Abstract
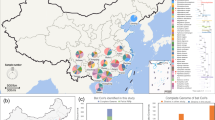
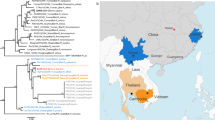
Coronaviruses, such as severe acute respiratory syndrome coronavirus and Middle East respiratory syndrome coronavirus, pose significant public health threats. Bats have been suggested to act as natural reservoirs for both these viruses, and periodic monitoring of coronaviruses in bats may thus provide important clues about emergent infectious viruses. The Eastern bent-wing bat Miniopterus fuliginosus is distributed extensively throughout China. We therefore analyzed the genetic diversity of coronaviruses in samples of M. fuliginosus collected from nine Chinese provinces during 2011–2013. The only coronavirus genus found was Alphacoronavirus. We established six complete and five partial genomic sequences of alphacoronaviruses, which revealed that they could be divided into two distinct lineages, with close relationships to coronaviruses in Miniopterus magnater and Miniopterus pusillus. Recombination was confirmed by detecting putative breakpoints of Lineage 1 coronaviruses in M. fuliginosus and M. pusillus (Wu et al., 2015), which supported the results of topological and phylogenetic analyses. The established alphacoronavirus genome sequences showed high similarity to other alphacoronaviruses found in other Miniopterus species, suggesting that their transmission in different Miniopterus species may provide opportunities for recombination with different alphacoronaviruses. The genetic information for these novel alphacoronaviruses will improve our understanding of the evolution and genetic diversity of coronaviruses, with potentially important implications for the transmission of human diseases.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Apweiler, R., Attwood, T.K., Bairoch, A., Bateman, A., Birney, E., Biswas, M., Bucher, P., Cerutti, L., Corpet, F., Croning, M.D., Durbin, R., Falquet, L., Fleischmann, W., Gouzy, J., Hermjakob, H., Hulo, N., Jonassen, I., Kahn, D., Kanapin, A., Karavidopoulou, Y., Lopez, R., Marx, B., Mulder, N.J., Oinn, T.M., Pagni, M., Servant, F., Sigrist, C.J., and Zdobnov, E.M. (2001). The InterPro database, an integrated documentation resource for protein families, domains and functional sites. Nucleic Acids Res 29, 37–40.
Bateman, A., Birney, E., Cerruti, L., Durbin, R., Etwiller, L., Eddy, S.R., Griffiths-Jones, S., Howe, K.L., Marshall, M., and Sonnhammer, E.L. (2002). The Pfam protein families database. Nucleic Acids Res 30, 276–280.
Bosch, B.J., van der Zee, R., de Haan, C.A., and Rottier, P.J. (2003). The coronavirus spike protein is a class I virus fusion protein: structural and functional characterization of the fusion core complex. J Virol 77, 8801–8811.
Chen, L.L., Ou, H.Y., Zhang, R., and Zhang, C.T. (2003). ZCURVE_CoV: a new system to recognize protein coding genes in coronavirus genomes, and its applications in analyzing SARS-CoV genomes. Biochem Biophys Res Commun 307, 382–388.
Chu, D.K., Peiris, J.S., Chen, H., Guan, Y., and Poon, L.L. (2008). Genomic characterizations of bat coronaviruses (1A, 1B and HKU8) and evidence for co-infections in Miniopterus bats. J Gen Virol 89, 1282–1287.
Cui, J., Han, N., Streicker, D., Li, G., Tang, X., Shi, Z., Hu, Z., Zhao, G., Fontanet, A., Guan, Y., Wang, L., Jones, G., Field, H.E., Daszak, P., and Zhang, S. (2007). Evolutionary relationships between bat coronaviruses and their hosts. Emerg Infect Dis 13, 1526–1532.
King, A.M.Q., Adams, M.J., Carstens, E.B. (2011). Virus Taxonomy, Classification and Nomenclature of Viruses. Ninth Report of the International Committee on Taxonomy of Viruses, International Union of Microbiological Societies, Virology Division. London: Elsevier Academic Press, 806–828.
