Abstract
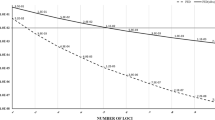
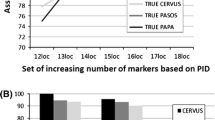
Captive populations provide a precious genetic resource for endangered animals and a source of individuals for reintroduction to depleted habitats. Therefore, accuracy in determining paternity is of vital importance for managing captive populations and in selecting representative individuals of known genetic characteristics for release. In this study, we established a fast and effective method to conduct paternity testing for captive giant pandas in the Wolong population. This technique uses two highly polymorphic microsatellites initially, subsequent use of five less polymorphic markers and then paternity exclusion testing carried out using the giant panda paternity exclusion program we have developed. Our results revealed that (1) both sets of markers successfully identified the real fathers in 25 cases of paternity testing and (2) the success rate of paternity exclusion varied with the degree of polymorphism of the markers used. Subsequently, we conducted correlation analysis between the success rates of paternity identification with these markers, parameters of genetic diversity and tests of neutrality. We found that the paternity exclusion power of microsatellites was significantly correlated with the number of alleles (Na), expected heterozygosity (H E) and observed homozygosity statistic (F O) (all P < 0.05). From this, we developed a new variable, Na×H E/F O, showing a highly significant positive correlation with the resolution power of microsatellites (P = 0.001). Moreover, the first two highly polymorphic loci gave a 100% success rate of excluding non-paternal males because they yielded higher values of Na×H E/F O than the other five less polymorphic markers. Thus, the Na×H E/F O parameter appears suitable to serve as a criterion for selecting microsatellite markers, which could be used for high-resolution molecular techniques of paternity determination among a range of captive animals besides giant pandas.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Frankham R, Ballou J D, Briscoe D A. Introduction to Conservation Genetics. New York: Cambridge University Press, 2002
Howard J G, Donoghue A M, Barone M A, et al. Successful induction of ovarian activity and laparoscopic intrauterine artificial insemination in the Cheetah (Acinonyx jubatus). J Zoo Wildl Med, 1992, 23: 288–300
Wolf K N, Wildt D E, Vargas A, et al. Age dependent changes in sperm production, semen quality and testicular volume in the blackfooted ferret (Mustela nigripes). Biol Reprod, 2000, 63: 179–187
Ungerfeld R, González-Pensado S, Bielli A, et al. Reproductive biology of the pampas deer (Ozotoceros bezoarticus): A review. Acta Vet Scand, 2008, 50: 16
Andrabi S M, Maxwell W M. A review on reproductive biotechnologies for conservation of endangered mammalian species. Anim Reprod Sci, 2007, 99: 223–243
Xie Z, Gipps J. The 2010 International Studbook for Giant Panda (Ailuropoda melanoleuca). Beijing: Chinese Association of Zoological Garden, 2010
State Forestry Administration of China. The Third National Survey Report on Giant Panda in China. Beijing: Science Press, 2006
Wan Q H, Yu J Q, Yong Y G, et al. Reliable genetic analyses based on formalin-fixed materials: A new noninvasive sampling perspective for the giant panda. Conserv Genet, 2006, 7: 467–471
Brock M K, Whiteo B N. Application of DNA fingerprinting to the recovery program of the endangered Puerto Rican parrot. Proc Natl Acad Sci USA, 1992, 89: 11121–11125
Questiau S, Eybert M C, Taberlet P. Amplified fragment length polymorphism (AFLP) markers reveal extra-pair parentage in a bird species: The bluethroat (Luscinia svecica). Mol Ecol, 1999, 8: 1331–1339
Huang C W, Cheng Y S, Rouvier R, et al. AFLP fingerprinting for paternity testing in ducks. Br Poul Sci, 2007, 48: 323–330
Reis S P, Gonçalves E C, Silva A, et al. Genetic variability and efficiency of DNA microsatellite markers for paternity testing in horse breeds from the Brazilian Marajó archipelago. Genet Mol Biol, 2008, 31: 68–72
Paredes M, Norambuena M C, Molina B. Genetic diversity analysis in 12 microsatellite loci, used in equine paternity test in Chile. Arch Zoo, 2009, 58: 111–116
Stevanovic J, Stanimirovic Z, Dimitrijevic V, et al. Evaluation of 11 microsatellite loci for their use in paternity testing in Yugoslav Pied cattle (YU Simmental cattle). Czech J Anim Sci, 2010, 55: 221–226
Wan Q H, Wu H, Fujihara T, et al. Which genetic marker for which conservation genetic issue? Electrophoresis, 2004, 25: 2165–2176
Zhang H M, Guo Y, Li D S, et al. Sixteen novel microsatellite loci developed for the giant panda (Ailuropoda melanoleuca). Conserv Genet, 2009, 10: 589–592
Zane L, Bargelloni L, Patarnello T. Strategies for microsatellite isolation: A review. Mol Ecol, 2002, 11: 1–16
Sambrook J, Russell D W. Molecular Cloning: A Laboratory Manual, 3rd ed. Cold Spring Harbor: Cold Spring Harbor Laboratory Press, 2001
Belkhir K P, Borsa P, Chikhi L, et al. GENETIX, logiciel sous Windows™ pour la génétique des populations. Montpellier: Laboratoire Génome, Populations, Interactions CNRS UMR 5000, Universitéde Montpellier II, 2001
Raymond M, Rousset F. GENEPOP (version 1.2): Population genetics software for exact tests and ecumenicism. J Hered, 1995, 86: 248–249
Lancaster A, Nelson M P, Meyer D, et al. PyPop: A software framework for population genomics: Analyzing large-scale multi-locus genotype data. In: Altman R B, Dunker A K, Hunter L, et al., eds. Pacific Symposium on Biocomputing 8 Singapore: World Scientific, 2003. 514–525
Lancaster A, Single R M, Solberg O D, et al. PyPop update—A software pipeline for large-scale multilocus population genomics. Tissue Antigens, 2007, 69: 192–197
SPSS Inc. SPSS 16.0 for Windows, Chicago: SPSS Inc., 2007
Fang S G, Wan Q H, Wu H, et al. Conservation Genetics of the Giant Panda. Beijing: Science Press, 2008
Kalinowski S T, Taper M L, Marshall T C. Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol Ecol, 2007, 16: 1099–1106
Gerber S, Chabrier P, Kremer A. FaMoz: A software for parentage analysis using dominant, codominant and uniparentally inherited markers. Mol Ecol Notes, 2003, 3: 479–481
Fung W K. User-friendly programs for easy calculations in paternity testing and kinship determinations. Forensic Sci Int, 2003, 136: 22–34
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Li, D., Cui, H., Wang, C. et al. A fast and effective method to perform paternity testing for Wolong giant pandas. Chin. Sci. Bull. 56, 2559–2564 (2011). https://doi.org/10.1007/s11434-011-4571-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-011-4571-3