sequence characteristics
For the six newly examined species, we obtained six ITS and four ETS sequences (Table S1). With respect to plastid sequences, we obtained all markers for D. microcarpum (≡ F. namanganica), two (rpoB-trnC and rps16 intron) for D. balchanorum and D. karataviense (= F. sabulosa), and we failed to sequence plastid markers for D. badhysi, D. sabulosum (≡ F. sabulosa), and D. kopedaghense.
The ETS region was the shortest albeit most variable among the analysed markers (Table 1). For example, the second most variable marker, ITS had 5.6% parsimony informative positions (34/608), while ETS had 9.7% (46/474). The combined alignment of nuclear markers had 7.3% parsimony informative sites (80/1082). Five accessions forming two groups (Ferula hyrcana (Koso-Pol.) Puchałka & al. 0071, D. balchanorum KEIB_AP_00802 and F. ovina (Boiss.) Boiss. 0124, F. ovina 0156, F. tschimganica Lipsky ex Korovin 0201) shared identical sequences. Each group was represented by a single terminal in the subsequent maximum likelihood phylogenetic analyses of nuclear data. Plastid markers were longer, less variable and had more indels than nuclear markers did (Table 1). The manual removal of indels and ambiguous sites had little effect on the number of parsimony informative positions. The concatenated plastid data sets had 32 and 30 parsimony informative sites in full and trimmed alignments, respectively. Each species analysed had an unique plastid haplotype.
PartitionFinder chose separate substitution models for ITS and ETS in the nuclear data set for both ML and BI analyses (Table 2). For BI, SYM + G and GTR + G were selected for ITS and ETS, respectively. Both full and trimmed plastid data sets were partitioned identically for ML analyses with rpoC1 and rps16 forming one partition, and rpoB-trnC having separate substitution model. The partitioning for BI analysis, on the other hand, resulted in two different divisions for full and trimmed data sets (Table 2). For the former, all markers were treated as separate partitions, while for the latter, rpoC1 and rps16 formed one partition. In general, the GTR model was selected for these partitioning schemes, although with various combinations of G and I parameters (Table 2). The same difference between full and trimmed data sets was also obtained for concatenated nuclear and plastid markers data sets (Table 2).
Phylogenetic position of the six species formerly described in Dorema
The incongruence between plastid and nuclear genomes recognised in Ferula did not influence the placement of the analysed species (Figs. 1, 2 and Fig. S1). The six analysed species were grouped in one clade with formerly studied congeners in the analyses of nuclear, plastid, and combined data sets. In the most informative combined analyses, all species were placed within well-supported clade representing section Peucedanoides (PP = 1.0/1.0, BS = 87/84%, for manually trimmed and full data sets, respectively; Figs. 1, 2). However, they did not form a monophyletic group, but they were intermingled with other species of Ferula in one highly supported clade (clade A′) within the section (PP = 0.92/0.93, BS = 97/96%). In general, the relationships in this clade were unresolved or poorly supported.
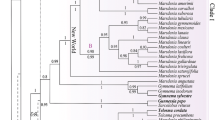
The Bayesian 50% majority-rule consensus tree of 46 representatives of Ferulinae inferred from analysis of combined nuclear (nrDNA ITS and ETS) and trimmed plastid sequence data (rpoB-trnC intergenic spacer, rps16, and rpoC1 introns). Posterior probability values are given along branches for both combined analyses, i.e. with full and trimmed plastid data. Members of former Dorema are indicated in boldface. Major clades representing subgenera (Ferula, Leutea, Narthex, Safinia, and Sinoferula) and sections (Euryangium, Ferula, Glaucoselinum, Macrorrhiza, Merwia, Pachycarpa, Peucedanoides, Scorodosma, Soranthus, and Stenocarpa) are bracketed and named according to Panahi et al. (2018). Clade A’ comprising all species from former Dorema, is also marked. For simplicity, outgroup taxa are omitted
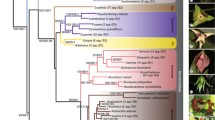
The maximum likelihood tree of 46 representatives of Ferulinae inferred from analysis of combined nuclear (nrDNA ITS and ETS) and trimmed plastid sequence data (rpoB-trnC intergenic spacer, rps16 and rpoC1 introns). Bootstrap support values are given along branches for both combined analyses, i.e. using full and trimmed plastid data. See Fig. 1 for details
Phylogenetic relationships within Ferula