Dveksler, G.S., Pensiero, M.N., Cardellichio, C.B., Williams, R.K., Jiang, G.S., Holmes, K.V., and Dieffenbach, C.W. (1991). Cloning of the mouse hepatitis virus (MHV) receptor: expression in humaaan and hamster cell lines confers susceptibility to MHV. J Virol 65, 6881–6891.
Gonzalez, J.M., Gomez-Puertas, P., Cavanagh, D., Gorbalenya, A.E., and Enjuanes, L. (2003). A comparative sequence analysis to revise the current taxonomy of the family Coronaviridae. Arch Virol 148, 2207–2235.
Graham, R.L., and Baric, R.S. (2010). Recombination, reservoirs, and the modular spike: mechanisms of coronavirus cross-species transmission. J Virol 84, 3134–3146.
Guindon, S., Dufayard, J.F., Lefort, V., Anisimova, M., Hordijk, W., and Gascuel, O. (2010). New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59, 307–321.
Herrewegh, A.A., Smeenk, I., Horzinek, M.C., Rottier, P.J., and de Groot, R.J. (1998). Feline coronavirus type IIstrains 79-1683 and 79-1146 originate from a double recombination between feline coronavirus type I and canine coronavirus. J Virol 72, 4508–4514.
Holmes, K.V., and Enjuanes, L. (2003). Virology. The SARS coronavirus: a postgenomic era. Science 300, 1377–1378.
Jonassen, C.M., Kofstad, T., Larsen, I.L., Lovland, A., Handeland, K., Follestad, A., and Lillehaug, A. (2005). Molecular identification and characterization of novel coronaviruses infecting graylag geese (Anser anser), feral pigeons (Columbia livia) and mallards (Anas platyrhynchos). J Gen Virol 86, 1597–1607.
Kosakovsky Pond, S.L., Posada, D., Gravenor, M.B., Woelk, C.H., and Frost, S.D. (2006). GARD: a genetic algorithm for recombination detection. Bioinformatics 22, 3096–3098.
Lai, M.M. (1990). Coronavirus: organization, replication and expression of genome. Annu Rev Microbiol 44, 303–333.
Lai, M.M., and Cavanagh, D. (1997). The molecular biology of coronaviruses. Adv Virus Res 48, 1–100.
Lai, M.M.C., and Holmes, K.V. (2001). Coronaviruses. In: Knipe, D.M., Howley, P.M., Griffin, D.E., Lamb, R.A., Martin, M.A., Roizman, B., and Straus, S.E., eds. Fields Virology. Philadelphia: Lippincott Williams & Wilkins 1163–1185.
Lau, S.K., Li, K.S., Tsang, A.K., Shek, C.T., Wang, M., Choi, G.K., Guo, R., Wong, B.H., Poon, R.W., Lam, C.S., Wang, S.Y., Fan, R.Y., Chan, K.H., Zheng, B.J., Woo, P.C., and Yuen, K.Y. (2012a). Recent trans mission of a novel alphacoronavirus, bat coronavirus HKU10, from Leschenault’s rousettes to pomona leaf-nosed bats: first evidence of interspecies transmission of coronavirus between bats of different suborders. J Virol 86, 11906–11918.
Lau, S.K., Woo, P.C., Li, K.S., Huang, Y., Tsoi, H.W., Wong, B.H., Wong, S.S., Leung, S.Y., Chan, K.H., and Yuen, K.Y. (2005). Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proc Natl Acad Sci USA 102, 14040–14045.
Lau, S.K., Woo, P.C., Li, K.S., Huang, Y., Wang, M., Lam, C.S., Xu, H., Guo, R., Chan, K.H., Zheng, B.J., and Yuen, K.Y. (2007). Complete genome sequence of bat coronavirus HKU2 from Chinese horseshoe bats revealed a much smaller spike gene with a different evolutionary lineage from the rest of the genome. Virology 367, 428–439.
Lau, S.K., Woo, P.C., Yip, C.C., Fan, R.Y., Huang, Y., Wang, M., Guo, R., Lam, C.S., Tsang, A.K., Lai, K.K., Chan, K.H., Che, X.Y., Zheng, B.J., and Yuen, K.Y. (2012b). Isolation and characterization of a novel Betacoronavirus subgroup A coronavirus, rabbit coronavirus HKU14, from domestic rabbits. J Virol 86, 5481–5496.
