Abstract
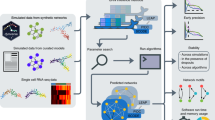
Identification of differential regulators is critical to understand the dynamics of cellular systems and molecular mechanisms of diseases. Several computational algorithms have recently been developed for this purpose by using transcriptome and network data. However, it remains largely unclear which algorithm performs better under a specific condition. Such knowledge is important for both appropriate application and future enhancement of these algorithms. Here, we systematically evaluated seven main algorithms (TED, TDD, TFactS, RIF1, RIF2, dCSA_t2t, and dCSA_r2t), using both simulated and real datasets. In our simulation evaluation, we artificially inactivated either a single regulator or multiple regulators and examined how well each algorithm detected known gold standard regulators. We found that all these algorithms could effectively discern signals arising from regulatory network differences, indicating the validity of our simulation schema. Among the seven tested algorithms, TED and TFactS were placed first and second when both discrimination accuracy and robustness against data variation were considered. When applied to two independent lung cancer datasets, both TED and TFactS replicated a substantial fraction of their respective differential regulators. Since TED and TFactS rely on two distinct features of transcriptome data, namely differential co-expression and differential expression, both may be applied as mutual references during practical application.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Zeng L, Yu J, Huang T, Jia H, Dong Q, He F, Yuan W, Qin L, Li Y, Xie L. Differential combinatorial regulatory network analysis related to venous metastasis of hepatocellular carcinoma. BMC Genomics, 2012, 13(Suppl 8): S14
Tu K, Yu H, Hua YJ, Li YY, Liu L, Xie L, Li YX. Combinatorial network of primary and secondary microRNA-driven regulatory mechanisms. Nucleic Acids Res, 2009, 37: 5969–5980
Ideker T, Krogan NJ. Differential network biology. Mol Syst Biol, 2012, 8: 565
de la Fuente A. From ‘differential expression’ to ‘differential networking’-identification of dysfunctional regulatory networks in diseases. Trends Genet, 2010, 26: 326–333
Fang G, Kuang R, Pandey G, Steinbach M, Myers CL, Kumar V. Subspace differential coexpression analysis: problem definition and a general approach. Pac Symp Biocomput, 2010, 15: 145–156
Zhang B, Li H, Riggins RB, Zhan M, Xuan J, Zhang Z, Hoffman EP, Clarke R, Wang Y. Differential dependency network analysis to identify condition-specific topological changes in biological networks. Bioinformatics, 2009, 25: 526–532
Zhou Q, Hong Y, Zhan Q, Shen Y, Liu Z. Role for Kruppel-like factor 4 in determining the outcome of p53 response to DNA damage. Cancer Res, 2009, 69: 8284–8292
Strano S, Dell’Orso S, Di Agostino S, Fontemaggi G, Sacchi A, Blandino G. Mutant p53: an oncogenic transcription factor. Oncogene, 2007, 26: 2212–2219
Essaghir A, Toffalini F, Knoops L, Kallin A, van Helden J, Demoulin JB. Transcription factor regulation can be accurately predicted from the presence of target gene signatures in microarray gene expression data. Nucleic Acids Res, 2010, 38: e120
Reverter A, Hudson NJ, Nagaraj SH, Perez-Enciso M, Dalrymple BP. Regulatory impact factors: unraveling the transcriptional regulation of complex traits from expression data. Bioinformatics, 2010, 26: 896–904
Hudson NJ, Reverter A, Dalrymple BP. A differential wiring analysis of expression data correctly identifies the gene containing the causal mutation. PLoS Comput Biol, 2009, 5: e1000382
Yang J, Yu H, Liu BH, Zhao Z, Liu L, Ma LX, Li YX, Li YY. DCGL v2.0: An R package for unveiling differential regulation from differential co-expression. PLoS One, 2013, 8: e79729
Huang CL, Lamb J, Chindelevitch L, Kostrowicki J, Guinney J, Delisi C, Ziemek D. Correlation set analysis: detecting active regulators in disease populations using prior causal knowledge. BMC Bioinformatics, 2012, 13: 46
Yu H, Liu BH, Ye ZQ, Li C, Li YX, Li YY. Link-based quantitative methods to identify differentially coexpressed genes and gene pairs. BMC Bioinformatics, 2011, 12: 315
Van den Bulcke T, Van Leemput K, Naudts B, van Remortel P, Ma H, Verschoren A, De Moor B, Marchal K. SynTReN: a generator of synthetic gene expression data for design and analysis of structure learning algorithms. BMC Bioinformatics, 2006, 7: 43
Schaffter T, Marbach D, Floreano D. GeneNetWeaver: in silico benchmark generation and performance profiling of network inference methods. Bioinformatics, 2011, 27: 2263–2270