The single representative of the genus Leutea was sister to all studied Ferula species including the six newly analysed species of Dorema (Figs. 1, 2). Monophyly of Ferula was strongly supported in BI analyses and moderately in ML analyses (PP = 1.0/1.0, BS = 71/67%). Ferula licentiana Hand.-Mazz., a representative of subgenus Sinoferula Spalik, Puchałka & M.Panahi, was sister to the remaining congeners. Subgenus Ferula was strongly supported in BI trees and moderately in ML (PP = 1.0/1.0, BS = 76/79%). Ferula koso-poljanskyi Korovin, representing subgenus Safinia Spalik, M.Panahi & Puchałka, was a sister in both BI and ML analyses (PP = 0.93/0.95, BS = 45/43%) to the members of subgenus Narthex, which in turn was well supported by BI and poorly by ML method (PP = 0.92/0.99, BS = 39/54%). Within the subgenus Narthex, four sections, Euryangium (Kauffm.) Pimenov, Merwia (B.Fedtsch.) Koso-Pol., Pachycarpa (Korovin) Banasiak, M.Panahi & Spalik, and Scorodosma (Bunge) Boiss., were clustered in a group highly supported in BI analyses and moderately in ML trees (PP = 0.94/0.94, BS = 61/62%). Moreover, Euryangium, Merwia, and Scorodosma formed a highly supported clade (PP = 1.0/1.0, BS = 92/93%). The remaining sections within the subgenus Narthex—Glaucoselinum (Schischk.) Pimenov, Macrorrhiza Korovin, and Soranthus (Ledeb.) Pimenov—had an uncertain position or were poorly supported by both BI and ML analyses.
Discussion
Phylogenetic position of Dorema
In spite of being described at the beginning of the nineteenth century by Scottish botanist D. Don, a comprehensive taxonomical treatment of the genus Dorema had not been accomplished until the monograph by M.G. Pimenov (Pimenov 1988). This author critically summarised all available information about the genus that was spread out across various floras, historical publications, and monographs. Importantly, knowing that herbarium material for Dorema is often incomplete, he also collected information from the field. He argued that the genus comprised 12 species, for which he generated a simple phenogram (although without providing technical details) based on four vegetative traits of stem and leaves and 12 generative traits of inflorescence, flowers, and fruits. The results were used to propose an infrageneric classification. Pimenov divided the genus into two sections: Dorema (four species) and Microcarpa Pimenov (eight species), which division constitutes the only hitherto proposed hypothesis on infrageneric relationships within Dorema. We included all species from both sections in the present study.
One of the main arguments for the separation of Dorema as a distinct genus was the presence of simple umbels arranged in a panicle-like inflorescence. Pimenov in his monograph emphasised several times the uniqueness of this feature among Apiaceae. For example, he criticised Koso-Poljansky (1925) for the inclusion of Haussknechtia elymaitica Boiss. into Dorema (Dorema elymaitica (Boiss.) Koso-Pol.) arguing that this species is only superficially similar to Dorema because of the compression of compound umbels into capitulum-like structures (Pimenov 1988). Contrary to the morphological arguments, molecular analyses showed unequivocally that Dorema is nested within Ferula (Panahi et al. 2015). The present study corroborated this result for all 12 species of Dorema recognised by Pimenov (1988), including six analysed for the first time. All species were placed in a well-supported clade corresponding to section Peucedanoides according to the new classification of Ferula (Panahi et al. 2018; Figs. 1, 2 and Fig. S1). This section, except Dorema, includes mostly species previously placed by Korovin (1947) in subgenera Peucedanoides and Dorematoides Korovin. Interestingly, within the section, highly supported clade A’ was formed comprising all 12 species formerly placed in Dorema (Figs. 1, 2) and three species—F. dubjanskyi Korovin, F. feruloides and F. caspica—which have proliferating inflorescences: a new compound umbel grows in place of a central umbellet forming therefore a pseudoverticillate inflorescence and restoring its monopodial growth. This type of inflorescence might be considered as transitory between terminal compound umbels typical for most Ferula species and paniculate inflorescence with lateral simple umbels characteristic for Dorema (Fig. 3). According to Ajani & Claβen-Bockhoff (2021), simple umbels of Dorema are homologous to umbellets, while the long branches with simple umbels are homologous to compound umbels. They speculated that this unique inflorescence architecture not only allows for the increase in the number of umbellets per umbel but also enhances their wide spatial exposition along primary and secondary branches forming a conspicuous floral display, which is adaptive in arid environments characterised by short flowering season and pollinator limitation.