Li, W., Shi, Z., Yu, M., Ren, W., Smith, C., Epstein, J.H., Wang, H., Crameri, G., Hu, Z., Zhang, H., Zhang, J., McEachern, J., Field, H., Daszak, P., Eaton, B.T., Zhang, S., and Wang, L.F. (2005). Bats are natural reservoirs of SARS-like coronaviruses. Science 310, 676–679.
Liu, S., Chen, J., Kong, X., Shao, Y., Han, Z., Feng, L., Cai, X., Gu, S., and Liu, M. (2005). Isolation of avian infectious bronchitis coronavirus from domestic peafowl (Pavo cristatus) and teal (Anas). J Gen Virol 86, 719–725.
Makino, S., Keck, J.G., Stohlman, S.A., and Lai, M.M. (1986). High-frequency RNA recombination of murine coronaviruses. J Virol 57, 729–737.
Miller-Butterworth, C.M., Jacobs, D.S., and Harley, E.H. (2003). Strong population substructure is correlated with morphology and ecology in a migratory bat. Nature 424, 187–191.
Rest, J.S., and Mindell, D.P. (2003). SARS associated coronavirus has a recombinant polymerase and coronaviruses have a history of host-shifting. Infect Genet Evol 3, 219–225.
Shirato, K., Maeda, K., Tsuda, S., Suzuki, K., Watanabe, S., Shimoda, H., Ueda, N., Iha, K., Taniguchi, S., Kyuwa, S., Endoh, D., Matsuyama, S., Kurane, I., Saijo, M., Morikawa, S., Yoshikawa, Y., Akashi, H., and Mizutani, T. (2012). Detection of bat coronaviruses from Miniopterus fuliginosus in Japan. Virus Genes 44, 40–44.
Song, H.D., Tu, C.C., Zhang, G.W., Wang, S.Y., Zheng, K., Lei, L.C., Chen, Q.X., Gao, Y.W., Zhou, H.Q., Xiang, H., Zheng, H.J., Chern, S.W., Cheng, F., Pan, C.M., Xuan, H., Chen, S.J., Luo, H.M., Zhou, D.H., Liu, Y.F., He, J.F., Qin, P.Z., Li, L.H., Ren, Y.Q., Liang, W.J., Yu, Y.D., Anderson, L., Wang, M., Xu, R.H., Wu, X.W., Zheng, H.Y., Chen, J.D., Liang, G., Gao, Y., Liao, M., Fang, L., Jiang, L.Y., Li, H., Chen, F., Di, B., He, L.J., Lin, J.Y., Tong, S., Kong, X., Du, L., Hao, P., Tang, H., Bernini, A., Yu, X.J., Spiga, O., Guo, Z.M., Pan, H.Y., He, W.Z., Manuguerra, J.C., Fontanet, A., Danchin, A., Niccolai, N., Li, Y.X., Wu, C.I., and Zhao, G.P. (2005). Cross-host evolution of severe acute respiratory syndrome coronavirus in palm civet and human. Proc Natl Acad Sci USA 102, 2430–2435.
Sonnhammer, E.L., von Heijne, G., and Krogh, A. (1998). A hidden Markov model for predicting transmembrane helices in protein sequences. Proc Int Conf Intell Syst Mol Biol 6, 175–182.
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., and Kumar, S. (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28, 2731–2739.
Tang, X.C., Zhang, J.X., Zhang, S.Y., Wang, P., Fan, X.H., Li, L.F., Li, G., Dong, B.Q., Liu, W., Cheung, C.L., Xu, K.M., Song, W.J., Vijaykrishna, D., Poon, L.L., Peiris, J.S., Smith, G.J., Chen, H., and Guan, Y. (2006). Prevalence and genetic diversity of coronaviruses in bats from China. J Virol 80, 7481–7490.
van Boheemen, S., de Graaf, M., Lauber, C., Bestebroer, T.M., Raj, V.S., Zaki, A.M., Osterhaus, A.D., Haagmans, B.L., Gorbalenya, A.E., Snijder, E.J., and Fouchier, R.A. (2012). Genomic characterization of a newly discovered coronavirus associated with acute respiratory distress syndrome in humans. MBio doi: 10.1128/mBio.00473-12.