Hammerman PS, Lawrence MS, Voet D, Jing R, Cibulskis K, Sivachenko A, Stojanov P, McKenna A, Lander ES, Gabriel S, et al. Comprehensive genomic characterization of squamous cell lung cancers. Nature, 2012, 489: 519–525
Mitra R, Edmonds MD, Sun J, Zhao M, Yu H, Eischen CM, Zhao Z. Reproducible combinatorial regulatory networks elucidate novel oncogenic microRNAs in non-small cell lung cancer. RNA, 2014, 20: 1356–1368
Nymark P, Guled M, Borze I, Faisal A, Lahti L, Salmenkivi K, Kettunen E, Anttila S, Knuutila S. Integrative analysis of microRNA, mRNA and aCGH data reveals asbestos- and histology-related changes in lung cancer. Genes Chromosomes Cancer, 2011, 50: 585–597
Smyth GK. Limma: linear models for microarray data. In: Gentleman R, Carey V, Dudoit S, Irizarry R, Huber W, eds. Bioinformatics and Computational Biology Solutions using R and Bioconductor. New York: Springer, 2005
Matys V, Kel-Margoulis OV, Fricke E, Liebich I, Land S, Barre-Dirrie A, Reuter I, Chekmenev D, Krull M, Hornischer K, Voss N, Stegmaier P, Lewicki-Potapov B, Saxel H, Kel AE, Wingender E. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res, 2006, 34: D108–110
Kel AE, Gossling E, Reuter I, Cheremushkin E, Kel-Margoulis OV, Wingender E. MATCH: a tool for searching transcription factor binding sites in DNA sequences. Nucleic Acids Res, 2003, 31: 3576–3579
Sun J, Gong X, Purow B, Zhao Z. Uncovering microRNA and transcription factor mediated regulatory networks in glioblastoma. PLoS Comput Biol, 2012, 8: e1002488
Rimmele P, Komatsu J, Hupe P, Roulin C, Barillot E, Dutreix M, Conseiller E, Bensimon A, Moreau-Gachelin F, Guillouf C. Spi-1/PU.1 oncogene accelerates DNA replication fork elongation and promotes genetic instability in the absence of DNA breakage. Cancer Res, 2010, 70: 6757–6766
Romero OA, Torres-Diz M, Pros E, Savola S, Gomez A, Moran S, Saez C, Iwakawa R, Villanueva A, Montuenga LM, Kohno T, Yokota J, Sanchez-Cespedes M. MAX inactivation in small cell lung cancer disrupts MYC-SWI/SNF programs and is synthetic lethal with BRG1. Cancer Discov, 2014, 4: 292–303
Chen L, Wei T, Si X, Wang Q, Li Y, Leng Y, Deng A, Chen J, Wang G, Zhu S, Kang J. Lysine acetyltransferase GCN5 potentiates the growth of non-small cell lung cancer via promotion of E2F1, cyclin D1, and cyclin E1 expression. J Biol Chem, 2013, 288: 14510–14521
An O, Pendino V, D’Antonio M, Ratti E, Gentilini M, Ciccarelli FD. NCG 4.0: the network of cancer genes in the era of massive mutational screenings of cancer genomes. Database (Oxford), 2014, 2014: bau015
Isalan M, Lemerle C, Michalodimitrakis K, Horn C, Beltrao P, Raineri E, Garriga-Canut M, Serrano L. Evolvability and hierarchy in rewired bacterial gene networks. Nature, 2008, 452: 840–845
Bhattacharyya M, Bandyopadhyay S. Studying the differential co-expression of microRNAs reveals significant role of white matter in early Alzheimer’s progression. Mol Biosyst, 2013, 9: 457–466
Libermann TA, Zerbini LF. Targeting transcription factors for cancer gene therapy. Curr Gene Ther, 2006, 6: 17–33
Darnell JE Jr. Transcription factors as targets for cancer therapy. Nat Rev Cancer, 2002, 2: 740–749
Ravasi T, Suzuki H, Cannistraci CV, Katayama S, Bajic VB, Tan K, Akalin A, Schmeier S, Kanamori-Katayama M, Bertin N, Carninci P, Daub CO, Forrest AR, Gough J, Grimmond S, Han JH, Hashimoto T, Hide W, Hofmann O, Kamburov A, Kaur M, Kawaji H, Kubosaki A, Lassmann T, van Nimwegen E, MacPherson CR, Ogawa C, Radovanovic A, Schwartz A, Teasdale RD, Tegnér J, Lenhard B, Teichmann SA, Arakawa T, Ninomiya N, Murakami K, Tagami M, Fukuda S, Imamura K, Kai C, Ishihara R, Kitazume Y, Kawai J, Hume DA, Ideker T, Hayashizaki Y. An atlas of combinatorial transcriptional regulation in mouse and man. Cell, 2010, 140: 744–752
Goncalves JP, Francisco AP, Mira NP, Teixeira MC, Sa-Correia I, Oliveira AL, Madeira SC. TFRank: network-based prioritization of regulatory associations underlying transcriptional responses. Bioin-formatics, 2011, 27: 3149–3157
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at springerlink.fh-diploma.de
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Yu, H., Mitra, R., Yang, J. et al. Algorithms for network-based identification of differential regulators from transcriptome data: a systematic evaluation. Sci. China Life Sci. 57, 1090–1102 (2014). https://doi.org/10.1007/s11427-014-4762-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-014-4762-7