An umbel architecture in section Peucedanoides: a main compound umbel and two lateral ones in the form of dichasium (e.g. F. canescens, F. potaninii in clade A′), b ‘proliferous’ inflorescence; a new compound umbel grows in place of a central umbellet forming a pseudoverticillate inflorescence and restoring its monopodial growth (in clade A’: F. dubjanskyi, F. feruloides and F. caspica), and c paniculate inflorescence with simple umbels arranged along the branches; this type characterizes all species formerly described in Dorema
Despite using a highly variable ETS marker, the relationships within the clade A′ remained unresolved. Our analyses did not give a definite answer whether the species formerly placed in Dorema form a monophyletic group. However, the species from Pimenov’s section Microcarpa, formed a clade in some analyses, although with poor support. For example, in maximum likelihood trees based on the combined nuclear and plastid data sets, a clade comprising eight species of which seven are from sect. Microcarpa is apparent (Fig. 2, Fig. S1). These species are distinguished by several characters such as glabrous ovaries, glabrous dorsal side of petals and the lack of secretory canals in ripe mericarps (Pimenov 1988). The only species from Pimenov’s sect. Microcarpa that did not cluster in this group is F. downieorum Spalik & al. (≡ D. aureum Stocks), while the only addition to this group from former sect. Dorema is F. michaelii M.Panahi & al. (≡ D. aitchisonii Korovin ex Pimenov). The four remaining species are intermingled with other species of Ferula and did not form a clade in any analysis. Likely, there is no sufficient variation in the used markers to place all Dorema species in one or several well-supported groups. Therefore, our results cannot be used, for example, to answer questions concerning synonymy of D. sabulosum and D. karataviense postulated by Sennikov and Tojibaev (2021).
The utility of nrDNA ETS marker to resolve phylogenetic relationships within Ferula
Generally, the addition of the ETS marker to the combined phylogenetic analyses confirmed the recent classification system of Ferula based on molecular data (Panahi et al. 2018) and increased support for some internal branches. In particular, the monophyly of subgenus Narthex is well supported by BI reinforcing the taxonomic system proposed by Panahi et al. (2018). Similarly, the division of the subgenus Ferula into two sections, Stenocarpa Puchałka & Spalik and Ferula, was strongly confirmed in BI and ML analyses (Figs. 1, 2). In Panahi et al. (2018), this relationship was poorly supported by combined marker analyses, most likely because of contradictory phylogenetic signals placing F. stenocarpa Boiss. & Hausskn. ex Boiss. as a sister to the representatives of section Ferula only in ITS analysis. The close relationship of these two sections is also supported by morphology as they were formerly placed in sect. Anatriches Korovin, nom. illeg. (≡ sect. Ferula) based on leaf morphology—linear-lanceolate ultimate segments of leaves (Korovin 1947).
In former studies, the relationships among three sections, Euryangium, Scorodosma, and Merwia, were unresolved (Figs. 3 and S5 in Panahi et al. 2018). Here, they form one strongly supported clade (Figs. 1, 2). The taxonomy of these three sections has practical significance, because their members such as F. assa-foetida L., F. gummosa Boiss., and F. sumbul (Kauffm.) Hook.f. are sources of gum resins rich in biologically active chemical compounds of potential medicinal importance.
As exemplified by the present study, the ETS marker is highly variable (Table 2) and easy to sequence even from old herbarium materials, thus constituting a useful supplement to the ITS marker. Both markers can be applied for the tentative placement of scarce herbarium specimens or new species of Ferula on the phylogenetic tree of the genus. Over ten species of Ferula have been described since 2000 (e.g. Brullo et al. 2018, Pimenov and Kljuykov 2013, Ahmad 2013, Akalın et al. 2020) but few, to our knowledge, used ITS as a supplementary source of evidence alongside morphology and anatomy (e.g. Ma et al. 2019; Tuncay et al. 2023). However, even in combination with plastid markers, ETS and ITS do not provide a sufficient phylogenetic signal to answer many questions concerning phylogenetic relationships within Ferula as exemplified by the present study. New genomic methods such as genome skimming or capture-based target enrichment are necessary to answer these questions.
Taxonomic treatment
Ferula badhysiensis Puchałka, Spalik & Piwczyński, nom. nov. ≡ Dorema badhysi Pimenov, Opred. Rast. Sred. Azii 7: 379. 1983, non Ferula badhysi Korovin, Ill. Monogr. Ferula 67. 1947.—TYPE: Afghanistan, Badghis, 14–16 May 1885, Aitchison 462 (holotype: LE; isotype: K000685784).