Vijaykrishna, D., Smith, G.J., Zhang, J.X., Peiris, J.S., Chen, H., and Guan, Y. (2007). Evolutionary insights into the ecology of coronaviruses. J Virol 81, 4012–4020.
Weiss, S.R., and Navas-Martin, S. (2005). Coronavirus pathogenesis and the emerging pathogen severe acute respiratory syndrome coronavirus. Microbiol Mol Biol Rev 69, 635–664.
Woo, P.C., Lau, S.K., Lam, C.S., Lai, K.K., Huang, Y., Lee, P., Luk, G.S., Dyrting, K.C., Chan, K.H., and Yuen, K.Y. (2009). Comparative analysis of complete genome sequences of three avian coronaviruses reveals a novel group 3c coronavirus. J Virol 83, 908–917.
Woo, P.C., Lau, S.K., Lam, C.S., Lau, C.C., Tsang, A.K., Lau, J.H., Bai, R., Teng, J.L., Tsang, C.C., Wang, M., Zheng, B.J., Chan, K.H., and Yuen, K.Y. (2012). Discovery of seven novel Mammalian and avian coronaviruses in the genus deltacoronavirus supports bat coronaviruses as the gene source of alphacoronavirus and betacoronavirus and avian coronaviruses as the gene source of gammacoronavirus and deltacoronavirus. J Virol 86, 3995–4008.
Woo, P.C., Lau, S.K., Li, K.S., Poon, R.W., Wong, B.H., Tsoi, H.W., Yip, B.C., Huang, Y., Chan, K.H., and Yuen, K.Y. (2006a). Molecular diversity of coronaviruses in bats. Virology 351, 180–187.
Woo, P.C., Lau, S.K., Yip, C.C., Huang, Y., Tsoi, H.W., Chan, K.H., and Yuen, K.Y. (2006b). Comparative analysis of 22 coronavirus HKU1 genomes reveals a novel genotype and evidence of natural recombination in coronavirus HKU1. J Virol 80, 7136–7145.
Woo, P.C., Lau, S.K., and Yuen, K.Y. (2006c). Infectious diseases emerging from Chinese wet-markets: zoonotic origins of severe respiratory viral infections. Curr Opin Infect Dis 19, 401–407.
Woo, P.C., Lau, S.K., Huang, Y., and Yuen, K.Y. (2009). Coronavirus diversity, phylogeny and interspecies jumping. Exp Biol Med 234, 1117–1127.
Woo, P.C., Lau, S.K., and Yuen, K.Y. (2006). Infectious diseases emerging from Chinese wet-markets: zoonotic origins of severe respiratory viral infections. Curr Opin Infect Dis 19, 401–407.
Wu, Z., Yang, L., Ren, X., He, G., Zhang, J., Yang, J., Qian, Z., Dong, J., Sun, L., Zhu, Y., Du, J., Yang, F., Zhang, S., and Jin, Q. (2015). Deciphering the bat virome catalog to better understand the ecological diversity of bat viruses and the bat origin of emerging infectious diseases. ISME J 10, 609–620.
Zeng, Q., Langereis, M.A., van Vliet, A.L., Huizinga, E.G., and de Groot, R.J. (2008). Structure of coronavirus hemagglutinin-esterase offers insight into corona and influenza virus evolution. Proc Natl Acad Sci USA 105, 9065–9069.
Author information
Authors and Affiliations
Corresponding authors
Additional information
This article is published with open access at springerlink.fh-diploma.de
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Du, J., Yang, L., Ren, X. et al. Genetic diversity of coronaviruses in Miniopterus fuliginosus bats. Sci. China Life Sci. 59, 604–614 (2016). https://doi.org/10.1007/s11427-016-5039-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-016-5039-0