Abstract
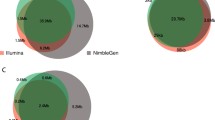
Here we present an adaptation of NimbleGen 2.1M-probe array sequence capture for whole exome sequencing using the Illumina Genome Analyzer (GA) platform. The protocol involves two-stage library construction. The specificity of exome enrichment was approximately 80% with 95.6% even coverage of the 34 Mb target region at an average sequencing depth of 33-fold. Comparison of our results with whole genome shot-gun resequencing results showed that the exome SNP calls gave only 0.97% false positive and 6.27% false negative variants. Our protocol is also well suited for use with whole genome amplified DNA. The results presented here indicate that there is a promising future for large-scale population genomics and medical studies using a whole exome sequencing approach.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Albert T J, Molla M N, Muzny D M, et al. Direct selection of human genomic loci by microarray hybridization. Nat Methods, 2007, 4: 903–905
Levy S, Sutton G, Ng P C, et al. The diploid genome sequence of an individual human. PLoS Biol, 2007, 5: e254
Wang J, Wang W, Li R, et al. The diploid genome sequence of an Asian individual. Nature, 2008, 456: 60–65
Wheeler D A, Srinivasan M, Egholm M, et al. The complete genome of an individual by massively parallel DNA sequencing. Nature, 2008, 452: 872–876
Porreca G J, Zhang K, Li J B, et al. Multiplex amplification of large sets of human exons. Nat Methods, 2007, 4: 931–936
Gnirke A, Melnikov A, Maguire J, et al. Solution hybrid selection with ultra-long oligonucleotides for massively parallel targeted sequencing. Nat Biotechnol, 2009, 27: 182–189
Hodges E, Xuan Z, Balija V, et al. Genome-wide in situ exon capture for selective resequencing. Nat Genet, 2007, 39: 1522–1527
Okou D T, Steinberg K M, Middle C, et al. Microarray-based genomic selection for high-throughput resequencing. Nat Methods, 2007, 4: 907–909
Li H, Durbin R. Fast and accurate short read alignment with Bur rows-Wheeler transform. Bioinformatics, 2009, 25: 1754–1760
Li R, Yu C, Li Y, et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics, 2009, 25: 1966–1967
Li R, Li Y, Fang X, et al. SNP detection for massively parallel whole-genome resequencing. Genome Res, 2009, 19: 1124–1132
Choi M, Scholl U I, Ji W, et al. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc Natl Acad Sci USA, 2009, 106: 19096–19101
McKenna A, Hanna M, Banks E, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res, 2010, 20: 1297–1303
Albers C A, Lunter G, Macarthur D G, et al. Dindel: Accurate indel calls from short-read data. Genome Res, 2010, 21: 961–973
Frazer K A, Murray S S, Schork N J, et al. Human genetic variation and its contribution to complex traits. Nat Rev, 2009, 10: 241–251
Ng S B, Turner E H, Robertson P D, et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature, 2009, 461: 272–276
Ng S B, Bigham A W, Buckingham K J, et al. Exome sequencing identifies MLL2 mutations as a cause of Kabuki syndrome. Nat Genet, 2010, 42: 790–793
Ng S B, Buckingham K J, Lee C, et al. Exome sequencing identifies the cause of a mendelian disorder. Nat Genet, 2010, 42: 30–35
Bilguvar K, Ozturk A K, Louvi A, et al. Whole-exome sequencing identifies recessive WDR62 mutations in severe brain malformations. Nature, 2010, 467: 207–210
Yi X, Liang Y, Huerta-Sanchez E, et al. Sequencing of 50 human exomes reveals adaptation to high altitude. Science, 2010, 329: 75–78
Li Y, Vinckenbosch N, Tian G, et al. Resequencing of 200 human exomes identifies an excess of low-frequency non-synonymous coding variants. Nat Genetics, 2010, 42: 969–972
Author information
Authors and Affiliations
Corresponding author
Additional information
Contributed equally to this work
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Jiang, T., Yang, L., Jiang, H. et al. High-performance single-chip exon capture allows accurate whole exome sequencing using the Illumina Genome Analyzer. Sci. China Life Sci. 54, 945–952 (2011). https://doi.org/10.1007/s11427-011-4232-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-011-4232-4