Etymology: The specific epithet badhysi is not available in Ferula because of F. badhysi, a species described from Turkmenistan and at present synonymised with F. oopoda. To keep the reference to Badghis (Badhys), a province in Afghanistan from which the species was described, we propose the epithet badhysiensis, i.e. from Badhys, retaining the original spelling of the name.
Ferula balchanorum (Pimenov) Puchałka, Spalik & Piwczyński, comb. nov. ≡ Dorema balchanorum Pimenov, Opred. Rast. Sred. Azii 7: 378. 1983.—TYPE: Turkmenistan, Uly Balkan, northern slope, ascent to Mt Arlan, Pat'ma, 10 May 1975, Pimenov et al. 351 (holotype: MW barcodes MW0594061 and MW0594063).
Ferula kopetdagi Spalik & Puchałka, nom. nov. ≡ Dorema kopetdaghense Pimenov, Opred. Rast. Sred. Azii 7: 378. 1983, non Ferula kopetdagensis Korovin, Ill. Monogr. Ferula, 61. 1947.— TYPE: Turkmenistan, Kopet Dag, Firuza ravine, 21 May 1972, MG Pimenov 151 (holotype: MW barcodes MW0594064 and MW0594067; isotypes: LE, MW barcodes MW0594065, MW0594066).
Etymology: The substitute epithet also refers to the Kopet Dag, from where the species was described.
References
Ahmad SA (2013) Ferula shehbaziana (Apiaceae), a new species from Kurdistan, Iraq. Harvard Pap Bot 18:99–100. https://doi.org/10.3100/025.018.0202
Ajani Y, Claβen-Bockhoff R (2021) The unique inflorescence structure of Dorema aucheri (Apiaceae): an adaptation to the arid environment. J Arid Environ 184:104194. https://doi.org/10.1016/j.jaridenv.2020.104194
Akalın E, Tuncay HO, Olcay B, Miski M (2020) A new Ferula (Apiaceae) species from southwest Anatolia: Ferula pisidica Akalın & Miski. Plants 9:740. https://doi.org/10.3390/plants9060740
Brullo S, Brullo C, Cambria S et al (2018) A new species of Ferula (Apiaceae) from Malta. Phytotaxa 382:74–88. https://doi.org/10.11646/phytotaxa.382.1.3
Drude CGO (1898) Umbelliferae. In: Engler A, Prantl K (eds) Die Naturlichen Pflanzenfamilien, vol 3. W. Engelmann, Lepzig
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucl Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Gouy M, Guindon S, Gascuel O (2010) SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Molec Biol Evol 27:221–224. https://doi.org/10.1093/molbev/msp259
Korovin EP (1939) Lignes principales de la systematique du genre Ferula (Tourn.) L. Bull Soc Imp Nat Moscou 48:5–15
Korovin EP (1947) Generis Ferula (Tourn.) L. monographia illustrata. Academiae Scientiarum UzRSS, Tashkent
Koso-Poljansky BM (1925) O Merwia Androssowi B.Fedtsch. 1924. Byull Voronezhsk Obshch Estestvoisp Gosud Univ 1:37–38
Kurzyna-Młynik R, Oskolski AA, Downie SR, Kopacz R, Wojewódzka A, Spalik K (2008) Phylogenetic position of the genus Ferula (Apiaceae) and its placement in tribe Scandiceae as inferred from nrDNA ITS sequence variation. Pl Syst Evol 274:47–66. https://doi.org/10.1007/s00606-008-0022-2
Lanfear R, Calcott B, Ho SYW, Guindon S (2012) PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Molec Biol Evol 29:1695–1701. https://doi.org/10.1093/molbev/mss020
Lanfear R, Calcott B, Kainer D, Mayer C, Stamatakis A (2014) Selecting optimal partitioning schemes for phylogenomic datasets. BMC Evol Biol 14:82. https://doi.org/10.1186/1471-2148-14-82
Logacheva MD, Valiejo-Roman CM, Degtjareva GV, Stratton JM, Downie SR, Samigullin TH, Pimenov M (2010) A comparison of nrDNA ITS and ETS loci for phylogenetic inference in the Umbelliferae: an example from tribe Tordylieae. Molec Phylogen Evol 57:471–476. https://doi.org/10.1016/j.ympev.2010.06.001
Ma X, Liao C, Chen Y, Xu H (2019) Ferula paeoniifolia sp. nov. (Apiaceae) from Sichuan, China. Nord J Bot 37:e01751. https://doi.org/10.1111/njb.01751
Panahi M, Banasiak Ł, Piwczyński M, Puchałka R, Oskolski AA, Spalik K (2015) Phylogenetic relationships among Dorema, Ferula and Leutea (Apiaceae: Scandiceae: Ferulinae) inferred from nrDNA ITS and cpDNA noncoding sequences. Taxon 64:99–112
