Abstract
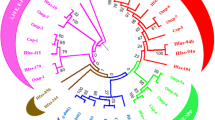
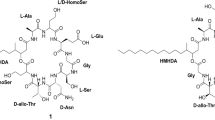
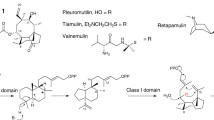
Beauvericin, a cyclohexadepsipeptide-possessing natural product with synergistic antifungal, insecticidal, and cytotoxic activities. We isolated and characterized the fpBeas gene cluster, devoted to beauvericin biosynthesis, from the filamentous fungus Fusarium proliferatum LF061. Targeted inactivation of the F. proliferatum genomic copy of fpBeas abolished the production of beauvericin. Comparative sequence analysis of the FpBEAS showed 74% similarity with the BbBEAS that synthesizes the cyclic trimeric ester beauvericin in Beauveria bassiana, which assembles N-methyl-dipeptidol monomer intermediates by the programmed iterative use of the nonribosomal peptide synthetase modules. Differences between the organization of the beauvericin loci in F. proliferaturm and B. bassiana revealed the mechanism for high production of beauvericin in F. proliferatum. Our work provides new insights into beauvericin biosynthesis, and may lead to beauvericin overproduction and creation of new analogs via synthetic biology approaches.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Zhang L, Yan K, Zhang Y, et al. High-throughput synergy screening identifies microbial metabolites as combination agents for the treatment of fungal infections. Proc Natl Acad Sci USA, 2007, 104: 4606–4611
Hoffman H L, Ernst E J, Klepser M E. Novel triazole antifungal agents. Exp Opin Invest Drugs, 2000, 9: 593–605
Hamill R L, Higgens C E, Boaz H E, et al. The structure of beauvericin, a new depsipeptide antibiotic toxic to Artemia salina. Tetrahedron Lett, 1969, 49: 4255–4258
Roeske R W, Isaac S, Steinrau L, et al. Synthesis and ion-transport properties of peptide antibiotic beauvericin. Fed Proc, 1971, 30: 1282–1288
Schwarzer D, Finking R, Marahiel M A. Nonribosomal peptides: from genes to products. Nat Prod Rep, 2003, 20: 275–287
Finking R, Marahiel M A. Biosynthesis of nonribosomal peptides. Ann Rev Microbiol, 2004, 58: 453–488
Kopp F, Marahiel M A. Macrocyclization strategies in polyketide and nonribosomal peptide biosynthesis. Nat Prod Rep, 2007, 24: 735–749
Pieper R, Haese A, Schroder W, et al. Arrangement of catalytic sites in the multifunctional enzyme enniatin synthetase. Eur J Biochem, 1995, 230: 119–126
Haese A, Schubert M, Herrmann M, et al. Molecular characterization of the enniatin synthetase gene encoding a multifunctional enzyme catalyzing n-methyldepsipeptide formation in fusarium-scirpi. Mol Microbiol, 1993, 7: 905–914
Peeters H, Zocher R, Kleinkauf H. Synthesis of beauvericin by a multifunctional enzyme. J Antibiotics, 1988, 41: 352–359
Xu Y, Orozco R, Wijeratne E M, et al. Biosynthesis of the cyclooligomer depsipeptide beauvericin, a virulence factor of the entomopathogenic fungus beauveria bassiana. Chem Biol, 2008, 15: 898–907
Zhang T, Jia X, Zhuo Y, et al. Cloning and characterization of a novel 2-ketoisovalerate reductase from the beauvericin producer fusarium proliferatum lf061. BMC Biotechnol, 2012, 12: 55
Moretti A, Mule G, Ritieni A, et al. Further data on the production of beauvericin, enniatins and fusaproliferin and toxicity to artemia salina by fusarium species of gibberella fujikuroi species complex. Int J Food Microbiol, 2007, 118: 158–163
Xu Y Q, Orozco R, Wijeratne E M K, et al. Biosynthesis of the cyclooligomer depsipeptide beauvericin, a virulence factor of the entomopathogenic fungus beauveria bassiana. Chem Biol, 2008, 15: 898–907
Lee H S, Song H H, Ahn J H, et al. Statistical optimization of growth medium for the production of the entomopathogenic and phytotoxic cyclic depsipeptide beauvericin from fusarium oxysporum kfcc11363p. J Microbiol Biotechnol, 2008, 18: 138–144
Moller E M, Bahnweg G, Sandermann H, et al. A simple and efficient protocol for isolation of high-molecular-weight DNA from filamentous fungi, fruit bodies, and infected-plant tissues. Nucleic Acids Res, 1992, 20: 6115–6116
Frandsen R J N, Andersson J A, Kristensen M B, et al. Efficient four fragment cloning for the construction of vectors for targeted gene replacement in filamentous fungi. BMC Mol Biol, 2008, 9: 70