Panahi M, Banasiak Ł, Piwczyński M, Puchałka R, Kanani MRMR, Oskolski AA, Modnicki D, Miłobędzka A, Spalik K (2018) Taxonomy of the traditional medicinal plant genus Ferula (Apiaceae) is confounded by incongruence between nuclear rDNA and plastid DNA. Bot J Linn Soc 188:173–189. https://doi.org/10.1093/botlinnean/boy055
Pimenov M (1988) Monographic revision of the genus Dorema D. Don (Umbelliferae). Bull Soc Imp Nat Moscou 93:76–90
Pimenov M, Leonov MV (1993) The genera of the Umbelliferae: a nomenclator. Royal Botanic Gardens, Kew
Pimenov MG, Kljuykov EV (2013) Ferula divaricata (Umbelliferae), a new species from Central Anatolia, Turkey. Phytotaxa 99:36–37
Piwczyński M, Wyborska D, Gołębiewska J, Puchałka R (2018) Phylogenetic positions of seven poorly known species of Ferula (Apiaceae) with remarks on the phylogenetic utility of the plastid trnH-psbA, trnS-trnG, and atpB-rbcL intergenic spacers. Syst Biodivers 16:428–440. https://doi.org/10.1080/14772000.2018.1442374
Poczai P, Hyvönen J (2010) Nuclear ribosomal spacer regions in plant phylogenetics: problems and prospects. Molec Biol Rep 37:1897–1912. https://doi.org/10.1007/s11033-009-9630-3
Rambaut A, Drummond A, Xie D, Baele G, Suchard M (2018) Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67:901–904. https://doi.org/10.1093/sysbio/syy032
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Sennikov AN, Tojibaev KS (eds)(2021) Checklist of vascular plants of the Tian-Shan Mountain System. Korea National Arboretum, Pocheon
Shneyer VS, Borschtschenko GP, Pimenov M (1995) Immunochemical appraisal of relationships within the tribe Peucedaneae (Apiaceae). Pl Syst Evol 198:1–16. https://doi.org/10.1007/BF00985104
Stamatakis A (2014) RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Tuncay HO, Akalın E, Doğru-Koca A, Eruçar FM, Miski M (2023) Two new Ferula (Apiaceae) species from Central Anatolia: Ferula turcica and Ferula latialata. Horticulturae 9:144. https://doi.org/10.3390/horticulturae9020144
Yi TS, Jin GH, Wen J (2015) Chloroplast capture and intra- and inter-continental biogeographic diversification in the Asian - New World disjunct plant genus Osmorhiza (Apiaceae). Molec Phylogen Evol 85:10–21. https://doi.org/10.1016/j.ympev.2014.09.028
Acknowledgements
The authors are deeply indebted to Dmitry Lyskov (Moscow State University) for providing material of Dorema species and for his comments on the manuscript. The authors also thank the curators of the herbaria listed in Table S1 for providing plant material; Liliana Szczeparska and Michał Gierek for help in laboratory procedures. This work was supported by the National Science Centre grants no. 2015/18/E/NZ8/00716 to MP and no. 2015/19/B/NZ8/00163 to KS.
Author information
Authors and Affiliations
Contributions
MP and RP contributed to conceptualisation; MP and PT contributed to methodology; MP contributed to formal analysis and investigation; MP, RP, and KS contributed to writing—original draft preparation; ŁB, MP, RP, KS and PT contributed to writing—review and editing; and MP and KS contributed to funding acquisition. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Handling Editor: Christian Parisod.
Supplementary Information
Below is the link to the electronic supplementary material.
Information on electronic supplementary material
Information on electronic supplementary material
Online Resource 1. The maximum likelihood and Bayesian 50% majority-rule consensus trees inferred from analyses of nuclear data, full and trimmed plastid data, and combined nuclear and full plastid data.
Online Resource 2. Accessions of Dorema, Ferula, and outgroups from which nrDNA and pDNA sequence data were obtained, with corresponding voucher information and GenBank reference numbers.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Puchałka, R., Spalik, K., Trzeciak, P. et al. Phylogenetic position of Dorema within Ferula (Apiaceae). Plant Syst Evol 309, 28 (2023). https://doi.org/10.1007/s00606-023-01857-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00606-023-01857-z