Stanke M, Morgenstern B. Augustus: a web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic Acids Res, 2005, 33: W465–W467
Stanke M, Steinkamp R, Waack S, et al. Augustus: a web server for gene finding in eukaryotes. Nucleic Acids Res, 2004, 32: W309–W312
Udwary D W, Merski M, Townsend C A. A method for prediction of the locations of linker regions within large multifunctional proteins, and application to a type I polyketide synthase. J Mol Biol, 2002, 323: 585–598
Rausch C, Weber T, Kohlbacher O, et al. Specificity prediction of adenylation domains in nonribosomal peptide synthetases (NRPS) using transductive support vector machines (TSVMs). Nucleic Acids Res, 2005, 33: 5799–5808
Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol, 1987, 4: 406–425
Tamura K, Dudley J, Nei M, et al. MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol, 2007, 24: 1596–1599
Sharom F J. Abc multidrug transporters: structure, function and role in chemoresistance. Pharmacogenomics, 2008, 9: 105–127
Xu Y Q, Rozco R, Wijeratne E M K, et al. Biosynthesis of the cyclooligomer depsipeptide bassianolide, an insecticidal virulence factor of beauveria bassiana. Fungal Genet Biol, 2009, 46: 353–364
Jirakkakul J, Punya J, Pongpattanakitshote S, et al. Identification of the nonribosomal peptide synthetase gene responsible for bassianolide synthesis in wood-decaying fungus xylaria sp. BCC1067. Microbiol-SGM, 2008, 154: 995–1006
Billich A, Zocher R. N-methyltransferase function of the multifunctional enzyme enniatin synthetase. Biochemistry, 1987, 26: 8417–8423
Glinski M, Urbanke C, Hornbogen T, et al. Enniatin synthetase is a monomer with extended structure: evidence for an intramolecular reaction mechanism. Arch Microbiol, 2002, 178: 267–273
Rausch C, Hoof I, Weber T, et al. Phylogenetic analysis of condensation domains in nrps sheds light on their functional evolution. BMC Evol Biol, 2007, 7: 78
Sieber S A, Marahiel M A. Molecular mechanisms underlying nonribosomal peptide synthesis: approaches to new antibiotics. Chem Rev, 2005, 105: 715–738
Fan Y H, Fang W G, Guo S J, et al. Increased insect virulence in beauveria bassiana strains overexpressing an engineered chitinase. Appl Environ Microbiol, 2007, 73: 295–302
Yu J H, Keller N. Regulation of secondary metabolism in filamentous fungi. Ann Rev Phytopathol, 2005, 43: 437–458
Brakhage A A. Regulation of fungal secondary metabolism. Nat Rev Microbiol, 2013, 11: 21–32
MacPherson S, Larochelle M, Turcotte B. A fungal family of transcriptional regulators: the zinc cluster proteins. Microbiol Mol Biol Rev MMBR, 2006, 70: 583–604
Challis G L, Ravel J, Townsend C A. Predictive, structure-based model of amino acid recognition by nonribosomal peptide synthetase adenylation domains. Chem Biol, 2000, 7: 211–224
von Dohren H, Keller U, Vater J, et al. Multifunctional peptide synthetases. Chem Rev, 1997, 97: 2675–2705
Palmero D, Iglesias C, de Cara M, et al. Species of fusarium isolated from river and sea water of southeastern spain and pathogenicity on four plant species. Plant Dis, 2009, 93: 377–385
Doss R P, Potter S W, Chastagner G A, et al. Adhesion of nongerminated botrytis-cinerea conidia to several substrata. Appl Environ Microbiol, 1993, 59: 1786–1791
Turgeon B G, Oide S, Bushley K. Creating and screening cochliobolus heterostrophus non-ribosomal peptide synthetase mutants. Mycol Res, 2008, 112: 200–206
Wang L, Tian X, Wang J, et al. Autoregulation of antibiotic biosynthesis by binding of the end product to an atypical response regulator. Proc Natl Acad Sci USA, 2009, 106: 8617–8622
Xu G, Wang J, Wang L, et al. Pseudo gamma-butyrolactone receptors respond to antibiotic signals to coordinate antibiotics biosynthesis. J Biol Chem, 2010, 285: 27440–27448
Author information
Authors and Affiliations
Corresponding authors
Additional information
Contributed equally to this work
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Zhang, T., Zhuo, Y., Jia, X. et al. Cloning and characterization of the gene cluster required for beauvericin biosynthesis in Fusarium proliferatum. Sci. China Life Sci. 56, 628–637 (2013). https://doi.org/10.1007/s11427-013-4505-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-013-4505-1